Abstract
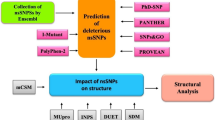
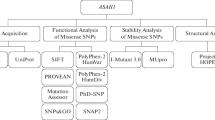
The 2-hydroxyglutaric aciduria (2-HGA) is a rare neurometabolic disorder that leads to the development of brain damage. It is classified into three categories: D-2-HGA, L-2-HGA, and combined D,L-2-HGA. The D-2-HGA includes two subtypes: type I and type II caused by the mutations in D2HGDH and IDH2 proteins, respectively. In this study, we studied six mutations, four in the D2HGDH (I147S, D375Y, N439D, and V444A) and two in the IDH2 proteins (R140G, R140Q). We performed in silico analysis to investigate the pathogenicity and stability changes of the mutant proteins using pathogenicity (PANTHER, PhD-SNP, SIFT, SNAP, and META-SNP) and stability (i-Mutant, MUpro, and iStable) predictors. All the mutations of both D2HGDH and IDH2 proteins were predicted as disease causing except V444A, which was predicted as neutral by SIFT. All the mutants were also predicted to be destabilizing the protein except the mutants D375Y and N439D. DSSP plugin of the PyMOL and Molecular Dynamics Simulations (MDS) were used to study the structural changes in the mutant proteins. In the case of D2HGDH protein, the mutations I147S and V444A that are positioned in the beta sheet region exhibited higher Root Mean Square Deviation (RMSD), decrease in compactness and number of intramolecular hydrogen bonds compared to the mutations N439D and D375Y that are positioned in the turn and loop region, respectively. While the mutants R140Q and R140QG that are positioned in the alpha helix region of the protein. MDS results revealed the mutation R140Q to be more destabilizing (higher RMSD values, decrease in compactness and number of intramolecular hydrogen bonds) compared to the mutation R140G of the IDH2 protein. This study is expected to serve as a platform for drug development against 2-HGA and pave the way for more accurate variant assessment and classification for patients with genetic diseases.




Similar content being viewed by others
Abbreviations
- D-2-HGA:
-
D-2-hydroxyglutaric aciduria
- L-2-HGA:
-
L-2-hydroxyglutaric aciduria
- D, L-2-HGA:
-
Combined D, L-2-hydroxyglutaric aciduria
- D2HGDH:
-
D-2-Hydroxyglutarate Dehydrogenase
- IDH2:
-
Isocitrate dehydrogenase
- ddG:
-
stability free energy change
- RMSD:
-
Root Mean Square Deviation
- Rg:
-
Radius of Gyration
- DSSP:
-
Database of secondary structure assignments (and much more) for all protein entries in the Protein Data Bank
- MDS:
-
Molecular Dynamics Simulation
References
Achouri Y, Noël G, Vertommen D et al (2004) Identification of a dehydrogenase acting on D-2-hydroxyglutarate. Biochem J 381:35–42
Agrahari AK, Sneha P, George Priya Doss C et al (2017) A profound computational study to prioritize the disease-causing mutations in PRPS1 gene. Metab Brain Dis. https://doi.org/10.1007/s11011-017-0121-2
Agrahari AK, Kumar A, Siva R et al (2018) Substitution impact of highly conserved arginine residue at position 75 in GJB1 gene in association with X-linked Charcot-Marie-tooth disease: a computational study. J Theor Biol 437:305–317
Akbay EA, Moslehi J, Christensen CL et al (2014) D-2-hydroxyglutarate produced by mutant IDH2 causes cardiomyopathy and neurodegeneration in mice. Genes Dev 28:479–490
Ali SK, Doss CGP, Kumar DT, Zhu H (2015) CoagVDb: a comprehensive database for coagulation factors and their associated SAPs. Biol Res 48:1–8
Ali SK, Sneha P, Priyadharshini Christy J et al (2016) Molecular dynamics-based analyses of the structural instability and secondary structure of the fibrinogen gamma chain protein with the D356V mutation. J BiomolStructDyn:1–11
Amiel J, de Lonlay P, Francannet C et al (1999) Facial anomalies in D-2-hydroxyglutaric aciduria. Am J Med Genet 86:124–129
Avnir Y, Prachanronarong KL, Zhang Z et al (2017) Structural determination of the broadly reactive anti-IGHV1-69 anti-idiotypic antibody G6 and its Idiotope. Cell Rep 21:3243–3255
Balss J, Pusch S, Beck A-C et al (2012) Enzymatic assay for quantitative analysis of (d)-2-hydroxyglutarate. Acta Neuropathol 124:883–891
Bayar A, Acun C, Dursun A, Verhoeven N, Bonafe L, Keser S, Superti-Furga A (2005) Metaphyseal enchondrodysplasia with 2-hydroxy-glutaric aciduria: observation of a third case and further delineation. Clin Dysmorphol 14:7–11
Berendsen HJC, Postma JPM, van Gunsteren WF, et al (1984) Molecular dynamics with coupling to an external bath. J Chem Phys 81:3684–3690
Biasini M, Bienert S, Waterhouse A et al (2014) SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res 42:W252–W258
Bulka B, desJardins M, Freeland SJ (2006) An interactive visualization tool to explore the biophysical properties of amino acids and their contribution to substitution matrices. BMC Bioinformatics 7:329
Capriotti E, Fariselli P, Casadio R (2005) I-Mutant2.0: predicting stability changes upon mutation from the protein sequence or structure. Nucleic Acids Res 33:W306–W310
Capriotti E, Calabrese R, Casadio R (2006) Predicting the insurgence of human genetic diseases associated to single point protein mutations with support vector machines and evolutionary information. Bioinformatics 22:2729–2734
Capriotti E, Altman RB, Bromberg Y (2013) Collective judgment predicts disease-associated single nucleotide variants. BMC Genomics 14:S2
Carlson B (2008) SNPs-A Shortcut to Personalized Medicine | GEN. https://www.genengnews.com/gen-articles/snpsa-shortcut-to-personalized-medicine/2507. Accessed 21 Feb 2018
Chalmers RA, Lawson AM, Watts RW et al (1980) D-2-hydroxyglutaric aciduria: case report and biochemical studies. J Inherit Metab Dis 3:11–15
Chen C-W, Lin J, Chu Y-W (2013) iStable: off-the-shelf predictor integration for predicting protein stability changes. BMC Bioinformatics 14 Suppl 2:S5
Cheng J, Randall A, Baldi P (2005) Prediction of protein stability changes for single-site mutations using support vector machines. Proteins StructFunctBioinforma 62:1125–1132
Clarke NF, Andrews I, Carpenter K et al (2003) D-2-hydroxyglutaric aciduria: a case with an intermediate phenotype and prenatal diagnosis of two affected fetuses. Am J Med Genet 120A:523–527
Dang L, White DW, Gross S et al (2009) Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature 462:739–744
Doss CGP, Alasmar DR, Bux RI et al (2016) Genetic epidemiology of Glucose-6-phosphate dehydrogenase deficiency in the Arab world. Sci Rep 6:37284
Essmann U, Perera L, Berkowitz ML et al (1995) A smooth particle mesh Ewald method. J Chem Phys 103:8577
García AE, Sanbonmatsu KY (2002) Alpha-helical stabilization by side chain shielding of backbone hydrogen bonds. Proc Natl AcadSci U S A 99:2782–2787
George Priya Doss C, Zayed H (2017) Comparative computational assessment of the pathogenicity of mutations in the Aspartoacylase enzyme. Metab Brain Dis 32:2105–2118
George Priya Doss C, Chakraborty C, Monford Paul Abishek N, et al (2014a) Application of evolutionary based in silico methods to predict the impact of single amino acid substitutions in Vitelliform macular dystrophy. In: Advances in protein chemistry and structural biology. pp 177–267
George Priya Doss C, Chakraborty C, Narayan V, Thirumal Kumar D (2014b) Computational approaches and resources in single amino acid substitutions analysis toward clinical research. Advances in protein chemistry and structural biology, In, pp 365–423
George Priya Doss C, Alasmar DR, Bux RI et al (2016) Genetic epidemiology of Glucose-6-dehydrogenase deficiency in the Arab world. Sci Rep 6:37284
Gibson KM, ten Brink HJ, Schor DSM et al (1993) Stable-isotope dilution analysis of D- and L-2-hydroxyglutaric acid: application to the detection and prenatal diagnosis of D- and L-2-hydroxyglutaric acidemias. Pediatr Res 34:277–280
Gregersen N, Kølvraa S, Rasmussen K et al (1980) Biochemical studies in a patient with defects in the metabolism of acyl-CoA and sarcosine: another possible case of glutaric aciduria type II. J Inherit Metab Dis 3:67–72
Guex N, Peitsch MC (1997) SWISS-MODEL and the Swiss-Pdb viewer: an environment for comparative protein modeling. Electrophoresis 18:2714–2723
Guruprasad K, Rajkumar S (2000) Beta-and gamma-turns in proteins revisited: a new set of amino acid turn-type dependent positional preferences and potentials. J Biosci 25:143–156
Hemachandran H, Jain F, Mohan S et al (2018) Glandular hair constituents of MallotusphilippinensisMuell. Fruit act as tyrosinase inhibitors: insights from enzyme kinetics and simulation study. Int J BiolMacromol 107:1675–1682
Hess B, Bekker H, Berendsen HJC, Fraaije JGEM (1997) LINCS: a linear constraint solver for molecular simulations. J Comput Chem 18:1463–1472
Honey EM, van Rensburg M, Knoll DP et al (2003) Spondyloenchondromatosis with D-2-hydroxyglutaric aciduria: a report of a second patient with this unusual combination. Clin Dysmorphol 12:95–99
Horbinski C (2013) What do we know about IDH1/2 mutations so far, and how do we use it? Acta Neuropathol 125:621–636
Johnson AD, Handsaker RE, Pulit SL et al (2008) SNAP: a web-based tool for identification and annotation of proxy SNPs using HapMap. Bioinformatics 24:2938–2939
Kaufman EE, Nelson T, Fales HM, Levin DM (1988) Isolation and characterization of a hydroxyacid-oxoacid transhydrogenase from rat kidney mitochondria. J Biol Chem 263:16872–16879
Kranendijk M, Struys EA, van Schaftingen E, et al (2010a) IDH2 mutations in patients with D-2-Hydroxyglutaric aciduria. Science (80-) 330:336–336
Kranendijk M, Struys EA, Van Schaftingen E, Gibson KM, Kanhai WA, Van der Knaap MS, Amiel J, Buist NR, Das AM, de Klerk JB, Feigenbaum AS (2010b) IDH2 mutations in patients with D-2-hydroxyglutaric aciduria. Science 330(6002):336–336
Kranendijk M, Salomons GS, Gibson KM et al (2011) A lymphoblast model for IDH2 gain-of-function activity in d-2-hydroxyglutaric aciduria type II: novel avenues for biochemical and therapeutic studies. Biochim Biophys Acta Mol basis Dis 1812:1380–1384
Lobanov MI, Bogatyreva NS, Galzitskaia O V (n.d.) [Radius of gyration is indicator of compactness of protein structure]. Mol Biol (Mosk) 42:701–6
Lovell SC, Davis IW, Arendall WB et al (2003) Structure validation by Calpha geometry: phi, psi and Cbeta deviation. Proteins 50:437–450
Mi H, Thomas P (2009) PANTHER pathway: an ontology-based pathway database coupled with data analysis tools. In: methods in molecular biology (Clifton, N.J.). Pp 123–140
Misra VK, Struys EA, O’Brien W et al (2005) Phenotypic heterogeneity in the presentation of d-2-hydroxyglutaric aciduria in monozygotic twins. Mol Genet Metab 86:200–205
Miyamoto S, Kollman PA (1992) Settle: an analytical version of the SHAKE and RATTLE algorithm for rigid water models. J Comput Chem 13:952–962
Mosaeilhy A, Mohamed MM, GPD C et al (2017) Genotype-phenotype correlation in 18 Egyptian patients with glutaricacidemia type I. Metab Brain Dis 32:1417–1426
Naine SJ, Devi CS, Mohanasrinivasan V et al (2015) Binding and molecular dynamic studies of sesquiterpenes (2R-acetoxymethyl-1,3,3-trimethyl-4t-(3-methyl-2-buten-1-yl)-1t-cyclohexanol) derived from marine Streptomyces sp. VITJS8 as potential anticancer agent. Appl Microbiol Biotechnol:1–14
Narang P, Bhushan K, Bose S, Jayaram B (2005) A computational pathway for bracketing native-like structures for small alpha helical globular proteins. Phys Chem Chem Phys 7:2364
Ng PC, Henikoff S (2003) SIFT: predicting amino acid changes that affect protein function. Nucleic Acids Res 31:3812–3814
Nota B, Hamilton EM, Sie D et al (2013) Novel cases of D-2-hydroxyglutaric aciduria with IDH1 or IDH2 mosaic mutations identified by amplicon deep sequencing. J Med Genet 50:754–759
Nyhan WL, Shelton GD, Jakobs C et al (1995) D-2-Hydroxyglutaric Aciduria. J Child Neurol 10:137–142
Oizel K, Gratas C, Nadaradjane A et al (2015) D-2-Hydroxyglutarate does not mimic all the IDH mutation effects, in particular the reduced etoposide-triggered apoptosis mediated by an alteration in mitochondrial NADH. Cell Death Dis 6:e1704
P S DKT, Tanwar H et al (2017) Structural analysis of G1691S variant in the human Filamin B gene responsible for Larsen syndrome: a comparative computational approach. J Cell Biochem 118:1900–1910
Pace CN, Fu H, Lee Fryar K et al (2014) Contribution of hydrogen bonds to protein stability. Protein Sci 23:652–661
Parrinello M, Rahman A (1981) Polymorphic transitions in single crystals: a new molecular dynamics method. J Appl Phys 52:7182–7190
Poux S, Arighi CN, Magrane M et al (2017) On expert curation and scalability: UniProtKB/Swiss-Prot as a case study. Bioinformatics 33:3454–3460
Pronk S, Páll S, Schulz R et al (2013) GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29:845–854
Rajasekaran R, Sudandiradoss C, George Priya Doss C, Sethumadhavan R (2007) Identification and in silico analysis of functional SNPs of the BRCA1 gene. Genomics 90(4):447–452
Rashed MS, AlAmoudi M, Aboul-Enein HY (2000) Chiral liquid chromatography tandem mass spectrometry in the determination of the configuration of 2-hydroxyglutaric acid in urine. Biomed Chromatogr 14:317–320
Rose GD, Gierasch LM, Smith JA (1985) Turns in peptides and proteins. Adv Protein Chem 37:1–109
Schmid N, Eichenberger AP, Choutko A et al (2011) Definition and testing of the GROMOS force-field versions 54A7 and 54B7. Eur Biophys J 40:843–856
Schneider JP, Kelly JW (1995) Templates that Induce. Alpha.-helical, .Beta.-sheet, and loop conformations. Chem Rev 95:2169–2187
Smeitink J (2010) Metabolism, gliomas, and IDH1. N Engl J Med 362:1144–1145
Sneha P, George Priya Doss C (2016) Chapter seven– molecular dynamics: new frontier in personalized medicine. In: Advances in Protein Chemistry and Structural Biology. pp 181–224
Sneha P, Thirumal Kumar D, Siva R et al (2017a) Determining the role of missense mutations in the POU domain of HNF1A that reduce the DNA-binding affinity: a computational approach. PLoS One 12(4):e0174953
Sneha P, Thirumal Kumar D, Tanwar H et al (2017b) Structural analysis of G1691S variant in the human Filamin B gene responsible for Larsen syndrome: a comparative computational approach. J Cell Biochem 118:1900–1910
Sneha P, Ebrahimi EA, Ghazala SA, Thirumal Kumar D, Siva R, George Priya Doss C, Zayed H (2018) Structural analysis of missense mutations in Galactokinase (GALK1) leading to Galactosemia type-2. J Cell Biochem. https://doi.org/10.1002/jcb.27097
Struys EA (2006) D-2-Hydroxyglutaric aciduria: unravelling the biochemical pathway and the genetic defect. J Inherit Metab Dis 2006 Feb 1;29(1):21–9
Struys EA, Korman SH, Salomons GS et al (2005a) D-2-Hydroxyglutarate dehydrogenase mutations in phenotypically mild D-2-hydroxyglutaric aciduria. Ann Neurol 58:626–630
Struys EA, Salomons GS, Achouri Y et al (2005b) Mutations in the d-2-Hydroxyglutarate dehydrogenase gene cause d-2-Hydroxyglutaric aciduria. Am J Hum Genet 76:358–360
Sudhakar N, George Priya Doss C, Thirumal Kumar D et al (2015) Deciphering the impact of somatic mutations in exon 20 and exon 9 of PIK3CA gene in breast tumors among Indian women through molecular dynamics approach. J BiomolStructDyn:1–13
Sugita K, Kakinuma H, Okajima Y et al (1995) Clinical and MRI findings in a case ofd-2-hydroxyglutaric aciduria. Brain Dev 17:139–141
Sujitha SP, Kumar DT, Doss CGP et al (2016) DNA repair gene (XRCC1) polymorphism (Arg399Gln) associated with schizophrenia in south Indian population: a genotypic and molecular dynamics study. PLoS One 11:e0147348
Talkhani IS, Saklatvala J, Dwyer J (2000) D-2-hydroxyglutaric aciduria in association with spondyloenchondromatosis. Skelet Radiol 29(5):289–292
Thirumal Kumar D, George Priya Doss C (2016a) Investigating the inhibitory effect of Wortmannin in the hotspot mutation at codon 1047 of PIK3CA kinase domain: a molecular docking and molecular dynamics approach. Adv Protein ChemStructBiol 102:267–297
Thirumal Kumar D, George Priya Doss C (2016b) Role of E542 and E545 missense mutations of PIK3CA in breast cancer: a comparative computational approach. J Biomol Struct Dyn:1–13. https://doi.org/10.1080/07391102.2016.1231082
Thirumal Kumar D, George Priya Doss C, Sneha P et al (2016) Influence of V54M mutation in giant muscle protein titin: a computational screening and molecular dynamics approach. J BiomolStructDyn:1–12
Thirumal Kumar D, Lavanya P, George Priya Doss C et al (2017) A molecular docking and dynamics approach to screen potent inhibitors against Fosfomycin resistant enzyme in clinical Klebsiella pneumoniae. J Cell Biochem. https://doi.org/10.1002/jcb.26064
Unissa AN, Doss CGP, Kumar T et al (2017) Analysis of interactions of clinical mutants of catalase-peroxidase (KatG) responsible for isoniazid resistance in Mycobacterium tuberculosis with derivatives of isoniazid. J Glob Antimicrob Resist 11:57–67. https://doi.org/10.1016/j.jgar.2017.06.014
Van der Knaap MS, Jakobs C, Hoffmann GF, Duran M, Muntau AC, Schweitzer S, Kelley RI, Parrot-Roulaud F, Amiel J, De Lonlay P, Rabier D (1999) D-2-hydroxyglutaric aciduria: further clinical delineation. J Inherit Metab Dis 22(4):404–413
Wade RC, Goodford PJ (1989) The role of hydrogen-bonds in drug binding. ProgClinBiol Res 289:433–444
Wajne M, Vargas CR, Funayama C et al (2002) D-2-Hydroxyglutaric aciduria in a patient with a severe clinical phenotype and unusual MRI findings. J Inherit Metab Dis 25:28–34
Wang Z, Moult J (2001) SNPs, protein structure, and disease. Hum Mutat 17:263–270
Wang Z, Moult J (2003) Three-dimensional structural location and molecular functional effects of missense SNPs in the T cell receptor V?? Domain. Proteins StructFunct Genet 53:748–757
Ward PS, Patel J, Wise DR et al (2010) The common feature of leukemia-associated IDH1 and IDH2 mutations is a Neomorphic enzyme activity converting α-ketoglutarate to 2-Hydroxyglutarate. Cancer Cell 17:225–234. https://doi.org/10.1016/j.ccr.2010.01.020
Xie X, Baird D, Bowen K et al (2017) Allosteric mutant IDH1 inhibitors reveal mechanisms for IDH1 mutant and isoform selectivity. Structure 25:506–513
Yagawa K, Yamano K, Oguro T et al (2010) Structural basis for unfolding pathway-dependent stability of proteins: Vectorial unfolding versus global unfolding. Protein Sci 19:693–702
Yun S, Guy HR (2011) Stability tests on known and misfolded structures with discrete and all atom molecular dynamics simulations. J Mol Graph Model 29:663–675
Zaki OK, Krishnamoorthy N, El Abd HS et al (2017a) Two patients with Canavan disease and structural modeling of a novel mutation. Metab Brain Dis 32:171–177
Zaki OK, Priya Doss CG, Ali SA et al (2017b) Genotype–phenotype correlation in patients with isovaleric acidaemia: comparative structural modelling and computational analysis of novel variants. Hum Mol Genet 12:e0174953
Zayed H (2015a) Propionic acidemia in the Arab world. Gene 564(2):119–124
Zayed H (2015b) Canavan disease: an Arab scenario. Gene 560(1):9–14
Zhang Z, Norris J, Schwartz C, Alexov E (2011) In silico and in vitro investigations of the mutability of disease-causing missense mutation sites in Spermine synthase. PLoS One 6:e20373
Zhao G, Winkler ME (1996) A novel alpha-ketoglutarate reductase activity of the serA-encoded 3-phosphoglycerate dehydrogenase of Escherichia coli K-12 and its possible implications for human 2-hydroxyglutaric aciduria. J Bacteriol 178:232–239
Acknowledgements
The authors acknowledge the management of Vellore Institute of Technology, Vellore, India and (BRAF) @ CDAC for providing the facilities required to perform this work.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Electronic supplementary material
ESM 1
(DOCX 203 kb)
Rights and permissions
About this article
Cite this article
Thirumal Kumar, D., Jerushah Emerald, L., George Priya Doss, C. et al. Computational approach to unravel the impact of missense mutations of proteins (D2HGDH and IDH2) causing D-2-hydroxyglutaric aciduria 2. Metab Brain Dis 33, 1699–1710 (2018). https://doi.org/10.1007/s11011-018-0278-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11011-018-0278-3