Abstract
The detailed understanding of the molecular features of a ligand binding to a target protein, facilitates the successful design of potent and selective inhibitors. We present a case study of ATP-competitive kinase inhibitors that include a pyradine moiety. These compounds have similar chemical structure, except for distinct terminal hydrophobic cyclopentyl or isopropyl groups, and block kinase activity of casein kinase 2 subunit α (CK2α), which is a target for several diseases, such as cancer and glomerulonephritis. Although these compounds display similar inhibitory potency against CK2α, the crystal structures reveal that the cyclopentyl derivative gains more favorable interactions compared with the isopropyl derivative, because of the additional ethylene moiety. The structural observations and biological data are consistent with the thermodynamic profiles of these inhibitors in binding to CK2α, revealing that the enthalpic advantage of the cyclopentyl derivative is accompanied with a lower entropic loss. Computational analyses indicated that the relative enthalpic gain of the cyclopentyl derivative arises from an enhancement of a wide range of van der Waals interactions from the whole complex. Conversely, the relative entropy loss of the cyclopentyl derivative arises from a decrease in the molecular fluctuation and higher conformational restriction in the active site of CK2α. These structural insights, in combination with thermodynamic and computational observations, should be helpful in developing potent and selective CK2α inhibitors.
Similar content being viewed by others
Introduction
In many cases of structure-based drug design (SBDD), insufficient understanding of the atomic resolution structure of the ligand–target protein complex restricts rational drug design, modification, and optimization. For e.g., the introduction of a methyl group to an adenosine deaminase inhibitor as a lead compound yielded a 70-fold gain in activity, which corresponds to a large binding free energy change of 11.6 kJ/mol; however, it was reported that the group contribution assigned to a methylene group was statistically estimated to be only 3–4 kJ/mol [1]. In contrast, the introduction of a large substituent composed of 13 atoms to the same lead compound yielded only a 80-fold increase in the inhibitory activity [2, 3]. Crystal structures of both complexes showed that the introduction of both groups in these derivatives were properly accommodated at the active site of the target protein, and computational analysis indicated that both modifications increased the interactions with the protein compared with the original lead compound [2, 3]. Therefore, successful design of potent and selective protein inhibitors, in terms of structure-based drug design, strongly depends on an appropriate understanding of the physicochemical features determining the ligand binding to the target protein. In the adenosine deaminase study, computational analyses of the binding free energy supported the design of the ligand [4]. In addition to computational evaluation, thermodynamic profiling of the protein–ligand complex plays a vital role in an accurate structural analysis. Thermodynamic profiling of a homologous pair of thrombin inhibitors, differing only by one methylene unit, showed that both inhibitors have the same Gibbs free energy of binding, yet showed a dramatic difference in the enthalpic and entropic contributions to the Gibbs free energy value [5]. The results provided valuable insights for correctly understanding the crystal structures of the ligand–target protein complexes for both inhibitors. Freire recently stated that structure–activity relationships (SARs) that explicitly incorporate the interplay between enthalpy and entropy accelerate the optimization process [6], and the experience of many pharmaceutical laboratories has shown that the simultaneous optimization of enthalpy and entropy without thermodynamic data is difficult to achieve.
Protein kinase CK2 is a highly conserved serine/threonine kinase, and more than 300 protein substrates of CK2 have been identified [7]. The CK2 holoenzyme consists of two catalytic subunits and two regulatory subunits [8]. CK2α is the catalytic subunit and has no phosphorylation site required for activation. Consequently, CK2 is constitutively active with or without the regulatory subunit, CK2β [9]. CK2 has important roles in growth, proliferation, and survival of cells via phosphorylation of substrates [7]. Since CK2 is expressed in a wide variety of tumors [10], it represents an important target protein for tumor therapy. Down-regulation of CK2 in tumor cell lines leads to cell death, because CK2 suppresses apoptosis in tumor cells [11]. We have recently reported that CK2 is also a target protein for glomerulonephritis (GN) therapy, supported by experiments showing that administration of an antisense oligodeoxynucleotide against CK2 or low-molecular weight CK2-specific inhibitors effectively prevented progression of the renal disease in a rat model of GN [12].
Several planar chemical or natural compounds have been identified as ATP-competitive, selective CK2 inhibitors, such as ellagic acid [13], emodin [14], apigenin [15], 4,5,6,7-tetrabromo-1H-benztriazole [16], 5-oxo-5,6-dihydroindolo[1,2-a]quinazolin-7-yl [17], and 5,6-dichloro-1-beta-D-ribofuranosylbenzimidazol (DRB) [18]. Other trials to produce potent and selective CK2 inhibitors are currently in progress [19, 20]. In the process of a lead optimization stage, expectant lead structures are modified by systemically changing functional groups or side-chains on a given core scaffold. The crystal structures of human CK2α complexed with ATP-competitive inhibitors, including emodin [14], DRB [21], AMPPNP as an ATP analog [22, 23], and ellagic acid [24] provide valuable starting points that guide the selection of an appropriate functional group to be attached or modified. There is a pair of homologous CK2 inhibitors with very high inhibitory activities (compounds 1 and 2 shown in Fig. 1), which differ only by an ethylene unit in the hydrophobic substituent for the 6-amino-indazole moiety. These compounds display almost the same potency to CK2 activity [19], although the additional ethylene moiety of compound-2 has more favorable interactions with CK2 than compound-1.
In order to establish the correct rationale for SBDD based on these compounds, we have evaluated the binding features of these two homologous compounds by calorimetric, structural, and computational analyses.
Methods
Preparation of human CK2α
Recombinant human CK2α was prepared according to the reported protocol [18]. The C-terminal truncated form of CK2α was cloned into the pGEX6P-1 expression vector (GE Healthcare) and expressed in E. coli strain HMS174 (DE3) as a GST-fused protein at the N-terminus. Cells were cultured in LB medium supplemented with ampicillin at 310 K. Expression was induced by the addition of 0.5 mM isopropyl-1-thio-beta-d-galactopyranoside when cells had reached an optical density of 600 nm reading of 0.5 at 298 K for 20 h. The cells were harvested, resuspended in a buffer consisting of 150 mM NaCl and 25 mM Tris–HCl, pH 7.4, and sonicated. After removing the cellular debris by centrifugation, the supernatant was loaded onto a GSTrap HP column (GE Healthcare, England) and incubated at 277 K for 1 h. The column was washed with cleavage buffer containing 50 mM Tris–HCl, pH 8.0, 150 mM NaCl, 1 mM EDTA, 5 mM dithiothreitol, and 0.1% Tween 20. The GST-fused CK2α protein was digested with 80 U/ml PreScission protease (GE Healthcare) for 4 h at 277 K. The GST-free CK2α protein was eluted using the MonoQ buffer consisting of 25 mM Tris–HCl, pH 8.0, and 5 mM dithiothreitol. CK2α was further purified by anion-exchange chromatography with a MonoQ column (GE Healthcare) using a linear NaCl gradient of 0–0.5 M, 50 CV of the MonoQ buffer at 277 K, and an AKTA explorer system (GE Healthcare).
Structure determination
The purified protein was concentrated to 5 mg/ml without buffer exchange. For the complex study, an excess amount of inhibitor was added to the protein solution and the centrifuged supernatant was used for crystallization. The prism-shaped crystals of the complexes and the apo-form were obtained by the sitting-drop vapor diffusion method using ethylene glycol as the precipitant. This condition was the same as that used for solving the ellagic acid-CK2α complex [18]. After dipping into Paratone-N oil (Hampton Research, Aliso Viejo, CA), the crystals were frozen using a nitrogen gas stream at 100 K. Diffraction data were collected at the BL6A and BL17A stations of the Photon Factory, or at the BL44XU station of the SPring-8. The data set was processed with the program HKL2000 [25]. The structures of the complex or the apo-form were solved by the molecular replacement method, carried out with the program MolRep [26] in the CCP4 suite [27]. All refinements and model modifications were performed using the program's DS Modeling and CNX (Accelrys, San Diego, CA). Data collection and refinement statistics are shown in Table 1. The final coordinates of the cyclopentyl-, isopropyl-derivative complexes, and the apo-form were deposited into the Protein Data Bank with the accession codes of 3at3, 3at4, and 3at2, respectively.
Calorimetric analyses with an isothermal titration calorimeter
The 28 μM inhibitor solution was prepared by dissolving the inhibitor in the experimental buffer containing 25 mM Tris–HCl, pH 8.5, and 250 mM NaCl. The buffer was used to measure the heats of dilution of the inhibitor solutions. The purified CK2α protein was dialyzed against the experimental buffer and concentrated to 4 μM. The concentrations of CK2α were determined by absorbance at 280 nm. The isothermal titration calorimeter (ITC) experiments were performed at 293 K on a VP-ITC (Microcal Inc., Northampton, MA). Inhibitor solutions were degassed before use and titrated into the stirred cell containing the prepared CK2α solution of 1.4 ml after a stable baseline had been achieved. The injection sequence consisted of 10 μl injections at 5 min intervals until complete saturation of the enzyme binding sites was achieved. The experimental data were analyzed using the Origin-ITC software package (Microcal Inc.). Heats of dilution were subtracted from the raw data before analysis.
Analyses of interaction energy and conformational space
Each complex structure was modeled on the basis of the crystal structures using the program MOE (Chemical Computing Group Inc, Montreal, Canada). The protonation states of the residues and the direction of the hydrogen atoms involved in hydrogen bonds were assigned by the Protonate3D algorithm [28]. The molecular dynamics (MD) simulations in a water environment for two ligands with and without protein were performed (four MD simulations). A water molecule sphere with a radius of 25 Å from the ligand was generated on both complex systems to consider the dynamics in an aqueous environment. After structure optimization, the MD simulations were performed by MOE. During the MD simulations, protein atoms far from the ligands were fixed to their original coordinates to focus on the ligand dynamics. In this case, atoms belonging to residues or water molecules with shortest distances of ≤20 Å from the ligand were allowed to move freely. The conditions and protocol for MD are as follows. After heating, the temperature was held at 300 K by the Nosé-Poincaré-Anderson method [29]. The cut-off distance for non-bonding interactions was 10 Å, the MMFF94x force field parameter [30] was assigned for every atom, all bond lengths including hydrogen atoms were fixed by the SHAKE procedure [31], and the time step for the geometry update was 1 fs. A total of 1,500 snapshots of the coordinates were taken from 750 ps production time after the 50 ps heating, and the 250 ps equilibration stages were analyzed. The residue-based interaction energy with the ligand was determined using the generalized Born/volume integral (GB/VI) solvation model [32] and was calculated for every snapshot.
To consider the dynamic structural effect, the RMSD values excluding hydrogen atoms were calculated for the residues (Arg47 and Asn161) near the substitution group during the MD simulations. Using the same MD scheme, the compounds in the solvated state without protein were also calculated. A 30 Å solvation shell around the compounds was used. The conformational flexibility of the compounds in the complex and that in the isolated state were then compared.
Furthermore, to explore the conformational space for the terminal hydrophobic group of the inhibitors, a rotational barrier between the hydrophobic group and the 6-amino-indazole moiety was calculated.
Results and discussion
Overall structures of CK2α
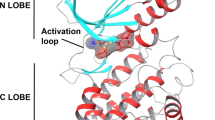
Based on the electron density maps, all amino acids of CK2α were well-ordered in the complexes and apo-form, including the entrance region of the ATP-binding site. The high resolution confers the rational assignment of the 11 ethylene glycol molecules as crystallization precipitants and 412 water molecules in the apo-form structure. On the other hand, the lower resolution of the complexes compared with the apo-form allows to assign no ethylene glycol molecule, and there are only 140 and 138 water molecules in the compound-1/and compound-2/CK2α complexes, respectively. Although the fewer atom-assignments of the complexes may yield the modestly high R-values compared to the apo-form (Table 1), the well-defined electron densities corresponding to the compounds as well as protein in the respective complex are observed sufficiently for the following discussion (Fig. 2b). The conformations of CK2α complexed with compound-1 and compound-2, with extension of the N-terminal segment (Met1-Asn35), were very similar to those previously reported [8, 15–18]. Both compounds did not induce a significant structural change to CK2α when compared with the apo-form. The structure of CK2α has a typical kinase fold consisting of N-terminal and C-terminal lobes, which are connected by a hinge region (Glu114-Asn118) (Fig. 2a). The N-terminal extension of CK2α was found to be tightly bound to the two lobes through an aromatic cluster, and a number of hydrogen bonds. At the interface of the N-terminal extension to the activation loop (Asp175-Ser194) in the C-lobe, an aromatic cluster was formed between Tyr23, Trp24, and Tyr26 on one side, and Phe181 and His183 on the other. The backbone nitrogen of Ala10 formed a hydrogen bond with the hydroxyl group of Tyr182 and the amino group of the Asn16 side-chain formed hydrogen bonds with the backbone oxygen atoms of Gly151 and Tyr182. In addition, the activation segment interacted with the αC-helix via three hydrogen bonds. The amino group of Lys77 formed a hydrogen bond with the backbone oxygen of Gly177, and the Nη1 atom of Arg80 made hydrogen bonds with the backbone oxygen of Ala179, and the carboxyl group of Asp180, which is also interacting with the N-terminal extension residue Tyr26. Thus, the activation segment and the αC-helix were in active conformation because of close contacts to the N-terminal extension. These overall structural features are conserved in the apo-form and the complexed forms.
Inhibitor binding to the ATP-binding region of CK2α
Two homologous inhibitors (Fig. 1) bound to the ATP-binding region in a similar manner, except for the terminal hydrophobic groups, is shown by the clear density map (Fig. 2b). Both inhibitors are bound to the CK2α with a horseshoe-shape conformation. The N2 atom of the indazole moiety, and the N1 atom of the pyrazine ring probably allow the inhibitors to adopt a compact and planar conformation, because the lone-pair electrons of these N atoms accept the C2 and C3 hydrogen atoms of the pyrrole ring, and of the pyrazine ring, to make a weak but favorable CH–N type intramolecular hydrogen bonds. Similar to the planar inhibitors previously reported [15], the planar conformation in these inhibitors gain attractive interactions from the van der Waals contacts with many hydrophobic residues, including Leu45, Val53, Val66, Ile95, Phe113, His160, Met163, and Ile174. The pyrazine ring interacts with Val66 in the β3 strand. The pyrrole ring was held in place by Ile174 in the activation segment. The indazole moiety was sandwiched between Leu45 in the glycine-rich loop located between the β1 and β2 strands, and Met163 in the extra β-strand that follows the activation segment. Furthermore, several hydrogen bonds were found between the CK2α and the inhibitor. The carboxyl group of the inhibitor formed a salt bridge, and hydrogen bonds with the Nζ atom of Lys68 and the backbone nitrogen of Asp175 in the activation loop, respectively. The inhibitors also interact with a highly conserved water molecule [8, 18] in the active site, which is ligated by Glu81. Lys68, Glu81, and Asp175 are essential residues for enzyme activity. One of the nitrogen atoms of the pyrazine ring forms a hydrogen bond with the backbone oxygen of Val116 in the hinge region connecting the N- and C-lobes. Most of the interactions were conserved in the two complexes, except for the remarkable local structural differences around the terminal hydrophobic groups of the inhibitors. The cyclopentyl groups of compound-2 had additional van der Waals interactions with Arg47 and Asn161, compared with the isopropyl group of compound-1, and released a water molecule between Asn161 and Asp175 observed in the compound-1/CK2α complex and the apo-form (Fig. 3).
Close-up views around the ATP-binding site of the three crystal structures. Residues surrounding the compounds are shown. Hydrogen bonds are displayed as dotted lines. a The compound-1 complex. b The compound-2 complex. The additional van der Waals interactions are shown as dashed lines. c The apo-form
Calorimetric data analysis
To clarify the differences between the binding properties of compound-1 and compound-2, we performed thermodynamic analyses using isothermal titration calorimetry (ITC). The standard binding enthalpy ΔH and the dissociation constant K D were determined by ITC experiments for both compounds. The entropic contribution to binding, −TΔS, has been calculated as the difference between the Gibbs free energy change, ΔG, derived from K D , and ΔH according to the Gibbs–Helmholtz equation. The ITC experiments yielded similar Gibbs energies of binding for compounds 1 and 2 (Table 2), reflecting the same inhibitory activities of the compounds. However, a dramatic difference was observed for the enthalpic and entropic contributions. For compound-2 binding to CK2α, the additional ethylene moiety at the terminal hydrophobic group resulted in an enthalpy gain of 11.1 kJ/mol (Table 2), which is in agreement with the statistical estimation derived by Kunz et al. [1]. However, the ethylene moiety makes only weak interactions through van der Waals contacts, giving rise to an enthalpy gain of no more than 1 kJ/mol. Therefore, other interactions contribute to the large enthalpy gain of compound-2.
On the other hand, the negative entropic term, −TΔS, of compound-1 turns positive for compound-2 and the observed entropy decrease of compound-2 was 9.2 kJ/mol (Table 2). The entropy loss of compound-2 is consistent with the enthalpy–entropy compensation rule. The restriction of compound-2 and/or protein motions in the complexed state prevails over the entropy gain, due to the release of the water molecule ligated to Asn161 and Asp175 upon compound-2 binding. The detailed analyses of the enthalpic and entropic contribution are discussed in the following sections.
Decomposition of the binding enthalpy to CK2α
The enthalpy gain of 11.1 kJ/mol for compound-2 compared with compound-1 was dissected by computational calculations of the average interaction energies of the inhibitor molecules during molecular dynamics (MD) simulations. The crystal structures intuitively suggested that the enthalpy gain of compound-2 was achieved by the additional ethylene moiety near Arg47 and Asn161. To evaluate the effect of the ethylene moiety in compound-2, the total interaction energy was decomposed into the individual interactions between residues of CK2α and the inhibitors. The difference of the calculated binding energies for the two compounds was similar to the measured enthalpy differences using ITC (Tables 2, 3), and thus the computational analysis provided a suitable approach to discuss the structural differences.
From the MD simulations, Arg47 and Asn161, interacting with the additional ethylene moiety of compound-2 (Fig. 3b), do not provide adequate energetic gain to account for the enthalpy gain observed by the ITC experiment (Table 3). Consequently, the computational analysis revealed that the favorable enthalpy change of compound-2 is mainly due to van der Waals interactions (Table 3), and widely distributed to the distal side of Arg47 and Asn161 in the ATP-binding site. Such interactions involve residues Leu45, Val53, Ile95, Phe113, Val116, Met163, Ile174, and Asp175, which are surrounding both compounds in the complex states (Fig. 3).
In addition to van der Waals interactions, the salt bridge of the inhibitor with Lys68, and hydrogen bonds formed between the inhibitor and His115, Val116, and Asp175 are well conserved in the crystal structures of the both protein-inhibitor complexes (Fig. 3a, b). The computational energy decomposition indicates that there was no enthalpy difference due to the salt bridge formed, and the hydrogen bonds between both compounds (data not shown). The strengths of the hydrogen bonds involving residues His115 and Val116 in the hinge region were equivalent for both compounds; however, the MD simulations suggest that compound-1 could move closer to the hinge region than compound-2. The enthalpy gain of compound-1 by the electrostatic term just counteracts the enthalpy loss due to the van der Waals term. The MD simulations also suggest that the carboxylic group of compound-1 moves away from Lys68, but closer to Asp175 compared with compound-2. The enthalpy gain of compound-1 by the interaction with Asp175 is offset by the loss of enthalpy due to the weaker interaction with Lys68. Therefore, the differences in the enthalpic contributions due to the interactions involving Lys68, His115, Val116, and Asp175 and compound-1 and compound-2 were computed to be negligible.
Compound-2 binding also gave rise to an additional enthalpic loss, because of the release of the structured-water molecule ligated to Asn161 and Asp175 observed in the crystal structure of the compound-1/CK2α complex and the apo-form. The energy loss due to the rupturing of the two hydrogen bonds with these residues was estimated to be 20–60 kJ/mol. This enthalpic loss would be partially canceled by the formation of new hydrogen bonds between the released water molecule and the bulk solvent. In total, the enthalpic loss by the structured-water release and the reduced electrostatic interactions is perhaps indicated in the calculated energy loss of 4.47 kJ/mol as the effective electrostatic term (Table 3).
This enthalpic augmentation of compound-2 binding probably reflects a transition from a state of loose interaction to that of tight binding, potentiated by the individual van der Waals interaction surpassing the enthalpic loss due to the release of water.
The relative rigidity of compound-2 in the complexed state results in an entropy loss
The decreased flexibility of compound-2 predicted by computational calculations was also observed in the crystal structure of the complex. The B-factor ratio of one atom against another atom defines the relative thermal mobility [33]. For compound-2, the B-factor ratio of the ligand atoms against the binding pocket residues averaged for Leu45, Val53, Val66, Lys68, Ile95, Phe113, His115, His160, Met163, Ile174, and Asp175 was calculated as 0.84, indicating that the ligand has lower thermal motion compared with the binding pocket atoms. On the other hand, this ratio increases significantly to 1.09 for compound-1. These results indicate that compound-1 confers higher mobility than compound-2 due to relatively loose binding at the ATP-binding site. Furthermore, the difference in the B-factor ratio between the cyclopentyl group of compound-2 and the isopropyl group of compound-1 increases with values of 0.68 and 1.52, respectively. This suggests that the thermal motion of the terminal isopropyl group of compound-1 is larger than the cyclopentyl group of compound-2. In other words, the flexibility/mobility of compound-2 is diminished when bound to the CK2α, whereas for compound-1 this is not as apparent.
Presumably, the decrease in the flexibility of compound-2 in the complexed state is derived from the potentiated van der Waals interactions, which gave rise to the enthalpy gain for the compound-2–kinase complex. However, simultaneously, this change contributes to the entropy loss of the compound-2 complex. A similar observation for the ligands that bind thrombin has been reported [33]. The tighter binding shown by the B-factor ratio represents the larger entropic loss.
An additional factor that leads to the entropy loss of the cyclopentyl derivative
The entropic loss of 9 kJ/mol of compound-2 observed by the ITC experiments compared with compound-1 is the summation of the gain by the water release, the losses due to the protein- and ligand conformational restrictions, and the losses due to the decrease in the vibrational freedom of the ligand and interacting residues.
As a contributor to the entropic gain, considerable attention should be focused on the release of the strongly-bound water molecule ligated to Asn161 and Asp175 in both the compound-1 complex and the apo-form (Fig. 3a, c). Dunitz postulated an upper limit of about 40 J/K as an entropic gain for the release of a tightly bound water molecule [34]. Conversely, a range of between 0 and 30 J/K for the entropy gain of transferring a water molecule from the protein to the bulk solvent has been suggested, and this corresponds to a free energy gain of between 0 and 9 kJ/mol at 300 K. The water ligated with these residues is primarily exposed to the solvent region, and thus a smaller entropic gain is postulated.
The conformational entropic loss of compound-2 upon binding to CK2α was evaluated by computational analysis. Cheng showed that most of the entropic loss arises from conformational losses, such as angular contributions from bonds between the sp3 atoms, rather than a drop in the number of accessible conformations [35]. Thus, the rotational barriers of the terminal hydrophobic groups, and the characteristic moieties between both compounds, were compared. In the isolated state, both inhibitors were equally stable in the range of 130–220° (compound-1) or 100–230° (compound-2) about the dihedral angle between the amino N atom and the terminal hydrophobic group (Fig. 4). In the complex state, the conformational window for compound-2 was narrowed to 190–215° (Fig. 4). The window range for compound-1 was slightly narrowed to 150–215° in the complexed state. Therefore, compound-2 is less favorable than compound-1 with respect to the conformational entropic term of binding to CK2α. On the other hand, the protein conformational entropy loss is identical based on the crystal structures, because the apo conformation, and the most free-state, remains in both complexes.
The perturbation loss of the interacting residues, as well as the ligand mentioned in the previous section, were observed based on the crystal structure data. The B-factor ratio of the residues interacting with the compound against the average of all residues in the compound-2 complex was further reduced compared to that observed for the compound-1 complex: in the apo-form: 0.88, in the compound-1 complex: 0.75, and in the compound-2 complex: 0.55.
Ultimately, the ITC-observed entropy loss of compound-2 compared with compound-1 primarily arises from the rotational restriction of the terminal hydrophobic group, and the fluctuation decrease of the ligand and protein in the complexed state. This surpasses the entropic gain due to the release of the bound water ligated at Asn161 and Asp175. Regrettably, the entropy loss by the conformational restriction is currently impossible to illustrate quantitatively. We must address this issue using more rigorous computational methods, such as the fragment molecular orbital method [36].
Overcoming the enthalpy–entropy compensation in CK2 drug discovery
The enthalpy–entropy compensation, composed of the enthalpic gain and entropic loss, was observed in the chemical modification of compound-1 to compound-2. Can we overcome this phenomenon by simultaneously gaining favorable enthalpy and entropy? Theoretical and experimental approaches suggest that the enthalpy–entropy compensation effect is not a general feature of weak associations [37, 38]. Reports revealed that successful chemical modification would be accomplished if the binding enthalpy and entropy were considered simultaneously, that is, maximizing the enthalpy gain and minimizing the entropy loss [23, 39–43]. Furthermore, keeping conformational rigidity allows us to realize enthalpic and entropic enhancement as recently shown by Freire [42]. Our results indicate that the entropy loss was mainly due to ligand perturbation fixation and ligand conformational restriction. Therefore, minor modification of compound-2, once in the rigid state, would likely confer lower entropy loss when compared with compound-1. The entropy gain involving desolvation would probably arise because of this minor modification, as shown by Kawasaki et al. [39]. The minor modification of the cyclopentyl moiety of compound-2, of course, is expected to produce further van der Waals interactions with Arg47, Asn161 and other residues, and thus it yields further enthalpy gains when compared with compound-2. The modification also reduces the enthalpy loss due to the release of bound water, which was a significant factor in our computational analyses. The crystal structure (Fig. 3b) suggests that the introduction of a small substituent such as halogen atoms in the cyclopentyl 3-position of compound-2 probably results in no bound water release. We therefore, believe that this minor modification would likely lead to further enthalpy energy gains and minimize the entropy loss (or obtain the entropy gain if fortunately). Such a modification may finally overcome the enthalpy–entropy compensation required for the inhibitor of CK2α.
Conclusions
X-ray crystal structures showed that two closely related inhibitors, with similar inhibitory activity for CK2α, formed similar complexes with the protein; however, the ITC data showed that the free energy decomposition of the enthalpy and entropy contributions differed for the two complexes. The energy calculations revealed that the enthalpic gain of compound-2 is largely because of van der Waals interactions, and is distributed not only in the proximal residues Arg47 and Asn161, but also widely in the distal residues that include Leu45, Val53, Ile95, Phe113, Val116, Met163, Ile174, and Asp175. Hydrogen bonds to His115, Val116, and Asp175 and the salt bridge with Lys68 were unaltered; although they were expected to be stronger. The enthalpy gain due to van der Waals interactions greatly exceeded the enthalpy loss due to the release of water. The total enthalpic gain, however, forfeits the entropic loss due to a decrease in fluctuation of the ligand and the ligand conformational restrictions of compound-2.
This study emphasizes the benefits of combining structural information about binding modes of protein-inhibitor complexes with thermodynamic data to characterize binding events. This combination offered an approach to rationally explain the observed magnitude of the binding affinities. Our study contributes to a better understanding of the details for inhibitor binding. The results presented should be valuable in structure-based drug design of CK2-derived disease therapy.
References
Kuntz ID, Chen K, Sharp KA, Kollman PA (1999) The maximal affinity of ligands. Proc Natl Acad Sci USA 96:9997–10002
Terasaka T, Kinoshita T, Kuno M, Nakanishi I (2004) A highly potent non-nucleotide adenosine deaminase inhibitor: efficient drug discovery by intentional lead hybridization. J Am Chem Soc 126:34–35
Terasaka T, Ohkumura H, Tsuji K, Kato T, Nakanishi I, Kinoshita T, Kato Y, Kuno M, Seki N, Naoe Y, Inoue T, Tanaka K, Nakamura K (2004) Structure-based design and synthesis of non-nucleoside, potent, and orally bioavailable adenosine deaminase inhibitors. J Med Chem 47:2728–2731
Kosugi T, Nakanishi I, Kitaura K (2009) Binding free energy calculation of adenosine deaminase inhibitor and the effect of methyl substitution in inhibitors. J Chem Inf Model 49:615–622
Gerlach C, Smolinski M, Steuber H, Sotriffer CA, Heine A, Hangauer DG, Klebe G (2007) Thermodynamic inhibition profile of a cyclopentyl and a cyclohexyl derivatives towards thrombin: the same but for different reasons. Angew Chem Int Ed 46:8511–8514
Freire E (2008) Do enthalpy and entropy distinguish first in class from best in class. Drug Discov Today 13:869–874
Meggio F, Pinna LA (2003) One-thousand-and-one substrates of protein kinase CK2alfa. FASEB J 17:349–368
Niefind K, Guerra B, Ermakowa I, Issinger OG (2001) Crystal structure of human protein kinase CK2: insights into basic properties of the CK2 holoenzyme. EMBO J 20:5320–5331
Mazzorana M, Pinna LA, Battistutta R (2008) A structural insight into CK2 inhibition. Mol Cell Biochem 316:57–62
Duncan JS, Litchfield DW (2008) Too much of a good thing: the role of protein kinase CK2 in tumorigenesis and prospects for therapeutic inhibition of CK2. Biochim Biophys Acta 1784:33–44
Ahmad KA, Wang G, Unger G, Slaton J, Ahmed K (2008) Protein kinase CK2—a key suppressor of apoptosis. Adv Enzyme Regul 48:179–187
Yamada M, Katsuma S, Adachi T, Hirasawa A, Shiojima S, Kadowaki T, Okuno Y, Koshimizu TA, Fujii S, Sekiya Y, Miyamoto Y, Tamura M, Yumura W, Nihei H, Kobayashi M, Tsujimoto G (2005) Inhibition of protein kinase CK2 prevents the progression of Glomerulonephritis. Proc Natl Acad Sci USA 102:7736–7741
Cozza G, Bonvini P, Zorzi E, Poletto G, Pagano MA, Sarno S, Sonellaa-Deana A, Zagotto G, Rosolen A, Pinna LA, Meggio F, Moro S (2006) Identification of ellagic acid as potent inhibitor of protein kinase CK2: a successful example of a virtual screening application. J Med Chem 49:2363–2366
Raaf J, Klopffleisch K, Issinger OG, Niefind K (2008) The catalytic subunit of human protein kinase CK2 structurally deviates from its maize homologue in complexed with the nucleotide competitive inhibitor emodin. J Mol Biol 377:1–8
Critchfield JW, Coligan JE, Folks TM, Butera ST (1997) Casein kinase II is a selective target of HIV-1 transcriptional inhibitors. Proc Natl Acad Sci USA 94:6110–6115
Sarno S, Reddy H, Meggio F, Ruzzene M, Davies SP, Donella-Deana A, Shugar D, Pinna LA (2001) Selectivity of 4, 5, 6, 7-tetrabromobenzotriazole, an ATP site-directed inhibitor of protein kinase CK2. FEBS Lett 496:44–48
Vangrevelinghe E, Zimmermann K, Schoepfer J, Portmann R, Fabbro D, Furet P (2003) Discovery of a potent and selective protein kinase CK2 inhibitor by high-throughput docking. J Med Chem 46:2656–2662
Zandomeni R, Zandomeni MC, Shugar D, Weinmann R (1986) Casein kinase type II is involved in the inhibition by 5, 6-dichloro-1-beta-D-ribofuranosylbenzimidazole of specific RNA polymerase II transcription. J Biol Chem 261:3414–3419
Suzuki Y, Cluzeau J, Hara T, Hirasawa A, Tsujimoto G, Oishi S, Ohno H, Fujii N (2008) Structure–activity relationship of pyrazine-based CK2 inhibitors: synthesis and evaluation of 2, 6-disubstituted pyrazines and 4.6-disubstituted pyrimidines. Arch Pharm 341:554–561
Nie Z, Perretta C, Erickson P, Margosiak S, Lu J, Averill A, Almassy R, Chu S (2008) Structure-based design and synthesis of novel macrocyclic pyrazolo[1,5-a][1,3,5]triazine compounds as potent inhibitors of protein kinase CK2 and their anticancer activities. Bioorg Med Chem Lett 18:619–623
Raaf J, Brunstein E, Issinger OG, Niefind K (2008) The Ck2 alpha/CK2 beta interface of human protein kinase CK2 harbors a binding pocket for small molecules. Chem Biol 15:111–117
Ermakova I, Boldyreff B, Issinger OG, Niefind K (2003) Crystal structure of a C-terminal deletion mutant of human protein kinase CK2 catalytic subunit. J Mol Biol 330:925–934
Niefind K, Yde CW, Ermakowa I, Issinger OG (2007) Evolved to be active: sulfate ions define substrate recognition sites of CK2alpha and emphasize its exceptional role within the CMGC family of eukaryotic protein kinases. J Mol Biol 370:427–438
Sekiguchi Y, Tetsuko N, Kinoshita T, Nakanishi I, Kitaura K, Hirasawa A, Tsujimoto G, Tada T (2009) Structural insight into human CK2alfa in complex with the potent inhibitor ellagic acid. Bioorg Med Chem Lett 19:2920–2923
Otwinoski Z, Minor W (1997) Processing of X-ray diffraction data collected in oscillation mode. Method in enzymology, vol 276. Academic press, New York, pp 307–326
Vagin A, Teplyakov A (2000) An approach to multi-copy search in molecular replacement. Acta Crystallogr Crystallogr D 56:1622–1624
Collaborative Computational Project, Number 4 (1994) The CCP4 suite: programs for protein crystallography. Acta Crytallogr D Biol Crystallogr 50:760–763
Labute P (2009) Protonate3D: assignment of ionization states and hydrogen coordinates to macromolecular structures. Proteins 75:187–205
Sturgeon JB, Laird BB (2000) Symplectic algorithm for constant-pressure molecular dynamics using a Nose-Poincare thermostat. J Chem Phys 112:3474–3482
Halgren TA, Nachbar RB (1996) Merck molecular force field. IV. Conformational energies and geometries for MMFF94. J Comput Chem 17:587–615
Ryckaert TA, Ciccotti G, Berendsen HJC (1977) Numerical integration of the Cartesian equation of motion of a system with constrains: molecular dynamics of n-alkanes. J Comput Phys 23:327–341
Labute P (2008) The generalized Born/volume integral implicit solvent model: estimation of the free energy of hydration using London dispersion instead of atomic surface area. J Comput Chem 29:1693–1698
Baum B, Mohamed M, Zayed M, Gerlach C, Heine A, Hangauer D, Klebe G (2009) More than a simple lipophilic contact: a detailed thermodynamic analysis of nonbasic residues in the S1 pocket of thrombin. J Mol Biol 390:56–69
Dunitz JD (1994) The entropic cost of bound water in crystals and biomolecules. Science 264:670
Chang CA, Chen W, Gilson MK (2007) Ligand conformational entropy and protein binding. Proc Natl Acad Sci USA 104:1534–1539
Fedorov DG, Ishimura K, Ishida T, Kitaura K, Pulay P, Nagase S (2007) Accuracy of the three-body fragment molecular orbital method applied to Moller-Plesset perturbation theory. J Comput Chem 28:1476–1484
Ford DM (2005) Enthalpy–entropy compensation is not a general feature of weak association. J Am Chem Soc 127:16167–16170
Starikov EB, Norden B (2007) Enthalpy–entropy compensation: a phantom or something useful? J Phys Chem B 111:14431–14435
Kawasaki Y, Chufan EE, Lafont V, Hidaka K, Kiso Y, Amzel LM, Freire E (2010) How much binding affinity can be gained by filling a cavity? Chem Biol Drug Des 75:143–151
Taylor JD, Ababou A, Fawaz RR, Hobbs CJ, Williams MA, Ladbury JE (2008) Structure, dynamics, and binding thermodynamic of the v-Src SH2 domain: implications for drug design. Proteins 73:929–940
Lafont V, Armstrong AA, Ohtaka H, Kiso Y, Amzel MA, Freire E (2007) Compensating enthalpic and entropic changes hinder binding affinity optimization. Chem Biol Drug Des 69:413–422
Freire E (2009) A thermodynamic approach to the affinity optimization of drug candidates. Chem Biol Drug Des 74:468–472
Liu C, Wrobleski ST, Lin J, Ahmed G, Metzger A, Wityak J, Gillooly KM, Shuster DJ, McIntyre KW, Pitt S, Shen DR, Zhang RF, Zhang H, Doweyko AM, Diller D, Henderson I, Barrish JC, Dodd JH, Schieven GL, Leftheris K (2005) 5-Cyanopyrimidine derivatives as a novel class of potent, selective, and orally active inhibitors of p38a MAP kinase. J Med Chem 48:6261–6270
Acknowledgments
These studies are supported by the Program of Fundamental Studies in Health Science of the National Institute of Biochemical Innovation (NIBIO). The synchrotron radiation experiments were done at the BL6A and BL17A stations in the Photon Factory, and at the BL44XU station in SPring-8 faculty with MX225-HE (Rayonix), which is financially supported by Academia Since and National Synchrotron Radiation Research Center (Taiwan, ROC).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Kinoshita, T., Sekiguchi, Y., Fukada, H. et al. A detailed thermodynamic profile of cyclopentyl and isopropyl derivatives binding to CK2 kinase. Mol Cell Biochem 356, 97–105 (2011). https://doi.org/10.1007/s11010-011-0960-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-011-0960-9