Abstract
Herein, we describe an open-source, Python-based, script to treat the output of differential scanning calorimetry (DSC) experiments, called pyDSC, available free of charge for download at https://github.com/leonardo-chiappisi/pyDSC under a GNU General Public License v3.0. The main aim of this program is to provide the community with a simple program to analyze raw DSC data. Key features include the correction from spurious signals, and, most importantly, the baseline is computed with a robust, physically consistent approach. We also show that the baseline correction routine implemented in the script is significantly more reproducible than different standard ones proposed by proprietary instrument control software provided with the microcalorimeter used in this work. Finally, the program can be easily applied to large amount of data, improving the reliability and reproducibility of DSC experiments.
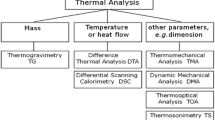
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Differential scanning calorimetry (DSC) is a powerful thermo-analytical technique which detects heat changes associated with physical and chemical transformation in biological and non-biological samples. Due to the simplicity of the technique, the relatively low-cost of the apparatus, the ease of data analysis, DSC found wide application in very diverse fields of both an academic and industrial research activities. Several excellent reviews covering the use of DSC can be found in the literature. With focus on colloidal and biophysical science, we address the reader to some covering topics such as protein conformation [16, 18, 31], lipid phase transitions [13], (bio)polymer stability [26], and on mixed systems [17, 25, 30].
In many cases, a DSC experiment is used to extract the enthalpy change \(\varDelta \text {H}\) of the studied process and the temperature of the transition. This analysis is straightforward, especially when the data exhibit a good signal-to-noise ratio. However, a DSC curve usually contains an abundance of information, which can be extracted by an in-depth analysis [7, 11, 12, 20, 24, 28, 32].
For every, even simple, analysis of a DSC curve, a correct evaluation of the baseline represents a critical step, whose importance is a matter of discussion since the origins of the technique [4, 14, 22, 27]. However, the baseline correction is mostly performed in a semiautomatic procedure by the proprietary software provided with the majority of commercial calorimeters. In fact, the correction consists in a spline interpolation between two arbitrary chosen regions before and after the transition region. While the procedure delivers correct results in the majority of cases, it is prone to artifacts—in particular, when the signal is close to the limits of the investigated temperature range and/or if a relevant heat capacity change occurs before and after the transition. In addition, most correction procedures are not based on a physical consistent model.
The baseline of a DSC curve reflects the difference between the heat capacity of the sample and of the reference. Provided spurious contributions to the signal from the empty cells and from a mismatch of buffer amount in sample and reference, besides the asymmetries of the measuring system, are taken into account, the baseline reflects the apparent heat capacity of the investigated compound. In the approximation of a two step transition, e.g., from a state A to a state B, the apparent heat capacity as a function of temperature T (\(C_{{\text {p}}}(T)\)) is given by:
with \(\alpha (T)\) being the degree of conversion of the process as a function of temperature; \(C_{{\text {P}}}^{\text{A}}\) and \(C_{{\text {P}}}^{\text{B}}\) are the apparent heat capacities of the compound in state A and B, respectively. \(C_{{\text {P}}}^{\text{A}}\) and \(C_{{\text {P}}}^{\text{B}}\) will be considered to be temperature independent within the investigated range in this work.
A physical consistent baseline subtraction, associated with a correction of residual signals, is simple steps which may significantly improve the data quality and the amount of information which can be extracted from the experiment. Herein, we describe a simple, freely accessible and modifiable tool, pyDSC, to treat DSC experiments. The program, which will be constantly updated, can be freely downloaded from the GitHub repository https://github.com/leonardo-chiappisi/pyDSC. At the same link, the up-to-date manual of the program can be found.
In the following, we provide a few experimental notes on a typical DSC experiment, a concise description of the operations performed by pyDSC, together with some examples.
Materials and methods
The DSC curves used to describe the functionalities of pyDSC were recorded on a micro-DSC III from Setaram, France. The instrument is provided with the SetSoft2000 software suite. To test the program, we have investigated a 8 mass% solution of the \(\hbox {EO}_{26}\)–\(\hbox {PO}_{{40}}\)–\(\hbox {EO}_{{26}}\) triblock copolymer Pluronic P85, bought from Sigma-Aldrich. The solutions were prepared using water of Millipore grade. Furthermore, pyDSC was applied to analyze DSC curves recorded on a Nano DSC, on a Multicell DSC and on a DSC 2920 from TA-Instruments.
Experimental
In order to work with the apparent heat capacity of the investigated compound, the DSC raw data need to be corrected for the contribution of the empty cells and of the buffer. Accordingly, three sets of measurements can be provided to the script:
-
The standard sample versus buffer run. For a proper normalization of the data, information such as the mass of sample and the concentration of the investigated compound needs to be known. Moreover, to minimize the contribution of the buffer to the baseline, the mass of buffer in the sample cell and in the reference cell should be as close as possible. We define the difference of mass of buffer in the sample cell and in the reference cell as \(\varDelta m_{{\text {BS}}}\).
-
The buffer versus buffer run. This run is used to determine the heat capacity of the buffer solution and will be used to correct the DSC data for the contribution of the buffer to the recorded signal. In contrast to the sample versus buffer run, where the cells are filled with possibly the exact amount of sample, in this run the amount of buffer in the two cells should differ, in order to get a good signal-to-noise. We recommend that the difference in mass between the two buffer cells, \(\varDelta m_{{\text {BB}}}\), is about 10% of the mass of buffer in the cell.
-
The empty cell versus empty cell run. This run is used to correct both the sample versus buffer and the buffer versus buffer run with the contribution arising from the different heat capacities of the empty cells and electronic asymmetries. In order to be effective, we recommend to use always the same reference and sample cells for all the measurements.
Program description
The script is written in Python 3, it makes use of the SciPy [19] and matplotlib [15] packages and is platform independent. Several operations on the data are performed, provided the required information is available. The following operations are executed on the raw data files:
-
1.
Data averaging and resizing The DSC raw data of the sample and, if provided, of the empty cell and buffer-buffer runs are read by the program and the points which fall outside the temperature region of interest are discarded. The reference measurements are averaged. If requested by the user, in order to facilitate any further data treatment, the number of points is reduced by a factor N, defined by the user.
-
2.
Heat flow conversion in heat capacity The heat flow is converted into a heat capacity upon division by the heating rate. The heating rate is either provided by the user, or it is determined from the raw data file if the time information is contained. The raw heat flows are plotted in the rawdata.pdf file while the heat flows normalized by the heating rate and the sample amount are plotted in corrected_data.pdf.
-
3.
Empty cell and buffer correction If the data file corresponding to the empty cell/empty cell and/or buffer/buffer runs are provided, the raw data are corrected for both contributions. In particular, the empty cell/empty cell run is used to take into account any non-constant contribution arising from the empty cells or the instrument. The buffer/buffer run is used to take into account any contribution arising from a mismatch in amount of buffer contained in the reference and in the sample cell. In detail, the apparent heat capacity of the sample run \(Cp_{{\text {s}}}\), is corrected for the apparent heat capacity of the empty cell \(Cp_{{\text {EC}}}\) and for the apparent heat capacity of the buffer/buffer run \(Cp_{{\text {bb}}}\) as:
$$\overline{Cp_{{\text {s}}}} = Cp_{{\text {s}}} - Cp_{{\text {EC}}} - \frac{\varDelta m_{{\text {BS}}}(Cp_{{\text {bb}}}-Cp_{{\text {EC}}})}{\varDelta m_{{\text {BB}}}}$$(2)with \(\varDelta m_{{\text {BS}}}\) and \(\varDelta m_{{\text {BB}}}\) being the mass difference between the buffer contained in the sample and in the reference in the sample run, and between the buffers in the buffer/buffer run. If no empty cell/empty cell or buffer/buffer runs are provided, this correction will not be performed and the non-corrected sample run will be further analyzed. In most of the cases, \(\varDelta m_{{\text {BS}}}\) should be so small that the last term might be negligible. For instruments working with non-removable sample cells with constant filling volume, i.e., \(\varDelta m_{{\text {BB}}}=0\), by loading in pyDSC a buffer-buffer titration instead of the empty cell/empty cell run.
-
4.
Normalization of apparent heat flow The apparent heat flow of the sample run is finally normalized by the amount of solute. In particular, the apparent heat capacity is normalized either by the mass of solute, resulting in a \(Cp_{{\text {s}}}\) given in J \(\hbox {K}^{-1}\,\hbox {g}^{-1}\), or by the moles of solute, if the molecular weight is known, thus resulting in a \(Cp_{{\text {s}}}\) given in J \(\hbox {K}^{-1}\,\hbox {mol}^{-1}\).
-
5.
Baseline correction The core task carried out by the program is to compute a physically meaningful baseline, as described earlier in the text. The baseline is constructed following an iterative procedure proposed elsewhere [22, 29]. In detail, the baseline \(CP_{{\text {bl}}}(T)\) is defined as follows:
$$CP_{{\text {bl}}}(T) = (1-\alpha )CP_{{\text {bl}}}^{{\text {pre}}}(T) + \alpha \cdot CP_{{\text {bl}}}^{{\text {post}}}(T)$$(3)with \(CP_{{\text {bl}}}^{{\text {pre}}}(T)\) and \(CP_{{\text {bl}}}^{{\text {post}}}(T)\) are the baselines determined in the regions at lower and higher temperatures than the peak, respectively. \(\alpha\) is the degree of conversion, defined as:
$$\alpha (T) = \frac{\int _0^{{\text {T}}} CP_{{\text {s}}}(T) - CP_{{\text {bl}}}(T) \text {d}T}{\int _0^{\infty } CP_{{\text {s}}}(T) - CP_{{\text {bl}}}(T) \text {d}T}$$(4)where \(CP_{{\text {s}}}(T)\) is the apparent heat capacity difference between sample and reference, after normalization and eventual correction. \(CP_{{\text {bl}}}^{{\text {pre}}}(T)\) and \(CP_{{\text {bl}}}^{{\text {post}}}(T)\) represent the temperature-dependent difference of heat capacity of sample and reference, eventually corrected from the contribution from the empty cells and solvent mismatch. They are assumed to exhibit a linear dependence from temperature. In Fig. 1, the baseline construction is schematically depicted.
At this stage of the program, the enthalpy change of the process, computed as \(\varDelta \text {H} = \int _0^{\infty }\! CP_{{\text {s}}}(T) - CP_{{\text {bl}}}(T)\, \text {d}T\), is evaluated and its uncertainty is estimated. The uncertainty in \(\varDelta \text {H}\) is estimated from the stochastic fluctuations of the signal in the regions where \(CP_{{\text {bl}}}^{{\text {pre}}}(T)\) and \(CP_{{\text {bl}}}^{{\text {post}}}(T)\) are determined. By definition, in these regions no processes take place, and the signal fluctuates around an apparent heat capacity of zero J \(\hbox {K}^{-1}\) is used to determine the standard deviation of the heat capacity. The heat exchanged during the examined process is computed by integration using the trapezoidal rule, and its uncertainty by standard error propagation.
Program usage
pyDSC performs all the aforementioned steps on the DSC raw data provided by the user, located in the folder rawdata. In addition, the user will provide all the necessary information to perform the corrections within the Files.txt and Input_params.txt text files. Full details are provided in the user manual and exemplary files at https://github.com/leonardo-chiappisi/DSC_correction. Finally, the pyDSC script can be executed within a terminal by invoking the Python3 interpreter as:
In addition to an output ASCII file for each sample run, the script will generate five output figures: Alpha.pdf, Baseline_data.pdf, Corrected_data.pdf, Final_data.pdf, and Rawdata.pdf.
In Rawdata.pdf, the recorded heat flows of the different runs are reported as a function of temperature.
In Corrected_data.pdf, the sample run heat capacity, after binning and normalization by sample amount, is reported as a function of temperature.
In Baseline_data.pdf, the heat capacity of each sample run, after binning, normalization by sample amount, and empty cell and buffer correction, is reported, together with the calculated baseline as a function of temperature. Moreover, the regions used for the determination of the baseline before and after the peak are highlighted.
In Final_data.pdf, the heat capacity of each sample run, after binning, normalization by sample amount, empty cell and buffer correction, and baseline subtraction, is reported as a function of temperature.
Finally, in alpha.pdf, the degree of conversion determined for each sample run is reported as a function of temperature.
The different plots allow the user to determine whether the correction procedures executed during each step are performed correctly and keep control over data treatment process.
In addition to the five figures, one ASCII file for each sample run is created. The file contains a summary of the information used to treat the data in the header, in addition to the computed values of enthalpy change and change in heat capacity at the peak temperature. Moreover, the file contains four data columns: temperature, the normalized, corrected, and binned heat capacity with baseline subtracted, without baseline subtraction, the baseline, and the degree of conversion of the process.
Results and discussion
The script is tested using a semidiluted solution of the triblock copolymer poly-(ethyleneoxide)–poly-(propyleneoxide)–poly-(ethyleneoxide) Pluronic P85 (\(\hbox {EO}_{{26}}\)–\(\hbox {PO}_{{40}}\)–\(\hbox {EO}_{{26}}\)). DSC was extensively employed to reveal the self-assembly behavior of this class of polymers in solutions [1,2,3, 10, 21], and the system was chosen due to its good signal-to-noise ratio, non-negligible baseline dependence from the temperature, and the large availability of reference data to compare with.
The proprietary software commonly used to perform a first analysis of the microcalorimetric data provide different routines for the baseline subtraction and the blank correction, which typically consists in a simple subtraction of the raw heat flow data. For what concerns the baseline evaluation and subtraction, different algorithms are available, depending on the vendor and on the software version. However, in all cases, the baseline under the transition peak is evaluated from, somehow arbitrary chosen, regions prior and after the process signal.
To test the robustness of the pyDSC script, the enthalpy of micellization recorded for a solution of the triblock copolymer \(\hbox {EO}_{{26}}\)–\(\hbox {PO}_{{40}}\)–\(\hbox {EO}_{{26}}\) was evaluated using pyDSC and two different algorithms provided by the Setaram data analysis software: “linear”—which uses a line from the starting to the end point arbitrarily selected by the user—and “curve”—which generates a spline curve from the selected initial/final points that are selected by user—using different regions for the baseline evaluation. For the sake of the comparison, we also report data analyzed with pyDSC without the correction for the buffer and empty cell contributions. The results are shown in Table 1 and two clear results stand out: the arbitrary choice of the baseline region has a non-negligible effect on the obtained values of \(\varDelta \text {H}_{{\text {m}}}\), with a variation of approx. 2 and 6% for the linear and the curve algorithm, respectively, versus only 0.7% found with pyDSC. The effect is even smaller when the contribution of the solvent and empty cells are taken into account. Furthermore, trending values are observed with a systematic variation of the obtained enthalpy change for linear integration. The DSC curves with the corresponding four baselines are shown in Fig. 2, further demonstrating that the arbitrary choice of the integration region as only a minimal effect on the results.
Apparent heat capacity recorded for a 8 mass% aqueous solution of \(\hbox {EO}_{{26}}\)–\(\hbox {PO}_{{40}}\)–\(\hbox {EO}_{{26}}\) during a heating ramp at \(0.5\,^\circ {\text {C}}\,{\text {min}}^{-1}\), with the corresponding baselines, determined assuming a peak temperature interval of 18–50 \(^\circ\)C (baseline 1), 17–51 \(^\circ\)C (baseline 2), 16–52 \(^\circ\)C (baseline 3), 15–53 \(^\circ\)C (baseline 4). The top insert shows a magnification of the baseline region
pyDSC was also applied to correct the calorimetric output from DSC experiments performed on very diverse systems on other instruments. In particular, in Fig. 3, pyDSC was applied for the correction of data stemming from the micro-DSC III from Setaram and from the DSC 2920, nano DSC, and Multicell DSC from TA-Instruments. The curves refer to the melting of isopropylpalmitate in a mixture of surfactant and alcohols [8], to the melting of Indium, to the thermal transition within self-assembled fibrils [5], and to the thermal denaturation of a protein [23] (See figure caption and corresponding references for further details).
Raw heat capacities (full lines) and calculated baselines (dashed lines) for different DSC curves. In particular, a the melting of Isopropylpalmitate in the presence of butanol and a polyoxyethylene glycol oleic ether nonionic surfactant, recorded on a multi-cell DSC from TA-Instruments (see Ref. [8] for full details on the system); b the melting of Indium, recorded on a DSC 2920 from TA-Instruments; c the melting of fibrils made from the supramolecular polymerization of a \(\pi\)-donor \(\pi\)-acceptor monomer, recorded on a Setaram micro-DSC III (see Ref. [5] for full details on the system); d thermal denaturation of the orange carotenoid protein recorded on a nano DSC from TA-Instruments (see Ref. [23] for full details on the system)
Finally, it is useful to recall that the trustworthiness of the microcalorimetric data does not depend only on the data correction and baseline subtraction procedures, but a regular instrument maintenance and calibration are equally essential [6, 9].
Conclusions
In this contribution, we present pyDSC a simple, python-based script to treat differential scanning calorimetry data. The software is platform independent and freely usable. It offers a simple but robust and physically meaningful approach to evaluate the baseline in DSC experiments. Moreover, the raw DSC data can be corrected for the contribution of the empty cells and the buffer, and their size reduced without loss of quality.
We anticipate that all these features will increase the robustness and reproducibility of the output of DSC experiments across very different communities. In addition, we commit ourselves to keep the software up to date, and ask the scientific community for feedback and suggestions for further implementations. The source code is available free of charge under a GNU General Public License v3.0 (GPLv3).
References
Alexandridis P, Hatton TA. Poly(ethylene oxide)–poly(propylene oxide)–poly(ethylene oxide) block-copolymer surfactants in aqueous-solutions and at interfaces—thermodynamics, structure, dynamics, and modeling. Colloids Surf A Physicochem Eng Asp. 1995;96(1–2):1–46.
Alexandridis P, Holzwarth JF, Hatton TA. Micellization of poly(ethylene oxide)–poly(propylene oxide)–poly(ethylene oxide) triblock copolymers in aqueous solutions: thermodynamics of copolymer association. Macromolecules. 1994;27(9):2414–25.
Barba AA, D’Amore M, Grassi M, Chirico S, Lamberti G, Titomanlio G. Investigation of Pluronic F127-water solutions phase transitions by DSC and dielectric spectroscopy. J Appl Polym Sci. 2009;114(2):688–95.
Brennan WP, Miller B, Whitwell JC. An improved method of analyzing curves in differential scanning calorimetry. Ind Eng Chem Fundam. 1969;8(2):314–8.
Cera L, Chiappisi L, Böttcher C, Schulz A, Schoder S, Gradzielski M, Schalley CA. PolyWhips: directional particle transport by gradient-directed growth and stiffening of supramolecular assemblies. Adv Mater. 2017;29(8):1604430.
Cerdeiriña CA, Míguez JA, Carballo E, Tovar CA, De La Puente E, Romaní L. Highly precise determination of the heat capacity of liquids by DSC: calibration and measurement. Thermochim Acta. 2000;347(1–2):37–44.
Chiappisi L, Lazzara G, Gradzielski M, Milioto S. Quantitative description of temperature induced self-aggregation thermograms determined by differential scanning calorimetry. Langmuir. 2012;28(51):17609–16.
Chiappisi L, Noirez L, Gradzielski M. A journey through the phase diagram of a pharmaceutically relevant microemulsion system. J Colloid Interface Sci. 2016;473:52–9.
Della Gatta G, Richardson MJ, Sarge SM, Stølen S. Standards, calibration, and guidelines in microcalorimetry. Part 2. Calibration standards for differential scanning calorimetry* (IUPAC Technical Report). Pure Appl Chem. 2006;78(7):1455–76.
Dintcheva NT, Catalano G, Arrigo R, Morici E, Cavallaro G, Lazzara G, Bruno M. Pluronic nanoparticles as anti-oxidant carriers for polymers. Polym Degrad Stabil. 2016;134:194–201.
Dlubek G, Pompe G, Pionteck J, Janke A, Kilburn D. Differential scanning calorimetry (DSC) for interdiffusion studies in PCV/PnBMA blend: a quantitative analysis. Macromol Chem Phys. 2003;204(9):1234–44.
Fessas D, Iametti S, Schiraldi A, Bonomi F. Thermal unfolding of monomeric and dimeric \(\beta\)-lactoglobulins. Eur J Biochem. 2001;268(20):5439–48.
Heerklotz H. The microcalorimetry of lipid membranes. J Phys Condens Matter. 2004;16(15):R441–67.
Höhne GWH, Hemminger WF, Flammersheim HJ. Differential scanning calorimetry. 2nd ed. Heidelberg: Springer; 2003.
Hunter JD. Matplotlib: a 2D graphics environment. Comput Sci Eng. 2007;9(3):90–5.
Ibarra-Molero B, Naganathan AN, Sanchez-Ruiz JM, Muñoz V. Modern analysis of protein folding by differential scanning calorimetry. In: Feig AL, editor. Methods in enzymology—calorimetry. Amstedam: Academic Press/Elsevier; 2016. p. 281–318.
Jelesarov I, Bosshard HR. Isothermal titration calorimetry and differential scanning calorimetry as complementary tools to investigate the energetics of biomolecular recognition. J Mol Recognit. 1999;12(1):3–18.
Johnson CM. Differential scanning calorimetry as a tool for protein folding and stability. Arch Biochem Biophys. 2013;531(1–2):100–9.
Jones E, Oliphant T, Paterson P. SciPy: open source scientific tools for Python
Leharne S. The physical chemistry of high-sensitivity differential scanning calorimetry of biopolymers. ChemTexts. 2017;3(1):1–12.
Lopes JR, Loh W. Investigation of self-assembly and micelle polarity for a wide range of ethylene oxidepropylene oxideethylene oxide block copolymers in water. Langmuir. 1998;14(4):750–6.
Malakhov DV, Abou Khatwa MK. Constructing a self-consistent integral baseline by using cubic splines. J Therm Anal Calorim. 2007;87(2):595–9.
Moldenhauer M, Sluchanko NN, Tavraz NN, Junghans C, Buhrke D, Willoweit M, Chiappisi L, Schmitt FJ, Vukojević V, Shirshin EA, Ponomarev VY, Paschenko VZ, Gradzielski M, Maksimov EG, Friedrich T. Interaction of the signaling state analog and the apoprotein form of the orange carotenoid protein with the fluorescence recovery protein. Photosynth Res. 2018;135(1–3):125–39.
Paterson IF, Armstrong JK, Chowdhry BBZ, Leharne SA. Thermodynamic model fitting of the calorimetric output obtained for aqueous solutions of oxyethyleneoxypropyleneoxyethylene triblock copolymers. Langmuir. 1997;13(8):2219–26.
Prenner E, Chiu M. Differential scanning calorimetry: an invaluable tool for a detailed thermodynamic characterization of macromolecules and their interactions. J Pharm Bioallied Sci. 2011;3(1):39.
Privalov PL, Dragan AI. Microcalorimetry of biological macromolecules. Biophys Chem. 2007;126(1–3):16–24.
Sandu C, Singh RK. Modeling differential scanning calorimetry. Thermochim Acta. 1990;159:267–98.
Schulthess T, Schönfeld HJ, Seelig J. Thermal unfolding of apolipoprotein A—1. Evaluation of methods and models. Biochemistry. 2015;54(19):3063–75.
Van der Plaats G. A theoretical evaluation of a heat-flow differential scanning calorimeter. Thermochim Acta. 1984;12:77–82.
Weber PC, Salemme FR. Applications of calorimetric methods to drug discovery and the study of protein interactions. Curr Opin Struct Biol. 2003;13(1):115–21.
Yang Z, Brouillette CG. A guide to differential scanning calorimetry of membrane and soluble proteins in detergents; 2016. pp. 319–58
Zehender F, Ziegler A, Schönfeld HJ, Seelig J. Thermodynamics of protein self-association and unfolding. The case of apolipoprotein A—I. Biochemistry. 2012;51(6):1269–80.
Acknowledgements
Open Access funding provided by Projekt DEAL. LC acknowledges the TU-Berlin and the ILL for postdoctoral funding though a three-year cooperation agreement. AC acknowledges the CFM foundation for funding her doctoral thesis. The partnership for soft condensed matter (PSCM) at the ILL Grenoble is acknowledged for providing the differential scanning calorimeter (DSC).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cisse, A., Peters, J., Lazzara, G. et al. PyDSC: a simple tool to treat differential scanning calorimetry data. J Therm Anal Calorim 145, 403–409 (2021). https://doi.org/10.1007/s10973-020-09775-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10973-020-09775-9