Abstract
The NAC (NAM, ATAF1/2 and CUC2) domain protein plays an important role in plant growth and development. The function of members of NAC gene family has been deeply studied in many plants. However, the evolutionary relationships and characteristics of NAC family genes in Dendrobium catenatum (D. candidum) unclear. In this study, we identified 33 NAC genes in D.catenatum, all contain NAM conservative domain. Subcellular localization predictions indicated that all the DcNAC proteins are localized to the nucleus. Phylogenetic analysis suggested that the DcNAC gene family could be divided into four groups. Then, the amino-acid composition, physicochemical properties, gene structure, motif, and promoter cis-acting elements were analyzed, the evolutionarily conservative gene DcNAC043-1 and DcNAC043-2 were found. Using qRT-PCR and phloroglucinol staining experiments, it was demonstrated that DcNAC043s can respond to drought stress in Dendrobium. Within a certain range, the longer the duration of drought stress, the higher the expression level of DcNAC043s, and the stronger the degree of plant lignification. We supplemented the relevant information of NAC gene family in D. catenatum. At the same time, the gene function of DcNAC043s and its contribution to the response of Dendrobium to drought stress were verified. These results provide a comprehensive evolutionary history of NAC genes in D. catenatum, and insight into the biological functions of DcNAC043s genes in response to drought stress.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants often suffer from a variety of natural stress, such as salt, drought, cold, pathogens, and insect pests (Shao et al. 2015; Khalil et al. 2022). To cope with the impact of these different environmental pressures on plant growth and development, plants have evolved a series of defense mechanisms to resile from these stresses (Aldhanhani et al. 2022). NAC transcription factor gene family is a large plant specific gene family, as a molecular switch that can activate or inactivate gene expression, it plays an important role in plant response to these stresses (Shao et al. 2015; Yu et al. 2016).
The NAC gene family is designated, after the initials of three genes, NAM (NO APICAL MERISTEM, petunia) (Souer et al. 1996), ATAF1/2 (ARABIDOPSIS TRANSCRIPTION ACTIVATION FACTOR, Arabidopsis) and CUC2 (CUP-SHAPE COTYLEDON, Arabidopsis) (Aida et al. 1997). The five N-terminal motifs (A, B, C, D, E) of NAC transcription factor protein form a conservative domain. The length is about 160 amino acids, and the flank contains α- Helix and β- Fold structure. The motif A related to the formation of dimers and the motifs C and D that have nuclear localization signals and participate in DNA binding are highly conserved. The substructure variation of motif B and E is large, which may be closely related to the functional diversity of NAC genes (Jia et al. 2019). On the contrary, C-terminal domains are very different, with transcriptional activation or inhibition activities, and participate in the regulation of multiple networks (Olsen et al. 2005).
At present, NAC transcription factor family has been widely studied in plants species. Under the induction of different plant development stages and various environmental factors, NAC can activate specific target proteins to regulate plant growth and development and plant secondary growth (Nakashima et al. 2012). For example, in Arabidopsis, 117 NAC genes were identified (Kim et al. 2006). Overexpression of the NAC family gene VNI2, which could inhibit the normal development of xylem vessels in roots and aboveground parts by inhibiting the expression of xylem vessel development genes (Yamaguchi et al. 2010). In rice (Oryza sative), 151 NAC genes were identified (Nuruzzaman et al. 2010). Among them, after OsNAC6 overexpression in the plants, the transgenic plants showed a semi dwarfing phenotype (Sakamoto et al. 2016). In Populus trichocarpa, 163 NAC genes were identified (Hu et al. 2010). It was reported that overexpression of PopNAC122 gene will reduce plant height, cell size and cell number (Grant et al. 2010).
In addition, NAC genes has also been proved to play an important role when plants were subjected to biotic and abiotic stresses (Lee et al. 2012). For example, many NAC genes have the function of enhancing plant drought resistance. AtNAC016 could improve plant drought resistance by inhibiting the expression of AREB1, a negative regulator of ABA signaling pathway in Arabidopsis (Sakuraba et al. 2015). In rice, after OsNAC10 gene overexpression, the drought resistance of the plant was enhanced, the results of field experiments showed that the rice yield under drought and control conditions was significantly higher than that of the wild type (WT) (Jeong et al. 2010). It was reported that after overexpression of TaNAC2, the resistance of plants to drought, salt and cold stress was enhanced in Triticum aestivum (Mao et al. 2012). To sum up, NAC gene family has been identified in most plants, and the functions of some genes have been verified. Nevertheless, little is known about this gene family in Dendrobium catenatum (D. candidum).
D. candidum is the third largest genus of Orchidaceae and contains approximately 1,450 species, with high ornamental and medicinal value in China (Zhang et al. 2016). NAC gene plays an important role in plant response to drought stress, which is crucial for plant growth and development (Hu et al. 2006; Zheng et al. 2009; Chen et al. 2018; Negi et al. 2018; Wu et al. 2023). However, few studies have focused on identifying NAC genes in D. candidum. We also examined that DcNAC043-1, DcNAC043-2 and DcMYB46 (the gene encoding DcNAC043 interacting protein) expression under drought stress. Simultaneously, the lignin content in the plant was detected under the condition of up regulation of DcNAC043s expression, to reveal the relationship) between DcNAC043s and drought stress responses. Our research provides insight into the evolution of NAC genes in plants and lays the foundation for elucidating the roles of NACs in regulating drought stress in D. candidum.
Materials and methods
Plant materials and treatments
The materials used in this study were collected by Ming-jin Huang, associate professor of Agricultural College/Dendrobium Institute of Guizhou University, in Yandang Mountain, Zhejiang Province, China, and then planted in Dendrobium Institute of Guizhou University (26°26′44″N/106°39′32"E). It was identified as Dendrobium candidum by the Department of Agriculture and Rural Affairs of Guizhou Province, named as Jinhu No. 1, and identified as “Qianren 20210012” (Non endangered species). To investigate the expression patterns of DcNAC043 and DcMYB46 genes under drought stress, the Dendrobium sown in nutritious soil in a growth chamber, with a 16 h light (25 °C)/8 h dark (20 °C) photoperiod, 60% humidity, and an 800 μmol m−2 s−1 light intensity. The photosynthetic photon flux density (PPFD) was measured using a quantum radiometer/photometer (LI 185 B; LI-COR Biosciences, Lincoln, NE, United States). One-years-old plants were transferred into new Nutrient soil for treatment. Dendrobium was treated with natural drought, three biological replicates of leaves were harvested after 0, 3, 6 and 9 day (d) of treatment (Fig. 1Aa–d). All samples were immediately frozen in liquid nitrogen and stored at – 80 ℃. All the samples were prepared for RNA extraction and quantitative real-time PCR (qRT-PCR) detection (Miao et al. 2021). The remaining dendrobium materials used in the experiment continue to grow in the Dendrobium Institute.
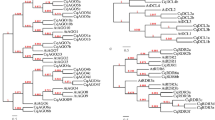
Phylogenetic relationships, gene structures of NAC proteins in A. thaliana and D.candidum, the amino acid sequences of AtNACs and DcNACs are aligned using ClustalW2. A Phylogenetic analysis of AtNAC and DcNAC proteins in Dendrobium. The phylogenetic tree is constructed by MEGA 7.0 using the neighbor-joining method with1000 bootstrap replicates and displayed using FigTree v1.4.0. At: A. thaliana; Dc: D. candidum. B Gene structures generated using the Gene Structure Display Server. Exons (CDS) and introns are shown as orange wedges and black lines, respectively
Identification of NAC family genes in D. catenatum
D. candidum Genomic data (BioProject ID PRJNA453230) were obtained from NCBI (http://www.ncbi.nlm.nih.gov/) publicly availabl (Miao et al. 2021). Pfam website (http://pfam.xfam.org/) was used to download the hidden Markov model (HMM) profile of the NAC domain (PF02365) (Mistry et al. 2021). To identify NAC proteins in these species, 117 NAC proteins sequences from Arabidopsis were used as queries in a reciprocal Basic Local Alignment Search Tool Protein (BLASTP) analysis (Camacho et al. 2009; Liu et al. 2018). Multiple sequence alignment of the NAC proteins was performed in ClustalW2 (http://www.ebi.ac.uk/Tools/msa/clustalw2) using the default parameters (E value < 1e-5) (Larkin et al. 2007). According to the aligned sequences, a new HMM in HMMER (HMMER 3.1, http://hmmer.org/) was constructedand and used as the query (E value < 0.01) to search against the D. candidum genome sequence data (Potter et al. 2018). NAC gene candidate was based on the gene encoding NAC domain protein (Shen et al. 2019).
Phylogenetic and protein property analysis of the NAC family in D. catenatum
Multiple sequence alignments of NAC coding sequences between D. candidum and A. thaliana sequences mentioned above were conducted using ClustalW2 with default parameters (http://www.genome.jp/tools bin/clustalw) (Liu et al. 2018). After screening the sequences that were closely related to Arabidopsis, based on these sequences (NAC031, NAC033, NAC043, NAC009, NAC018, NAC002 and NAC029), the related sequences of species that were closely related to D. candidum were obtained from NCBI (http://www.ncbi.nlm.nih.gov/) (Table S1), and then ClustalW2 Performs was usd multiple sequence alignments, with default parameters. The phylogenetic trees were generated using the Molecular Evolutionary Genetics Analysis (MEGA) 7 program (Tokyo Metropolitan University, Tokyo, Japan) (Kumar et al. 2016), with the NJ method, the p-distance + G substitution model, 1000 bootstrap replications, and conserved sequences with a coverage of 80% (Liu et al. 2018). Using FigTree v1.4. (http://tree.bio.ed.ac.uk/software/figtree/) to realize the phylogenetic trees visualization (Chen et al. 2015).
The physicochemical properties of each DcNAC protein were predicted using the ExPASy online program (http://web.expasy.org/translate/) (Duvaud et al. 2021). The transmembrane transport peptides were predicted using Tmpred (http://www.ch.embnet.org/software/TMPRED_form.html), and signal peptides were predicted using SignalP4.1 (http://www.cbs.dtu. dk/services/SignalP/) (Cruz et al. 2017). The subcellular localization of proteins were predicted using Plant-mPLoc online program (http://www.csbio.sjtu.edu.cn/bioinf/plant-multi/) (Li et al. 2020). The coding sequence alignments were imported into KaKs_calculator3.0 (Chinese Academy of Sciences, Beijing, China) to calculate the synonymous mutation rate (Ks) and non-synonymous mutation rate (Ka) using the NG method (Zhang et al. 2021). STRING datebase (https://string-db.org/) was used to display the protein–protein networks of NAC proteins (Szklarczyk et al. 2021). Gene Structure Display Server (GSDS 2.0, http://gsds.cbi.pku.edu.cn) was used to display the exon/intron structures of DcNAC genes (He et al. 2019).
Motif identification and cis-acting elements analysis
Conserved motifs in the proteins were identified using the Expectation Maximization for Motif Elucidation program (MEME v4.12.0, http://meme-suite.org) with the following parameter settings: The maximum number of motifs was 15, and the optimum width was set from 6 to 200. Only motifs with an e-value of 1e-10 were retained for further analysis (Nystrom et al., 2021). The results were visualized using TBtools (Chen et al. 2020). PlantCARE software (http://bioinformatics.psb.ugent.be/webtools/plantcare) was used to predict the cis-acting elements within 2000 bp upstream of all DcNAC genes (Sun et al. 2022a, b).
Expression analysis of the DcNAC043 and DcMYB46 by qRT-PCR
Total RNA was extracted from the Dendrobium leaf using an RNA Extraction Kit (Tiangen, Beijing, China), and cDNA was synthesized from 1 µg of total RNA using M-MLV transcriptase (TaKaRa Biotechnology, Dalian, China) according to the manufacturer’s instructions. And cDNA was synthesized using the PrimeScript™ 1st Strand cDNA Synthesis Kit (Takara, Japan) according to the manufacturer’s instructions (Ge et al. 2018). These reactions were executed under the following conditions: 37 °C for 15 min, 85 °C for 5 s, and finally ended at 4 °C. The reaction system was 2 × SYBR Green Pro Taq HS Premix 10 μL, primer F 0.4 μL, primer R 0.4 μL, cDNA 2 μL, ddH2O 7.2 μL (Pan et al. 2023).
Gene-specific primer sequences for the DcNAC043 and DcMYB46 were designed using Geneious Prime (Version. 2022.2) (Zhang et al. 2021) (Table S2). Each reaction was carried out in biological triplicate in a reaction volume of 20 µL containing 1.6 µL of gene-specific primers (1.0 µM), 2.0 µL of cDNA, 10 µL of SYBR green, 0.2 µL of ROX Reference Dye II, and 6 µL of sterile distilled water. The PCR program was as follows: 95 ℃ for 3 min; 45 cycles of 10 s at 95 ℃ and 30 s at 60 ℃; and then melt curve 65–95 ℃, increment 0.5 ℃ for 5 s (Liu et al. 2018). Melting curves were generated to estimate the specificity of these reactions. Relative expression levels were calculated using the 2−∆∆Ct method, with ACT2 used as internal controls (Wu et al. 2015).
Observation on lignin in cell wall of stem in D. catenatum
Stems of Dendrobium after 0, 3, 6 and 9 d of natural drought stress were sliced into 20 μM slice sections. Thin sections were stained for lignin with a HCl-phloroglucinol solution [2% (w/v) phloroglucinol (dissolved in 95% (v/v) ethanol): 2 M HCl = 1:1, fresh preparation] and observed under a Zeiss Axioscope A1 microscope (Leica) with a × 0.5 optical adapter (Patten et al. 2005).
Statistical analysis
All experiments were assessed using at least three independent repeats, and all data are represented as the mean ± SD. The statistical analysis was performed using a Student’s t test, and the significance was indicated using asterisks: *(p value < 0.05) or **(p value < 0.01).
Results
Identification of NAC proteins in D. candidum and other plants
Using the 117 AtNAC protein sequences in Arabidopsis as queries, 33 NAC proteins were identified from D. candidum. All DcNAC proteins share a conserved domain: The NAM domain. Relationships based on the phylogenetic tree; the nomenclature used for DcNAC protein was based on the corresponding AtNAC orthologs (Table 1). Based on the phylogenetic tree, eight relatively conservative NAC members were selected for subsequent analysis (Fig. 1A). Using the nine DcNAC protein sequences in D. candidum as queries, other 12 NAC043, 1 NAC033, 13 NAC031, 13 NAC029, 5 NAC018, 10 NAC009 and 9 NAC002 proteins were identified from 37 crops (P. equestris. and others) (Table 1).
The number of amino acid residues in the DcNAC proteins ranged from 220 (TuNAC018) to 449 (AsNAC043), with an average of 336. The relative molecular weights (MWs) ranged from 24.48 (TuNAC018) to 49.93 kDa (PtNAC031). The isoelectric points (pIs) were predicted to range from 5.32 (DcaNAC002) to 9.06 (PeNAC029), and 28 members had pI values > 7 and others had pI values < 7. No transmembrane helix or signal peptide was found in any NAC protein (Table 1). All the NAC proteins were predicted to localize in the nucleus, which were consistent with the general nuclear localization of TFs.
Phylogenetic analysis and classification of the NAC proteins in D. candidum and other plants
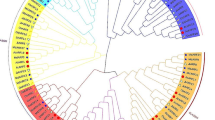
To clarify the evolutionary relationships between NAC proteins from Arabidopsis and D. candidum, we aligned the sequences of 33 NAC proteins using AtNACs. Based on affinities presented by the evolutionary tree, and we removed groups with only AtNAC or DcNAC, NAC proteins were divided into four groups (Fig. 2A). Although different copies of the same species were still more closely related in evolutionary relationship than the same copies of different species, we can still find the closely related copies of Arabidopsis and D. candidum in these groups. Including the NAC31 clade (class II), the NAC033 and NAC043 clade (class III), and the NAC009, NAC029, NAC018 and NAC002 clade (class IV).
Phylogenetic relationships and conserved motifs between DcNAC and other plant NAC proteins. A The phylogenetic tree is constructed using MEGA 7.0 with the neighbor-joining method (1000 bootstrap replicates) and displayed using FigTree v1.4.0. NAC proteins in the phylogenetic tree are clustered into four distinct groups. At: A. thaliana, Dc: D. candidum (marked in red), Cl: Carex littledalei, Jr: Juglans regia, Pt: Populus trichocarpa, Zm: Zea mays, Dca: Dioscorea cayenensis, Pe: Phalaenopsis equestris, Dn: Dendrobium nobile, Ma: Musa acuminata; Ac: Ananas comosus, Eg: Elaeis guineensis, Cn: Cocos nucifera, Pd: Phoenix dactylifera, Qs: Quercus suber, Pa: Prunus avium, Me: Manihot esculenta, Tc: Theobroma cacao, Vv: Vitis vinifera, Os: Oryza sativa, As: Apostasia shenzhenica, Ql: Quercus lobata, Zj: Ziziphus jujuba, Hb: Hevea brasiliensis, Ao: Asparagus officinalis, Tu: Triticum urartu, Mn: Morus notabilis, Dk: Diospyros kaki, Nt: Nicotiana tomentosiformis, Bn: Brassica napus, Ss: Spatholobus suberectus, Rc: Ricinus communis, Csa: Cannabis sativa, Tg: Tulipa gesneriana, Pj: Phtheirospermum japonicum, Sp: Solanum pennellii, Cc: Citrus clementina, Cs: Citrus sinensis. B Conservative motifs of DcNAC proteins identified by MEME. The motifs were indicated by colored boxes and their numbers are shown in the scale below the diagram. C Sequences of the 15 conserved motifs in DcNACs identified in this study
To further investigate the evolutionary relationships of NAC proteins in plants, we constructed a larger neighbor-joining (NJ) tree based on 8 NAC protein sequences identified from D. candidum, 63 NAC proteins from other 37 species (Phalaenopsis equestris et al.) were found and constructed a phylogenetic tree together. According to kinship, they could be divided into 4 classes (Fig. 2A). Class I and III were the two most conservative groups, class I contains only NAC031 clade and class III contains only NAC009 clade. Class II contains NAC033 and NAC043 two clade, probably because NAC033 had only two copies of AtNAC033 and DcNAC033. Class IV as the largest group, including NAC002, NAC018 and NAC029, with a total of 30 members (Fig. 2A). Interestingly, both DcNAC043, DcNAC031 and DcNAC009 appeared to be more closely related to Phalaenopsis equestris, probably because they were both orchids.
Gene structure and conserved Motif analyses
To compare the exon/intron structures of AtNACs and DcNACs genes, the coding sequences with their corresponding genomic sequences were aligned. 74 genes (43 AtNACs and 31 DcNACs) were mapped using the GSDS 2.0 software package. The results showed that all AtNACs used for analysis contained no introns, while almost all DcNACs contained at least one intron (except DcNAC002, DcNAC033, DcNAC070 and DcNAC074) (Fig. 1B). Among DcNACs, there were 6 copies of NAC100 and 3 copies of NAC098 in class I, except for NAC100-1 and NAC098-1, the rest of the copies contain an intron and 2 CDS regions. Similarly, two copies of DcNAC43 in class III, two copies of DcNAC009 in class IV, and three copies of DcNAC068 in class IV have similar gene structures (2 introns, 3 CDS regions; 1 intron, 2 CDS regions; 2 introns, 2 CDS regions, respectively) (Fig. 1B). It suggested that the gene structures of different copies of these genes were relatively conserved in the evolutionary process, and their functions may also be similar. In contrast, there seemed to be no regularity in the gene structure between different copies of other DcNAC genes, maybe there was a big difference in functionality.
To further understand the structural diversity of DcNAC proteins, we identified 15 putative motifs (motif 1–15) in the proteins using the MEME/MAST program (Fig. 2B, C). As shown in Fig. 2, all NAC protein sequences involved in the analysis contained motif 1, motif 2, motif 3, motif 4 and motif 5. However, each subgroup had its own unique motif, for example, in class I, the specific motif 14 of individual members, the specific motif 9 and motif 12 of class II, the specific motif 7, 11, 13 of class III, and the specific motif 6 of group IV. The finding of similar gene structures and conserved motifs within the same subfamily further supports the accuracy of the phylogenetic tree. The other side of the shield, the motif differences between different subfamilies also indicate functional diversity of the NAC gene family in D. candidum.
Responsive elements in DcNAC promoters
To further investigate the potential regulatory mechanisms of DcNAC cduring the abiotic stress response, a sequence 2 kb upstream from the translation initiation site of the DcNAC gene was submitted to PlantCARE to detect cis-elements. All genes involved in the analysis of promoter elements (DcNAC008-5, DcNAC009-1, DcNAC009-2, DcNAC018, DcNAC029, DcNAC031, DcNAC033, DcNAC043-1, DcNAC043-2, DcNAC068-1, DnNAC068-2, DcNAC068-3, DcNAC070, DcNAC074, DcNAC078, DcNAC094, DcNAC098-1, DcNAC098-3, DcNAC100-2, DcNAC100-3, DcNAC100-4, DcNAC100-5, DcNAC100-6) contain CAAT-box and TATA-box elements, which proved that all the genes can be transcribed normally and thus participate in plant growth and development.
Abiotic stress response elements and phytohormone response elements were analyzed. DcNAC009-1 and DcNAC043-2 contained 20 and 18 response elements (Hormones and Abiotic Stresses), respectively, which were the two genes with the highest element content (Fig. 3). The promoters of the two genes, DcNAC009-2 and DcNAC078, had the least stress response elements, and both had only five. Among them, MYB and MYC elements (response to drought and ABA induction) were the most numerous (49 and 66, respectively), and TCA-element (response to salicylic acid induction) was the least numerous (only 4). Among the promoters of 23 genes, drought stress response elements appeared most frequently, accounting for 48%.
Ka/Ks ratio calculation
The genomic information obtained from National Center for Biotechnology Information (NCBI) does not include the chromosomal location of the genes. Therefore, chromosomal location and collinearity analysis were not performed in this study. To explore the selective pressure on DcNAC genes, the non-synonymous/synonymous mutation ratio (Ka/Ks) was calculated for the genes involved in the evolutionary analysis, and values of Ka/Ks > 1, = 1 and < 1 indicate positive selection, neutral selection and purifying selection, respectively (Nekrutenko et al. 2002).
After removing gene pairs with no numerical results, the remaining genes were counted. As shown in Table 2, the Ka/Ks ratio for all DcNAC genes were < 1, ranging from 0.0807 (AtNAC087-2/DcNAC100-6) to 0.4293 (AtNAC003/DnNAC100-4), indicating the genes were negative selection during evolution and functionally conserved, which reduced the rate of change in aa profile. A comparation of the Ka/Ks ratio of DcNAC genes among the class I, II, III, IV showed that the average Ka/Ks ratio was higher in the class I (0.2040) than in the class III, II and IV (0.1534, 0.1376, and 0.1327, respectively), suggesting that DcNAC genes in the class I experienced higher selection pressure during the evolutionary history of D. candidum. However, the overall results showed that most DcNAC genes were slowly evolving in D. candidum.
Protein–protein networks analysis of DcNAC043s
The previous analysis found that NAC043 gene was relatively conservative in the evolutionary process. Therefore, based on D candidum, with Arabidopsis and Populus protein database were refered, the protein interacting with DcNAC043s were predicted by STRING software. As the Fig. 4 showed that the two copies of DcNAC043 were consistent with Arabidopsis, and both predicted to interact with DcMYB46 (Zhong et al. 2007a, b). Except Cellulose synthase A catalytic subunit 8 (AtCESA8), Homeobox protein knotted-1-like 3 (DcHOS66) and DcNAC031, the proteins appearing in the interaction network belonged to R2R3-MYB family. These proteins have been reported to participate in the growth and development of plant secondary cell wall (Fang et al. 2020; Im et al. 2021; Ye et al. 2021). Interestingly, although the proteins interacting with PtNAC043 did not belong to the R2R3-MYB family, they have also been reported to directly or indirectly regulate the growth and development of plant secondary cell wall (Xing et al. 2008; Min et al. 2020).
DcNAC043 responds to drought stress and regulates lignin synthesis
Previous studies showed that NAC043 responded to drought stress in ryegrass (Cheng et al. 2022). Based on the previous analysis, NAC043 had proved that its function was conservative. Therefore, we treated seedling of D. candidum under drought stress, and detected the expression of DcNAC043s during drought stress by qRT-PCR. The results showed that the change of DcNAC043s expression in leaves was related to the drought degree suffered by plants (Fig. 5B). With the increase of drought stress time, the expression of DcNAC043-1 and DcNAC043-1 in leaves also increased significantly (PDcNAC043-1 = 0.024, 0.017, 0.009 and PDcNAC043-2 = 0.03, 0.021, 0.017) (Fig. 5B). Consistent with the change trend of DcNAC043s, the expression of DcMYB46 also increased with the increase of drought time, which again proved that DcMYB46 was involved in regulating drought stress.
Morphology and staining of Dendrobium and expression of genes related to drought stress response. A a, b, c and d-I Morphology of dendrobium plants under drought stress for 0, 3, 6 and 9 days respectively. a-II, III and IV, lignin staining results of dendrobium plants under drought stress for 0 d, b-II, III and IV, lignin staining results of dendrobium plants under drought stress for 3 d, c-II, III and IV, lignin staining results of dendrobium plants under drought stress for 6 d, d-II, III and IV, lignin staining results of dendrobium plants under drought stress for 9 d. B Expression analysis of DcNAC043 and DcMYB46 under drought stress at 0,3,6 and 9 d. CK: Drought stress 0 day, as control, Ds_3d: Drought stress 3 days, Ds_6d: Drought stress 6 days, Ds_9d: Drought stress 9 days. ** denotes a significant difference at P < 0.01, as determined using Student’s t test, Bars represent SD of the average, mean ± SD
In Citrus grandis, CgNAC043 was found to regulate lignin synthesis, thereby affecting the development of secondary cell wall (Li et al. 2022). Therefore, we used the lignin specific stay HCl—phloroglucinol to dye it for analysis. We observed that the longer the drought stress time was, the more serious the lignification of the plant was, which indicates that the more lignin was accumulated (Fig. 5A). It was indirectly proved that DcNAC043s also participates in the synthesis of lignin in Dendrobium.
Discussion
NAC-TF has been found and classified in many plants, but there is no detailed evidence about the NAC-TFs family of Dendrobium. In this study, although the published full genome information of Dendrobium showed that there were 75 annotated NAC genes, which was similar to the number of NAC genes identified in grapes (74) and sugarcane (88) (Wang et al. 2013; Manimekalai et al. 2017). However, by comparing BLASTP with 117 AtNAC proteins and based on conservation analysis, this study ultimately obtained 33 DcNAC proteins. It was far less than potato (110) (Singh et al. 2013), soybean (101) (Shang et al. 2016), cacao tree (102) (Shen et al. 2019), rice (151) (Nuruzzaman et al. 2010) and poplar (163) (Hu et al. 2010). The difference in the number of NAC members among these species may be caused by genome wide polyploidy events during the evolution and differentiation of each species.
Interestingly, analysis of phylogenetic tree, gene structure, conservative motif and Ka/Ks value showed that NAC was unevenly distributed in Arabidopsis and Dendrobium, indicating that NAC gene family existed before the differentiation of the two species. It was different from the ancient and inactive evolutionary nature of NAC gene family in other species (Hu et al. 2010; Nuruzzaman et al. 2010; Wang et al. 2013; Singh et al. 2013, 2016; Shang et al. 2016; Manimekalai et al. 2017; Shen et al. 2019). It may be that Dendrobium and Arabidopsis were exposed to different environments during evolution, so gene differentiation occurred, and the number of NAC genes in their subfamilies was also different.
However, among all DcNAC genes, DcNAC009s and DcNAC043s are different from other DcNAC genes and are relatively conserved in terms of gene structure and physicochemical properties. Therefore, to further prove the evolutionary conservation of NAC009 and NAC043, we analyzed the gene structures of NAC009 in 10 different species and NAC043 in 15 different species, respectively. The results showed that NAC009 and NAC043 in all species (except Arabidopsis) have similar gene structures; what is more, closely related species have very similar gene lengths (Fig. 6). Meanwhile, motif analysis of phylogenetic tree and gene structure based on multiple species shows that NAC043 genes were relatively conservative, which is similar to the existing research results on the gene structure of NAC043 (Sun et al. 2023). All the NAC043 proteins contained not only motif 1, motif 2, motif 3, motif 4 and motif 5 that all the NAC proteins had, but also had their own unique motif 9 (SSITADTKTQMFHSSEGALDQILQYMGRSCKZESEAISNS) and motif 12 (MSISVNGQSQV) (Fig. 2B, C). Therefore, we infered that genes in this subfamily shared a common parent and may have similar functions.
Phylogenetic relationship and gene structure of DcNAC009 and DcNAC043 in different species. The amino acid sequences of AtNACs and DcNACs are aligned using ClustalW2. The phylogenetic tree is constructed using MEGA 7.0 with the neighbor-joining method (1000 bootstrap replicates) and displayed using FigTree v1.4.0. Exons (CDS) and introns are shown as orange wedges and black lines, respectively. A Gene structure differences of NAC009 among different species, B Gene structure differences of NAC043 among different species
In previous studies, NAC043 was shown to Ryegrass can respond to drought stress (Cheng et al. 2022). Based on this, we selected mature Dendrobium plants for drought stress treatment, and extracted plant leaves for qRT-PCR detection. The results showed that the expression of DcNAC043-1 and DcNAC043-2 in plants were up-regulated after drought stress induction, and the up-regulated trend was related to the duration of drought stress. The staining results of HCl-phloglucinol not only proved that DcNAC043s in Dendrobium responded to drought stress by regulating lignin content, but also further proved that DcNAC043s, like CgNAC043 gene in other species, had the function of regulating plant secondary cell wall growth (Li et al. 2022; Wu et al. 2023). Moreover, they were similar to the protein interaction network of AtNAC043 and CgNAC043. In the network, DcNAC043-1 and DcNAC043-2 also act as the master switch that activates the expression of many lignin biosynthetic genes (Zhong et al. 2006; Zhong et al. 2007a, b; McCarthy et al. 2009; Zhou et al. 2009; Hussey et al. 2011; Li et al. 2022) (Fig. 4). It was proved that DcNAC043s were involved in the synthesis of lignin again. However, whether DcNAC043-1 and DcNAC043-2 have functional redundancy needs further experimental verification.
Conclusion
Our research shows that there were significant differences in the evolution of NAC genes between Dendrobium and the model plant Arabidopsis. Therefore, the functions of many members of the DcNAC family need to refer to other species for further research. But DcNAC043-1 and DcNAC043-2, which had relatively conservative function, has proved its contribution to the drought stress of Dendrobium through our verification. And we believe that DcNAC043 plays a very important role in regulating the lignin synthesis of Dendrobium. However, further research is needed on their upstream and downstream regulatory relationships and interaction networks in lignin synthesis pathways, as well as their molecular mechanisms in response to drought stress.
Availability of data and materials
The datasets analyzed during the current study could be queried and obtained in the National Biotechnology Information Center repository (NCBI https://www.ncbi.nlm.nih.gov/genome/?条款=). The gene BioProject numbers and relevant references of 38 species including D. candidum are presented in Table S1.
References
Aida M, Ishida T, Fukaki H, Fujisawa H, Tasaka M (1997) Genes involved in organ separation in Arabidopsis: an analysis of the cup-shaped cotyledon mutant. Plant Cell 9(6):841–857. https://doi.org/10.1105/tpc.9.6.841
Aldhanhani AR, Ahmed ZF, Tzortzakis, Nikolaos SZ (2022) Maturity stage at harvest influences antioxidant phytochemicals and antibacterial activity of jujube fruit (Ziziphus mauritiana Lamk. and Ziziphus spina-christi L.). Ann. Agric. Sci 67:2. https://doi.org/10.1016/J.AOAS.2022.12.003
Bausher MG, Singh ND, Lee SB, Jansen RK, Daniell H (2006) The complete chloroplast genome sequence of Citrus sinensis (L.) Osbeck var “Ridge Pineapple”: organization and phylogenetic relationships to other angiosperms. BMC Plant Biol 6:21. https://doi.org/10.1186/1471-2229-6-21
Beinecke FA, Grundmann L, Wiedmann DR, Schmidt FJ, Caesar AS, Zimmermann M, Lahme M, Twyman RM, Prüfer D, Noll GA (2018) The FT/FD-dependent initiation of flowering under long-day conditions in the day-neutral species Nicotiana tabacum originates from the facultative short-day ancestor Nicotiana tomentosiformis. Plant J 96(2):329–342. https://doi.org/10.1111/tpj.14033
Bolger A, Scossa F, Bolger ME, Lanz C, Maumus F, Tohge T, Quesneville H, Alseekh S, Sørensen I, Lichtenstein G, Fich EA, Conte M, Keller H, Schneeberger K, Schwacke R, Ofner I, Vrebalov J, Xu Y, Osorio S, Aflitos SA, Schijlen E, Jiménez-Goméz JM, Ryngajllo M, Kimura S, Kumar R, Koenig D, Headland LR, Maloof JN, Sinha N, van Ham RC, Lankhorst RK, Mao L, Vogel A, Arsova B, Panstruga R, Fei Z, Rose JK, Zamir D, Carrari F, Giovannoni JJ, Weigel D, Usadel B, Fernie AR (2014) The genome of the stress-tolerant wild tomato species Solanum pennellii. Nat Genet 46:1034–1038. https://doi.org/10.1038/ng.3046
Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL (2009) BLAST+: architecture and applications. MC Bioinform 10:421. https://doi.org/10.1186/1471-2105-10-421
Caruso M, Merelo P, Distefano G, La Malfa S, Lo Piero AR, Tadeo FR, Talon M, Gentile A (2012) Comparative transcriptome analysis of stylar canal cells identifies novel candidate genes implicated in the self-incompatibility response of Citrus clementina. BMC Plant Biol 12:20. https://doi.org/10.1186/1471-2229-12-20
Chen SC, Ogata A (2015) MixtureTree annotator: a program for automatic colorization and visual annotation of MixtureTree. PLoS One 10(3):e0118893. https://doi.org/10.1371/journal.pone.0118893
Chen D, Chai S, McIntyre CL, Xue GP (2018) Overexpression of a predominantly root-expressed NAC transcription factor in wheat roots enhances root length, biomass and drought tolerance. Plant Cell Rep 37:225–237. https://doi.org/10.1007/s00299-017-2224-y
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, Xia R (2020) TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant 13(8):1194–1202. https://doi.org/10.1016/j.molp.2020.06.009
Cheng SB, Yang XZ, Zou L, Wu DD, Lu JL, Cheng YR, Wang Y, Zeng J, Kang HY, Sha LN, Fan X, Ma X, Zhang XQ, Zhou YH, Zhang HQ (2022) Comparative physiological and root transcriptome analysis of two annual ryegrass cultivars under drought stress. J Plant Physiol 277:153807. https://doi.org/10.1016/j.jplph.2022.153807
Cruz LM, Trefflich S, Weiss VA, Castro M (2017) Protein function prediction. Methods Mol Biol 1654:55–75. https://doi.org/10.1007/978-1-4939-7231-95
Daniell H, Wurdack KJ, Kanagaraj A, Lee SB, Saski C, Jansen RK (2008) The complete nucleotide sequence of the cassava (Manihot esculenta) chloroplast genome and the evolution of atpF in Malpighiales: RNA editing and multiple losses of a group II intron. Theor Appl Genet 116(5):723–737. https://doi.org/10.1007/s00122-007-0706-y
Duvaud S, Gabella C, Lisacek F, Stockinger H, Ioannidis V, Durinx C (2021) Expasy, the Swiss Bioinformatics Resource Portal, as designed by its users. Nucleic Acids Res 49(W1):W216–W227. https://doi.org/10.1093/nar/gkab225
Fang Y, Wu H, Zhang T, Yang M, Yin Y, Pan L, Yu X, Zhang X, Hu S, Al-Mssallem IS, Yu J (2012) A complete sequence and transcriptomic analyses of date palm (Phoenix dactylifera L.) mitochondrial genome. PLoS One 7(5):e37164. https://doi.org/10.1371/journal.pone.0037164
Fang S, Shang X, Yao Y, Li W, Guo W (2020) NST- and SND-subgroup NAC proteins coordinately act to regulate secondary cell wall formation in cotton. Plant Sci 301:110657. https://doi.org/10.1016/j.plantsci.2020.110657
Gao S, Wang B, Xie S, Xu X, Zhang J, Pei L, Yu Y, Yang W, Zhang Y (2020) A high-quality reference genome of wild Cannabis sativa. Hortic Res 7(1):73. https://doi.org/10.1038/s41438-020-0295-3
Ge M, Liu Y, Jiang L, Wang Y, Lv Y, Zhou L, Liang S, Bao H, Zhao H (2018) Genome-wide analysis of maize NLP transcription factor family revealed the roles in nitrogen response. Plant Growth Regul 84:95–105. https://doi.org/10.1007/s10725-017-0324-x
Grant EH, Fujino T, Beers EP, Brunner AM (2010) Characterization of NAC domain transcription factors implicated in control of vascular cell differentiation in Arabidopsis and Populus. Planta 232(2):337–352. https://doi.org/10.1007/s00425-010-1181-2
Handa H (2003) The complete nucleotide sequence and RNA editing content of the mitochondrial genome of rapeseed (Brassica napus L.): comparative analysis of the mitochondrial genomes of rapeseed and Arabidopsis thaliana. Nucleic Acids Res 31(20):5907–5916. https://doi.org/10.1093/nar/gkg795
He H, Liang G, Lu S, Wang P, Liu T, Ma Z, Zuo C, Sun X, Chen B, Mao J (2019) Genome-wide identification and expression analysis of GA2ox, GA3ox, and GA20ox are related to gibberellin oxidase genes in grape (Vitis vinifera L.). Genes (Basel) 10(9):680. https://doi.org/10.3390/genes10090680
Hu H, Dai M, Yao J, Xiao B, Li X, Zhang Q, Xiong L (2006) Overexpressing a NAM, ATAF, and CUC (NAC) transcription factor enhances drought resistance and salt tolerance in rice. Proc Natl Acad Sci USA 103(35):12987–12992. https://doi.org/10.1073/pnas.0604882103
Hu R, Qi G, Kong Y, Kong D, Gao Q, Zhou G (2010) Comprehensive analysis of NAC domain transcription factor gene family in Populus trichocarpa. BMC Plant Biol 10:145. https://doi.org/10.1186/1471-2229-10-145
Hubert O, Piral G, Galas C, Baurens FC, Mbéguié-A-Mbéguié D (2014) Changes in ethylene signaling and MADS box gene expression are associated with banana finger drop. Plant Sci 223:99–108. https://doi.org/10.1016/j.plantsci.2014.03.008
Hussey SG, Mizrachi E, Spokevicius AV, Bossinger G, Berger DK, Myburg AA (2011) SND2, a NAC transcription factor gene, regulates genes involved in secondary cell wall development in Arabidopsis fibres and increases fibre cell area in Eucalyptus. BMC Plant Biol 11:173. https://doi.org/10.1186/1471-2229-11-173
Im JH, Ko JH, Kim WC, Crain B, Keathley D, Han KH (2021) Mitogen-activated protein kinase 6 negatively regulates secondary wall biosynthesis by modulating MYB46 protein stability in Arabidopsis thaliana. PLoS Genet 17(4):e1009510. https://doi.org/10.1371/journal.pgen.1009510
Ishida JK, Yoshida S, Ito M, Namba S, Shirasu K (2011) Agrobacterium rhizogenes-mediated transformation of the parasitic plant Phtheirospermum japonicum. PLoS One 6(10):e25802. https://doi.org/10.1371/journal.pone.0025802
Jeong JS, Kim YS, Baek KH, Jung H, Ha SH, Do Choi Y, Kim M, Reuzeau C, Kim JK (2010) Root-specific expression of OsNAC10 improves drought tolerance and grain yield in rice under field drought conditions. Plant Physiol 153(1):185–197. https://doi.org/10.1104/pp.110.154773
Jia D, Jiang Q, van Nocker S, Gong X, Ma F (2019) An apple (Malus domestica) NAC transcription factor enhances drought tolerance in transgenic apple plants. Plant Physiol Biochem 139:504–512. https://doi.org/10.1016/j.plaphy.2019.04.011
Jing Z, Ruan X, Wang R, Yang Y (2018) Genetic diversity and relationships between and within persimmon (Diospyros L.) wild species and cultivated varieties by SRAP markers. Plant Syst Evol. https://doi.org/10.1007/s00606-013-0810-1
Kawahara Y, de la Bastide M, Hamilton JP, Kanamori H, McCombie WR, Ouyang S, Schwartz DC, Tanaka T, Wu J, Zhou S, Childs KL, Davidson RM, Lin H, Quesada-Ocampo L, Vaillancourt B, Sakai H, Lee SS, Kim J, Numa H, Itoh T, Buell CR, Matsumoto T (2013) Improvement of the Oryza sativa Nipponbare reference genome using next generation sequence and optical map data. Rice (NY) 6(1):4. https://doi.org/10.1186/1939-8433-6-4
Khalil HA, ElAnsary DO, Ahmed ZF (2022) Mitigation of Salinity Stress on Pomegranate (Punica granatum L. cv. Wonderful) Plant Using Salicylic Acid Foliar Spray. Hortic. https://doi.org/10.3390/HORTICULTURAE8050375
Kim YS, Kim SG, Park JE, Park HY, Lim MH, Chua NH, Park CM (2006) A membrane-bound NAC transcription factor regulates cell division in Arabidopsis. Plant Cell 18(11):3132–3144. https://doi.org/10.1105/tpc.106.043018
Kumar S, Stecher G, Tamura K (2016) MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol Biol Evol 33(7):1870–1874. https://doi.org/10.1093/molbev/msw054
Lantican DV, Strickler SR, Canama AO, Gardoce RR, Mueller LA, Galvez HF (2019) De Novo Genome Sequence Assembly of Dwarf Coconut (Cocos nucifera L. 'Catigan Green Dwarf') Provides Insights into Genomic Variation Between Coconut Types and Related Palm Species. G3 (Bethesda) 9(8): 2377–2393. https://doi.org/10.1534/g3.119.400215
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23(21):2947–2948. https://doi.org/10.1093/bioinformatics/btm404
Lee S, Seo PJ, Lee HJ, Park CM (2012) A NAC transcription factor NTL4 promotes reactive oxygen species production during drought-induced leaf senescence in Arabidopsis. Plant J 70(5):831–844. https://doi.org/10.1111/j.1365-313X.2012.04932.x
Li Q, Zhang N, Zhang L, Ma H (2015) Differential evolution of members of the rhomboid gene family with conservative and divergent patterns. New Phytol 206(1):368–380. https://doi.org/10.1111/nph.13174
Li H, Liang J, Chen H, Ding G, Ma B, He N (2016) Evolutionary and functional analysis of mulberry type III polyketide synthases. BMC Genomics 17:540. https://doi.org/10.1186/s12864-016-2843-7
Li H, Guan H, Zhuo Q, Wang Z, Li S, Si J, Zhang B, Feng B, Kong LA, Wang F, Wang Z, Zhang L (2020) Genome-wide characterization of the abscisic acid-, stress- and ripening-induced (ASR) gene family in wheat (Triticum aestivum L.). Biol Res 53(1):23. https://doi.org/10.1186/s40659-020-00291-6
Li X, Wang N, She W, Guo Z, Pan H, Yu Y, Ye J, Pan D, Pan T (2022) Identification and Functional Analysis of the CgNAC043 Gene Involved in Lignin Synthesis from Citrusgrandis “San Hong.” Plants (Basel) 11(3):403. https://doi.org/10.3390/plants11030403
Lin CS, Chen JJW, Chiu CC, Hsiao HCW, Yang CJ, Jin XH, Leebens-Mack J, de Pamphilis CW, Huang YT, Yang LH, Chang WJ, Kui L, Wong GK, Hu JM, Wang W, Shih MC (2017) Concomitant loss of NDH complex-related genes within chloroplast and nuclear genomes in some orchids. Plant J 90(5):994–1006. https://doi.org/10.1111/tpj.13525
Ling HQ, Ma B, Shi X, Liu H, Dong L, Sun H, Cao Y, Gao Q, Zheng S, Li Y, Yu Y, Du H, Qi M, Li Y, Lu H, Yu H, Cui Y, Wang N, Chen C, Wu H, Zhao Y, Zhang J, Li Y, Zhou W, Zhang B, Hu W, van Eijk MJT, Tang J, Witsenboer HMA, Zhao S, Li Z, Zhang A, Wang D, Liang C (2018) Genome sequence of the progenitor of wheat A subgenome Triticum urartu. Nature 557(7705):424–428. https://doi.org/10.1038/s41586-018-0108-0
Liu M, Chang W, Fan Y, Sun W, Qu C, Zhang K, Liu L, Xu X, Tang Z, Li J, Lu K (2018) Genome-wide identification and characterization of NODULE-INCEPTION-like protein (NLP) family genes in Brassica napus. Int J Mol Sci 19(8):2270. https://doi.org/10.3390/ijms19082270
Lowe TM, Eddy SR (1997) tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res 25(5):955–964. https://doi.org/10.1093/nar/25.5.955
Lu J, Pan C, Fan W, Liu W, Zhao H, Li D, Wang S, Hu L, He B, Qian K, Qin R, Ruan J, Lin Q, Lü S, Cui P (2022) A Chromosome-level Genome Assembly of Wild Castor Provides New Insights into its Adaptive Evolution in Tropical Desert. Genom Proteom Bioinform 20(1):42–59. https://doi.org/10.1016/j.gpb.2021.04.003
Manimekalai R, Jini N, Gokul M, Selvi AM, Gomathi R, Bakshi R (2017) Genome wide analysis of NAC gene family ‘sequences’ in sugarcane and its comparative phylogenetic relationship with rice, sorghum, maize and Arabidopsis for prediction of stress associated NAC genes. Agri Gene 3:2352–2151. https://doi.org/10.1016/j.aggene.2016.10.003
Mao X, Zhang H, Qian X, Li A, Zhao G, Jing R (2012) TaNAC2, a NAC-type wheat transcription factor conferring enhanced multiple abiotic stress tolerances in Arabidopsis. J Exp Bot 63(8):2933–2946. https://doi.org/10.1093/jxb/err462
Mariac C, Scarcelli N, Pouzadou J, Barnaud A, Billot C, Faye A, Kougbeadjo A, Maillol V, Martin G, Sabot F, Santoni S, Vigouroux Y, Couvreur TL (2014) Cost-effective enrichment hybridization capture of chloroplast genomes at deep multiplexing levels for population genetics and phylogeography studies. Mol Ecol Resour 14(6):1103–1113. https://doi.org/10.1111/1755-0998.12258
McCarthy RL, Zhong R, Ye ZH (2009) MYB83 is a direct target of SND1 and acts redundantly with MYB46 in the regulation of secondary cell wall biosynthesis in Arabidopsis. Plant Cell Physiol 50(11):1950–1964. https://doi.org/10.1093/pcp/pcp139
Miao L, Wei C, Mengna Y, Yonghai F, Guoxia S, Yuanfang X, Yue N, Xumei L, Hong Z, Lishi D, Zhanglin T, Kai Z, Liezhao L, Cunmin Q, Jiana L, Kun L (2021) Overexpression of DEFECTIVE IN ANTHER DEHISCENCE 1 increases rapeseed silique length through crosstalk between JA and auxin signaling. Ind Crops Prod 168:113576. https://doi.org/10.1016/j.indcrop.2021.113576
Min K, Yi G, Lee JG, Kim HS, Hong Y, Choi JH, Lim S, Lee EJ (2020) Comparative transcriptome and metabolome analyses of two strawberry cultivars with different storability. PLoS One 15(12):e0242556. https://doi.org/10.1371/journal.pone.0242556
Ming R, VanBuren R, Wai CM, Tang H, Schatz MC, Bowers JE, Lyons E, Wang ML, Chen J, Biggers E, Zhang J, Huang L, Zhang L, Miao W, Zhang J, Ye Z, Miao C, Lin Z, Wang H, Zhou H, Yim WC, Priest HD, Zheng C, Woodhouse M, Edger PP, Guyot R, Guo HB, Guo H, Zheng G, Singh R, Sharma A, Min X, Zheng Y, Lee H, Gurtowski J, Sedlazeck FJ, Harkess A, McKain MR, Liao Z, Fang J, Liu J, Zhang X, Zhang Q, Hu W, Qin Y, Wang K, Chen LY, Shirley N, Lin YR, Liu LY, Hernandez AG, Wright CL, Bulone V, Tuskan GA, Heath K, Zee F, Moore PH, Sunkar R, Leebens-Mack JH, Mockler T, Bennetzen JL, Freeling M, Sankoff D, Paterson AH, Zhu X, Yang X, Smith JA, Cushman JC, Paull RE, Yu Q (2015) The pineapple genome and the evolution of CAM photosynthesis. Nat Genet 47(12):1435–1442. https://doi.org/10.1038/ng.3435
Mistry J, Chuguransky S, Williams L, Qureshi M, Salazar GA, Sonnhammer ELL, Tosatto SCE, Paladin L, Raj S, Richardson LJ, Finn RD, Bateman A (2021) Pfam: The protein families database in 2021. Nucleic Acids Res 49(D1):D412–D419. https://doi.org/10.1093/nar/gkaa913
Nakashima K, Takasaki H, Mizoi J, Shinozaki K, Yamaguchi-Shinozaki K (2012) NAC transcription factors in plant abiotic stress responses. Biochim Biophys Acta 1819(2):97–103. https://doi.org/10.1016/j.bbagrm.2011.10.005
Negi S, Tak H, Ganapathi TR. A banana NAC transcription factor (MusaSNAC1) impart drought tolerance by modulating stomatal closure and H2O2 content (2018) A banana NAC transcription factor (MusaSNAC1) impart drought tolerance by modulating stomatal closure and H2O2 content. Plant Mol Biol 96(4-5): 457-471. https://doi.org/10.1007/s11103-018-0710-4
Nekrutenko A, Makova KD, Li WH (2002) The K(A)/K(S) ratio test for assessing the protein-coding potential of genomic regions: an empirical and simulation study. Genome Res 12(1):198–202. https://doi.org/10.1101/gr.200901
Nuruzzaman M, Manimekalai R, Sharoni AM, Satoh K, Kondoh H, Ooka H, Kikuchi S (2010) Genome-wide analysis of NAC transcription factor family in rice. Gene 465(1–2):30–44. https://doi.org/10.1016/j.gene.2010.06.008
Nystrom SL, McKay DJ (2021) Memes: A motif analysis environment in R using tools from the MEME Suite. PLoS Comput Biol 17(9):e1008991. https://doi.org/10.1371/journal.pcbi.1008991
Olsen AN, Ernst HA, Leggio LL, Skriver K (2005) NAC transcription factors: structurally distinct, functionally diverse. Trends Plant Sci 10(2):79–87. https://doi.org/10.1016/j.tplants.2004.12.010
Pan R, Wang Y, An F, Yao Y, Xue J, Zhu W, Luo X, Lai H, Chen S (2023) Genome-wide identification and characterization of 14-3-3 gene family related to negative regulation of starch accumulation in storage root of Manihot esculenta. Front Plant Sci 14:1184903. https://doi.org/10.3389/fpls.2023.1184903
Patten AM, Cardenas CL, Cochrane FC, Laskar DD, Bedgar DL, Davin LB, Lewis NG (2005) Reassessment of effects on lignification and vascular development in the irx4 Arabidopsis mutant. Phytochemistry 66(17):2092–2107. https://doi.org/10.1016/j.phytochem.2004.12.016
Peng S, Yang G, Liu C, Yu Z, Zhai M (2017) The complete chloroplast genome of the Juglans regia (Juglandales: Julandaceae). Mitochondrial DNA A DNA Mapp Seq Anal 28(3):407–408. https://doi.org/10.3109/19401736.2015.1127367
Potter SC, Luciani A, Eddy SR, Park Y, Lopez R, Finn RD (2018) HMMER web server: 2018 update. Nucleic Acids Res 46(W1):W200–W204. https://doi.org/10.1093/nar/gky448
Qu G, Bao Y, Liao Y, Liu C, Zi H, Bai M, Liu Y, Tu D, Wang L, Chen S, Zhou G, Can M (2022) Draft genomes assembly and annotation of Carex parvula and Carex kokanica reveals stress-specific genes. Sci Rep 12(1):4970. https://doi.org/10.1038/s41598-022-08783-z
Ramos AM, Usié A, Barbosa P, Barros PM, Capote T, Chaves I, Simões F, Abreu I, Carrasquinho I, Faro C, Guimarães JB, Mendonça D, Nóbrega F, Rodrigues L, Saibo NJM, Varela MC, Egas C, Matos J, Miguel CM, Oliveira MM, Ricardo CP, Gonçalves S (2018) The draft genome sequence of cork oak. Sci Data 5:180069. https://doi.org/10.1038/sdata.2018.69
Sakamoto S, Takata N, Oshima Y, Yoshida K, Taniguchi T, Mitsuda N (2016) Wood reinforcement of poplar by rice NAC transcription factor. Sci Rep 6:19925. https://doi.org/10.1038/srep19925
Sakuraba Y, Kim YS, Han SH, Lee BD, Paek NC (2015) The Arabidopsis transcription factor NAC016 promotes drought stress responses by repressing AREB1 transcription through a trifurcate feed-forward regulatory loop involving NAP. Plant Cell 27(6):1771–1787. https://doi.org/10.1105/tpc.15.00222
Shang H, Wang Z, Zou C, Zhang Z, Li W, Li J, Shi Y, Gong W, Chen T, Liu A, Gong J, Ge Q, Yuan Y (2016) Comprehensive analysis of NAC transcription factors in diploid Gossypium: sequence conservation and expression analysis uncover their roles during fiber development. Sci China Life Sci 59(2):142–153. https://doi.org/10.1007/s11427-016-5001-1
Shao H, Wang H, Tang X (2015) NAC transcription factors in plant multiple abiotic stress responses: progress and prospects. Front Plant Sci 6:902. https://doi.org/10.3389/fpls.2015.00902
Shen S, Zhang Q, Shi Y, Sun Z, Zhang Q, Hou S, Wu R, Jiang L, Zhao X, Guo Y (2019) Genome-Wide Analysis of the NAC Domain Transcription Factor Gene Family in Theobroma cacao. Genes 11(1):35. https://doi.org/10.3390/genes11010035
Shen LY, Luo H, Wang XL, Wang XM, Qiu XJ, Liu H, Zhou SS, Jia KH, Nie S, Bao YT, Zhang RG, Yun QZ, Chai YH, Lu JY, Li Y, Zhao SW, Mao JF, Jia SG, Mao YM (2021) Chromosome-Scale Genome Assembly for Chinese Sour Jujube and Insights Into Its Genome Evolution and Domestication Signature. Front Plant Sci 12:773090. https://doi.org/10.3389/fpls.2021.773090
Shirasawa K, Isuzugawa K, Ikenaga M, Saito Y, Yamamoto T, Hirakawa H, Isobe S (2017) The genome sequence of sweet cherry (Prunus avium) for use in genomics-assisted breeding. DNA Res 24(5):499–508. https://doi.org/10.1093/dnares/dsx020
Singh AK, Sharma V, Pal AK, Acharya V, Ahuja PS (2013) Genome-wide organization and expression profiling of the NAC transcription factor family in potato (Solanum tuberosum L.). DNA Res 20(4):403–423. https://doi.org/10.1093/dnares/dst019
Singh S, Grover A, Nasim M (2016) Biofuel Potential of Plants Transformed Genetically with NAC Family Genes. Front Plant Sci 7:22. https://doi.org/10.3389/fpls.2016.00022
Sloan DB, Wu Z, Sharbrough J (2018) Correction of Persistent Errors in Arabidopsis Reference Mitochondrial Genomes. Plant Cell 30(3):525–527. https://doi.org/10.1105/tpc.18.00024
Soderlund C, Descour A, Kudrna D, Bomhoff M, Boyd L, Currie J, Angelova A, Collura K, Wissotski M, Ashley E, Morrow D, Fernandes J, Walbot V, Yu Y (2009) Sequencing, mapping, and analysis of 27,455 maize full-length cDNAs. PLoS Genet 5(11):e1000740. https://doi.org/10.1371/journal.pgen.1000740
Sork VL, Fitz-Gibbon ST, Puiu D, Crepeau M, Gugger PF, Sherman R, Stevens K, Langley CH, Pellegrini M, Salzberg SL (2016) First Draft Assembly and Annotation of the Genome of a California Endemic Oak Quercus lobata Née (Fagaceae). G3 (Bethesda) 6(11): 3485–3495. https://doi.org/10.1534/g3.116.030411
Souer E, van Houwelingen A, Kloos D, Mol J, Koes R (1996) The no apical meristem gene of Petunia is required for pattern formation in embryos and flowers and is expressed at meristem and primordia boundaries. Cell 85(2):159–170. https://doi.org/10.1016/s0092-8674(00)81093-4
Sun H, Ren M, Zhang J (2022a) Genome-wide identification and expression analysis of fibrillin (FBN) gene family in tomato (Solanum lycopersicum L.). PeerJ 10:e13414. https://doi.org/10.7717/peerj.13414
Sun Q, Zhang B, Yang C, Wang W, Xiang L, Wang Y, Chan Z (2022b) Jasmonic acid biosynthetic genes TgLOX4 and TgLOX5 are involved in daughter bulb development in tulip (Tulipa gesneriana). Hortic Res 9 uhac006. https://doi.org/10.1093/hr/uhac006 (Advance online publication)
Sun S, Li X, Nie N, Chen Y, Gao S, Zhang H, He S, Liu Q, Zhai H (2023) Sweet potato NAC transcription factor NAC43 negatively regulates plant growth by causing leaf curling and reducing photosynthetic efficiency. Sun S, Li X, Nie N, Chen Y, Gao S, Zhang H, He S, Liu Q, Zhai H. 14: 1095977. https://doi.org/10.3389/fpls.2023.1095977
Szklarczyk D, Gable AL, Nastou KC, Lyon D, Kirsch R, Pyysalo S, Doncheva NT, Legeay M, Fang T, Bork P, Jensen LJ, von Mering C (2021) The STRING database in 2021: customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res 49(D1):D605–D612. https://doi.org/10.1093/nar/gkaa1074
Tang C, Yang M, Fang Y, Luo Y, Gao S, Xiao X, An Z, Zhou B, Zhang B, Tan X, Yeang HY, Qin Y, Yang J, Lin Q, Mei H, Montoro P, Long X, Qi J, Hua Y, He Z, Sun M, Li W, Zeng X, Cheng H, Liu Y, Yang J, Tian W, Zhuang N, Zeng R, Li D, He P, Li Z, Zou Z, Li S, Li C, Wang J, Wei D, Lai CQ, Luo W, Yu J, Hu S, Huang H (2016) The rubber tree genome reveals new insights into rubber production and species adaptation. Nat Plants 2(6):16073. https://doi.org/10.1038/nplants.2016.73
van Bakel H, Stout JM, Cote AG, Tallon CM, Sharpe AG, Hughes TR, Page JE (2011) The draft genome and transcriptome of Cannabis sativa. Genome Biol 12(10):R102. https://doi.org/10.1186/gb-2011-12-10-r102
Venturini L, Ferrarini A, Zenoni S, Tornielli GB, Fasoli M, Dal Santo S, Minio A, Buson G, Tononi P, Zago ED, Zamperin G, Bellin D, Pezzotti M, Delledonne M (2013) De novo transcriptome characterization of Vitis vinifera cv. Corvina Unveils varietal diversity. BMC Genom 14:41. https://doi.org/10.1186/1471-2164-14-41
Wang N, Zheng Y, Xin H, Fang L, Li S (2013) Comprehensive analysis of NAC domain transcription factor gene family in Vitis vinifera. Plant Cell Rep 32(1):61–75. https://doi.org/10.1007/s00299-012-1340-y
Wu Q, Ma X, Zhang K, Feng X (2015) Identification of reference genes for tissue-specific gene expression in Panax notoginseng using quantitative real-time PCR. Biotechnol Lett 37(1):197–204. https://doi.org/10.1007/s10529-014-1643-x
Wu J, Kong B, Zhou Q, Sun Q, Sang Y, Zhao Y, Yuan T, Zhang P (2023) SCL14 Inhibits the Functions of the NAC043-MYB61 Signaling Cascade to Reduce the Lignin Content in Autotetraploid Populus hopeiensis. Int J Mol Sci 24(6):5809. https://doi.org/10.3390/ijms24065809
Xing DH, Lai ZB, Zheng ZY, Vinod KM, Fan BF, Chen ZX (2008) Stress- and pathogen-induced Arabidopsis WRKY48 is a transcriptional activator that represses plant basal defense. Mol Plant 1(3):459–470. https://doi.org/10.1093/mp/ssn020
Xiong Y, Xiong Y, He J, Yu Q, Zhao J, Lei X, Dong Z, Yang J, Peng Y, Zhang X, Ma X (2020) The Complete Chloroplast Genome of Two Important Annual Clover Species, Trifolium alexandrinum and T. resupinatum: Genome Structure, Comparative Analyses and Phylogenetic Relationships with Relatives in Leguminosae. Plants (Basel) 9(4):478. https://doi.org/10.3390/plants9040478
Xu Q, Niu SC, Li KL, Zheng PJ, Zhang XJ, Jia Y, Liu Y, Niu YX, Yu LH, Chen DF, Zhang GQ (2022) Chromosome-Scale Assembly of the Dendrobium nobile Genome Provides Insights Into the Molecular Mechanism of the Biosynthesis of the Medicinal Active Ingredient of Dendrobium. Front Genet 13:844622. https://doi.org/10.3389/fgene.2022.844622
Yamaguchi M, Ohtani M, Mitsuda N, Kubo M, Ohme-Takagi M, Fukuda H, Demura T (2010) VND-INTERACTING2, a NAC domain transcription factor, negatively regulates xylem vessel formation in Arabidopsis. Plant Cell 22(4):1249–1263. https://doi.org/10.1105/tpc.108.064048
Ye JH, Lv YQ, Liu SR, Jin J, Wang YF, Wei CL, Zhao SQ (2021) Effects of light intensity and spectral composition on the transcriptome profiles of leaves in shade grown tea plants (Camellia sinensis L.) and regulatory network of flavonoid biosynthesis. Molecules 26(19):5836. https://doi.org/10.3390/molecules26195836
Yeoh KA, Othman A, Meon S, Abdullah F, Ho CL (2013) Sequence analysis and gene expression of putative oil palm chitinase and chitinase-like proteins in response to colonization of Ganoderma boninense and Trichoderma harzianum. Mol Biol Rep 40(1):147–158. https://doi.org/10.1007/s11033-012-2043-8
Yue TQ, Mei D, Zhen QZ, Jiang LD, Tao W (2016) Overexpression of the Medicago falcata NAC transcription factor MfNAC3 enhances cold tolerance in Medicago truncatula. Environ Exp Bot 129:67–76. https://doi.org/10.1016/j.envexpbot
Zeng L, Zhang N, Zhang Q, Endress PK, Huang J, Ma H (2017) Resolution of deep eudicot phylogeny and their temporal diversification using nuclear genes from transcriptomic and genomic datasets. New Phytol 214(3):1338–1354. https://doi.org/10.1111/nph.14503
Zhang Z (2022) KaKs_calculator 3.0: calculating selective pressure on coding and non-coding sequences. Genom Proteom Bioinform 20(3):536–540. https://doi.org/10.1016/j.gpb.2021.12.002
Zhang GQ, Xu Q, Bian C, Tsai WC, Yeh CM, Liu KW, Yoshida K, Zhang LS, Chang SB, Chen F, Shi Y, Su YY, Zhang YQ, Chen LJ, Yin Y, Lin M, Huang H, Deng H, Wang ZW, Zhu SL, Zhao X, Deng C, Niu SC, Huang J, Wang M, Liu GH, Yang HJ, Xiao XJ, Hsiao YY, Wu WL, Chen YY, Mitsuda N, Ohme-Takagi M, Luo YB, Van de Peer Y, Liu ZJ (2016) The Dendrobium catenatum Lindl genome sequence provides insights into polysaccharide synthase, floral development and adaptive evolution. Sci Rep 6:19029. https://doi.org/10.1038/srep19029
Zhang GQ, Liu KW, Li Z, Lohaus R, Hsiao YY, Niu SC, Wang JY, Lin YC, Xu Q, Chen LJ, Yoshida K, Fujiwara S, Wang ZW, Zhang YQ, Mitsuda N, Wang M, Liu GH, Pecoraro L, Huang HX, Xiao XJ, Lin M, Wu XY, Wu WL, Chen YY, Chang SB, Sakamoto S, Ohme-Takagi M, Yagi M, Zeng SJ, Shen CY, Yeh CM, Luo YB, Tsai WC, Van de Peer Y, Liu ZJ (2017) The Apostasia genome and the evolution of orchids. Nature 549(7672):379–383. https://doi.org/10.1038/nature23897
Zhang J, Weng Y, Ye D, You Y, Shi J, Chen J (2021) The complete chloroplast genome sequence of Casuarina equisetifolia. Mitochondrial DNA B Resour 6(10):3046–3048. https://doi.org/10.1080/23802359.2021.1967803
Zheng X, Chen B, Lu G, Han B (2009) Overexpression of a NAC transcription factor enhances rice drought and salt tolerance. Biochem Biophys Res Commun 379(4):985–989. https://doi.org/10.1016/j.bbrc.2008.12.163
Zhong R, Demura T, Ye ZH (2006) SND1, a NAC domain transcription factor, is a key regulator of secondary wall synthesis in fibers of Arabidopsis. Plant Cell 18(11):3158–3170. https://doi.org/10.1105/tpc.106.047399
Zhong R, Richardson EA, Ye ZH (2007a) The MYB46 transcription factor is a direct target of SND1 and regulates secondary wall biosynthesis in Arabidopsis. Plant Cell 19(9):2776–2792. https://doi.org/10.1105/tpc.107.053678
Zhong R, Richardson EA, Ye ZH (2007b) Two NAC domain transcription factors, SND1 and NST1, function redundantly in regulation of secondary wall synthesis in fibers of Arabidopsis. Planta 225(6):1603–1611. https://doi.org/10.1007/s00425-007-0498-y
Zhou J, Lee C, Zhong R, Ye ZH (2009) MYB58 and MYB63 are transcriptional activators of the lignin biosynthetic pathway during secondary cell wall formation in Arabidopsis. Plant Cell 21(1):248–266. https://doi.org/10.1105/tpc.108.063321
Acknowledgements
We thank Ming-jin Huang for providing the Dendrobium candidum. And grateful to Hong Yi from the Instrument and Equipment Sharing Platform of Large Scale Instruments & Equipments of Guizhou University, China.
Funding
This work was supported by the Major Special Project of Science and Technology Program in Guizhou (2017-5411-06); the Science and Technology Support Project of Guizhou (2018-2797); the Guizhou University Cultivation Project (702029201101); Guizhou Science and Technology Plan Project ([2020]4Y071).
Author information
Authors and Affiliations
Contributions
MZ and ML: conceptualization, methodology and funding acquisition. Y-qL, LT, Y-hT: analysis. Y-jZ, L-ySh, YH: validation. ML and TT: writing-original draft preparation. M-jH and M-sZ: discussion, writing-reviewing and editing. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Ethical approval
All experimental studies on plants were complied with relevant institutional, national, and international guidelines and legislation.
Consent to participate
Not applicable.
Consent for publication
Yes.
Additional information
Communicated by Hong-Xia Zhang.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, M., Tian, T., Li, Yq. et al. Genome-wide analysis of the NAC gene family and functional verification of the DcNAC043s in Dendrobium catenatum. Plant Growth Regul 102, 571–588 (2024). https://doi.org/10.1007/s10725-023-01077-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-023-01077-y