Abstract
Global tomato productivity is threatened by biotic and abiotic stressors. To support and guarantee an adequate yield of tomato crops, agricultural practices have been based on the intensive use of fertilisers with negative impacts on the environment. This study presents a simple and effective strategy of functional bioaugmentation, suitable for different varieties, to replace chemical fertilisation. A tailored microbial formula composed by eight indigenous strains (including the genera Delftia, Pseudomonas, Paenarthrobacter, Phyllobacterium, Bacillus, and Acinetobacter) was developed as biofertilizer. Strains were selected from native soil for their plant growth-promoting (PGP) functions, and combined respecting the taxonomic composition of the original PGP heterotrophic community structure. The effect of the bio-fertilisation vs chemical fertilisation was tested in three successive field trials in the company greenhouse, with different tomato varieties (Camone, Oblungo, Cherry). When bio-fertilisation was applied only twice during the Camone’s life cycle, tomato yield was significantly reduced (0.8 vs 2.1 kg per plant, p = 0.0003). However, monthly inoculation during plant growth led to a fruit yield comparable to that obtained with chemical fertilisers (about 1.5 kg per plant for Oblungo, and about 2 kg per plant for Cherry variety, p = 0.9999). Bio-fertilization did not significantly affect plant height; only during the last growing period of the Cherry variety, a significantly higher average plant height (p < 0.0001) was observed with chemical fertiliser. The results indicate that a knowledge-based bacterial formula and monthly inoculation during the plant growth can be a successful bio-fertilisation strategy. These findings may pave the way towards more sustainable tomato production, since farming practices are becoming increasingly crucial, in accordance with Agenda 2030 and the UE “Farm to Fork” strategy.
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The tomato (Solanum lycopersicum L.) represents one of the most important crops due to its great economic and nutritional value (Cordero et al. 2018; Čechura et al. 2021). The global annual tomato production has increased by 300% over the last four decades (Giuliani et al. 2019) and is currently around 180 million tons (Wu et al. 2022). In 2021, Italy produced over 6 million tonnes of processing tomatoes on a surface of over 71 thousand hectares, becoming the second largest producing country in the world after the United States and ahead of China (ANICAV Italy’s National Industrial Association of Vegetable Food Preserves 2021). Although the world scenario is characterized by stable production over the last year, world tomato consumption is steadily increasing (François-Xavier Branthôme 2022).
To remain competitive, Italian tomato growers need to improve their productivity and efficiency. Until now, the strategy to obtain high yield was a very intensive agroecosystem, based on a large exploitation of chemical fertilisers, pesticides, herbicides, and water (Giuliani et al. 2019; Morra et al. 2021). However, this intensive production system is very expensive and energy-consuming, thus it could have high costs in terms of environmental impact and reduction of agricultural yields. Indeed, intensive agriculture contributes to the degradation of soil quality, changes its structure and its water holding capacity and increases surface runoff and loss of nutrients, causing eutrophication of water sources and posing risks to public health (Manfredi et al. 2019; Ye et al. 2020; Kumar et al. 2022). Also, greenhouse farming, frequently used to meet tomato growing demand particularly during colder seasons, should shift its focus from maximising total production to minimising chemical fertilizer use (Wu et al. 2022), using more efficient and low-cost management.
Within the new agricultural technologies, the inoculation of crops with plant growth-promoting bacteria (PGPB) as bio-fertilisers has emerged as a sustainable and environmentally friendly method to improve soil fertility and plant growth, while simultaneously reducing the application of synthetic fertilisers, and maximising the efficiency in the use of resources (Flores-Félix et al. 2021; Kumar et al. 2022). Moreover, the cost of biological fertilisers can be competitive compared to chemical fertilisers (Lobo et al. 2019). Many studies report the application of beneficial microbiomes as a valid tool to combat the deterioration of cultivable topsoil caused by traditional agricultural practices, like overgrazing and tilling, and by the excessive use of chemical fertilisers throughout the years (Cordero et al. 2018; Sah et al. 2021).
Soil quality and rich microbial diversity are crucial to support the growth of high-quality plants. According to Sah et al. 2021, one gram of soil contains about 4000 distinct bacterial genomes. Among these microbes, PGPB exert their growth-promoting effect through different mechanisms that may be direct, such as the improvement of nutrient availability and the phytohormone production, or indirect, as the competition with harmful soil microorganisms, the enhancement of symbiotic relations, and the contribution to the mitigation of abiotic and biotic stresses (Cordero et al. 2018; Oleńska et al. 2020; Kumar et al. 2022).
The beneficial effects of PGPB have been demonstrated in many studies on species of agronomic interest including cucumber (Gamalero et al. 2008), pepper (del Amor and Cuadra-Crespo 2012), pumpkin, corn, broccoli, lettuce (Angulo et al. 2020). However, tomato is one of the most used crops in PGPB application, due to its high economic value and the fast growth which allows the evaluation of the bioaugmentation effects in a relatively short time (Cordero et al. 2018). Nonetheless, a small number of studies have compared the effects of the bio-fertilisation and chemical fertilisers, and even fewer have included a stress factor in the comparison (Cordero et al. 2018; Angulo et al. 2020). Moreover, previous research on different crops concluded that PGPB inoculation must be combined with chemical treatment to achieve the best results on plant growth (Bona et al. 2018; Cordero et al. 2018; Angulo et al. 2020). For tomato species as well, the few studies that compared bio- and chemical fertilisation, described lower fruit yields when the chemicals were completely replaced by PGPB inoculations (Adesemoye et al. 2009; Ye et al. 2020). These results do not encourage tomato growers to apply bio-fertilisation.
There is a great debate on the development of microbial inoculants for agriculture (Kaminsky et al. 2019). Anyway, the success of bio-fertilisation may depend on various experimental factors such as the application of a single strain or consortium (Compant et al. 2019), the choice of introducing autochthonous or allochthonous microorganisms (Ortiz et al. 2015), the composition of the chosen inoculum (Adesemoye et al. 2009), frequency and period of application (He et al. 2019; Riva et al. 2021). The present study aims to propose a customised, knowledge-based, and effective strategy of functional bioaugmentation, that could be advantageous for the farmers. The experimentation described in this paper was carried out in the commercial greenhouse of the “Cooperativa Santa Margherita Terra e Sole'' farm (SW Sardinia, Italy). The goal was to improve the sustainability of the agroecosystem by using indigenous bacteria as bio-fertilisers to replace chemical fertilisation. Three tomato varieties (Camone, Oblungo, Cherry) were tested in three consecutive growing seasons, to compare the chemical (F) fertilization currently in use with the biological (B) fertilisation performed with a microbial formula obtained according to the strategy that will be described.
Material and methods
Soil sampling
Samples from the bulk soil of 10 tomato plants (about 100 g from each plant) were randomly collected in the greenhouse of the farm “Cooperativa Santa Margherita Terra e Sole” (Pula, SW Sardinia—Italy) in September 2018. The soil samples were stored in sterile tubes at RT (room temperature) and transported to the laboratory where they were mixed to create a composite sample for microbiological and chemical analyses. The soil was used both for chemical and microbiological analyses.
Chemical and mineralogical analysis of soil
Mineralogy of the agricultural soil was investigated by powder X-rays diffraction (PXRD). Samples were dried at RT, then were manually ground using an agate mortar, and the analysis were carried out using laboratory θ–2θ equipment (Panalytical, Almelo, Netherlands) with Ni-filtered Cu Kα1 radiation (λ = 1.54060 Å), operating at 40 kV and 40 mA, and an X′celerator detector. Diffraction patterns were analyzed with X’Pert HighScore Plus 2.1 (Panalytical, Almelo, The Netherlands) using the PDF-2 database (International Centre for Diffraction Data) to identify the crystallographic phases in the samples. Chemistry and trace chemistry of the samples were investigated by X-rays fluorescence (XRF) (Potts and Webb 1992). Soil samples were grinded and pressed into solid pellets. Soil analyses were performed with a Rigaku ZSX Primus II WDXRF spectrometer at the Structural Crystallography Centre (CRIST), Florence, Italy. The contents of C, H and N in soil samples were determined by Perkin Elmer 2400 CHN analyser (Hitachi, Tokyo, Japan).
Community-level physiological profile (CLPP)
To proceed with the microbiological study, the microbial community was extracted from the sampled soil. From the composite sample, 50 g were taken in triplicate, and each placed in 500 mL of 0.1% sodium pyrophosphate in sterile bottles and agitated in an orbital shaker at 180 rpm for 90 min at RT. The soil suspension was inoculated into Biolog ECOPlates (Biolog Inc., Hayward, CA, USA), according to Sprocati et al. (2014) to analyse the Community-Level Physiological Profile (CLPP). Briefly, soil suspension was diluted 1:10 and dispensed into the 96 wells of the microplates (100 μL per well) containing 31 different substrates in triplicate. The plates were incubated at 28 °C in the dark and read every 24 h in the microplate reader using a double wavelength (OD590–OD750). Kinetic analysis was performed using average well colour development (AWCD) as a parameter that captures an integral fingerprinting of carbon sources utilisation. AWCD was calculated as the arithmetic mean of the OD values of all the wells in the plate per reading time (Garland 1996). Patterns of substrates utilisation for each replicate were recorded and analysed with the Microlog software (version 5.0). Heatmap was generated with Excel (Windows Office 13). Data are the mean of six individual OD values.
Isolation of Bacteria and identification
The total heterotrophic bacterial community was enumerated by plating serial dilutions (up to 10−5) of the same soil suspension, on different agar media: Tryptic Soy Agar (Laboratorios Conda, Madrid, Spain) as general medium, Mineral Medium (Schmidt and Schlegel 1989) for oligotrophic bacteria selection, and Nitrogen free (NF) Agar (Dobereiner J et al. 1976) to select the nitrogen fixing bacteria (N-fix). The plates were incubated at RT for several weeks until microbial growth was observed, and heterotrophic bacterial strains were isolated. NF plates were incubated in a microaerophilic atmosphere at RT up to 14 days before proceeding with the isolation of the N-fix isolates. Chemicals for media preparation and for analytical assays were purchased from Carlo Erba (Milano, Italy) unless otherwise specified.
For strain identification, single colony 16S r-DNA amplification was performed by polymerase chain reaction (PCR) with the Euroclone Gradient One thermocycler (Euroclone, Milano, Italy) using the universal Eubacteria primers 9bmf (5’- GAG TTT GAT YHT GGC TCA G -3’) and 1512r (5’- ACG GHT ACC TTG TTA CGA CTT -3’) to amplify the 16S rRNA gene (ca. 1500 bp), according to the procedure described by Mühling et al. (2008). Sequence similarity searches were conducted using the BLAST network service of the NCBI database (http://www.ncbi.nlm.nih.gov/BLAST/) to identify the nearest relatives of the partially sequenced 16S rRNA genes. Phylogenetic analysis was performed using MEGA X (Kumar et al. 2018). The phylogenetic tree of the aligned sequences was constructed using the Neighbour-Joining method (Tamura et al. 2004). The sequences generated in this study have been deposited in the GenBank database under accession numbers OP159876 to OP159914. All strains were stored in 20% glycerol at − 80 °C for long-term storage.
PGP assays and bacterial formula assembling
The isolated strains were tested for specific plant growth-promoting activities such as nitrogen fixation (N-fix), phosphate and potassium solubilisation (P-sol and K-sol), indoleacetic acid (IAA) and siderophore production (O-CAS).
Nitrogen fixers were selected or tested on NF solid medium (described in the previous paragraph) on plates incubated at RT and in a microaerophilic environment (Mirza and Rodrigues 2012). The Pikovskaya’s medium (PVK) was used to select strains able to solubilise phosphates (Gupta et al. 1994). Potassium solubilisation was assessed according to the protocol suggested by Zhang (Zhang and Kong 2014), by using solid Aleksandrov medium (Hu et al. 2006) and adding 0.2% of potassium and aluminium silicate in different form: ignimbrite, trachytic lava, microcline (Setiawati and Mutmainnah 2016). The capacity of the strains to produce indoleacetic acid was evaluated with the protocol suggested by Patten and Glick (2002). To detect siderophores production, the isolated strains were tested by chrome azurol S (O-CAS) assay, prepared according to Schwyn and Neilands (1987) following Pérez-Miranda et al. (2007) modification.
The T-S tailored microbial formula was assembled by choosing the PGP strains with multiple traits and complementary functions. The strains of the formula were repeatedly tested over time, as described above, to verify the stability of PGP properties.
Field experiments
Three field experiments were carried out using three varieties of tomato with different growing seasons: Camone DRW7723 (Bayer,Leverkusen, Germany); Oblungo Artù (Sementiera Medhermes, Ragusa, Italy); and Cherry Dantesco (Vilmorin Italia, Bologna, Italy). The plants were grown in a portion of a commercial greenhouse (“Santa Margherita Terra e Sole”) in Santa Margherita di Pula (SW Sardinia, Italy). The greenhouse has a surface of 3000 m2, with natural light and no cooling/heating supplies, provided with drip irrigation. The distance between rows was 1 m, while the distance between adjacent plants within a row was 30–50 cm depending on the tomato variety. In each experimentation, two entire rows (about 200 plants) were treated with conventional chemical fertiliser (F) and were separated by two rows of plants inoculated with the bacterial Formula T–S (B). According to the previous analysis (data not shown) the greenhouse soil was classified as sandy-loam (silt, 22%; clay, 13%; and sand, 65%) with pH 7.3.
The home-made chemical fertiliser was composed of ammonium nitrate 25 g/L, potassium nitrate 25 g/L, calcium nitrate 75 g/L (YaraTera, Italia), magnesium sulphate 25 g/L (DRT, Turkey), potassium sulphate 25 g/L (Haifa Hiberia, Spain), iron chelates -EDDHA 3,8% (Valagro, Italy), phosphoric acid and microelements (Jisa, Spain). The fertiliser mixture was provided by fertirrigation, according to the variety needs.
The 8 strains composing the bacterial Formula T-S were grown individually in TSB up to the stationary phase (38 to 48 h depending on the strain), then centrifuged to eliminate the culture broth. The cell pellets were suspended in sodium pyrophosphate 0.1% (w/v) to a cell density of 109 CFU/mL. Before the application in the field, equal amounts of the strain suspensions were mixed and diluted with tap water to a final concentration of about 107 CFU/mL and inoculated into the soil by watering each plant with 400 mL of mixed suspension.
The three experimental campaigns were carried out with tomato varieties having different growing season and different water and temperature requirements. In order to establish the optimal number of microbial applications (bio-fertilisation), the output of the first campaign was used to adjust the following field trials. The first experimentation was performed in the period October 2019-June 2020, on Camone tomato plants. Microbial inoculation was applied twice on the two selected rows (once in October 2019 and once in February 2020), while the chemical fertiliser was provided daily by fertigation on the other two rows. The second experimental campaign was performed between October 2020 and February 2021, on the Oblungo variety. The microbial inoculations were provided monthly, while the chemical fertiliser was applied daily, following the established farm practice. The last experimental campaign was performed in the period May 2021-October 2021, and involved the study of Cherry tomatoes, the application of monthly microbial inoculations, and daily chemical fertilisation. Irrigation and chemical fertilisation followed the normal farm plan.
Tomato growth parameters and productivity analysis
The parameters of plant growth and productivity (plant height, fruit weight and number) were monitored according to the physiological growth of the plant varieties, in order to compare the effectiveness of the two treatments: chemical fertilisation and microbial inoculation (bio-fertilisation). The parameters were expressed as an average of 30 plants per treatment.
The evaluation of the weight and number of tomatoes collected per plant was carried out: (i) for the first campaign from May to June 2020; (ii) for the second campaign from November 2020 to February 2021; (iii) for the third campaign from August to September 2021.
Statistical analyses
The experimental data (average plant height, average weight of the harvest of tomatoes, average number of collected tomatoes) were subjected to statistical analyses, one-way analysis of variance (ANOVA) and Tukey HSD-test, through Xlstat software, to evaluate any statistically significant difference between the groups of microbial inoculation and chemical fertilisation, at a significance level α = 0.05.
The PGP traits of the isolated strains were subjected to principal component analysis (PCA), to evaluate any difference among bacterial strains with regard to the metabolic characteristics. Results were obtained by considering for each bacterial strain the relative percentage of the expressed variables.
Results
Soil characterization
The chemical composition of the farm soil (Table 1) showed high amounts of silica (73.82%) and aluminium oxides (14.29%), the presence of abundant K (K2O, 6.82%), and P in traces around 600 mg.kg−1. The mineralogical association, determined by XRD analyses, consists of quartz (SiO2)K-feldspar (orthoclase/microcline, KAlSi3O8), plagioclase (NaAlSi3O8) and phyllosilicates such as phlogopite (KMg3(Si3Al)O10(OH)2), muscovite (KAl2(AlSi3O10)(OH)2), illite (K0.65Al2.0[Al0.65Si3.35O10](OH)2) and kaolinite (Al2Si2O5(OH)4). These results reflect the geological setting of the area, characterised by the widespread occurrence of granitic rocks (Barca et al. 2009). Quartz, feldspars, and muscovite are primary components of granitoids, whereas kaolinite and illite are weathering products of feldspars and muscovite, respectively. CHN analysis results indicate C values in the low range. The technique detected total C from carbonate minerals and organic materials: as the soil is poor in carbonate this amount must be referred to organic carbon. Moreover, the soil is poor in hydrogen, while it contains appreciable amounts of N.
Functional diversity analysis of microbial soil communities
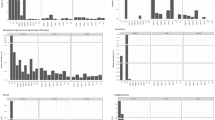
Average Well Colour Development (AWCD) was used as an indicator of microbial activity in Biolog analysis. The AWCD showed a lag phase of 24 h and increased rapidly after 48 h of incubation, reaching the plateau after 7 days of incubation in ECOPlates, indicating a high metabolic activity (Fig. 1).
All groups of substrates (carbohydrates, P-sugars, carboxylic acids, aminoacids, amines and polymers) contained in the ECOplates were efficiently used by the microbial community (Fig. 1), showing a high functional diversity (96%, percentage of the number of utilized carbon substrates on the overall number). Among the substrates, only D, L α-glycerol-phosphate was below the threshold value (OD 0.2). The carbon sources readily metabolised, with OD values higher than 1.0 after 48 h of incubation, were β-Methyl-D-Glucoside, D-Mannitol, D-Cellobiose, N-Acetyl-D-Glucosamine, Itaconic Acid, d-Malic Acid, 4-Hydroxybenzoic Acid, l-Asparagine, l-Serine, and Glycogen. After 72 h of incubation α-D-Lactose, d-Galactonic Acid Lactone, l-Arginine, l-Phenylalanine, Phenylethylamine, Tween 40, Tween 80, and α -Cyclodextrin were metabolised. The slowest kinetic reaction was with D-Xylose, i-Erythritol, α -D-Glucose-1-Phosphate, d-Galacturonic Acid, Pyruvic Acid Methyl Ester, d-Glucosaminic Acid, α-Ketobutyric Acid, Glycyl-L-Glutamic Acid, 2-Hydroxybenzoic Acid, and l-Threonine. The substrates γ-Hydroxybutyric Acid and Putrescine were poorly metabolised and reached OD values lower than 1.0.
Isolation and identification of bacteria
The microbial load of the heterotrophic population was 3 × 107 CFU g−1 of soil. Based on differential colony morphologies, a total of 40 bacterial strains were isolated and identified using 16S rRNA gene sequencing to species level (Table 2 and Table S1 in Supplementary Material).
The 40 strains isolated from the cultivable fraction of the heterotrophic population were examined together with the reference strains of related taxa and the phylogenetic status is shown in the cladogram compiled by Neighbour-Joining methods (Fig. 2). The most represented class was Bacilli, with seven genera and 11 species; 13 strains belong to 3 genera of the class γ-Proteobacteria (Pseudomonas, Enterobacter, Acinetobacter); 6 strains belong to 4 genera of the class Actinobacteria (Streptomyces, Microbacterium, Paenarthrobacter and Arthrobacter); 2 strains of Delftia lacustris belong to the class β-Proteobacteria; 4 strains belong to 3 genera of the class α-Proteobacteria (Ensifer, Sphingobium, and Phyllobacterium).
Evolutionary relationships of taxa. The evolutionary history was inferred using the Neighbour-Joining method (Tamura et al 2004). The optimal tree with the sum of branch length = 3.73998697 is shown. The tree is drawn to scale, with branch lengths in the same units as those of the evolutionary distances used to infer the phylogenetic tree. The evolutionary distances were computed using the Maximum Composite Likelihood method and are in the units of the number of base substitutions per site. The analysis involved 101 nucleotide sequences. Codon positions included were 1st + 2nd + 3rd + Noncoding. All ambiguous positions were removed for each sequence pair. There were a total of 1651 positions in the final dataset. Evolutionary analyses were conducted in MEGA X (Kumar et al 2018). Bacteria selected for Formula T–S are marked with red dots
Characterization of strains for PGP traits
After isolation and identification, all strains were characterised for PGP traits (Table 2). Within the 40 isolated, about 93% were positive for nitrogen-fixation; 53% for phosphate solubilisation, 58% for potassium solubilisation, 58% were able to produce IAA and 33% siderophores. Eight isolates (IN11, ITA5, IN3, ITA14, ITA15, ITA17, IN1, and IN4) were selected to be part of the bacterial Formula T-S, because they exhibited multiple PGP activities. According to the ecology-based approach, the Formula T-S was assembled reflecting as much as possible the original composition of the PGP-native community, including 8 species belonging to the genera Delftia, Pseudomonas, Paenarthrobacter, Phyllobacterium, Bacillus, and Acinetobacter (indicated by red dots in Fig. 2).
In the Formula T-S (Table 2 and Table S2 in Supplementary material) all the strains are nitrogen-fixers. Most of the eight strains were able to produce different amounts of IAA. Ps. plecoglossicida IN11 was the best IAA producer and the most efficient phosphate solubilizer, while Ps. plecoglossicida ITA14 and Bacillus tequilensis ITA17 were excellent potassium-solubilizing bacteria. Except for Acinetobacter venetianus (IN4), all the selected bacteria can produce siderophores. This common feature was particularly marked in Bacillus subtilis (ITA15).
The distribution of PGP traits [i.e., the ability to fix atmospheric nitrogen (N-fix), to solubilise phospates (P-sol), to solubilise potassium (K-sol), to produce IAA and siderophores (O-CAS)] among the 40 isolated strains was analyzed by principal components analysis (PCA). Figure 3a shows the biplot of PCA obtained for all the strains, in which the first two components (PC1 and PC2) account for the 42.68% and 33.86% of the variance, respectively. The strains selected for the Formula T-S (black circles) included bacteria with higher capacity to express the overall plant growth promoting traits than the rest of the isolated community. In fact, they are mainly distributed in the right side of the biplot (positive values of PC1), influenced by the contribution of most of the variables considered (Fig. 3a). On the contrary, the rest of microbial isolates (white circles) is mainly distributed in the second quadrant of the biplot (negative PC1 values). In the biplot in Fig. 3b, (55.76% and 30.90% of the variance accounted by PC1 and PC2, respectively), bacteria of the microbial consortium showed a wide distribution in five different clusters, as result of their multiple PGP traits, and can be grouped based on their metabolic similarities. Among the eight selected bacteria, ITA 15 strain showed higher capacity to produce siderophores than the others. The formula T-S, assembled with the best-performing PGP strains by choosing those with multiple traits and complementary functions, was prepared for bioaugmentation treatment in field experiments.
a Biplot obtained by Principal Component Analysis (PCA) of all isolated strains, considering the first two components (PC1 42.68%, PC2 33.86%); b biplot of bacteria selected for the microbial formula, obtained by Principal Component Analysis (PCA) considering the first two components (PC1 55.76%, PC2 30.90%)
Field experiments
Figure 4 shows the average plant height, measured at different stages of plant growth, during the three field experiments in two different soil treatments: bio-fertilisation (B) and chemical fertilisation (F). In the first and in the second field experiment, the average trend of plant growth was similar for both bio- and chemical fertilisation (Fig. 4a, b). No significant differences were observed between treatment with bacteria and chemical fertilisation, either for the first (p = 0.952; p = 0.801; p = 0.904; p = 0.997 after 2, 4, 8 and 10 weeks from transplanting-WAT, respectively) or for the second experiment (p = 0.999; p = 0.999; p = 0.996; p = 0.998 after 7, 8, 11 and 14 WAT, respectively). In the third field experiment (Fig. 4c), the average trend of plant growth was similar in the first period, with no significant difference after 6 weeks (p = 1.00) and slight difference after 8 weeks (p = 0.037). However, during the last period (10–11 weeks after transplanting), the average plant height was significantly higher in the case of chemical fertiliser (p < 0.0001).
The average plant height measured during the first, second and third field experiment (WAT: weeks after transplanting). a first experiment, from March to June 2020; b second experiment, from October to December 2020; c third experiment, from June to July 2021. Two different soil treatments were compared: the bio-fertilisation (B) and the application of chemical fertiliser (F). The error bar corresponds to standard deviation. The significance of the differences between groups was evaluated by one-way analysis of Variance (ANOVA) and Tukey HSD-test. The asterisk (*) corresponds to a statistically significant difference (p < 0.05)
The average weight of tomatoes per plant in the three field experiments is shown in Fig. 5. The first field experiment revealed a significant difference of the average weight (p < 0.0001) between bio-fertilisation (B) and chemical fertilisation (F) groups, with higher values for F. In the following experiments, the average weight was very similar between B and F groups, with no significant difference in both the second experiment (p = 1.00) and third experiment (p = 0.989). Likewise, in the first experiment a higher number of tomatoes was harvested in group F (p = 0.0003), while similar results were observed during the second and third experiments for B and F groups (p = 0.9999) (data not shown).
Evaluation of the average weight of tomatoes per plant during the first, second and third field experiment, in different periods: 1st experiment, from May to June 2020; 2nd experiment, from November 2020 to February 2021; 3rd experiment, from August to September 2021.Two different soil treatments were compared: the bio-fertilisation (B) and the application of chemical fertiliser (F). Three different varieties of tomatoes were considered: Camone (1st Exp), Oblungo (2nd Exp) and Cherry (3rd Exp). The error bar corresponds to standard deviation. The significance of the differences between groups was evaluated by one-way analysis of variance (ANOVA) and Tukey HSD-test. The asterisk (*) corresponds to a statistically significant difference (p < 0.0001)
Although B, compared to F, significantly reduced the height in Cherry variety, the plant productivity was not significantly modified: F and B groups showed similar results on the average weight of tomatoes (p > 0.05). The increase in the frequency of inoculation (monthly repetition) gave positive results in terms of fruits yield in the two experiments run with Oblungo and Cherry varieties.
Discussion
This work assessed for the first time the effectiveness of the bioaugmentation strategy for a full substitution of chemical fertilization on a tomato farm in Sardinia. When the research began, it was known that the goal was very ambitious: Sardinia Island is among the hotspots for climate change (Marras et al. 2021) and is therefore experiencing soil fertility loss due to rising temperatures and changing changes in precipitation regimes. (Mondal 2021; Caloiero and Guagliardi 2021). As a result, there has been an increased demand for chemical fertilisers to cope with the decrease in agricultural yields. Therefore, considering the unsustainability of the traditional intensive production system, the far-sighted farmers involved in this project recognized the importance of testing alternative methods to produce tomatoes, reducing environmental impacts and facing the effects of climate change.
The choice to apply the emerging bioaugmentation approach based on a combination of microorganisms with different PGP traits was based on findings from various studies showing that microbial consortia have the potential to increase plant growth much more than inoculants with a single bacterial species (He et al. 2019; Compant et al. 2019; Mitter et al. 2021). However, a smart and knowledge-driven selection of consortia and strains is required (Tosi et al. 2020). Indeed, as previously reported in literature, no microbial inoculant can be universally effective for all systems, and efficacy may be affected by many factors such as the ability of inoculated microorganisms to persist in soil, depending on their compatibility with the environmental characteristics and the degree of spatial competition with other organisms in the target niche (Mannino et al. 2020) or the interactions between a specific plant type and the selected PGP strains (Adesemoye et al. 2009; Mitter et al. 2021). This study investigated the culturable indigenous microbial community, functionally linked with chemical characteristics of native soil and with the needs of the target tomato species, to select and to locally isolate pre-adapted bacteria to be used as bioaugmentation strategy (Sprocati et al. 2014). This approach aimed to widen as much as possible the biodiversity of PGP bacteria to be included in the inoculum, in order to establish a successful bioaugmentation practice.
The bacterial community seemed to preserve a peculiar taxonomic (Fig. 2 and Table S1) and functional biodiversity (Fig. 1) and the observed bacterial density (3 × 107 CFU g −1) was about an order of magnitude higher than what normally found in other agricultural soils sampled in summer (Bevivino et al. 2014; Bhowmik et al. 2019). While taking in account that microorganism load may vary within and between different soil types and conditions (Vieira and Nahas 2005), the observed bacterial load could indicate a good quality of soil microbiome.
In vitro tests were carried out to assess the PGP potential of all the isolates and to choose the most effective strains for field bioaugmentation. The rationale of the choice for the bacterial formula composition was the combination of microorganisms with the highest number of PGP traits, but also with different and complementary capacities likely to induce positive effects on plant physiology and fruit yield. (Fig. 3). Moreover, according to the ecology-based approach suggested by Dejonghe et al. (2001) and applied in our previous works, to improve the survival and functioning of bacteria in the tomato soil microbiome, the formula was assembled including different species reflecting as much as possible the original microbial community biodiversity (Fig. 2).
All the eight bacteria included in Formula T–S were diazotrophic microbes, able to convert atmospheric nitrogen into ammonia (Table 2). The biologically fixed nitrogen, more sustainable than chemical fertilisers and less accessible for leaching and volatilization, allows the replenishment of soil total nitrogen content and regulates the crop growth and yield. In addition, an increase of the root system development and a more efficient nutrient uptake by the plant are also due to the production of IAA (Kumar et al. 2020). This capacity was very high in Pseudomonas plecoglossicida (IN11), Paenarthrobacter nitroguajacolicus (ITA5) and Phyllobacterium phragmitis (IN3) belonging to genera already tested as efficient IAA producers (Menéndez et al. 2020; Pérez-Rodriguez et al. 2020; Riva et al. 2021) and, to a lesser extent, in all the other isolates, except Bacillus tequilensis (ITA17) and Acinetobacter venetianus (IN4). These last two bacteria, and in particular Bacillus tequilensis (ITA17) were chosen, together with Pseudomonas plecoglossicida (ITA14), for the excellent ability to solubilise K (K-sol) in laboratory tests, as already reported in literature (Ahmad et al. 2016; Etesami et al. 2017; Saxena et al. 2020; Ashfaq et al. 2020). Soil analysis showed the presence of microcline, muscovite and phlogopite, an important source of insoluble K originating from weathering processes of granite minerals (Table 1). The presence of K-sol bacteria in Formula T–S was able to increase the bioavailability of K to meet the requirement of the tomato plants.
Phosphorus (P) is an essential nutrient required for diverse plant metabolic processes such as respiration, biosynthesis, photosynthesis, energy transfer, and signal transduction. However, it is slightly bioavailable in many agricultural lands (Kumar et al. 2022) and in our experimental field the amount of P was very scarce (Table 1). Bacteria belonging to different genera such as Arthrobacter, Bacillus, Beijerinckia, Burkholderia, Enterobacter, Pseudomonas, Erwinia, Mesorhizobium, Flavobacterium, Rhodococcus and many others, can solubilize phosphates converting them into a bioavailable form (Lobo et al. 2019; Shilev 2020). For this PGP trait, two strains of P. plecoglossicida (IN11 and ITA14) were chosen as the most efficient phosphate-solubilizing microbes among all the isolates. Another important nutrient for plant cells is Fe but, despite its abundance on Earth, it is not widely accessible in soils when present in complexes of hydroxides and oxyhydroxides. Different PGPB, such as Pseudomonas, Bacillus, and Phyllobacterium found in this study, possess the ability to synthesise siderophores. These Fe-chelating compounds have a high affinity to Fe3+ and form complexes that lead to Fe-mobilisation (reducing Fe3+to Fe2+) and make it bioavailable for plant roots (Shilev 2020; Pérez-Rodriguez et al. 2020; Flores-Félix et al. 2021). Moreover, it is known that siderophore-producing bacteria play a crucial role not only in growth promotion but also in biocontrol activity, by competing for Fe3+ with the pathogens in the rhizosphere (Kumar et al. 2022). For instance, several strains of Bacillus subtilis have been reported to suppress fungal pathogens in plants using siderophores (Manasa et al. 2021; Kumar et al. 2022). In agreement with these literature data, the highest siderophores production observed in this study was performed by two strains of Bacillus (ITA15, ITA17). Thus, all the eight strains tested as possible bio-fertiliser are phylogenetically affiliated to bacterial species that present a PGP potential. Moreover, in terms of biosafety, all the selected isolates belong to the risk group 1, as stated in the reference document provided by the German Committee on Biological Agents—ABAS, TRBA 466 (2020), and their use in field does not imply particular concern for human health. A further advantage of using native microorganisms is to avoid the risk of introducing foreign strains, which could prove dangerous once in contact with the indigenous community, as also reported by Mahmud et al. (2021).
After the positive results obtained by in vitro screening of PGP bacteria, in agreement with the bottom-up approach suggested by Riva et al. (2021), we tested the real effect of these bacterial inoculants on plant production of three different tomato varieties. The experiments were carried out in a commercial greenhouse, with the aim to generate meaningful information for the researchers and, at the same time, to minimise eventual loss of profit for the farmers. In fact, to avoid any reduction in tomato production, the experimental design did not include negative controls, represented by plants without fertilisation. Anyway, the greenhouse experimentations were essential because laboratory screening and small-scale experiments provide only limited information, and success in open fields is often variable. In other studies, some consortia with multiple PGP-activities showed low efficiency when applied in field experiments (Cardinale et al. 2015; Compant et al. 2019; Riva et al. 2021), while other bacteria, that did not display a promising set of PGP activities in in vitro assays, proved to be the best growth promoters when tested directly on plants (Cardinale et al. 2015). Therefore, if field experiments are fundamental to select the most efficient bioinoculant, it is equally important to perform long-term experiments of bio-fertilisation and evaluate the PGP effect exerted throughout the plant life cycle and especially in fruit production (Riva et al. 2021). In the present work, although bio-fertilization affected the vegetative growth in the Cherry variety, the plant productivity of all varieties was comparable to chemical fertilization when the number of microbial inoculations was optimised (from two initial bio-fertilisation to monthly applications per growing season). Previous works (Heuvelink 1999; Massa et al. 2019) have also observed that vegetative growth in tomato plants does not necessarily lead to higher yield.
As highlighted in a recent study by He et al. (2019), plants require distinct types of microbial activities at different stages of growth. Therefore, not only the co-inoculation of bacterial strains with different properties, but also the frequency of PGPB application, could influence plant performance. This can explain the different conclusions found in Adesemoye et al. (2009) and Ye et al. (2020) that indicated microbial inoculation as a promising complement of synthetic fertilisers but not as a valid substitute: the strategy used to assemble the microbial formula and the frequency of inoculation can be crucial in the success of bio-fertilization.
Conclusions
The present work demonstrates the feasibility of replacing chemical fertilizers to sustain and guarantee adequate tomato yield with functional bioaugmentation, aimed at implementing the best PGP functions already present within the soil microbial community. The relevant point is the strategy used to assemble the bioaugmentation formula including different bacterial strains, native to the farm soil. The strains were selected for the best and complementary PGP traits, and combined fitting as closely as possible the taxonomic composition of the indigenous PGP microbial community structure. The formula T-S was composed by 8 strains with a balanced mix of the O-CAS trait, followed by IAA, N-fix, K-sol, and P-sol. By inoculating the selected strains at high concentrations in the agricultural soil, the PGP functions of the whole native community are strengthened, without creating relevant alteration in the community structure. Furthermore, the frequency of bio-fertilisation was also crucial: a functional bioaugmentation by monthly inoculation was at least equal to chemical fertilisation, although the procedure is still open to further optimisation. In light of the above, these results may pave the way towards a more sustainable production of Solanum lycopersicum, also applicable to other agronomic crops. In agreement with Agenda 2030 and the UE “Farm to Fork” strategy, such environmentally-sound practices are becoming increasingly crucial.
Still, some important aspects remain to be explored: (1) an in-depth investigation of the microbial ecology of the target soil to enrich the bioaugmentation formula with microbial inoculants not considered in this work, i.e. cyanobacteria and mycorrhiza, (2) the effect of the bio-fertiliser on tomatoes quality in terms of organoleptic properties, Vitamin C content, and nitrate accumulation as suggested by Ye et al. (2020); (3) the effect of Formula T-S on the chemical and biological properties of soil in a continuous cropping system and (4) the potential of this approach in the scenario of increasing drought and salinity caused by climate change.
Data availability
Supplementary data are provided in a file.
Abbreviations
- PGP:
-
Plant growth promoting
- PGPB:
-
Plant growth promoting bacteria
- RT:
-
Room temperature
- CLPP:
-
Community-level physiological profile
- AWCD:
-
Average well colour development
- PXRD:
-
Powder X-rays diffraction
- XRF:
-
X-rays fluorescence
- CRIST:
-
Structural crystallography centre
- CFUs:
-
Colony-forming units
- NF:
-
Nitrogen free
- N-fix:
-
Nitrogen fixing
- PCR:
-
Polymerase chain reaction
- P-sol:
-
Phosphate solubilisation
- K-sol:
-
Potassium solubilisation
- IAA:
-
Indole-acetic acid
- PVK:
-
Pikovskaya’s medium
- O-CAS:
-
Overlaid chrome azurol S
- F:
-
Chemical fertiliser
- B:
-
Bacterial formula T-S (biological fertiliser)
- ANOVA:
-
Analysis of variance
References
Adesemoye AO, Torbert HA, Kloepper JW (2009) Plant growth-promoting rhizobacteria allow reduced application rates of chemical fertilizers. Microb Ecol 58:921–929. https://doi.org/10.1007/s00248-009-9531-y
Ahmad M, Nadeem SM, Naveed M, Zahir ZA (2016) Potassium-solubilizing bacteria and their application in agriculture. Potassium solubilizing microorganisms for sustainable agriculture. Springer, India, pp 293–313
Angulo J, Martínez-Salgado MM, Ortega-Blu R, Fincheira P (2020) Combined effects of chemical fertilization and microbial inoculant on nutrient use efficiency and soil quality indicators. Scientia Agropecuaria 11:375–380. https://doi.org/10.17268/sci.agropecu.2020.03.09
ANICAV Italy’s National Industrial Association of Vegetable Food Preserves (2021) News: “Pomodoro da industria: campagna 2021 da record”. https://anicav.it/2021/11/16/campagna2021. Accessed 23/01/2023
Ashfaq M, Hassan HM, Ghazali AHA, Ahmad M (2020) Halotolerant potassium solubilizing plant growth promoting rhizobacteria may improve potassium availability under saline conditions. Environ Monit Assess 192:697. https://doi.org/10.1007/s10661-020-08655-x
Barca S, Serri R, Rizzo R, et al (2009) ISPRA Servizio Geologico d’Italia, Regione Autonoma della Sardegna: note Illustrative della Carta Geologica d’Italia alla scala 1:50.000. Foglio 565 “Capoterra”
Bevivino A, Paganin P, Bacci G et al (2014) Soil bacterial community response to differences in agricultural management along with seasonal changes in a Mediterranean region. PLoS One. https://doi.org/10.1371/journal.pone.0105515
Bhowmik A, Singh Kukal S, Saha D et al (2019) Potential indicators of soil health degradation in different land use-based ecosystems in the Shiwaliks of Northwestern India. Sustainability 11:3908. https://doi.org/10.3390/su11143908
Bona E, Todeschini V, Cantamessa S et al (2018) Combined bacterial and mycorrhizal inocula improve tomato quality at reduced fertilization. Sci Hortic 234:160–165. https://doi.org/10.1016/j.scienta.2018.02.026
François-Xavier Branthôme (2022) Consumption: 2021 in the wake of 2020. In: 14TH WPTC Congress. https://www.tomatonews.com/en/consumption-2021-in-the-wake-of-2020_2_1618.html. Accessed 23/01/2023.
Caloiero T, Guagliardi I (2021) Climate change assessment: seasonal and annual temperature analysis trends in the Sardinia region (Italy). Arab J Geosci 14:2149. https://doi.org/10.1007/s12517-021-08527-9
Cardinale M, Ratering S, Suarez C et al (2015) Paradox of plant growth promotion potential of rhizobacteria and their actual promotion effect on growth of barley (Hordeum vulgare L.) under salt stress. Microbiol Res 181:22–32. https://doi.org/10.1016/j.micres.2015.08.002
Čechura L, Kroupová ZŽ, Samoggia A (2021) Drivers of productivity change in the italian tomato food value chain. Agriculture (switzerland) 11:996. https://doi.org/10.3390/agriculture11100996
Committee on Biological Agents - ABAS (2020) TRBA 466: Classification of prokaryotes (bacteria and archaea) into risk groups. http://regelwerke.vbg.de/vbg_trba/trba466/trba466_0_.html. Accessed 23/01/2023.
Compant S, Samad A, Faist H, Sessitsch A (2019) A review on the plant microbiome: ecology, functions, and emerging trends in microbial application. J Adv Res 19:29–37
Cordero I, Balaguer L, Rincón A, Pueyo JJ (2018) Inoculation of tomato plants with selected PGPR represents a feasible alternative to chemical fertilization under salt stress. J Plant Nutr Soil Sci 181:694–703. https://doi.org/10.1002/jpln.201700480
Dejonghe W, Boon N, Seghers D, Top EM, Verstraete W (2001) Bioaugmentation of soils by increasing microbial richness: missing links. Environ Microbiol 3(10):649–657. https://doi.org/10.1046/j.1462-2920.2001.00236.x
del Amor FM, Cuadra-Crespo P (2012) Plant growth-promoting bacteria as a tool to improve salinity tolerance in sweet pepper. Funct Plant Biol 39:82–90. https://doi.org/10.1071/FP11173
Dobereiner J, Marriel IE, Nery M (1976) Ecological distribution of Spirillum lipoferum Beijerinck. Can J Microbiol 22:1464–1473
Etesami H, Emami S, Alikhani HA (2017) Potassium solubilizing bacteria (KSB): Mechanisms, promotion of plant growth, and future prospects-a review. J Soil Sci Plant Nutr 17:897–911. https://doi.org/10.4067/S0718-95162017000400005
Flores-Félix JD, Velázquez E, Martínez-Molina E et al (2021) Connecting the lab and the field: genome analysis of phyllobacterium and rhizobium strains and field performance on two vegetable crops. Agronomy 11:1124. https://doi.org/10.3390/agronomy11061124
Gamalero E, Berta G, Massa N et al (2008) Synergistic interactions between the ACC deaminase-producing bacterium Pseudomonas putida UW4 and the AM fungus Gigaspora rosea positively affect cucumber plant growth. FEMS Microbiol Ecol 64:459–467. https://doi.org/10.1111/j.1574-6941.2008.00485.x
Garland JL (1996) Analytical approaches to the characterization of samples of microbial communities using patterns of potential C source utilization. Soil Biol Biochem 28:213–221. https://doi.org/10.1016/0038-0717(95)00112-3
Giuliani MM, Gatta G, Cappelli G et al (2019) Identifying the most promising agronomic adaptation strategies for the tomato growing systems in Southern Italy via simulation modeling. European J Agron 111:125937. https://doi.org/10.1016/j.eja.2019.125937
Gupta R, Singal R, Shankar A et al (1994) Short communication a modified plate assay for screening phosphate solubilizing microorganisms. J Gen App Microbiol 40:255–260. https://doi.org/10.2323/jgam.40.255
He Y, Pantigoso HA, Wu Z, Vivanco JM (2019) Co-inoculation of Bacillus sp. and Pseudomonas putida at different development stages acts as a biostimulant to promote growth, yield and nutrient uptake of tomato. J Appl Microbiol 127:196–207. https://doi.org/10.1111/jam.14273
Heuvelink E (1999) Evaluation of a dynamic simulation model for tomato crop growth and development. Ann Bot 83(4):413–422
Hu X, Chen J, Guo J (2006) Two phosphate- and potassium-solubilizing bacteria isolated from Tianmu Mountain, Zhejiang, China. World J Microbiol Biotechnol 22:983–990. https://doi.org/10.1007/s11274-006-9144-2
Kaminsky LM, Trexler Rv, Malik RJ et al (2019) The inherent conflicts in developing soil microbial inoculants. Trends Biotechnol 37:140–151. https://doi.org/10.1016/J.TIBTECH.2018.11.011
Kumar S, Stecher G, Li M et al (2018) MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Kumar BP, Reddy NS, Sparjanbabu DS (2020) Role of microbial communities to mitigate climate change in agriculture. Int J Curr Microbiol Appl Sci 9:1477–1483. https://doi.org/10.20546/ijcmas.2020.910.176
Kumar S, Diksha Sindhu SS, Kumar R (2022) Biofertilizers: An ecofriendly technology for nutrient recycling and environmental sustainability. Curr Res Microb Sci 3:100094. https://doi.org/10.1016/j.crmicr.2021.100094
Lobo CB, Juárez Tomás MS, Viruel E et al (2019) Development of low-cost formulations of plant growth-promoting bacteria to be used as inoculants in beneficial agricultural technologies. Microbiol Res 219:12–25
Mahmud K, Missaoui A, Lee K et al (2021) Rhizosphere microbiome manipulation for sustainable crop production. Curr Plant Biol 27:100210. https://doi.org/10.1016/J.CPB.2021.100210
Manasa M, Ravinder P, Gopalakrishnan S et al (2021) Co-inoculation of bacillus spp. for growth promotion and iron fortification in sorghum. Sustainability 13:12091. https://doi.org/10.3390/su132112091
Manfredi P, Cassinari C, Gatti M, Trevisan M (2019) Growth and yield response of tomato (Solarium lycopersicum L.) to soil reconstitution technology. Agrochimica 63:73–83. https://doi.org/10.12871/00021857201916
Mannino G, Nerva L, Gritli T et al (2020) Effects of different microbial inocula on tomato tolerance to water deficit. Agronomy 10:170. https://doi.org/10.3390/agronomy10020170
Marras PA, Lima DCA, Soares PMM et al (2021) Future precipitation in a Mediterranean island and streamflow changes for a small basin using EURO-CORDEX regional climate simulations and the SWAT model. J Hydrol (Amst) 603:127025. https://doi.org/10.1016/j.jhydrol.2021.127025
Massa D, Bonetti A, Cacini S, Faraloni C, Prisa D, Tuccio L, Petruccelli R (2019) Soilless tomato grown under nutritional stress increases green biomass but not yield or quality in presence of biochar as growing medium. Horticult Environ Biotechnol 60:871–881. https://doi.org/10.1007/s13580-019-00169-x
Menéndez E, Pérez-Yépez J, Hernández M et al (2020) Plant growth promotion abilities of phylogenetically diverse mesorhizobium strains: effect in the root colonization and development of tomato seedlings. Microorganisms 8:412. https://doi.org/10.3390/microorganisms8030412
Mirza BS, Rodrigues JLM (2012) Development of a direct isolation procedure for free-living diazotrophs under controlled hypoxic conditions. Appl Environ Microbiol 78:5542–5549. https://doi.org/10.1128/AEM.00714-12
Mitter EK, Tosi M, Obregón D et al (2021) Rethinking crop nutrition in times of modern microbiology: innovative biofertilizer technologies. Front Sustain Food Syst 5:606815. https://doi.org/10.3389/fsufs.2021.606815
Mondal S (2021) Impact of climate change on soil fertility. In: Choudhary DK, Mishra A, Varma A (eds) Climate change and the microbiome: sustenance of the ecosphere. Springer International Publishing, Cham, pp 551–569
Morra L, Cozzolino E, Salluzzo A et al (2021) Plant growth, yields and fruit quality of processing tomato (Solanum lycopersicon L.) as affected by the combination of biodegradable mulching and digestate. Agronomy 11:100. https://doi.org/10.3390/agronomy11010100
Mühling M, Woolven-Allen J, Murrell JC, Joint I (2008) Improved group-specific PCR primers for denaturing gradient gel electrophoresis analysis of the genetic diversity of complex microbial communities. ISME J 2:379–392. https://doi.org/10.1038/ismej.2007.97
Oleńska E, Małek W, Wójcik M et al (2020) Beneficial features of plant growth-promoting rhizobacteria for improving plant growth and health in challenging conditions: a methodical review. Sci Total Environ 743:140682. https://doi.org/10.1016/j.scitotenv.2020.140682
Ortiz N, Armada E, Duque E et al (2015) Contribution of arbuscular mycorrhizal fungi and/or bacteria to enhancing plant drought tolerance under natural soil conditions: effectiveness of autochthonous or allochthonous strains. J Plant Physiol 174:87–96. https://doi.org/10.1016/J.JPLPH.2014.08.019
Patten CL, Glick BR (2002) Role of pseudomonas putida indoleacetic acid in development of the host plant root system. Appl Environ Microbiol 68:3795–3801. https://doi.org/10.1128/AEM.68.8.3795-3801.2002
Pérez-Miranda S, Cabirol N, George-Téllez R et al (2007) O-CAS, a fast and universal method for siderophore detection. J Microbiol Methods 70:127–131. https://doi.org/10.1016/J.MIMET.2007.03.023
Pérez-Rodriguez MM, Piccoli P, Anzuay MS et al (2020) Native bacteria isolated from roots and rhizosphere of Solanum lycopersicum L. increase tomato seedling growth under a reduced fertilization regime. Sci Rep 10:15642. https://doi.org/10.1038/s41598-020-72507-4
Potts PJ, Webb PC (1992) X-ray fluorescence spectrometry. J Geochem Explor 44:251–296. https://doi.org/10.1016/0375-6742(92)90052-A
Riva V, Mapelli F, Dragonetti G et al (2021) Bacterial inoculants mitigating water scarcity in tomato: the importance of long-term in vivo experiments. Front Microbiol 12:675552. https://doi.org/10.3389/fmicb.2021.675552
Sah S, Krishnani S, Singh R (2021) Pseudomonas mediated nutritional and growth promotional activities for sustainable food security. Curr Res Microb Sci 2:100084. https://doi.org/10.1016/j.crmicr.2021.100084
Saxena AK, Kumar M, Chakdar H et al (2020) Bacillus species in soil as a natural resource for plant health and nutrition. J Appl Microbiol 128:1583–1594. https://doi.org/10.1111/jam.14506
Schmidt T, Schlegel HG (1989) Nickel and cobalt resistance of various bacteria isolated from soil and highly polluted domestic and industrial wastes. FEMS Microbiol Lett 62:315–328. https://doi.org/10.1111/j.1574-6968.1989.tb03386.x
Schwyn B, Neilands JB (1987) Universal chemical assay for the detection and determination of siderophores. Anal Biochem 160:47–56. https://doi.org/10.1016/0003-2697(87)90612-9
Setiawati TC, Mutmainnah L (2016) Solubilization of potassium containing mineral by microorganisms from sugarcane rhizosphere. Agric Sci Procedia 9:108–117. https://doi.org/10.1016/j.aaspro.2016.02.134
Shilev S (2020) Plant-growth-promoting bacteria mitigating soil salinity stress in plants. Appl Sci 10:1–20. https://doi.org/10.3390/app10207326
Sprocati AR, Alisi C, Tasso F et al (2014) Bioprospecting at former mining sites across Europe: microbial and functional diversity in soils. Environ Sci Pollut Res 21:6824–6835. https://doi.org/10.1007/s11356-013-1907-3
Tamura K, Nei M, Kumar S (2004) Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc Natl Acad Sci USA 101:11030–11035
Tosi M, Mitter EK, Gaiero J, Dunfield K (2020) It takes three to tango: The importance of microbes, host plant, and soil management to elucidate manipulation strategies for the plant microbiome. Can J Microbiol 66:413–433. https://doi.org/10.1139/cjm-2020-0085
Vieira FCS, Nahas E (2005) Comparison of microbial numbers in soils by using various culture media and temperatures. Microbiol Res 160:197–202. https://doi.org/10.1016/j.micres.2005.01.004
Wu Y, Yan S, Fan J et al (2022) Combined effects of irrigation level and fertilization practice on yield, economic benefit and water-nitrogen use efficiency of drip-irrigated greenhouse tomato. Agric Water Manag 262:107401. https://doi.org/10.1016/j.agwat.2021.107401
Ye L, Zhao X, Bao E et al (2020) Bio-organic fertilizer with reduced rates of chemical fertilization improves soil fertility and enhances tomato yield and quality. Sci Rep 10:177. https://doi.org/10.1038/s41598-019-56954-2
Zhang C, Kong F (2014) Isolation and identification of potassium-solubilizing bacteria from tobacco rhizospheric soil and their effect on tobacco plants. Appl Soil Ecol 82:18–25. https://doi.org/10.1016/j.apsoil.2014.05.002
Acknowledgements
The authors are grateful to Salvatore Vacca, Giulia Siclari and the Siclari family for their contribution in running the field experiments.
Funding
Open access funding provided by Ente per le Nuove Tecnologie, l'Energia e l'Ambiente within the CRUI-CARE Agreement. This work was supported by “SUPREME: developing tools for Sustainable food Production in mediterranean area using MicrobEs”-Id: ERANETMED2—72—094.
Author information
Authors and Affiliations
Contributions
GDG coordinated the project. ARS, CA coordinated the research activity. GDG, ARS, GM, PP, CA, and FT contributed to the conception and design of the study. GDG, DM, PAM, CA and PP collected soil samples. DM ED, and PAM performed mineralogical and chemical analyses and data elaboration. TC, PP, CA, GM and FT performed bacteria isolation and characterization. PP and FT performed sequence analysis. TC, CI, PP, CA, GM and FT prepared the inocula. TC, ARS, GDG, ED, DM and PAM set up the greenhouse experiment and data collection.CI performed statistical analysis and plant data interpretation. PP and CI wrote the first draft of the manuscript. CA, ED, and GDG wrote sections of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests. The authors have no relevant financial interests to disclose.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent to publication
Not applicable.
Additional information
Communicated by Jupei Shen.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Paganin, P., Isca, C., Tasso, F. et al. A bacterial formula with native strains as alternative to chemical fertiliser for tomato crop. Plant Growth Regul 102, 251–266 (2024). https://doi.org/10.1007/s10725-023-00993-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-023-00993-3