Abstract
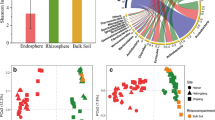
The bacterial microbiota inhabiting the endosphere and rhizoplane regulate plant growth. The mutualistic interaction between sweet sorghum and soil bacteria has drawn increasing research attention. Nevertheless, the root-inhabiting bacterial microbiota of sweet sorghum’s perennial analog have rarely been characterized. Here, the root-inhabiting bacterial microbiota of the perennial sweet sorghum cultivar NaPBS778 (N778 simply) and its control TP60 were discovered at the flowering and maturing stages under field growth by high-throughput amplicon sequencing of the 16S rRNA gene via Illumina MiSeq. Nearly all alpha diversity indices of aerial and primary root samples of N778 were not significantly distinct from those of TP60 at the maturing stage, except for the observed species (Sobs) and phylogenetic diversity indices. The beta diversity of aerial and primary root samples showed no significant differences between N778 and its control TP60 at the maturing stage. Moreover, the bacterial microbiota in N778 aerial and primary roots was not only predominated by Proteobacteria, Actinobacteria, and Bacteroidetes at the phylum level but also strikingly distinct from the bacterial microbiota in rhizosphere soil at the genus level. Additionally, the root samples of N778 at the maturing stage were considerably enriched with OTU1262 being a potential cold-adapted bacterium belonging to Pseudarthrobacter, OTU434 plus OTU1304 belonging to Streptomyces and associated with crop nitrogen stress-tolerance, and OTU836 belonging to the family Oxalobacteraceae and potentially promoting crop growth. Our findings suggest that the perennial sweet sorghum cultivar N778 may recruit potentially cold-tolerant, plant growth-promoting, and nitrogen stress-tolerant bacterial taxa into roots at the maturing stage.






Similar content being viewed by others
References
Abd El-Daim IA, Bejai S, Meijer J (2019) Bacillus velezensis 5113 Induced Metabolic and Molecular Reprogramming during Abiotic Stress Tolerance in Wheat. Sci Rep 9:16282. https://doi.org/10.1038/s41598-019-52567-x
Ali S, Duan J, Charles TC, Glick BR (2014) A bioinformatics approach to the determination of genes involved in endophytic behavior in Burkholderia spp. J Theor Biol 343:193–198. https://doi.org/10.1016/j.jtbi.2013.10.007
Appiah-Nkansah NB, Li J, Rooney W, Wang DH (2019) A review of sweet sorghum as a viable renewable bioenergy crop and its techno-economic analysis. Renew Energy 143:1121–1132. https://doi.org/10.1016/j.renene.2019.05.066
Balota M, Payne WA, Veeragom SK, Stewart BA, Rosenow DT (2010) Respiration and Its Relationship to Germination, Emergence, and Early Growth Under Cool Temperatures in Sorghum. Crop Sci 50(4):1414–1422. https://doi.org/10.2135/cropsci2009.08.0448
Bandara AY, Weerasooriya DK, Bell TH, Esker PD (2021) Prospects of alleviating early planting-associated cold susceptibility of soybean using microbes: New insights from microbiome analysis. J Agron Crop Sci 207(2):171–185. https://doi.org/10.1111/jac.12476
Beckers B, De Beeck MO, Thijs S, Truyens S, Weyens N, Boerjan W, Vangronsveld J (2016) Performance of 16S rDNA Primer Pairs in the Study of Rhizosphere and Endosphere Bacterial Microbiomes in Metabarcoding Studies. Front Microbiol 7:650. https://doi.org/10.3389/fmicb.2016.00650
Beirinckx S, Viaene T, Haegeman A, Debode J, Amery F, Vandenabeele S, Nelissen H, Inze D, Tito R, Raes J, De Tender C, Goormachtig S (2020) Tapping into the maize root microbiome to identify bacteria that promote growth under chilling conditions. Microbiome 8(1):54. https://doi.org/10.1186/s40168-020-00833-w
Bekele WA, Fiedler K, Shiringani A, Schnaubelt D, Windpassinger S, Uptmoor R, Friedt W, Snowdon RJ (2014) Unravelling the genetic complexity of sorghum seedling development under low-temperature conditions. Plant Cell Environ 37(3):707–723. https://doi.org/10.1111/pce.12189
Bulgarelli D, Rott M, Schlaeppi K, van Themaat EVL, Ahmadinejad N, Assenza F, Rauf P, Huettel B, Reinhardt R, Schmelzer E, Peplies J, Gloeckner FO, Amann R, Eickhorst T, Schulze-Lefert P (2012) Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota. Nature 488(7409):91–95. https://doi.org/10.1038/nature11336
Bulgarelli D, Schlaeppi K, Spaepen S, van Themaat EVL, Schulze-Lefert P (2013) Structure and Functions of the Bacterial Microbiota of Plants. Annu Rev Plant Biol 64:807–838. https://doi.org/10.1146/annurev-arplant-050312-120106
Bulgarelli D, Garrido-Oter R, Munch PC, Weiman A, Droge J, Pan Y, McHardy AC, Schulze-Lefert P (2015) Structure and Function of the Bacterial Root Microbiota in Wild and Domesticated Barley. Cell Host Microbe 17(3):392–403. https://doi.org/10.1016/j.chom.2015.01.011
Burke JJ, Emendack Y, Hayes C, Xin ZG, Burow G (2019) Registration of Five Grain Sorghum Lines with Early-Season Seedling Cold Tolerance. J Plant Regist 13(3):422–426. https://doi.org/10.3198/jpr2019.02.0010crg
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ, Fierer N, Knight R (2011) Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci USA 108:4516–4522. https://doi.org/10.1073/pnas.1000080107
Casto AL, Murphy KM, Gehan MA (2021) Coping with cold: Sorghum cold stress from germination to maturity. Crop Sci 61(6):3894–3907. https://doi.org/10.1002/csc2.20609
Chen SF, Zhou YQ, Chen YR, Gu J (2018) Fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34(17):884–890. https://doi.org/10.1093/bioinformatics/bty560
Chopra R, Burow G, Burke JJ, Gladman N, Xin ZG (2017) Genome-wide association analysis of seedling traits in diverse Sorghum germplasm under thermal stress. BMC Plant Biol 17:12. https://doi.org/10.1186/s12870-016-0966-2
Chumpitaz-Segovia C, Alvarado D, Ogata-Gutierrez K, Zuniga-Davila D (2020) Bioprospection of native psychrotolerant plant-growth-promoting rhizobacteria from Peruvian Andean Plateau soils associated with Chenopodium quinoa. Can J Microbiol 66(11):641. https://doi.org/10.1139/cjm-2020-0036
Cloutier M, Chatterjee D, Elango D, Cui J, Bruns MA, Chopra S (2021) Sorghum Root Flavonoid Chemistry, Cultivar, and Frost Stress Effects on Rhizosphere Bacteria and Fungi. Phytobiomes J 5(1):39–50. https://doi.org/10.1094/pbiomes-01-20-0013-fi
Cox S, Nabukalu P, Paterson AH, Kong W, Nakasagga S (2018) Development of Perennial Grain Sorghum. Sustainability 10(1):172. https://doi.org/10.3390/su10010172
da Silva JF, da Silva TR, Escobar IEC, Fraiz ACR, dos Santos JWM, do Nascimento TR, dos Santos JMR, Peters SJW, de Melo RF, Signor D, Fernandes PI (2018) Screening of plant growth promotion ability among bacteria isolated from field-grown sorghum under different managements in Brazilian drylands. World J Microbiol Biotechnol 34(12):186. https://doi.org/10.1007/s11274-018-2568-7
Dong W, Su S, You L, Huang S, Qi J, Lu G, Huang Y, Yang Y (2013) QTLs analysis of tillers number in F_6 sorghum population. J Nanjing Forestry Univ (NSE) 37(2):55–58
Edgar RC (2013) UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10(10):996–998. https://doi.org/10.1038/Nmeth.2604
Edwards J, Johnson C, Santos-Medellin C, Lurie E, Podishetty NK, Bhatnagar S, Eisen JA, Sundaresan V (2015) Structure, variation, and assembly of the root-associated microbiomes of rice. Proc Natl Acad Sci USA 112(8):E911–920. https://doi.org/10.1073/pnas.1414592112
Fazal A, Wen ZL, Lu YT, Hua XM, Yang MK, Yin TM, Han HW, Lin HY, Wang XM, Lu GH, Qi JL, Yang YH (2020) Assembly and shifts of the bacterial rhizobiome of field grown transgenic maize line carrying mcry1Ab and mcry2Ab genes at different developmental stages. Plant Growth Regul 91(1): 113–126. https://doi.org/10.1007/s10725-020-00591-7
Fernandez O, Theocharis A, Bordiec S, Feil R, Jacquens L, Clement C, Fontaine F, Barka EA (2012) Burkholderia phytofirmans PsJN Acclimates Grapevine to Cold by Modulating Carbohydrate Metabolism. Mol Plant-Microbe Interact 25(4):496–504. https://doi.org/10.1094/mpmi-09-11-0245
Habyarimana E, Lorenzoni C, Redaelli R, Alfieri M, Amaducci S, Cox S (2018) Towards a perennial biomass sorghum crop: A comparative investigation of biomass yields and overwintering of Sorghum bicolor x S-halepense lines relative to long term S-bicolor trials in northern Italy. Biomass Bioenergy 111: 187–195. https://doi.org/10.1016/j.biombioe.2017.03.004
Heijo G, Taule C, Mareque C, Stefanello A, Souza EM, Battistoni F (2021) Interaction among endophytic bacteria, sweet sorghum (Sorghum bicolor) cultivars and chemical nitrogen fertilization. FEMS Microbiol Ecol 97(2):fiaa245. https://doi.org/10.1093/femsec/fiaa245
Hu HL, Li M, Wang GX, Drosos M, Li Z, Hu ZY, Xi BD (2019) Water-soluble mercury induced by organic amendments affected microbial community assemblage in mercury-polluted paddy soil. Chemosphere 236:12445. https://doi.org/10.1016/j.chemosphere.2019.124405
Huse SM, Dethlefsen L, Huber JA, Mark WD, Relman DA, Sogin ML (2008) Exploring microbial diversity and taxonomy using SSU rRNA hypervariable tag sequencing. PLoS Genet 4(11):e1000255. https://doi.org/10.1371/journal.pgen.1000255
Iniguez AL, Dong YM, Carter HD, Ahmer BMM, Stone JM, Triplett EW (2005) Regulation of enteric endophytic bacterial colonization by plant defenses. Mol Plant-Microbe Interact 18(2):169–178. https://doi.org/10.1094/mpmi-18-0169
Irizarry I, White JF (2017) Application of bacteria from non-cultivated plants to promote growth, alter root architecture and alleviate salt stress of cotton. J Appl Microbiol 122(4):1110–1120. https://doi.org/10.1111/jam.13414
Irizarry I, White JF (2018) Bacillus amyloliquefaciens alters gene expression, ROS production and lignin synthesis in cotton seedling roots. J Appl Microbiol 124(6):1589–1603. https://doi.org/10.1111/jam.13744
Kakar KU, Ren XL, Nawaz Z, Cui ZQ, Li B, Xie GL, Hassan MA, Ali E, Sun GC (2016) A consortium of rhizobacterial strains and biochemical growth elicitors improve cold and drought stress tolerance in rice (Oryza sativa L.). Plant Biol 18(3):471–483. https://doi.org/10.1111/plb.12427
Lamers J, van der Meer T, Testerink C (2020) How Plants Sense and Respond to Stressful Environments. Plant Physiol 182(4):1624–1635. https://doi.org/10.1104/pp.19.01464
Lata R, Chowdhury S, Gond SK, White JF (2018) Induction of abiotic stress tolerance in plants by endophytic microbes. Lett Appl Microbiol 66(4):268–276. https://doi.org/10.1111/lam.12855
Li MY, Wang JL, Yao T, Wang ZL, Zhang HR, Li CN (2021) Isolation and Characterization of Cold-Adapted PGPB and Their Effect on Plant Growth Promotion. J Microbiol Biotechnol 31(9):1218–1230. https://doi.org/10.4014/jmb.2105.05012
Liu HW, Carvalhais LC, Crawford M, Singh E, Dennis PG, Pieterse CMJ, Schenk PM (2017) Inner Plant Values: Diversity, Colonization and Benefits from Endophytic Bacteria. Front Microbiol 8. https://doi.org/10.3389/fmicb.2017.02552. Article 2552
Lopes LD, Chai YN, Marsh EL, Rajewski JF, Dweikat I, Schachtman DP (2021) Sweet Sorghum Genotypes Tolerant and Sensitive to Nitrogen Stress Select Distinct Root Endosphere and Rhizosphere Bacterial Communities. Microorganisms 9(6):e1329. https://doi.org/10.3390/microorganisms9061329
Lu G-H, Tang C-Y, Hua X-M, Cheng J, Wang G-H, Zhu Y-L, Zhang L-Y, Shou H-X, Qi J-L, Yang Y-H (2018a) Effects of an EPSPS-transgenic soybean line ZUTS31 on root-associated bacterial communities during field growth. PLoS ONE 13(2):e0192008. https://doi.org/10.1371/journal.pone.0192008
Lu G-H, Na Z, Zijian Z, Cao R, Fazal A, Huang S, Zhang H, Chen J, Cheng R, Li Y, Yang Y, Yang H, Na Z-Y (2022) The perennial sweet sorghum cultivar and its recruiting rhizosphere dominant bacterial taxa under field growth. Pak J Bot 54(3):865–878. https://doi.org/10.30848/PJB2022-3(46)
Lundberg DS, Lebeis SL, Paredes SH, Yourstone S, Gehring J, Malfatti S, Tremblay J, Engelbrektson A, Kunin V, del Rio TG, Edgar RC, Eickhorst T, Ley RE, Hugenholtz P, Tringe SG, Dangl JL (2012) Defining the core Arabidopsis thaliana root microbiome. Nature 488(7409):86–90. https://doi.org/10.1038/nature11237
Magoc T, Salzberg SL (2011) FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27(21):2957–2963. https://doi.org/10.1093/bioinformatics/btr507
Mareque C, da Silva TF, Vollu RE, Beracochea M, Seldin L, Battistoni F (2018) The Endophytic Bacterial Microbiota Associated with Sweet Sorghum (Sorghum bicolor) Is Modulated by the Application of Chemical N Fertilizer to the Field. Int J Genomics 2018: 7403670. https://doi.org/10.1155/2018/7403670
Marla SR, Shiva S, Welti R, Liu SZ, Burke JJ, Morris GP (2017) Comparative Transcriptome and Lipidome Analyses Reveal Molecular Chilling Responses in Chilling-Tolerant Sorghums. Plant Genome 10(3):25. https://doi.org/10.3835/plantgenome2017.03.0025
Marla SR, Burow G, Chopra R, Hayes C, Olatoye MO, Felderhoff T, Hu ZB, Raymundo R, Perumal R, Morris GP (2019) Genetic Architecture of Chilling Tolerance in Sorghum Dissected with a Nested Association Mapping Population. G3-Genes. Genomes Genet 9(12):4045–4057. https://doi.org/10.1534/g3.119.400353
Maulana F, Tesso TT (2013) Cold Temperature Episode at Seedling and Flowering Stages Reduces Growth and Yield Components in Sorghum. Crop Sci 53(2):564–574. https://doi.org/10.2135/cropsci2011.12.0649
Menamo T, Kassahun B, Borrell AK, Jordan DR, Tao Y, Hunt C, Mace E (2021) Genetic diversity of Ethiopian sorghum reveals signatures of climatic adaptation. Theor Appl Genet 134(2):731–742. https://doi.org/10.1007/s00122-020-03727-5
Mishra PK, Bisht SC, Ruwari P, Selvakumar G, Joshi GK, Bisht JK, Bhatt JC, Gupta HS (2011) Alleviation of cold stress in inoculated wheat (Triticum aestivum L.) seedlings with psychrotolerant Pseudomonads from NW Himalayas. Arch Microbiol 193(7):497–513. https://doi.org/10.1007/s00203-011-0693-x
Naveed M, Hussain MB, Zahir ZA, Mitter B, Sessitsch A (2014) Drought stress amelioration in wheat through inoculation with Burkholderia phytofirmans strain PsJN. Plant Growth Regul 73(2):121–131. https://doi.org/10.1007/s10725-013-9874-8
Ortiz D, Hu JY, Fernandez MGS (2017) Genetic architecture of photosynthesis in Sorghum bicolor under non-stress and cold stress conditions. J Exp Bot 68(16):4545–4557. https://doi.org/10.1093/jxb/erx276
Peiffer JA, Spor A, Koren O, Jin Z, Tringe SG, Dangl JL, Buckler ES, Ley RE (2013) Diversity and heritability of the maize rhizosphere microbiome under field conditions. Proc Natl Acad Sci USA 110(16):6548–6553. https://doi.org/10.1073/pnas.1302837110
Price MN, Dehal PS, Arkin AP (2010) FastTree 2-Approximately Maximum-Likelihood Trees for Large Alignments. PLoS ONE 5(3):e9490. https://doi.org/10.1371/journal.pone.0009490
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glockner FO (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41(D1):D590–D596. https://doi.org/10.1093/Nar/Gks1219
Reddy PS, Gomashe S (2017) Can cold tolerance bring in hybrids on commercial front in winter sorghum? Curr Sci 113(2):307–313. https://doi.org/10.18520/cs/v113/i02/307-313
Rutayisire A, Lubadde G, Mukayiranga A, Edema R (2021) Response of Sorghum to Cold Stress at Early Developmental Stage. Int J Agron 2021: 8875205. https://doi.org/10.1155/2021/8875205
Schultz CR, Brantley KM, Wallace JG (2022) The role of genetic variation in Zea mays response to beneficial endophytes. Plant Growth Regul 98(1):167–177. https://doi.org/10.1007/s10725-022-00842-9
Shin Y, Lee BH, Lee KE, Park W (2020) Pseudarthrobacter psychrotolerans sp. nov., a cold-adapted bacterium isolated from Antarctic soil. Int J Syst Evol Microbiol 70(12):6106–6114. https://doi.org/10.1099/ijsem.0.004505
Subramanian P, Krishnamoorthy R, Chanratana M, Kim K, Sa T (2015) Expression of an exogenous 1-aminocyclopropane-1-carboxylate deaminase gene in psychrotolerant bacteria modulates ethylene metabolism and cold induced genes in tomato under chilling stress. Plant Physiol Biochem 89:18–23. https://doi.org/10.1016/j.plaphy.2015.02.003
Sun AQ, Jiao XY, Chen QL, Wu AL, Zheng Y, Lin YX, He JZ, Hu HW (2021) Microbial communities in crop phyllosphere and root endosphere are more resistant than soil microbiota to fertilization. Soil Biol Biochem 153:108113. https://doi.org/10.1016/j.soilbio.2020.108113
Tapia-Vazquez I, Sanchez-Cruz R, Arroyo-Dominguez M, Lira-Ruan V, Sanchez-Reyes A, Sanchez-Carbente MD, Padilla-Chacon D, Batista-Garcia RA, Folch-Mallol JL (2020) Isolation and characterization of psychrophilic and psychrotolerant plant-growth promoting microorganisms from a high-altitude volcano crater in Mexico. Microbiol Res 232:126394. https://doi.org/10.1016/j.micres.2019.126394
Theocharis A, Bordiec S, Fernandez O, Paquis S, Dhondt-Cordelier S, Baillieul F, Clement C, Barka EA (2012) Burkholderia phytofirmans PsJN Primes Vitis vinifera L. and Confers a Better Tolerance to Low Nonfreezing Temperatures. Mol Plant-Microbe Interact 25(2):241–249. https://doi.org/10.1094/mpmi-05-11-0124
Tiryaki D, Aydin I, Atici O (2019) Psychrotolerant bacteria isolated from the leaf apoplast of cold-adapted wild plants improve the cold resistance of bean (Phaseolus vulgaris L.) under low temperature. Cryobiology 86:111–119. https://doi.org/10.1016/j.cryobiol.2018.11.001
Verma SK, Kingsley K, Irizarry I, Bergen M, Kharwar RN, White JF (2017) Seed-vectored endophytic bacteria modulate development of rice seedlings. J Appl Microbiol 122(6):1680–1691. https://doi.org/10.1111/jam.13463
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microb 73(16):5261–5267. https://doi.org/10.1128/Aem.00062-07
White JF, Kingsley KI, Kowalski KP, Irizarry I, Micci A, Soares MA, Bergen MS (2018) Disease protection and allelopathic interactions of seed-transmitted endophytic pseudomonads of invasive reed grass (Phragmites australis). Plant Soil 422(1–2):195–208. https://doi.org/10.1007/s11104-016-3169-6
Woldesemayat AA, Modise DM, Gemeildien J, Ndimba BK, Christoffels A (2018) Cross-species multiple environmental stress responses: An integrated approach to identify candidate genes for multiple stress tolerance in sorghum (Sorghum bicolor (L.) Moench) and related model species. PLoS ONE 13(3):e0192678. https://doi.org/10.1371/journal.pone.0192678
Xu N, Tan GC, Wang HY, Gai XP (2016) Effect of biochar additions to soil on nitrogen leaching, microbial biomass and bacterial community structure. Eur J Soil Biol 74:1–8. https://doi.org/10.1016/j.ejsobi.2016.02.004
Yu JM, Tuinstra MR (2001) Genetic analysis of seedling growth under cold temperature stress in grain sorghum. Crop Sci 41(5):1438–1443. https://doi.org/10.2135/cropsci2001.4151438x
Yu J, Tuinstra MR, Claassen MM, Gordon WB, Witt MD (2004) Analysis of cold tolerance in sorghum under controlled environment conditions. Field Crop Res 85(1):21–30. https://doi.org/10.1016/s0378-4290(03)00125-4
Yu P, He XM, Baer M, Beirinckx S, Tian T, Moya YAT, Zhang XC, Deichmann M, Frey FP, Bresgen V, Li CJ, Razavi BS, Schaaf G, von Wiren N, Su Z, Bucher M, Tsuda K, Goormachtig S, Chen XP, Hochholdinger F (2021) Plant flavones enrich rhizosphere Oxalobacteraceae to improve maize performance under nitrogen deprivation. Nat Plants 7(4):481–499. https://doi.org/10.1038/s41477-021-00897-y
Zegada-Lizarazu W, Luna DF, Monti A (2016) Differential characteristics of photochemical acclimation to cold in two contrasting sweet sorghum hybrids. Physiol Plant 157(4):479–489. https://doi.org/10.1111/ppl.12430
Zheng L-Y, Guo X-S, He B, Sun L-J, Peng Y, Dong S-S, Liu T-F, Jiang S, Ramachandran S, Liu C-M, Jing H-C (2011) Genome-wide patterns of genetic variation in sweet and grain sorghum (Sorghum bicolor). Genome Biol 12(11):R114. https://doi.org/10.1186/gb-2011-12-11-r114
Acknowledgements
We are grateful to Yunting Lu (Nanjing University, NJU), Yinsong Wang (NJU), Zijian Zhang (HNU), Jialing Chen (HNU), Yan Li (HNU) and Yingming Pu (YEARI) for assisting with sampling. We thank the data supporting from “Soil Science Data Center, National Earth System Science Data Sharing Infrastructure, National Science & Technology Infrastructure of China. (http://soil.geodata.cn/). Analysis of amplicon data was conducted via the online Majorbio Cloud Platform (www.majorbio.com).
Funding
This work was funded by the Xiangyu Talent research launch project (31LGH00 and 31CR001).
Author information
Authors and Affiliations
Contributions
Gui-Hua Lu and Zhong-Yuan Na conceived and designed the experiments; Gui-Hua Lu, Rui Cao, and Zhiye Na performed the experiments; Gui-Hua Lu and Kezhi Zheng analyzed the data; Yonghua Yang, Bo Sun, and Hongjun Yang provided resources; and Gui-Hua Lu and Aliya Fazal wrote the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest. The funding sponsors had no impact on the design of the experiments, the study, the data interpretation, the manuscript’s writing, or the decision to publish the results.
Ethical approval
This article does not contain any research on human participants or animals performed by any authors.
Additional information
Communicated by Hang-Wei Hu.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lu, GH., Cao, R., Fazal, A. et al. Composition and diversity of root-inhabiting bacterial microbiota in the perennial sweet sorghum cultivar at the maturing stage. Plant Growth Regul 99, 567–582 (2023). https://doi.org/10.1007/s10725-022-00929-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-022-00929-3