Abstract
Chlorophyll (Chl) content is an important agronomic trait directly affecting the photosynthetic rate. Using a high-density genetic map of 132 recombinant inbred lines (RILs) derived from the cross between 93-11 and PA64s, we detected the quantitative trait loci (QTLs) for Chl content of the top three leaves under two nitrogen (N) conditions at two developmental stages. A total of 32 main-effect QTLs located on chromosomes 1, 4, 5, 6, 7, 8, and 12 were identified, and these QTLs individually accounted for 6.0–20.8 % of the total phenotypic variation. A major QTL qFCC7 L affecting the Chl content under low N condition was identified, and its positive allele came from PA64s. This QTL might be associated with the ability to tolerate low-N stress in rice. The chromosomal segment substitution line (CSSL) with the corresponding segment from PA64s had a higher SPAD value and photosynthetic rate than 93-11 and showed a lower specific leaf area (SLA). We performed a fine-mapping using a BC4F2 population via marker-assisted backcross and finally mapped this QTL to a 124.5 kb interval on the long arm of chromosome 7. Candidate gene analysis showed that there were sequence variations and expression differences in the predicted candidate gene between the two parents. These results suggest that the QTL qFCC7 L may be useful for breeding the rice varieties with higher photosynthetic rate and grain yield.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chlorophyll (Chl) is the most important photosynthetic pigment, and Chl content is a critical breeding trait because it is positively correlated with photosynthesis (Xu et al. 1997; Teng et al. 2004; Hu et al. 2009; Zhang et al. 2009; Wang et al. 2016). Previous reports have demonstrated that high-yielding varieties have a higher leaf Chl content and photosynthetic rate than the previously released low-yielding lines, particularly at heading stage (Peng et al. 2008). Increasing Chl content may be an effective way to increase dry matter production and grain yield in rice. Chl content is strongly affected by nitrogen (N) supply because N is a major component of Chl. In recent years, to obtain the maximum grain yield, rice varieties are selected to allow high chemical fertilizers input, including N input (Tong et al. 2006). However, excessive N supply will result in lower N use efficiency, which causes higher cost for agricultural production and environmental hazards (Tong et al. 2006; Kaushal et al. 2011; Hashim et al. 2015; Zhou et al. 2016). Insufficient N supply may lead to low Chl content, small leaves and low biomass production, which will consequently reduce grain yield and quality (Prasertsak and Fukai 1997; Adhikari et al. 1999). Increasing leaf Chl content under low N supply condition may be an effective way to improve N utilization efficiency.
Previous studies suggested that Chl content is controlled by multi-genes, usually called quantitative trait loci (QTL) (Wang et al. 2008). In recent years, with the rapid development of molecular markers and the application of high-resolution linkage map, some QTLs associated with Chl content have been identified. Wu and Luo (1996) detected three main-effect QTLs on chromosomes 2, 4, and 7 and one epistatic locus controlling Chl content using an F2 population of Palawan/IR42. Wang et al. (2003) discovered six QTLs associated with the contents of Chl a and Chl b on chromosomes 1, 4, and 11. Teng et al. (2004) detected three QTLs for Chl content on chromosomes 1, 3, and 8 using a double-haploid population of ZYQ8/JX17. Seven QTLs related to Chl content at tillering and heading stage were identified and mapped to chromosomes 1, 2, 3, 4, 6, and 8 using backcross-recombinant inbred lines (Dong et al. 2007). In other studies, Zuo et al. (2007) identified ten QTLs associated with Chl content at three different growth stages of rice on chromosomes 1, 3, 4, 5, 7, 11, and 12. Hu et al. (2009) detected 13 main QTLs controlling Chl content under well-watered and water stressed conditions on chromosomes 1, 2, 3, 4, 5, 6, and 10. Jiang et al. (2012) identified 22 QTLs related to Chl content using 94 recombinant inbred lines (RILs) derived from a cross between two japonica cultivars, Lijiangxintuanheigu (LTH) and Shennong 265 (SN265), and these QTLs distributed over all chromosomes except chromosome 5. Zhang et al. (2014) discovered five QTLs controlling Chl content on chromosomes 1, 4, 7, 12 and two QTLs on chromosomes 4 and 12 at heading and maturity stages, respectively. On the whole, these QTLs distributed on all chromosomes, mainly on chromosomes 1, 2, 3, 4 and 11. In these previous researches, some QTLs can be detected at several growth stages but the other QTLs can only be detected at a specific developmental stage (Wang et al. 2003; Zuo et al. 2007).
In a previous study, a high-density SNP-based genetic linkage map has been constructed using 132 RILs derived from a cross between 93-11 and Pei-Ai 64s (PA64s) (Gao et al. 2013). 93-11 is an indica restorer line, and PA64s is a photo-thermo-sensitive male sterile line with higher Chl content than 93-11. The pioneer two-line super hybrid rice variety, Liangyoupeijiu (LYP9), is derived from the cross between these two parents and has greatly increased yield.
The top three leaves of rice are the most important organs for photosynthesis and the main source of carbohydrates production (Foyer 1987). In the current study, using the genetic linkage map, we discovered 32 QTLs related to the Chl content of the top three leaves under two N conditions at two developmental stages. Among these QTLs, we identified a major-effect QTL, qFCC7 L controlling Chl content under low N condition. A chromosomal segment substitution line (CSSL7) was used to confirm this putative QTL. We performed fine mapping using a BC4F2 population. We also analyzed the candidate genes, which could provide a reference for cloning the target gene related to Chl content.
Materials and methods
Plant materials and genetic map
A total of 132 RILs from LYP9 and the high-density linkage map for this population were used in this study. The linkage map was constructed by using single nucleotide polymorphism (SNP) markers by Gao et al. (2013). For the development of CSSL7, an RIL containing the PA64s alleles at qFCC7 L region was selected by target markers and then successively backcrossed with 93-11 for four generations. Three markers M1, RM234, and RM21981 were used for marker-assisted selection of each generation. To avoid the interference from other QTLs, we developed some markers to identify the genotypes of the regions which also contained our target QTLs.
Field experiment and trait measurement
The 132 RILs and two parental lines were grown in Hangzhou, China (May, 2010) under two N conditions (LN, without N fertilizer; HN, with 150 kg/hm2 N fertilizer). The two parental lines and CSSL7 were grown in Hainan, China in 2013. The BC4F2 population with 3562 individuals was grown in Hangzhou (May, 2013) at LN condition. The plants of each line at the 25-day-old seedling age were transplanted into a four-row plot with six plants per row and a spacing of 20 cm × 20 cm.
Rice leaf Chl content exhibited dynamic change. Previous reports demonstrate that the growth rate around heading stage is significantly associated with the final yield in rice (Takai et al. 2006; Horie et al. 2006). Heading stage is an important period in the life cycle of rice because photosynthetic products are transported from source to the bank at this period. Recent studies discovered some QTLs controlling Chl content only at specific developmental stages (Zuo et al. 2007). Based on these research findings, we measured leaf Chl content at initial heading stage (a period between the end of booting stage and heading stage, when the flag leaf is fully expanded and flag leaf sheath is opened but the tip of inflorescence is not emerged from sheath) and heading stage. The Chl contents of the first leaf (FL), second leaf (SL) and third leaf (TL) from the top were evaluated by Soil and Plant Analysis Development (SPAD) meter (SPAD-502, Minolta Camera Co., Osaka, Japan), a portable device which provides a nondestructively method for estimating leaf Chl content (Chubachi et al. 1986). In some studies, leaf Chl content is presented as SPAD value which can also be used in estimating leaf photosynthesis (A-JIA et al. 2008; Takai et al. 2010). Three randomly selected plants from the middle of a row in each rice line were tested. For each observed leaf, SPAD values were measured in the upper, middle, and base sections of the leaf on main panicle, and the mean SPAD value of the leaf sections served as the phenotypical value of SPAD (Zhang et al. 2014). The Chl content (mg dm−2) of leaves was calculated as follows: Y = 0.0996 × −0.152 (Zhang et al. 2014), where X is the mean SPAD value measured by the Chl meter, and Y is the actual leaf Chl content. Photosynthetic rate of leaves was measured by a portable photosynthesis measuring system (LI-6499XT, http://www.ecotek.com.cn). The specific leaf area (SLA, cm2 g−1) of leaves was calculated as follows (Li et al. 2005a): SLA = S/G, S meant the leaf area measured by the leaf area meter, and G represented leaf dry weight.
Data analysis and QTL analysis
Phenotypic variations were analyzed using the SAS (Statistical Analysis System) V8.01 software. QTL analysis was performed using the MultiQTL1.6 package (http://www.multiqtl.com) with maximum likelihood interval mapping. The LOD threshold for each dataset was set based on a permutation test (1000 permutations, P = 0.05). Major-effect QTLs were accepted when their LOD scores were larger than 2.5. QTLs were named according to McCouch et al. (1997).
Mapping population
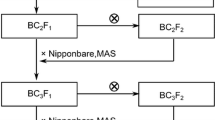
To fine map the major QTL qFCC7 L , a backcross population was constructed in our study. The RILs with PA64s genotype in the qFCC7 L region and most of the genetic background from 93 to 11 (based on the genotype of RILs) were selected to successively backcross with 93-11. Additionally, two markers M1 and RM234 (Table S1) were used for marker-assisted selection of each generation. As a result, a BC4F1 line with 93-11 genetic background exhibiting heterozygous (identified by the markers M1 and RM234) at the qFCC7 L region was constructed in Hainan. After self-crossing, we obtained a BC4F2 population for fine mapping of qFCC7 L .
Fine mapping of qFCC7 L
SSR markers were obtained from the Gramene database (http://www.gramene.org/). Additionally, we designed the insertions/deletions (InDels) markers M1, M2, and M3 within the target region on chromosome 7 using the InDels identified between 93-11 and PA64s (Gao et al. 2013) by two software, Primer Premier 5.0 and Oligo 7. All relevant primers are listed in Table S1. Total genomic DNA was extracted from young leaves using the CTAB method (Murray and Thompson 1980; Doyle and Doyle 1987).
RNA extraction and expression pattern analysis
Quantitative real-time polymerase chain reaction (qRT-PCR) was performed to analyze the expression pattern of the peptide transporter gene OsPTR4. Total RNA was isolated at heading stage using an RNA extraction kit (Axygen), and the first-strand cDNA was synthesized using the ReverTra Ace qRT-PCR RT Kit (TOYOBO CO. LTD.; http://www.toyobo.cn/) following the manufacturer’s instructions. The primers used in qRT-PCR analysis were designed using an online tool (https://www.genscript.com/ssl-bin/app/primer). The primers for amplification of the internal reference OsActin1 were 5′-CCATTGGTGCTGAGCGTTT-3′ (forward) and 5′-CGCAGCTTCCATTCCTATGAA-3′ (reverse). The primers for amplification of OsPTR4 were 5′-CAAGATTGGCTCACTCTCCA-3′ (forward) and 5′-AACCCAGAATGTCACGCATA-3′ (reverse). Each 10 μL reaction system consisted of (1) μL total cDNA, 5 μL of 2× SYBR Green PCR Master Mix (TaKaRa Co. LTD., Otsu, Japan; http://www.takara.com.cn), (2) μL of each 1 μM primer. The analysis of each sample was performed with four replicates. Reactions were run on the ABI PRISM®7900HT Sequence Detector (Applied Biosystems). The 2− ∆∆CT method (Livak and Schmittgen 2001) was adopted to analyze the relative expression level, and the two-tailed t test was used to analyze statistically significant difference.
Results
Performance of SPAD value in RILs and parents
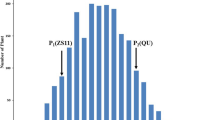
The phenotypic differences between 93-11 and PA64s as well as the leaf SPAD value variations in the RIL population under two N conditions at two developmental stages are summarized in Fig. 1 and Table S2. The top three leaves of 93-11 showed lower SPAD values when compared with those of PA64s (Table S2). The SPAD values of the top three leaves in RILs all showed continuous distribution under LN and HN conditions at two different developmental stages. In addition, some lines showed apparent transgressive variations in the RIL population (Fig. 1). These results indicated that the Chl contents of the top three leaves were controlled by multiple genes, and the data satisfied the demands for QTL analysis.
Distribution of SPAD values of the top three leaves in the RIL population. a–c SPAD values of the top three leaves surveyed at initial heading stage. d–f SPAD values of the top three leaves surveyed at heading stage. White columns indicate the plants grown at low N condition. Grey columns indicate the plants grown at high N condition. Arrows with smooth end indicate 93-11, arrows with notch end indicate PA64s
QTL analysis for Chl content
A total of 32 QTLs related to Chl content of the top three leaves under two N levels at two stages were detected (Fig. 2; Table S3). These QTLs were distributed on chromosomes 1, 4, 5, 6, 7, 8, and 12, with LOD scores between 2.51 and 9.46 and accounted for 6.0–20.8 % of the total phenotypic variation. Among these QTLs, 19 QTLs were clustered in three intervals on chromosomes 4, 6, and 7 (Fig. 2). Six QTLs were detected between the SNP markers SNP4-246 and SNP4-271, explaining 10.3–20.8 % of the phenotypic variations; seven QTLs were located in the region between SNP6-90 and SNP6-106, explaining 8.6–20.5 % of the phenotypic variations; six QTLs were detected between SNP7-160 and SNP7-192, explaining 6.0–13.9 % of the phenotypic variations. We considered that these 19 QTLs were actually consisted of three loci. The positive effect of most QTLs came from the alleles of PA64s, the parent with high Chl content (Table S3).
Locations of main-effect QTLs for chlorophyll content (SPAD value) in the RIL population of 93-11 and PA64s. QTLs detected in low N condition are designated with shapes used light color and QTLs identified in high N condition are marked with shapes used dark color. The number on the left of each chromosome is the marker’s genetic distance (cM)
Fine mapping of qFCC7 L
Among these QTLs, we discovered that the QTL qFCC7 L controlled the Chl contents of the top three leaves under LN condition at initial heading stage and the Chl contents of FL and TL under LN condition at heading stage. This QTL explained 13.9 % of the total phenotypic variation of the FL Chl content, and the PA64s allele could increase the Chl content. We confirmed this QTL by the CSSL7 with 93-11 genetic background but containing a PA64s chromosome segment at qFCC7 L region (Fig. S1). The CSSL7 had a higher SPAD value and photosynthetic rate than 93-11 (Fig. 3a, b) and showed a lower SLA (Fig. 3c), suggesting that this locus may also influence photosynthesis and leaf thickness. To fine map this major-effect QTL, we constructed a BC4F2 population. Previous studies demonstrated that at least 50 % of the photosynthetic products for final yield are provided by flag leaf (Li et al. 1998). So we used the SPAD value of FL in BC4F2 population as the reference indicator for mapping. Three InDels and six SSR markers were used for fine mapping of qFCC7 L . After analyzing the genotypes and phenotypes of 3562 individuals in BC4F2 population, we finally positioned qFCC7 L between the markers M3 and RM21938 in a 124.5 kb interval on the long arm of chromosome 7 (Fig. 4). According to the Rice Genome Annotation Project Website (http://rice.plantbiology.msu.edu/), there were 16 open reading frames (ORFs) with different predicted biological functions in this region. LOC_Os07g41200 encodes a homolog of Arabidopsis thaliana LONGIFOLIA proteins; LOC_Os07g41240 encodes a putative cytochrome P450 protein; LOC_Os07g41250 encodes a putative peptide transporter protein; LOC_Os07g41260 encodes a putative PPR repeat domain-containing protein. The functions of these four genes were predicted to be related to yield, leaf color and N transport based on previous studies (Wang et al. 2015b; Yang et al. 2013; Fan et al. 2014; Tan et al. 2014).
Agronomic traits of leaves at heading stage. a Chl content of FL, SL, and TL of 93-11, CSSL7, and PA64s. Data was measured from six plants at Hainan. b Photosynthetic rate (PR) of 93-11, CSSL7, and PA64s. c Specific leaf area (SLA) of 93-11, CSSL7, and PA64s. Each bar indicates the mean ± SD of three independent replicates. Asterisks indicate significant difference (Student’s t test, *P < 0.05, **P < 0.01)
Fine mapping for qFCC7 L . The black and white boxes indicate the chromosomal segments from PA64s or 93-11, respectively. ** means the SPAD values of lines L2, L3, L6, and PA64s which contained the qFCC7 L candidate region, have significant difference compared with 93-11 (P < 0.01). Student’s t test was used for statistical analysis, data are shown as mean ± SD, n = 3
Candidate gene identification and expression analysis
The encoded peptide transporter belongs to the nitrate transporter 1/peptide transporter (NRT1/PTR) family which is involved in the uptake and transport of nitrate/peptide and has the functions related to N use efficiency (Fan et al. 2014; Li et al. 2015a). There was a predicted peptide transporter (PTR4) in the mapped interval, so we performed a further analysis of this gene. Deep resequencing showed that OsPTR4 contained five exons and four introns, and sequence variations existed between the parents but mainly in the introns (Fig. 5a). A total of 15 base variations were detected between the parents, including 4 base insertions and 11 base substitutions. Among these variations, four base substitutions caused the changes in three amino acids at the fourth and fifth exons (Fig. 5a). qRT-PCR detected obvious expression variations of OsPTR4 between 93-11 and PA64s during heading stage. The expression levels of OsPTR4 in 93-11 were lower than those in PA64s in all plant tissues (Fig. 5b). The OsPTR4 transcript level in the leaves was the highest, and that in the roots was the second, which were consistent with its uptake and transport capacities. Based on these results, we speculate that the differences in gene structure and expression level lead to the different functions between the parents.
Sequence differences and expression analysis of OsPTR4 between 93-11 and PA64s. a Sequence differences of OsPTR4 between 93-11 and PA64s. ATG and TAA denote start and stop codon. Black boxes indicate exons, discontinuous straight lines between black boxes represent introns. Sequence discrepancies are labeled as shown. Red alphabets represent changed amino acid residues. b Expression analysis of OsPTR4 between 93-11 and PA64s. Each bar indicates the mean ± SD of three independent replicates. Student’s t test was used for statistical analysis. ∗∗ represents significant difference at 0.01 level
Discussion
Chl content is influenced by N supply because there is a certain relationship between N content and Chl content (Wang et al. 2002; Huang et al. 2008). Interestingly, there are very little differences between the Chl contents under zero N input and 150 kg/hm2 N input in the parental lines. The cross between 93-11 and PA64s produces a strong heterosis hybrid rice variety, LYP9, which can absorb more N fertilizer and has a slow reaction to N level. So the parental lines showed little differences in Chl content between two N levels. On the other hand, Chl content is controlled by multi-genes which have some interactions and can reduce the final effect.
To improve fertilizer use efficiency, we hope to dissect its genetic basis using the parents and the derived RILs. For genetic analysis and QTL mapping, we usually choose the parents with obvious phenotype differences. But in this study, the Chl content difference between parents was not very significant. Thus, the selection of parental lines seemed to not match well with the objective of the experiment. However, Li et al. (2005b) have confirmed that QTLs can be identified using the genetic populations without obvious parental differences, and some minor but useful QTLs are thus detected and can be mined via the construction of backcrossing lines (Li et al. 2015b). We have identified 32 such QTLs at two different growth stages. Although we did not observe much difference in Chl content for the parental lines, the presence of such a large number of QTLs indicates that Chl content, particularly in PA64s, is controlled by the interactions of multiple genes. Our study can supplement the discovery of Li et al. (2005b) that minor phenotypic variation may also result from multiple QTL identification. All these findings indicate that the parent PA64s is an elite sterile material for genetic dissection and breeding application.
Photosynthesis is the fundamental factor of dry matter production and grain yield, and high photosynthetic ability is one of the major breeding targets for further improving rice yield. Some studies have demonstrated a positive relationship between leaf photosynthesis and leaf Chl content (Kura-Hotta et al. 1987; Xu et al. 1997; Hu et al. 2009). Using different populations, many QTLs related to Chl content have been reported (Wang et al. 2003; Teng et al. 2004; Jiang et al. 2012). In this study, a mapping population developed from two parental lines, one with high Chl content but low yield potential (PA64s) and another with low Chl content but high yield (93-11), provided a unique opportunity to identify QTLs for the Chl content which contributed to the high heterosis of the pioneer hybrid rice LYP9 developed by crossing these two parents and would benefit the molecular breeding of new hybrid rice genotypes. A total of 32 main-effect QTLs for Chl content were detected (Fig. 2; Table S3), including 13 QTLs discovered under low N level and 19 QTLs identified under high N level. Previous reports have demonstrated that rice yield will remain stagnant with the increasing N utilization. The main reasons causing this phenomenon are unreasonable fertilization and the low use efficiency of N fertilizer (Borrell et al. 1998; Peng et al. 1996). Chl content is closely related to N supply. Under N-stress condition, the variation of Chl content can reflect the difference in N use efficiency of different genotypes. Therefore, the QTLs detected at low N level may contribute to the high tolerance to low N stress. They are useful for understanding the genetic mechanism underlying N use efficiency and reducing the excessive input of N fertilizer. Under N deprivation condition, nitrate reductase activity which affects N use efficiency is associated with the expression level of HvNar1 in barley (Xu et al. 2016). Some QTLs detected in this study were in or near the regions which have been reported previously. The QTLs between SNP1-10 and SNP1-27 are near to the region of qCHM1 (RM490–RM580) reported by Jiang et al. (2012) and HSPAD-1b (RM6902–RM8111) identified by Zhang et al. (2014). The QTLs between SNP4-246 and SNP4-271 are corresponding to qCH4, qCM4b and qCTH4 (RM255–RM559) found by Jiang et al. (2012) and qLSCHL4 (STS4-5–RM349) detected by Zhang et al. (2014). We also detected some new loci, the qSCC5 H between SNP5-203 and SNP5-226 controlling the Chl content of second leaf in high N level at heading stage, the qFCC8 L between SNP8-139 and SNP8-164 explaining 9.4 % of phenotypic variations of the FL Chl content at initial heading stage. We also discovered that the expressions of some QTLs had spatial-temporal specificity, and they could only be detected at a specific time. For example, the QTL between SNP6-106 and SNP6-137 only affected the Chl content at initial heading stage, whereas the QTL located between SNP5-203 and SNP5-226 on chromosome 5 was detected at heading stage. The Chl content of leaves might be affected by the variation of days-to-heading. Wang et al. (2015a) reported that Ghd7 is a major locus for the natural variation of Chl content. We discovered that some genes related to heading time were on the corresponding regions of QTLs for Chl content. In our study, qSCC6 IL /qTCC6 IL was near Se5/OsHY1/OsHO1 which also located on chromosome 6. Another major-effect QTL controlling heading time, DTH8/Ghd8/LHD1, is on the corresponding region of qFCC8 L . Actually, Gao et al. (2013) also detected this QTL for heading date on DTH8 using the same RILs in this study. Recently, Ding et al. (2016) discovered that the transcription levels of two photoperiod-sensitive regulators, Heading date 3a (Hd3a) and RICE FLOWERING LOCUS T 1 (RFT1), were down-regulated in leaves by nitrogen fertilizer. These findings suggested that Chl content and nitrogen fertilizer are closely related to heading time. Liu et al. (2015) identified 27 QTLs related to leaf shape (length and width) and the Chl content of top three leaves across five different environments using a set of CSSLs derived from the cross of 93-11 and PA64s, the same parental lines used in this study. Undoubtedly, some QTLs detected in Liu’s study are at the same regions identified in this research. For example, the QTLs between SNP1-10 and SNP1-27 are at the same region of qFLC1. The QTL between SNP8-38 and SNP8-62 is on the corresponding region of qFLC8-1. However, we also detected some QTLs different from the loci identified in Liu’s study. These results demonstrate that QTL detection is not only influenced by the parents but also limited by the construction of populations, developmental stages and environments.
Among these QTLs, only one QTL affected the Chl content of the top three leaves under low N level at initial heading stage, and it also controlled the Chl contents of FL and TL at heading stage (Fig. 2). Actually, Wu and Luo (1996) also discovered that this QTL can control leaf Chl content under nitrogen stressed condition. Therefore, qFCC7 L may be the same QTL as the previously identified QTLs. The CSSL7 which contained a PA64s chromosome segment in the qFCC7 L region with 93-11 genetic background showed a higher SPAD value than 93-11 (Fig. 3a). High Chl content per unit leaf area may reflect the presence of a larger number of chloroplasts per cell and/or high Chl content per chloroplast when assuming that leaf thicknesses do not change (Takai et al. 2010). However, previous studies have shown that SPAD value may sometimes reflect the variation in leaf thickness because the readings are based on the leaf Chl’s absorption of specific spectral bands which may be influenced by leaf thickness (Peng et al. 1993; Li et al. 2009). SLA is considered to be correlated with leaf thickness and was significantly lower in CSSL7 compared with that in 93-11 (Fig. 3c). These results suggest that the higher SPAD value of CSSL7 may result from its thicker leaves, and the relationship between leaf thickness and SPAD value needs further in-depth studies. Thicker leaves are considered to be important for increasing leaf photosynthesis because they can capture light energy efficiently by more Chl per unit leaf area, and they have sufficient mesophyll surfaces, allowing more efficient CO2 diffusion and transport (Terashima et al. 2006). Thus, the higher photosynthetic rate of CSSL7 may result from the increased Chl content or leaf thickness, or both (Fig. 3b). Although this QTL was only detected in one-year data, CSSL7 also showed a higher SPAD value than 93-11 at Hainan, which demonstrates that this QTL can be stably expressed at different conditions. Using a BC4F2 population, we fine mapped this QTL to a 124.5 Kb region between the markers M3 and RM21938 (Fig. 4), and this interval contained 16 ORFs. In fact, our team has detected a QTL controlling leaf width in this interval (Zhang et al. 2015). But whether this QTL is identical to qFCC7 L needs further study. We also performed a deep analysis of the candidate gene OsPTR4 and discovered the sequence variations and different expression levels between the parents which may lead to the different functions between parents (Fig. 5).
In modern cultivation conditions, the rice varieties with high yield always rely on the condition of high N input. Here, we have detected a major effect QTL qFCC7 L controlling Chl content at low N condition. Therefore, it may be useful for breeding the rice varieties capable of maintaining high leaf Chl content under low N condition. This is a good method to improve N use efficiency. Further researches should be carried out to confirm whether OsPTR4 contributes to Chl content. Identifying the locations of these QTLs can help further understanding the molecular genetic mechanisms of Chl content and photosynthesis. Therefore, mapping the QTLs controlling Chl content and leaf-related traits can accelerate the breeding new rice varieties with high photosynthetic capacity and yield via genome selection (Guo et al. 2014).
References
Adhikari C, Bronson KF, Panuallah GM, Regmi AP, Saha PK, Dobermann A, Olk DC, Hobbs PR, Pasuquin E (1999) On-farm soil N supply and N nutrition in the rice-wheat system of Nepal and Bangladesh. Field Crops Res 64:273–286
A-JIA, Zeng LJ, Xue DW, Hu J, Zeng DL, Gao ZY, Guo LB, Li SG, Qian Q (2008) QTL analysis for chlorophyll content in four grain-filling stage in rice (in Chinese with an English abstract). Acta Agron Sin 34:61–66
Borrell AK, Garside AL, Fukai S, Reid DJ (1998) Season, nitrogen rate, and plant type affect nitrogen uptake and nitrogen use efficiency in rice. Aus J Agric Res 49:829–843
Chubachi T, Asano I, Oikawa T (1986) The diagnosis of nitrogen nutrition of rice plants (Sasanishiki) using chlorophyll meter. Soil Sci Plant Nutr 57:190–193
Ding CQ, Wang Y, You SL, Liu ZH, Wang SH, Ding YF (2016) Digital gene expression analysis reveals nitrogen fertilizer increases panicle size by repressing Hd3a signaling in rice. Plant Growth Regul 79:47–54
Dong YJ, Yang Z, Xu J, Lin D, Sugimoto Y, Luo L, Mei H (2007) Quantitative trait loci for leaf chlorophyll content at two developmental stages of rice (Oryza sativa L.). Commun Biometry Crop Sci 1:1–7
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Fan XR, Xie D, Chen JG, Lu HY, Xu YL, Ma C, Xu GH (2014) Over-expression of OsPTR6 in rice increased plant growth at different nitrogen supplies but decreased nitrogen use efficiency at high ammonium supply. Plant Sci 227:1–11
Foyer CH (1987) The basis for source-sink interaction in leaves. Plant Physiol Biochem 25:649–657
Gao ZY, Zhao SC, He WM et al (2013) Dissecting yield-associated loci in super hybrid rice by resequencing recombinant inbred lines and improving parental genome sequences. Proc Natl Acad Sci USA 110:14492–14497
Guo LB, Gao ZY, Qian Q (2014) Application of resequencing to rice genomics, functional genomics and evolutionary analysis. Rice 7:4
Hashim MM, Yusop MK, Othman R, Wahid SA (2015) Characterization of nitrogen uptake pattern in Malaysian rice MR219 at different growth stages using 15N isotope. Rice Sci 22:250–254
Horie T, Matsuura S, Takai T, Kuwasaki K, Ohsumi A, Shiraiwa T (2006) Genotypic difference in canopy diffusive conductance measured by a new remote-sensing method and its association with the difference in rice yield potential. Plant Cell Environ 29:653–660
Hu SP, Zhou Y, Zhang L, Zhu XD, Li L, Luo LJ, Liu GL, Zhou QM (2009) Correlation and quantitative trait loci analyses of total chlorophyll content and photosynthetic rate of rice (Oryza sativa) under water stress and well-watered conditions. J Integr Plant Biol 51:879–888
Huang JL, He F, Cui KH, Buresh RJ, Xu B, Gong WH, Peng SB (2008) Determination of optimal nitrogen rate for rice varieties using a chlorophyll meter. Field Crops Res 105:70–80
Jiang S, Zhang X, Zhang F, Xu Z, Chen W, Li Y (2012) Identification and fine mapping of qCTH4, a quantitative trait loci controlling the chlorophyll content from tillering to heading in rice (Oryza sativa L.). J Hered 103:720–726
Kaushal SS, Groffman PM, Band LE, Elliott EM, Shields CA, Kendall C (2011) Tracking nonpoint source nitrogen pollution in human-impacted watersheds. Environ Sci Technol 45:8225–8232
Kura-Hotta M, Satoh K, Katoh S (1987) Relationship between photosynthesis and chlorophyll content during leaf senescence of rice seedlings. Plant Cell Physiol 28:1321–1329
Li ZK, Pinson SR, Stansel JW, Paterson AH (1998) Genetic dissection of the source-sink relationship affecting fecundity and yield in rice (Oryza sativa L.). Mol Breed 4:419–426
Li YL, Johnson DA, Su YZ, Cui JY, Zhang TH (2005a) Specific leaf area and leaf dry matter content of plants growing in sand dunes. Bot Bull Acad Sin 46:127–134
Li ZK, Fu BY, Gao YM, Xu JL, Ali J, Lafitte HR, Jiang YZ, Rey DJ, Vijayakumar CHM, Maghirang R, Zheng TQ, Zhu LH (2005b) Genome-wide introgression lines and their use in genetic and molecular dissection of complex phenotypes in rice. Plant Mol Biol 59:33–52
Li JW, Yang JP, Fei PP, Song JL, Li DS, Ge CS, Chen WY (2009) Responses of rice leaf thickness, SPAD readings and chlorophyll a/b ratios to different nitrogen supply rates in paddy field. Field Crops Res 114:426–432
Li Y, Ouyang J, Wang YY, Hu R, Xia K, Duan J, Wang Y, Tsay YF, Zhang M (2015a) Disruption of the rice nitrate transporter OsNPF2.2 hinders root-to-shoot nitrate transport and vascular development. Sci Rep. doi:10.1038/srep09635
Li YY, Tao HJ, Xu J, Shi ZY, Ye WJ, Wu LW, Zeng DL, Gao ZY, Guo LB (2015b) QTL analysis for cooking traits of super rice with a high-density SNP genetic map and fine mapping of a novel boiled grain length locus. Plant Breed 134:535–541
Liu X, Liu LL, Xiao YH, Liu SJ, Tian YL, Chen LG, Wang ZQ, Jiang L, Zhao ZG, Wan JM (2015) Genetic dissection of leaf-related traits using 156 chromosomal segment substitution lines. J Plant Biol 58:402–410
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−∆∆CT method. Methods 25:402–408
McCouch SR, Cho YG, Yano M, Paul E, Blinstrub M, Morishima H, Kinoshita T (1997) Report on QTL nomenclature. Rice Genet Newslett 14:11–13
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4326
Peng S, Garcia FV, Laza RC, Cassman KG (1993) Adjustment for specific leaf weight improves chlorophyll meter’s estimate of rice leaf nitrogen concentration. Agron J 85:987–990
Peng S, Garcia FV, Laza RC, Sanico AL, Visperas RM, Cassman KG (1996) Increased N-use efficiency using a chlorophyll meter on high yielding irrigated rice. Field Crops Res 47:243–252
Peng SB, Khush GS, Virk P, Tang QY, Zou YB (2008) Progress in ideotype breeding to increase rice yield potential. Field Crops Res 108:32–38
Prasertsak A, Fukai S (1997) Nitrogen availability and water stress interaction on rice growth and yield. Field Crops Res 52:249–260
Takai T, Matsuura S, Nishio T, Ohsumi A, Shiraiwa T, Horie T (2006) Rice yield potential is closely related to crop growth rate during late reproductive period. Field Crops Res 96:328–335
Takai T, Kondo M, Yano M, Yamamoto (2010) A quantitative trait locus for chlorophyll content and its association with leaf photosynthesis in rice. Rice 3:172–180
Tan JJ, Tan ZH, Wu FQ, Sheng PK, Heng YQ, Wang XH, Ren YL, Wang JL, Guo XP, Zhang X, Cheng ZJ, Jiang L, Liu XM, Wang HY, Wan JM (2014) A novel chloroplast-localized pentatricopeptide repeat protein involved in splicing affects chloroplast development and abiotic stress response in rice. Mol Plant 7:1329–1349
Teng S, Qian Q, Zeng DL, Kunihiro Y, Fujimoto K, Huang DN, Zhu LH (2004) QTL analysis of leaf photosynthetic rate and related physiological traits in rice (Oryza sativa L.). Euphytica 135:1–7
Terashima I, Hanba YT, Tazoe Y, Vyas P, Yano S (2006) Irradiance and phenotype: comparative eco-development of sun and shade leaves in relation to photosynthetic CO2 diffusion. J Exp Bot 57:343–354
Tong HH, Mei HW, Yu XQ, Xu XY, Li MS, Zhang SQ, Luo LJ (2006) Identification of related QTLs at late developmental stage in rice (Oryza sativa L.) under two nitrogen levels. Acta Genet Sin 33:458–467
Wang SH, Cao WX, Wang QS, Ding YF, Huang PS, Ling QH (2002) Positional distribution of leaf color and diagnosis of nitrogen nutrition in rice plant (in Chinese). Sci Agric Sin 35:1461–1466
Wang B, Lan T, Wu WR, Li WM (2003) Mapping of QTLs controlling chlorophyll content in rice. Acta Genet Sin 30:1127–1132
Wang FH, Wang GX, Li XY, Huang JL, Zheng JK (2008) Heredity, physiology and mapping of a chlorophyll content gene of rice (Oryza sativa L.). J Plant Physiol 165:324–330
Wang Q, Xie W, Xing H, Yan J, Meng X, Li X, Fu X, Xu J, Lian X, Yu S, Xing Y, Wang G (2015a) Genetic architecture of natural variation in rice chlorophyll content revealed by a genome-wide association study. Mol Plant 8:946–957
Wang YX, Xiong GS, Hu J, Jiang L, Yu H, Xu J, Fang YX, Zeng LJ, Xu EB, Xu J, Ye WJ, Meng XB, Liu RF, Chen HQ, Jing YH, Wang YH, Zhu XD, Li JY, Qian Q (2015b) Copy number variation at the GL7 locus contributes to grain size diversity in rice. Nat Genet 47:944–948
Wang L, Xu J, Nian JQ, Shen NW, Lai KK, Hu J, Zeng DL, Ge CW, Fang YX, Zhu L, Qian Q, Zhang GH (2016) Characterization and fine mapping of the rice gene OsARVL4 regulating leaf morphology and leaf vein development. Plant Growth Regul 78:345–356
Wu P, Luo AC (1996) Investigation on genetic background of leaf chlorophyll content variation in rice under nitrogen stressed condition via molecular markers. Acta Genet Sin 23:431–438
Xu YF, Ookawa T, Ishihara K (1997) Analysis of the photosynthetic characteristics of the high-yielding rice cultivar Takanari. Jpn J Crop Sci 66:616–623
Xu HW, Liu CH, Lu RJ, Guo GM, Chen ZW, He T, Gao RH, Li YB, Huang JH (2016) The difference in responses to nitrogen deprivation and re-supply at seedling stage between two barely genotypes differing nitrogen use efficiency. Plant Growth Regul 79:119–126
Yang WB, Gao MJ, Yin X, Liu JY, Xu YH, Zeng LJ, Li Q, Zhang SB, Wang JM, Zhang XM, He ZH (2013) Control of rice embryo development, shoot apical meristem maintenance, and grain yield by a novel cytochrome P450. Mol Plant 6:1945–1960
Zhang GH, Xu Q, Zhu XD, Qian Q, Xue HW (2009) SHALLOT-LIKE1 is a KANADI transcription factor that modulates rice leaf rolling by regulating leaf abaxial cell development. Plant Cell 21:719–735
Zhang GH, Li SY, Wang L, Ye WJ, Zeng DL, Rao YC, Peng YL, Hu J, Yang YL, Xu J, Ren DY, Gao ZY, Zhu L, Dong GJ, Hu XM, Yan MX, Guo LB, Li CY, Qian Q (2014) LSCHL4 from japonica cultivar, which is allelic to NAL1, increases yield of indica super rice 93-11. Mol Plant 7:1350–1364
Zhang B, Ye WJ, Ren DY, Tian P, Peng YL, Gao Y, Ruan BP, Wang L, Zhang GH, Guo LB, Qian Q, Gao ZY (2015) Genetic analysis of flag leaf size and candidate genes determination of a major QTL for flag leaf width in rice. Rice 8:1–10
Zhou CH, Zhao ZK, Pan XH, Huang S, Tan XM, Wu JF, Shi QH (2016) Integration of growing milk vetch in winter and reducing nitrogen fertilizer application can improve rice yield in double-rice cropping system. Rice Sci 23:132–143
Zuo HL, Xiao K, Dong YJ, Xu JL, Li ZK, Luo LJ, Mei HW (2007) Molecular detection of quantitative trait loci for leaf chlorophyll content at different growth-stages of rice (Oryza sativa L.). Asian J Plant Sci 6:518–522
Acknowledgments
This work was supported by the grants from the Natural Science Fund of China (Grant Nos. 31521064 and 31271700) and the Chinese 973 Program (Grant No. 2013CBA01405).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declared that they have no conflict of interest.
Additional information
Weijun Ye and Shikai Hu have contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Ye, W., Hu, S., Wu, L. et al. Fine mapping a major QTL qFCC7 L for chlorophyll content in rice (Oryza sativa L.) cv. PA64s. Plant Growth Regul 81, 81–90 (2017). https://doi.org/10.1007/s10725-016-0188-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-016-0188-5