Abstract
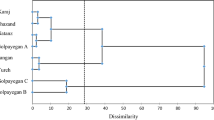
Genetic variation, as a multi-scalar variable, is important to genetic resource and conservation programs. M. mozaffarianiiMentha mozaffarianii Jamzad is an aromatic, medicinal and endemic plant in southern Iran. However, the genetic information of M. mozaffarianii is lacking. The study presented here is one of the first investigations to survey the genetic diversity of this valuable plant using Inter simple sequence repeat (ISSR) markers and morphological traits. This study also performed an association analysis to provide any correlation between the markers mentioned above. Individuals revealed by morphological features were divided into two major groups using cluster analysis based on Ward’s method. Results of ISSR analysis revealed genetic variations within and between populations to be 54% and 46%, respectively. In addition, 24 selected ISSR primers produced 343 polymorphic bands (Nei’s genetic diversity 0.146–0.427, Shannon’s information index 0.240–0.614). Cluster analysis using Unweighted Pair Group Method with Arithmetic Mean divided all individuals into two major groups and a high co-phenetic correlation coefficient (r = 0.850) was observed. The study of population structure as a prerequisite for correlation analysis showed two subgroups (K = 2). In mixed linear model analysis, 249 sites were identified to be related to the studied traits (p ≤ 0.02). The site ISSR42-20 was found to be associated with six features. This characterization could improve our understanding of the relationships among the evaluated traits. Flower diameter had the highest number of related markers (17 locations). Results showed that ISSRs were successful markers to show genetic diversity among populations. Results of the current study provided valuable information regarding M. mozaffarianii, which could be helpful in genetic studies, conservation, reproduction, breeding programs, and production of cultivars with high biomass contents.









Similar content being viewed by others
Data availability
The data sets generated and analyzed were available to the authors during the current study.
Abbreviations
- AMOVA:
-
Analysis of molecular variance
- AFLP:
-
Amplified fragment length polymorphism
- ANOVA:
-
Analysis of variance
- GLM and MLM:
-
General and mixed linear models
- UPGMA:
-
Unweighted pair group method with arithmetic mean
- MCMC:
-
Markov chain Monte Carlo
- GST :
-
Genetic differentiation coefficient between populations
- Nm:
-
Gene flow
- HS :
-
Genetic diversity index within populations
- ISSR:
-
Inter-simple sequence repeat
- Nm:
-
Gene flow
- H:
-
Nei’s gene diversity
- Ne:
-
Number of effective alleles
- PCAmix:
-
Principal component analysis of mixed data
- PCoA:
-
Principal coordinates analysis
- PIC:
-
Polymorphism information content
- RAPD:
-
Random amplified polymorphic DNA
- RFLP:
-
Restriction fragment length polymorphism
- I:
-
Shannon’s information index
- SSR:
-
Simple sequence repeat
- Ba:
-
Bastack
- Bo:
-
Bokhoun
- D:
-
Damtang
- H:
-
Hemag
- Ge:
-
Geno
- Bu:
-
Khaeez
- F:
-
Komarj
- S:
-
Sikhuran
- Z:
-
Zakin
- GT:
-
Grow type
- NL:
-
Number of leave in 15 cm
- LL:
-
Leaf length
- LW:
-
Leaf width
- R1:
-
Leaf L/W rate
- LC:
-
Leaf color
- LS:
-
Leaf shape
- LT:
-
Leaf tipe
- LM:
-
Leaf margin
- TL:
-
Thickness of leaf
- VN:
-
Veins number
- VC:
-
Vein conditions
- HL:
-
Heart-shaped leaf in shoot
- P:
-
Petiole
- PL:
-
Petiole length
- PC:
-
Petiole color
- NS:
-
Number of the main stem
- SML:
-
Stem the main length
- SC:
-
Stem color
- SDS:
-
Stem diagonal near the soil
- IML:
-
Internodes of the main length
- NNS:
-
Number nodes in the main stem
- NIS:
-
Number internodes in the main stem
- LNN:
-
Leafe number in node
- SSN:
-
Secondary stem number
- SSD:
-
Secondary stem diagonal
- PH:
-
Plant height
- PD:
-
Plant diagonal
- R2:
-
Plant diagonal/height rate
- FBNSL:
-
Flowering branches number in smooth leaf shoot
- FBNCL:
-
Flowering branches number in congress leaf shoot
- FSL:
-
Flowering stem length
- PF:
-
Place of emergence flowering branch
- FSC:
-
Flowering stems conditions
- LF:
-
Leaf size on flowering branch
- IL:
-
Inflorescence length
- ID:
-
Inflorescence diagonal
- DII:
-
Distance of internodes in inflorescence
- NNI:
-
Number nodes in inflorescence
- NF:
-
Number of flower at full flowering in inflorescence nods
- FCI:
-
Flower conditions at the end of inflorescence
- FD:
-
Flower diameter
- FFC:
-
Flower flag color
- FC:
-
Flower color
- FS:
-
Flowering season
References
Akagi H, Yokozeki Y, Inagaki A, Nakamura A, Fujimura T (1996) A codominant DNA marker closely linked to the rice nuclear restorer gene, RF-1, identified with inter-SSR fingerprinting. Genome 39:1205–1209. https://doi.org/10.1139/g96-152
Arman M, Yousefzadi M, Khademi SZ (2011) Antimicrobial activity and composition of the essential oil from Mentha mozaffarianii. J Essent Oil Bear Pl 14:131–135. https://doi.org/10.1080/0972060X.2011.10643912
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES (2007) TASSEL: software for association mapping of complex traits in diverse samples. J Bioinform 23:2633–2635. https://doi.org/10.1093/bioinformatics/btm308
Chaudhray P, Gauchan DRB, Rana B, Sthapit R, Jarvis DI (2004) Potential loss of rice landraces from a terai community in Nepal: a case study from Kachrowa, Bara. PGR Newsletter 137:14–21
Chavent M, Kuentz-Simonet V, Labenne A, Saracco J (2017) Multivariate analysis of mixed data: the PCA mixdata R package. arXiv:1411.4911. https://arxiv.org/abs/1411.4911
Choupani A, Shojaeiyan AA, Maleki M (2019) Genetic relationships of Iranian endemic mint species, Mentha mozaffarianii Jamzad and some other mint species revealed by ISSR markers. Biotechnologia 100:19–28. https://doi.org/10.5114/bta.2019.83208
Culley TM, Wolfe AD (2001) Population genetic structure of the cleistogamous plant species Viola pubescens Aiton (Violaceae), as indicated by allozyme and ISSR molecular markers. J Hered 86:545–556. https://doi.org/10.1046/j.1365-2540.2001.00875.x
Delnevo N, Piotti A, Carbognani M, Etten EJ, Stock WD, Field DL, Byrne M (2021) Genetic and ecological consequences of recent habitat fragmentation in a narrow endemic plant species within an urban context. Biodivers Conserv 30:3457–3478. https://doi.org/10.1007/s10531-021-02256-x
Earl DA, vonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–436. https://doi.org/10.1007/s12686-011-9548-7
Engelhardt KAM, Lloyd ML, Neel MC (2014) Effects of genetic diversity on conservation and restoration potential at individual, population, and regional scales. Biol Conserv 179:6–16. https://doi.org/10.1016/j.biocon.2014.08.011
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Fang DQ, Roose ML (1997) Identification of closely related citrus cultivars with inter-simple sequence repeat markers. Theor Appl Genet 95:408–417. https://doi.org/10.1007/s001220050577
Fischer M, Husi R, Prati D, Peintinger M, vanKleunen M, Schmid B (2000) RAPD variation among and within small and large populations of the rare clonal plant Ranunculus reptans (Ranunculaceae). Am J Bot 87:1128–1137. https://doi.org/10.2307/2656649
Ghani A, Rowshan V (2017) Comparative study of volatile constituents from two Iranian endemic mint (Mentha mozaffarianii Jamzad) ecotypes. Taylor Fr TEOP 20:1293–1301. https://doi.org/10.1080/0972060X.2017.1397552
Ghani A, Neamati SH, Azizi M, Saharkhiz MJ, Farsi M (2014) Artificial autotetraploidy induction possibility of two Iranian endemic mint (Mentha mozaffarianii) ecotypes. Not Sci Biol 6:185–191. https://doi.org/10.15835/nsb629129
Griffiths SM, Taylor-Cox ED, Behringer DC, Butler MJ IV, Preziosi RF (2020) Using genetics to inform restoration and predict resilience in declining populations of a keystone marine sponge. Biodivers Conserv 29:1383–1410. https://doi.org/10.1007/s10531-020-01941-7
Grimalt M, García-Martínez S, Carbonell P et al (2022) Relationships between chemical composition, antioxidant activity and genetic analysis with ISSR markers in flower buds of caper plants (Capparis spinosa L.) of two subspecies spinosa and rupestris of Spanish cultivars. Genet Resour Crop Evol 69:1451–1469. https://doi.org/10.1007/s10722-021-01312-3
Gupta AK, Mishra R, Singh AK, Srivastava A, Lal RK (2017) Genetic variability and correlations of essential oil yield with agro-economic traits in Mentha species and identification of promising cultivars. Ind Crops Prod 95:726–732. https://doi.org/10.1016/j.indcrop.2016.11.041
Hassanpouraghdam MB, Mohammadzadeh A, Morshedloo MR, Asadi M, Rasouli F, Vojodi Mehrabani L, Najda A (2022) Mentha aquatica L. populations from the hyrcanian hotspot: volatile oil profiles and morphological diversity. J Agron. https://doi.org/10.3390/agronomy12102277
Hosseini M, Ebrahimi M, Samsampour D, Abadía J, Khanahmadi M, Amirian R, Naseh Ghafoori I, Ghaderi-Zefrehei M, Gogorcena Y (2021) Association analysis and molecular tagging of phytochemicals in the endangered medicinal plant licorice (Glycyrrhiza glabra L.). Phytochemistry 183:112629. https://doi.org/10.1016/j.phytochem.2020.112629
Jedrzejczyk I, Rewers M (2018) Genome size and ISSR markers for Mentha L. (Lamiaceae) genetic diversity assessment and species identification. Ind Crops Prod 120:171–179. https://doi.org/10.1016/j.indcrop.2018.04.062
Khan MMH, Rafi M, Ramlee SI, Jusoh M, Mamun M, Halidu J (2021) DNA fngerprinting, fxation index (Fst), and admixture mapping of selected Bambara groundnut (Vigna subterranea L. Verdc.) accessions using ISSR markers system. Nature portfolio. Sci Rep 11:14527. https://doi.org/10.1038/s41598-021-93867-5
Kohler H, Friedt W (1999) Genetic variability as identified by AP_PCR and reaction to Sclerotinia sclerotiorum among interspecific sunflower (Helianthus annuus L.) hybrid progenies. Crop Sci 39:1456–1463. https://doi.org/10.2135/cropsci1999.3951456x
Kumar A, Mishra P, Singh SC, Sundaresan V (2014) Efficiency of ISSR and RAPD markers in genetic divergence analysis and conservation management of Justicia adhatoda L., a medicinal plant. Plant Syst Evol 300:1409–1420. https://doi.org/10.1007/s00606-013-0970-z
Kumar A, Pratap B, Kumar Gautam D et al (2019) Variability, heritability and genetic advance studies in French marigold (Tagetes patula L.). J Pharmacogn Phytochem 8:1046–1048
Kumar R, Das Munshi A, Behera TK, Jat GS, Choudhary H, Talukdar A, Dash P, Singh D (2022) Association mapping, trait variation, interaction and population structure analysis in cucumber (Cucumis sativus L.). Genet Resour Crop Evol. https://doi.org/10.1007/s10722-022-01352-3
Lamare A, Rao SR (2015) Efficacy of RAPD, ISSR and DAMD markers in assessment of genetic variability and population structure of wild Musa acuminata colla. Physiol Mol Biol Plants 21:349–358. https://doi.org/10.1007/s12298-015-0295-1
Mohammadi F, Sharifi-Sirchi G, Samsampour D (2017) Morphological, genetic and pigment diversity of Nerium indicum Mill in Iran. Cell Mol Biol 63:64–70. https://doi.org/10.14715/cmb/2017.63.9.12
Nei M (1972) Genetic distance between populations. Am Nat 106:283–392. https://doi.org/10.1086/282771
Peakall R, Smouse PE (2012) GeneAlex 6.5: genetic analysis in Excel. Population genetic software for teaching and research–an update. J Bioinform 28:2537–2539. https://doi.org/10.1093/bioinformatics/bts460
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959. https://doi.org/10.1534/genetics.116.195164
Rao VR, Hodgkin T (2002) Genetic diversity and conservation and utilization of plant genetic resources. PCTOC 68:1–19
Robins JG, Campbell T, Bauchan GR, He C, Viands DR, Hansen JL, Brummer EC (2007) Genetic mapping of biomass production in tetraploid alfalfa. Crop Sci 47(1–10):32. https://doi.org/10.2135/cropsci2005.11.0401
Rodrigues L, van den Berg C, Póvoa O, Monteiro A (2013a) Low genetic diversity and significant structuring in the endangered Mentha cervina populations and its implications for conservation. Biochem Syst Ecol 50:51–61. https://doi.org/10.1016/j.bse.2013.03.007
Rodrigues L, Póvoa O, van den Berg C, Figueiredo AC, Moldão M, Monteiro A (2013b) Genetic diversity in Mentha cervina based on morphological traits, essential oils profile and ISSRs markers. Biochem Syst Ecol 51:50–59. https://doi.org/10.1016/j.bse.2013.08.014
Rohlf FJ (1998) On applications of geometric morphometrics to studies of ontogeny and phylogeny. Syst Biol 47:147–158. https://doi.org/10.1080/106351598261094
Sabboura D, Yacoub R, Lawand S (2016) Assessment of genetic relationships among mint species. Int J Chemtech Res 9:462–468
Sam-Daliri H, Mousavi Z, Naderi N, Asgarpanah J (2016) Chemical composition and analgesic activity of the essential oil of Mentha mozaffarianii Jamzad leaves. Bulg Chem Commun 48(4):641–645
Semagn K, Bjornstad A, Ndjiondjop MN (2006) An overview of molecular market methods for plants. Afr J Biotechnol 5:2540–2568. https://doi.org/10.5897/AJB
Shannon CE, Weaver W (1949) The mathematical theory of communication. University of Illinois Press, Urbana
Singh P, Pandey AK (2018) Prospective of essential oils of the genus mentha as biopesticides: a review. Front Plant Sci 9:1295. https://doi.org/10.3389/fpls.2018.01295
Soilhi Z, Trindade H, Vicente S, Gouiaa S, Khoudi H, Mekki M (2019) Assessment of the genetic diversity and relationships of a collection of Mentha spp. in Tunisia using morphological traits and ISSR markers. Hortic Sci Biotechnol. https://doi.org/10.1080/14620316.2019.1702482
Sullivan ER, Barker C, Powell I, Ashton PA (2019) Genetic diversity and connectivity in fragmented populations of Rhinanthus minor in two regions with contrasting land-use. Biodivers Conserv 28:3159–3181. https://doi.org/10.1007/s10531-019-01811-x
Tafrihi M, Imran M, Tufail T, Gondal TA, Caruso G, Sharma S, Sharma R, Atanassova M, Atanassov L, Valere Tsouh Fokou P, Pezzani R (2021) The wonderful activities of the genus mentha: not only antioxidant properties. Molecules 26:1118. https://doi.org/10.3390/molecules26041118
Xlstat (2021) https://www.xlstat.com/en/news/xlstat-version-2021-1
Yeh FC, Yang RC, Boyle TBJ, Ye Z, Xiyan JM, Yang R, Boyle TJ (2000) PopGene32, microsoft windows-based freeware for population genetic analysis, version 1.32. Molecular biology and biotechnology center. University of Alberta, Canada
Acknowledgements
This project was carried out with the cooperation of the Environmental Department of Hormozgan Province. We wish to express our gratitude for the use of their facilities. We would also like to thank Dr. Tahmineh Alidadi, MSc Fariba Mohammadi, Dr. Gholamreza Sharifi-Sirchi, and Dr. Siavash Samavii for their cooperation in implementing this project.
Funding
The authors declare that no funds, Grants, or other support were received during the arrangement of this manuscript.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design; FR, DS, MAS prepared Materials. FR performed Data collection and analysis. FR wrote the first draft of the manuscript then, other authors commented on previous manuscript versions. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare they have no known financial interests or personal relationships with competitors that affect the work reported in this article.
Human or animal rights
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Roshanibakhsh, F., Samsampour, D., Askari Seyahooei, M. et al. Strong relationship between molecular and morphological attributes in Iranian mentha populations (Mentha mozaffarianii Jamzad). Genet Resour Crop Evol 70, 1721–1745 (2023). https://doi.org/10.1007/s10722-022-01532-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-022-01532-1