Abstract
Stephania tetrandra S. Moore (family: Menispermaceae), a dioecious herbaceous vine and the only species in the subgenus Botryodiscia of the genus Stephania of the family Menispermaceae, is mainly distributed in hilly areas south of the Huaihe River in China and found in many provinces of China, showing a high genetic diversity. This paper aimed to study genetic diversity of and genetic relationship among individuals of S. tetrandra within China to provide a basis for evaluation, exploitation, and utilization of S. tetrandra by using simple sequence repeat (SSR) molecular markers. Our results show that effective products were amplified from the 26 screened SSR gene loci, a total of 183 alleles amplified (2–16 alleles amplified by each pair of primers). Among the 26 loci, 16 had a PIC value higher than 0.5, indicating a high level of polymorphism. For most of the loci, the number of effective alleles was lower than that of the observed alleles, and the observed heterozygosity was lower than the expected heterozygosity. The genetic differentiation coefficient (0.021–0.547) was lower than 0.05 (low level of genetic differentiation) for 7 loci and higher than 0.25 (very high level of genetic differentiation) for 2 loci, and had a value representing a medium level of genetic differentiation for the remaining 17 loci. The intra-population inbreeding coefficient had a positive value for 21 loci, suggesting the presence of inbreeding and homozygous excess. The gene flow value was bigger than 1, indicating that genetic drift and natural selection played an unimportant role in population genetic differentiation of S. tetrandra. Based on discriminant analysis of principal components and Bayesian Information Criterion, K-means clustering was performed on 620 samples. These samples were divided into 9 genetic clusters, whose similarity coefficients and genetic distances were 0.755–0.918 and 0.067–0.280, respectively, indicating that these clusters were highly similar and short-distanced. The Bayesian clustering analysis was implemented in the STRUCTURE software to analyze the genetic structure of S. tetrandra and it was found that the 620 samples could be clustered into 5 ancestor groups; the 9 clusters and 40 natural populations inherited genes from the 5 groups to varying degrees, but the proportion of genes inherited from the 5 groups by each cluster and natural population differed. S. tetrandra was characterized by the presence of population structure and pronounced genetic subdivision, which, together with the presence of gene flow, may indicate a relatively stable recent state of these populations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Stephaniae Tetrandrae Radix, the dried root of S. tetrandra in the family Menispermaceae, has been recorded in most editions of Chinese Pharmacopoeia (National Pharmacopoeia Committee 2015). It has the effect of inducing diuresis to alleviate edema, dispelling wind, and relieving pain, and is widely used clinically (Kang et al. 2014; Wang 2014; Wang et al. 2017). It showed good clinical effects on chronic hepatitis, early liver cirrhosis, allergic dermatosis, diabetes, prostatitis, seminal vesiculitis, and other diseases (Xue and Zhu 2012; Liu et al. 2017b; Wang et al.2017). Tetrandrine, its main active substance, has been demonstrated to have a good effect in anti-inflammation, analgesia, blood pressure lowering, anti-silicosis, blood sugar lowering, anti-free radical injury, anti-liver fibrosis, and anti-tumor (Kang et al. 2014; Wu et al. 2014; Qiu et al. 2014). S. tetrandra is mainly distributed in hilly areas south of the Huaihe River in China and found in many provinces of China, including Jiangxi, Anhui, Zhejiang, Fujian, Hunan, and Guangdong. S. tetrandra is a dioecious species (Chinese Academy of Sciences Institute of Botany 1972). According to our resource survey, the ratio of female and male individuals in wild populations of S. tetrandra is 1:3, resulting in a low level of seed production. We also found that dispersal of the fruit of this species by the wind was difficult because of its high mass, and it is slightly bitter in taste and maybe birds do not like eating it, restricting the habitat expansion of wild S. tetrandra. This species, however, possesses a strong ability to reproduce asexually, and in nature, it shows both seed propagation and vegetative propagation (Guo et al. 2007). In the process of sexual reproduction of plants, mutations such as deletion, duplication, inversion, translocation, and chromosome doubling can occur in the course of chromosome exchange, which can cause genetic variation (Li et al. 2023) and genetic diversity of species (Zhang et al. 2019). Asexual reproduction has less effect on genetic variation than sexual reproduction. At the same time, in the process of evolution, genetic variation and genetic drift in the germplasm may occur in plants in different distribution regions in response to environmental changes, resulting in differences in germplasm in different distribution regions (Peng and Tang, 2017; Lu et al. 2019). The study of genetic diversity can reveal the genetic structure and relationship of plant germplasm resources, and can be used for germplasm identification and high-quality resource evaluation (Xie and Yun 2000). So far, very few studies on the gender differentiation and genetic structure of S. tetrandra have been conducted. Whether there is any difference in the germplasm of S. tetrandra plant individuals from different distribution areas as well as the information on their genetic structure and evolutionary relationship remains to be disclosed.
Simple sequence repeats (SSRs) are widely distributed in the whole genome of eukaryotes and have highly polymorphic characteristics among individuals and populations (Chen et al. 2009). SSR sequences from different species showed a high allelic diversity in composition, length, mutation rate, and their distribution in chromosomes (Chen et al. 2009). Such sequences have relatively conservative flanking sequences, based on which SSR primers with high specificity can be designed and used for effective amplification of specific SSRs (Zhang 2014). This method has become a common approach for the development of SSR primers due to a large number of primers developed, simplicity, and low cost (Xie and Yun 2000; Chen et al. 2009; Ao et al. 2009; Zhang 2014). SSR molecular markers are codominant, in line with the Mendelian law, easy to operate, and highly repeatable, reliable, and polymorphic, so they have been widely employed in the study of the correlation between genomic differences and character variation, plant genetics, and breeding (Xie and Yun 2000; Liu et al. 2017a, b; Xie et al. 2017; Su et al. 2018). Therefore, we used SSR markers to analyze the genetic diversity and relationship of S. tetrandra in this paper.
S. tetrandra populations are present in different geographical locations. Their germplasms may differ, leading to the discrepancy in the quality of crude drug derived from them. The difference in germplasms is reflected by the difference in SSR markers. Therefore, in the present study, we tried to disclose the genetic diversity of the species S. tetrandra and the genetic relationship among different populations of S. tetrandra to provide a basis for resource conservation and utilization of that species.
Materials and Methods
Plant Materials and DNA Extraction
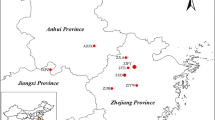
We gathered wild samples from 93 villages in 40 cities or counties in Guangdong, Anhui, Jiangxi, Zhejiang, Fujian, and Hunan provinces of China (Fig. 1). Sampling cities or counties (each one represents one population) were at least 50 km apart, and some (basically more than 5) plant individuals (Fig. 1) were chosen from populations in each city or county (depending on the distribution density of S. tetrandra populations). Fresh, healthy young leaves of 620 randomly chosen individuals of S. tetrandra (148 ones from Anhui, 53 from Fujian, 44 from Guangdong, 54 from Hunan, 279 from Jiangxi, and 42 from Zhejiang) were collected in the same period of time and quickly dried off for subsequent use. At the same time, specimens were collected from all sampling sites and confirmed by professor Songji Wei, a botanist from Guangxi University of Chinese Medicine, China, as genuine S. tetrandra, and grown at Xianhu Campus of Guangxi University of Chinese Medicine to establish a gene bank of S. tetrandra. The geographical and climatic information of the cities and counties where sampling sites were located are shown in Table 1. Genomic DNA was extracted using the magnetic bead–based kit for isolating genomic DNA from TIANGEN Biotech (Beijing), 2 to 3 μL of it was subjected to testing by 1.5% agarose gel electrophoresis, and the rest was stored at − 20 °C for PCR test.
A map showing the village-level sampling sites (all in China) selected for this study. The horizontal axis represents the longitude and the vertical axis represents the latitude. Each red five-pointed star is a village-level sampling site. The village-level sampling sites in each county or city are not too far apart and the cities and counties are at least 50 km apart
Primer Screening
Three samples were selected randomly for transcriptome sequencing using Illumina next-generation sequencing technology. A total of 45,024 SSR markers were mined from transcriptome sequences. Two hundred SSR markers were screened from marker loci which had 2 to 6 bases each and were present in more than 90% of samples and primers were designed according to conserved sequences on both ends of these SSR marker loci. Four individual plant samples from different populations were randomly selected and used to analyze the amplification effect and polymorphism of the 200 primers to select primers for polymorphic SSR markers for subsequent study (for each of the 4 plant samples, the amplified fragment size of the same SSR marker primer was different). The primers were synthesized by Tianyi Huiyuan Co., Ltd. in China.
Analysis of Population Genetic Diversity
The selected primers for polymorphic SSR markers were used to amplify DNA of 620 samples, in order to analyze SSR and population genetic diversity based on amplification results. The SSR fluorescent primer system used in PCR amplification (15μL in total) was as follows: mix, 7.5 μL; ddH2O, 4.5 μL; forward primer, 1 μL (10 μmol/L); reverse primer, 1 μL(10 μmol/L); DNA template, 1 μL. The amplification procedure is shown in the Supplementary Materials. Post-amplification SSRs were detected by fluorescence-labeled capillary electrophoresis. Formamide and relative molecular weight internal standard were mixed at a ratio of 250:1 (v/v). A total of 9 μL of the resultant mixture was loaded into the microplate, to which 1 μL of PCR product diluted tenfold was added. The 3730XL sequencer was used for capillary electrophoresis, and the Fragment (Plant) fragment analysis software in GeneMarker was used to analyze the original data obtained by the sequencer. The position of the internal standard in each lane and the peak position of each sample were compared and analyzed to obtain the size of fragments.
Statistical Analysis
GenAlEx software was used to estimate various genetic parameters, including effective number of alleles per locus (Ne), observed number of alleles per locus (Na), Shannon information index (I), heterozygosity (Ht), observed heterozygosity (Ho), expected heterozygosity (He), fixation index (F), genetic differentiation coefficient (Fst), and gene flow value (Nm) (Xu et al. 2015). The polymorphism information content (PIC) of each locus was calculated using Cervus software (Slate et al. 2000). GenAlEx software was employed to calculate pairwise genetic differentiation coefficients (Fst) and perform molecular variance analysis of clusters. Using find.cluster in the R Package (adegenet), the K-means algorithm was run to generate a graph with different K values as the horizontal coordinate and the obtained BIC values as the vertical coordinate; the K value at the first inflection point was selected to determine the number of clusters (Jombart et al. 2010). The UPGMA (Unweighted Pair Group Method using Arithmetic Averages) clustering was performed, and tree maps were constructed using Phylip software based on Nei’s genetic distances of clusters. The principal coordinate analysis (PCoA) was performed using GenAlEx software. Bayesian statistics and Markov chain Monte Carlo Simulation were used to estimate the assigned proportion of each individual in each group (membership coefficient, Q). Data on all K values (from 1 to 15) were input into STRUCTURE Harvester software v0.6.1 (available at http://taylor0.biology.ucla.edu/structureHarvester/) to estimate delta K and prepare the indfiles for the software CLUMPP (v1.1.2), which was used to perform alignments from the Q values of each K STRUCTURE group. distruct (v1.1) was used to visualize the resulting bar plots, and individuals were assigned to the cluster for which they showed the highest Q value. STRUCTURE software (v2.3.4) was used to infer the group structure with the following parameters: BURNIN 10000, NUMREPS 30000.
Results
Primer Screening
A total of 45,024 SSR markers were mined from the transcriptome sequences, of which 23,189 ones were single base repeats and 9272 ones were double bases. From the marker sites with 2–6 base repeats each which were present in more than 90% of the samples, 200 SSR markers were selected (see Supplementary Materials) and primers were designed. Twenty-six pairs of primers for the selected polymorphic SSR markers were randomly chosen and used to amplify the DNA of 620 samples of S. tetrandra (Table 2).
Genetic Diversity of SSR Loci
Statistical results of genetic diversity analysis of 26 SSR loci are shown in Table 3. The results show that effective products were amplified from all of the 26 SSR loci, including 183 alleles, 2–16 alleles per pair of primers. UG-29652-FAM had the highest Na (16 alleles), whereas UG-10578-HEX, UG-20112-FAM, and UG-33750-FAM had the lowest Na (only 2 for each). The Ne was 1.471–8.099 (3.162 on average). For most of the loci, the effective number of alleles was lower than the observed number of alleles, indicating the interaction between alleles. The I was 0.521–2.217 (1.202). The Ho was 0.128–0.804 (0.498). The Ne was 0.320–0.877 (0.603), UG-30904-HEX having the highest value and UG-19736-FAM having the lowest value. For all the 26 loci, except UG-10578-HEX and UG-53337-HEX, the Ho was lower than the He, indicating a departure from the Hardy–Weinberg equilibrium. The PIC was 0.272–0.864 (0.551); this parameter had a value bigger than 0.5 for 16 loci which were highly polymorphic and a value between 0.25 and 0.5 for 10 loci which were middle-level polymorphic.
F-Statistics and Gene Flow
As shown in Table 3, the intra-population inbreeding coefficient (Fis) was − 0.121 to 0.493 (0.101 on average); this parameter had a positive value for 21 loci, suggesting the presence of inbreeding and homozygous excess. The inter-population inbreeding coefficient (Fit) was − 0.027 to 0.748 (0.185). The genetic differentiation coefficient (Fst) was 0.021–0.547 (0.102); this parameter had a value lower than 0.05 (indicating a low level of genetic differentiation) for 7 loci, a value higher than 0.25 (indicating a very high level of genetic differentiation) for 2 loci, and a value indicating middle-level genetic differentiation for the remaining 17 loci. The gene flow (Nm) values varied from 0.207 to 11.440, with an average of 4.082; this parameter had a value higher than 1 for 24 loci, suggesting that inter-population genetic differentiation of these 24 loci was not a result of genetic drift.
Genetic Similarity, Distance, and Differentiation
K-means clustering was performed on amplified fragment length data of SSR primers for 620 plant individual samples collected from 40 cities or counties in 6 provinces of China. Bayesian Information Criterion (BIC) values were obtained by choosing different K values (the number of clusters). It was found that the value of K at the first inflection point was 9 (see Fig. 2); thus, it seemed most appropriate to divide the 620 samples into 9 clusters. The first cluster comprised 52 samples from Anhui, Fujian, Guangdong, Hunan, and Jiangxi; the second cluster consisted of 66 samples from Anhui, Hunan, Jiangxi, and Zhejiang; the 3rd, 6th, and 9th clusters had 81, 61, and 78 samples, respectively, which were collected from all the provinces; Clusters 4 and 8 each contained 73 samples from Anhui, Hunan, Jiangxi, Fujian, and Zhejiang. Cluster 5 had 61 samples, collected from Fujian, Guangdong, Hunan, and Jiangxi, whereas Cluster 7 had 75 samples from Anhui, Fujian, Jiangxi, and Zhejiang (see Fig. 3). It is clear that almost all clusters comprised samples from different geographical locations, indicating that geographical locations pose little impact on the genetic differentiation of S. tetrandra and grouping based on geographical regions is not reasonable.
Similarity test was performed on the 9 clusters to obtain their similarity coefficients and genetic distances (see Table 4). A UPGMA-based tree map was generated as shown in Fig. 4, in which the length of branch denotes Nei’s genetic distance. As can be seen in Table 4, similarity coefficients of 9 clusters of S. tetrandra were 0.755–0.918, and genetic distances was 0.067–0.280, indicating that these clusters had a high level of similarity and short genetic distances.
Cluster analysis of Nei’s genetic distances. The scale on the upper left corner denotes genetic distance which increases from right to left starting from zero. The lowermost branch point represents the genetic distance at which samples are divided into Clusters 3 and 4, and the branch point immediately to the left of it represents the distance where samples are divided into Clusters 2, 3, and 4, and other branch points can be interpreted in a similar fashion
Between-cluster genetic differentiation coefficients (Fst) were calculated for the 9 clusters, and a heat map (Fig. 5) was created. It can be seen in Fig. 5 that the genetic differentiation between Cluster 1 and Cluster 2 and between Cluster 1 and Cluster 4 was higher (circles having deeper color), Fst being above 0.095, whereas Clusters 3 and 4 showed the least genetic differentiation (the circle having the lightest color). F-statistics analysis results (Table 5) showed that Fis, Fit, Fst, and adjusted Fst were 0.172, 0.248, 0.092, and 0.219, respectively, indicating a moderate level of genetic differentiation. The gene flow value was 2.461, bigger than 1.
Principal Co-ordinates Analysis (PCoA) Result
PCoA can reveal the difference between two or two groups of samples by intuitively comparing the linear distance between the samples on coordinate axes. The shorter the linear distance between two or two groups of samples, the smaller the genetic difference. The PCoA result of the 9 clusters is shown in Fig. 6, from which we know that the PCoA result was basically consistent with that of the clustering analysis.
Genetic Structure Analysis Result
For the purpose of further exploration of the genetic relationship between the 40 natural populations and between the 9 clusters, the STRUCTURE software was used to perform Bayesian clustering to compare their genetic structure. STRUCTURE is a calculation method based on the Bayesian model. A corresponding maximum likelihood is generated for the simulation of each K value. In STRUCTURE, the maximum likelihood is output as a ln-transformed value (ln likelihood, L(K)). The larger the L(K) is, the more the K value is approximate to the true level. The L(K) generally increases with the K value but will slowly plateau out. The goal of selecting the optimal K value is to find the inflection point. Using this method, the change of L(K) with K and that of delta K with K were disclosed as shown in Figs. 7 and 8, respectively. In Fig. 8, delta K reached the maximum level when K was 5, suggesting that the 620 samples could be divided into 5 groups. Genetic maps of group structure were generated with K = 5 (see Fig. 9A–C). In Fig. 9A, all 620 plant samples from sampling sites in different cities or counties contained the genetic lineage of the 5 groups, and every group made contributions to the hereditary basis of the samples, resulting from crossing among the 5 groups. This indicates that the distribution of natural habitats had little influence upon the division of the five groups. In Fig. 9B, all the 9 clusters inherited genes from the 5 groups to varying degrees, but the proportion of genes inherited from the 5 groups by each cluster differed. Clusters 1, 2, and 8 inherited more genes from Group 1 than from other groups, and Cluster 3 was more influenced genetically by Group 5 than by other groups. In Fig. 9C, all natural populations inherited genes from the 5 groups to varying degrees; the proportions of genes inherited from the 5 groups were similar for all the natural populations except that GDNXS, GDSXX, HNXXX, and JXQNX inherited more genes from Group 2 than from other groups.
Population structure analysis results. A Assigned proportion of each individual in the groups when K = 5. B Assigned proportion of each cluster in the groups when K = 5. C Assigned proportion of each natural population in the groups when K = 5. In C, the abbreviations stand for sampling cities and counties, the first two letters representing the name of a province, and those followed being the name of a city or a county (X means a county and S means a city; e.g., AHDZX is “Dongzhi County, Anhui Province” (see Table 1))
Discussion
SSR Genetic Diversity Analysis
Genetic diversity refers to the sum of genetic variations among individuals within a species or within a population. Intraspecific diversity is not only an important source of species genetic diversity, but also a decisive factor for a species to respond successfully to external disturbance, and an important index used for the evaluation of germplasm resources (Nafees 2015; Zhen 2017). S. tetrandra is the only species in the subgenus Botryodiscia of the genus Stephania of the family Menispermaceae. Retrieval in TIMETREE (http://timetree.org/) using terms “Menispermaceae” and “Stephania” showed that the evolution time of plants in the genus Stephania was about 36.4 MYA, and the evolution time of S. tetrandra, about 4.96MYA, was shorter than that of other species in the genus Stephania. This short evolution time may have a certain influence on the degree of species differentiation of S. tetrandra. In our experiment, the genetic diversity of SSRs was analyzed on 620 samples from Anhui, Fujian, Guangdong, Hunan, Jiangxi, and Zhejiang of China. The amplification results of 26 SSR loci showed that the values of Na varied from 2 to 16. For all loci, there were great differences in PIC, Ne, I, Ho, and He (Table 3), consistent with findings of other species (Huang 2019; Ismail et al. 2019). These differences may be due to the fact that the EST-SSRs used in our study are transcribed regions of the genomes; therefore, they can be either neutral markers or included into genome regions associated with adaptively important traits (i.e., being subject to selection) and thus can determine the ability of S. tetrandra to adapt to any environmental stress factors; meanwhile, they are of great significance to the evaluation of germplasm resources (An et al. 2009). PIC values of 16 loci among the 26 loci were bigger than 0.5, indicative of a high level of polymorphism; and the values of PIC of the remaining 10 loci were greater than 0.25 (a medium level of polymorphism). According to Purvis and Franklin (2005), loci with PIC > 0.7 are the best genetic markers. In the present work, 6 loci had a PIC value bigger than 0.7, suggesting that it was possible to select SSR loci for efficiently investigating the genetic diversity of S. tetrandra from the loci tested. From the results of our experiment, it was also found that Ho for all loci except for UG-10578-HEX and UG-53337-HEX was lower than He, suggesting that most loci in daughter populations violated the Hardy–Weinberg equilibrium, maybe due to the fact that parental populations have been affected by non-random factors in the process of gene transmission. Fst can be used to assess the population differentiation degree and genetic structure; a value between 0 and 0.05 indicates no or low-level differentiation, a value between 0.05 and 0.15 indicates a medium level of differentiation, a value between 0.15 and 0.25 indicates a high level of differentiation, and a value greater than 0.25 means differentiation is at a very high level (Wright 1977). In our study, the value of Fst was bigger than 0.05 for most loci, suggesting that there was a medium level of genetic differentiation. The gene flow Nm were 0.102 and 4.082 (individuals per generation), greater than 1 for most loci, meaning that the current genetic structure has been maintained and genetic differentiation due to genetic drift has been overcome (Ellstrand and Elam 1993). Moreover, the results of Fst and Nm suggested that there was a moderate level of subdivision of populations (Wright 1965). A high heterozygosity across the whole species may make it more resilient and enable its sustainable development, since individuals with greater heterozygosity within a population have increased developmental stability (Leary et al. 1983; Novicic et al. 2011). From the point of view of breeding, however, heterozygosity may bring some difficulties to the selection and breeding of good germplasms (Liu 2005); that is to say, a low level of heterozygosity may be beneficial to the selection and breeding of high-quality germplasms.
Genetic Variation and Structure Analyses
All samples collected in the present study came from wild populations of S. tetrandra. Prior to the analysis of genetic differentiation and genetic structure, DAPC (discriminant analysis of principal components)–based genotyping on the 620 samples was performed. We found that the resulting genetic clusters were not closely associated with natural populations (refer to the Supplementary Materials for the genotyping results: detection based on BIC and cluster.Neis_distance). In the present study, we aimed to analyze the genetic diversity and genetic structure of the species S. tetrandra, and since there were great differences among individuals from the same natural population, we carried out further analyses using genetic clusters obtained from the genotyping instead of natural populations. According to the genotyping results, optimally the 620 samples should be divided into 9 clusters, each one of which contained individuals from different geographical locations (i.e., from different natural populations), suggesting that genetic clustering based on natural populations was suboptimal. In our study, the similarity coefficients and genetic distances of the 9 genetic clusters were 0.755–0.918 and 0.067–0.280, respectively, genetically highly similar and short-distanced. Furthermore, as demonstrated by F-statistics analysis, there was a medium level of differentiation between the 9 clusters, and a value of gene flow greater than 1 indicated that gene exchange occurred frequently between the clusters, so the clusters were highly similar (the similarity coefficients being 0.755–0.918), suggesting that genetic drift and natural selection may play an unimportant role in S. tetrandra; germplasm differentiation of S. tetrandra may have mainly resulted from individual variation. For the purpose of further exploration of the genetic relationships between populations of S. tetrandra, the Bayesian clustering analysis was implemented in the STRUCTURE software to compare the genetic structure of 620 samples of S. tetrandra (Porras-Hurtado et al. 2013; Rosenberg 2004). Results of the STRUCTURE analysis showed that the 620 samples could be divided into 5 groups. Due to the influence from gene flow, the 9 clusters and 40 natural populations inherited genes from the 5 groups to varying degrees.
Factors Associated with the Genetic Differentiation of S. tetrandra
S. tetrandra is a dioecious species with unisexual flowers and reproductive characteristics typical of the family Menispermaceae (Cheng 2022). Self-crossing is not possible since male and female reproductive organs are located in different plant individuals; thus, the homozygous excess should be mainly due to inbreeding between closely related individuals. We conducted a comprehensive field survey on the distribution, habitat, and individual biological characteristics of wild S. tetrandra in China, and found that the ratio of female and male individuals in wild populations of S. tetrandra was 1:3, severe gender bias existing. Moreover, dispersal of the fruit of this species by the wind is difficult because of its high mass (about 0.789 g per fruit) and that it is not easy to shed from the plant, and it is slightly bitter in taste and birds do not like eating it, resulting in difficult dispersal of its seeds. Therefore, the geographical distribution of S. tetrandra is limited to a certain range. From the distribution map of the sampling sites as shown in Fig. 1 of our study, Jiangxi had the highest distribution density of S. tetrandra populations, and the distribution density in other provinces decreased with the distance between them and Jiangxi. The roots and rhizomes of S. tetrandra, however, possess a strong ability to reproduce, and in nature, this species shows both seed propagation and vegetative propagation (Guo et al. 2007). In the present study, all plant individuals instead of natural populations from different geographical locations were used in cluster analysis and genetic structure analysis because individuals from the same natural population could not be grouped into the same genetic cluster, indicating the presence of population structure and pronounced genetic subdivision.
S. tetrandra is mainly distributed in Guangdong, Hunan, Jiangxi, Anhui, Zhejiang, and Fujian in China. Affected by various factors, germplasms and phenotypic characteristics of S. tetrandra from different habitats differ (Yang et al. 2020). The difference in medicinal quality of traditional Chinese drugs has an adverse effect on the development of their resources and their clinical use, and the quality difference is mainly affected by germplasms (Meng et al. 2013). The analysis of genetic differences of germplasms from different geographical locations can provide a basis for the selection of excellent germplasms (Zawedde et al. 2015) and the selection of suitable environmental conditions of producing areas as well as the development and utilization of medicinal resources (Ma and Xiao 1998).
Conclusion
Altogether, the species studied is characterized by the presence of population structure and pronounced genetic subdivision, which, together with the presence of gene flow, may indicate a relatively stable recent state of these populations. However, the overexploitation of S. tetrandra resources, as well as other anthropogenic factors, can disturb the existing equilibrium. In this regard, the obtained results can be used for monitoring, rational using, and restoration of these populations. Moreover, developed markers will also be useful for the design of plantation cultivation, and since they mark the transcribed genome, they can be applied to research on increasing the productivity of S. tetrandra and developing alternative sources of target biologically active substances (for example, via in vitro cultivation).
Availability of Data and Materials
All relevant data are within the manuscript and its Supporting Information files.
References
An N, Guo HB, Ke WD (2009) Genetic variation in rhizome lotus (Nelumbo nucifera Gaertn. ssp. nucifera) germplasms from China assessed by RAPD markers. Agric Sci China 8(1):31–3
Ao RGL, Jia X, Ge TM (2009) Strategies for developing SSR molecular markers. J Hubei Minzu University (natural Science Edition) 27(4):462–467
Chen HQ, Sui C, Wei JH (2009) Summary of strategies for developing SSR primers. Mol Plant Breed 7(4):845–851
Cheng XL (2022) Propagation and cultivation techniques for Stephania tetrandra. Anhui For Sci Technol 48(3):26–28
Chinese Academy of Sciences Institute of Botany (1972) Illustrated handbook of higher plants in China (Book One). Science Press, Beijing
Ellstrand NC, Elam DR (1993) Population genetic consequences of small population size: implications for plant conservation[J]. Ann Rev Ecol Syst 24:217–242. https://doi.org/10.1146/annurev.es.24.110193.001245
Guo GS, Liu LH, Zeng XC, Zhang YP, Yuan HS (2007) Study on reproductive materials of Stephania tetrandra. Res Pract Chin Med 2:17–19
Huang YQ (2019) Genetic diversity analysis and core germplasm construction of Phoebe bournei based on SSR molecular markers. Master’s Thesis
Ismail NA, Rafii MY, Mahmud TMM, Hanafi MM, Miah G (2019) Genetic diversity of torch ginger (Etlingera elatior) germplasm revealed by ISSR and SSR markers. Biomed Res Int. https://doi.org/10.1155/2019/5904804
Jombart T, Devillard S, Balloux F (2010) Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genet 11:94
Kang OH, An HJ, Kim SB, Mun SH, Seo YS, Joung DK, Choi JG, Shi DW, Kwon DY (2014) Tetrandrine suppresses pro-inflammatory mediators in PMA plus A23187-induced HMC-1 cells. Int J Mol Med 33:1335–1340
Li SJ, Pei HD, Liu XX, Chen J, Jiang J, Zhang CW (2023) Progress on phenotypic and genetic effects of chromosome-doubling in polyploid plants. J Cold-Arid Agric Sci 2(12):1079–1085
Liu G, Zhang DQ, Xie TJ, Chen HP, Luo JZ, Wu ZH (2017a) Microsatellites analysis on the genetic diversity of 39 Eucalyptus Species. Mol Plant Breed 15(9):3767–3779
Liu JH (2005) Protoplast isolation and culture of woody plants. Protocol for Somatic Embryogenesis in Woody Plants 77:553–566
Liu JQ, Zhang YN, Zhao W, Kuang HX, Wang QH (2017b) Progress in chemical constituents and pharmacology research of Stephania tetrandra. Acta Chin Med Pharm 45(3):100–103
Lu XL, Chen HL, Wei SJ, Bin XY, Ye QQ, Tang SQ (2019) Chloroplast and nuclear DNA analyses provide insight into the phylogeography and conservation genetics of Camellia nitidissima (Theaceae) in southern Guangxi. Tree Genetics & Genomes, China. https://doi.org/10.1007/s11295-019-1390-1
Leary RF, Allendorf FW, Knudsen KL (1983) Developmental stability and enzyme heterozygosity in rainbow trout. Nature 301:71–72
Ma XJ, Xiao PG (1998) The significance of genetic diversity of germplasm resources in the development of medicinal plants. China J Chin Materia Medica. 23(10):579–581, 600 (without English abstract).
Meng XC, Cao WL, Song Q, Wang XJ (2013) The significance of germplasm in the exploitation of Traditional Chinese medicine resources. Modern Chin Med 15(1):29–32 (without English abstract)
Nafees M (2015) Estimation of morphological, biochemical and genetic diversity in pomegranate (Punica granatum L.) germplasm. Doctoral Thesis
National Pharmacopoeia Committee (2015) Pharmacopoeia of the People’s Republic of China. China Medical Science and Technology Press, Beijing
Novicic ZK, Stamenkovic-Radak M, Pertoldi C, Jelic M, Veselinovic MS, Andjelkovic M (2011) Heterozygosity maintains developmental stability of sternopleural bristles in Drosophila subobscura interpopulation hybrids. J Insect Sci 11(113):1–21
Peng GQ, Tand SQ (2017) Fine-scale spatial genetic structure and gene flow of Camellia flavida, a shade-tolerant shrub in karst. Acta Ecol Sin 37(21):7313–7322. https://doi.org/10.5846/stxb201608191696
Porras-Hurtado L, Ruiz Y, Santos C, Phillips C, Carracedo A, Lareu MV (2013) An overview of STRUCTURE: applications, parameter settings, and supporting software. Front Genet 4:98. https://doi.org/10.3389/fgene.2013.00098
Purvis IW, Franklin IR (2005) Major genes and QTL influencing wool production and quality: a review. Genet Sel Evol 37(Suppl1):S97–S107
Qiu W, Su M, Xie F (2014) Tetrandrine blocks autophagic flux and induces apoptosis via energetic impairment in cancer cells. Cell Death Dis 5:84
Rosenberg NA (2004) DISTRUCT: a program for the graphical display of population structure. Mol Ecol Notes 4:137–138
Slate J, Marshall T, Pemberton J (2000) A retrospective assessment of the accuracy of the paternity inference program CERVUS. Mol Ecol 9(6):801–808
Su YJ, Wang J, Dai XB, Tang XB, Zhao DL, Zhang A, Zhou ZL, Cao QH (2018) Genetic diversity and population structure analysis of 303 sweet potato landraces using SSR markers. J Plant Genet Res 19(2):202–210
Wang GQ (2014) National compilation of Chinese herbal medicine. People’s Medical Publishing House, Beijing
Wang R, Ma TM, Liu F, Gao GQ (2017) Progress in pharmacological action and clinical application research of Radix Stephaniae Tetrandrae. China J Chin Materia Med 42(4):634–639
Wright S (1977) Evolution and the genetics of populations. Experimental Results and Evolutionary Deductions 59:815–826
Wright SW (1965) The interpretation of population structure by F-statistics with special regard to systems of mating. Evolution 19(3):395–420. https://doi.org/10.2307/2406450
Wu ZC, Wang GZ, Xu SQ (2014) Effects of tetrandrine on glioma cell malignant phenotype via inhibition of ADAM17. Tumour Biol 35:2205–2210. https://doi.org/10.1007/s13277-013-1293-y
Xie XM, Yun JF (2000) Genetic diversity and detective methods of plant. Chin J Grassland 6:51–59
Xie YJ, Li YY, Yan GY, He ZP, Bai YY (2017) SSR analysis on genetic diversity in Picria felterrae Lour. breeding materials. J South Agric 48(1):20–25
Xu YL, Cai NH, Xu Y, He CZ, Wang DW, Chen S, Duan AA (2015) Genetic variation and conservation units analysis in Pinus yunnanensis natural populations. For Res 28(6):883–891
Xue MH, Zhu LC (2012) Examples of experience in clinical application of Han Fangji. J Modern Integr Traditional Chin Western Med 21(15):1626
Yang F, Wu ML, Gao H, He QM, Xie YJ (2020) Quality regionalization and environmental impact factors analysis based on the main active ingredients of S. tetrandra. Chin J Exp Traditional Med Formulae 26(12):170–176
Zawedde BM, Ghislain M, Magembe E, Amaro GB, Grumet R, Hancock J (2015) Characterization of the genetic diversity of Uganda’s sweet potato (Ipomoea batatas) germplasm using microsatellites markers. Genet Resour Crop Evol 62:501–513
Zhang X (2014) Generation of molecular marker technology and primer design. Jiangsu Agri Sci 42(6):47–51
Zhang XX, Wang X, Hu Y, Zhou W, Chen XY, Hu XS (2019) Advances in the study of population genetic diversity at plant species’ margins. Chin J Plant Ecol 43(5):383–395
Zhen S (2017) A study on the phenotypic genetic diversity of Camellia germplasm resources in Macheng City. Asian Agric Res 9(4):77–80
Acknowledgements
Our thanks go to the editor and two anonymous reviewers for their generous comments.
Funding
This paper was supported by the National Natural Science Foundation of China (Project Numbers: 82360749, 81760681), “Gui School Young TCM Talents” project of Guangxi University of Chinese Medicine (2022C036), 2021 Basic Scientific Research Capability Promotion Project for Young and Middle-Aged Teachers in Guangxi Universities and Colleges (2021KY0288), and 2022 Innovation Project of Guangxi Graduate Education (YCSW2022357).
Author information
Authors and Affiliations
Contributions
M. W. and Y. B. wrote the main manuscript text, Z. D. prepared all tables and figures, and H. L. conducted statistical analyses. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Ethical Approval
Not applicable.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wu, M., Bai, Y., Huang, L. et al. SSR Marker–Based Genetic Diversity and Relationship Analyses of Stephania tetrandra S. Moore. Plant Mol Biol Rep (2024). https://doi.org/10.1007/s11105-024-01449-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11105-024-01449-2