Abstract
Genetic programming and artificial life systems commonly use tag matching to decide interactions between system components. However, the implications of criteria used to determine affinity between tags with respect evolutionary dynamics have not been directly studied. We investigate differences between tag-matching criteria with respect to geometric constraint and variation generated under mutation. In experiments, we find that tag-matching criteria can influence the rate of adaptive evolution and the quality of evolved solutions. Better understanding of the geometric, variational, and evolutionary properties of tag-matching criteria will facilitate more effective incorporation of tag matching into genetic programming and artificial life systems. By showing that tag-matching criteria influence connectivity patterns and evolutionary dynamics, our findings also raise fundamental questions about the properties of tag-matching systems in nature.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Computer programs ultimately translate into sequences of individual operations. These operations must specify the identities of registers and memory addresses they read from and write to. Most modern programming languages introduce a layer of abstraction that specifies operands indirectly in terms of named variables and unnamed literal values. As instances of computer programs by their very nature, genetic programs and digital artificial life systems must also specify computational operands on which to act. These computational operands include
-
program modules in genetic programs [35],
-
virtual hardware analogs like registers, memory addresses, stacks, or jump addresses in genetic programs [20, 26, 28],
-
molecules in artificial chemistries [2],
-
genes in artificial gene regulatory networks [3],
-
individual neurons or neural modules in neuroevolution [29], and
-
agents in agent-based models of complex systems [30].
In genetic programming, it is often essential for operations to undergo evolutionary adjustment to which computational elements they act on. This capability can tweak the semantics of existing evolved code, critical in particular for duplication and divergence processes commonly highlighted in discussions of evolvability [1]. This capability also allows for incorporation of new computational elements and for removal of existing computational elements. In genetic programming, dynamic reorganization of code modules can facilitate hierarchical problem-solving in genetic programming [16]. Similarly, reorganization and extension of existing computational elements is critical in the context of artificial life, where novelty and change are often of key interest [37].
Tag-based referencing, sometimes also termed “pattern matching” or “inexact referencing,” provides a practical solution for deciding computational operands. This approach attaches a tag to each computational operand that may be selected and a tag for each querying operation. Operand(s) are then selected for each query through a tag-matching process. A querying operation’s tag is compared to available operand tags. Then, typically, either:
-
the best-matching operand is selected (e.g., [33]),
-
all operands with match quality exceeding a threshold are selected (e.g., [30]),
-
operands are activated to continuously-varying degrees based on match qualities (e.g., [3]), or
-
operands are selected probabilistically based on match quality (e.g., [32]).
Inexact referencing facilitates orderly growth, shrinkage, and reconfiguration of a system’s sets of operands and operations. If an operand is deleted, it does not invalidate any existing operations, as other well-matching operands will fill its place. Likewise, new operations can be created or existing operations can be altered freely without concern for potentially invalid operands.
Indeed, inexact referencing techniques find common use in agent-based modeling [30], neuroevolution [29], artificial gene regulatory networks [3], genetic programming [19, 35], artificial chemistry [5], and artificial immunology [38]. These systems typically either use tagging schemes based on
However, other nonlinear tag-matching systems, such as Downing’s streak metric [7] and de Boer’s adjacency match metric [4], have been proposed. More exotic tag-matching extensions have explored, too, such as incorporation of a wildcard “match-any” character into tag alphabets [14] or automatic generation of operand tags based on the underlying functional behavior of each particular tagged component [23].
Although some efforts have been made to distinguish certain tag-matching criteria in terms of narrative explanations of their evolutionary properties and appeals to biological analogy [7, 31], no work has yet provided systematic, quantitative, and empirical insight into the ramifications of commonly used tag-matching criteria.
We hypothesize that properties of tag-matching criteria could affect evolvability through mechanisms including
-
bias of certain queries or operands against tight-affinity matches (i.e., tunable specificity)
-
bias to the stability of certain connections under mutation (i.e., tunable robustness),
-
bias to the likelihood of connections arising between subsets of queries and operands (i.e., modularity), and
-
mitigation of disruption under duplication of queries and operands (i.e., gene duplication [22, 27]).
In this work, we survey five tag-matching schemes: two based on integer representations, one based on Hamming distance, a “streak” metric based on the maximum length of identical substrings, and a control metric that uses a hashing algorithm to compute completely arbitrary match distances.
We explore how these tag-matching schemes differ with respect to
-
1.
geometric structure that biases or limits the patterns of connectivity that form among queries and operands (Sect. 3),
-
2.
variational properties that influence changes to connectivity observed under mutation (Sect. 4), and
-
3.
evolutionary consequences such as the rate of adaptive evolution and the quality of evolved solutions (Sect. 5).
Across several geometric analyses, we found the geometric structure of the integer metrics to be most restrictive, followed by the Hamming and then streak metrics. We observed large-effect one step mutations under the integer metrics and streak metrics, but not under the Hamming metric. Except for the control hash metric, match affinity decayed most rapidly along mutational walks under the integer metrics. Match affinity decayed slowest along mutational walks under the Hamming metric.
Evolutionary experiments also showed significant differences between tag-matching schemes. We found that rigid, one-dimensional geometric structure of the integer metrics impeded satisfaction of multiple simultaneous tag-matching requirements in scenarios where a query tag was required to closely match to more than one operand. Integer metrics also fared poorly in genetic programming experiments, even when the selected-for tag-matching scenario only involved matching queries to a single operand. Evolutionary conditions in these experiments were configured to emphasize duplication and divergence by restricting sources of variation, namely initial tag variation and ongoing insertion of randomly-generated tags. The Hamming and streak metrics generally fared best, with the streak metric outperforming the Hamming metric in some scenarios.
Although the Hamming and streak metrics generally matched or outperformed the integer metrics, confirming the extent to which these findings generalize across tag-matching application domains beyond those surveyed—particularly with respect to mutation operator used—necessitates further research. Improved understanding of the implications of tag-matching rules will directly enable more effective genetic programming practice. To a more theoretical bent, this work also provides a foundation for inquiry into the properties and mechanisms of tag-matching systems in nature.
In support of further investigations, all tag-matching techniques compared here were incorporated into the open-source Empirical C++ library [25] as interchangeable components of the MatchBin tool suite.
2 Tags and tag-matching metrics
Rigorous comparison of disparate tag-matching schemes required careful standardization of tag representation, mutation, and match quality calculation. This section summarizes the tag representation, mutation, and match quality calculation procedures used in our experiments and provides formal definitions for each of the surveyed tag-matching schemes within that framework.
In all experiments, we used 32-bit bitstrings as tags.Footnote 1 Formally, we define a tag t as a fixed-length binary vector,
where
In experiments where mutations were applied to tags, individual bits were toggled stochastically at a uniform per-bit rate.
We call the algorithm used to calculate match quality between two tags a tag-matching metric. A tag-matching metric takes two tags as operands and calculates a match distance between them. Low match distance indicates a “good” or “strong” match. High match distance indicates a “poor” or “weak” match. For consistency between metrics, we bound all match distances so that a distance of 0 is the “best” possible match and a distance of 1 is the “worst” possible match.
Occasionally, it is convenient to discuss match quality in terms of closeness instead of distance. So, we also employ a closeness terminology (which is inverse to the distance terminology). Low match closeness corresponds a “poor” or “weak” match. High match closeness corresponds to a “good” or “strong” match. Again, for consistency, we restrict closeness values between 0 and 1. A closeness of 0 corresponds to the “worst” possible match. A closeness of 1 corresponds to the “best” possible match.
We compared five tag-matching metrics: Hamming, hash, integer, bidirectional integer, and streak. The Hamming and bidirectional integer metrics are included because of their ubiquity in genetic programming and artificial life systems. The integer metric is included due to its use in seminal work exploring tag-matching in genetic programs [33,34,35]. The streak metric was proposed to model large-effect mutations observed in biology but, to our knowledge, has not yet been formally studied in an evolving system. The hash metric is introduced in this work as a control due to its completely geometrically-unstructured tag-matching scheme. Table 1 compares summary descriptions for each metric.
Sections 2.1, 2.2, 2.3, 2.4, and 2.5 provide formal definitions for each metric.
2.1 Hash metric
To our knowledge, the hash metric is original to this work. The metric produces an arbitrary, but deterministic, match distance between any two tags. In other words, the tag matching space is completely unstructured. We include it primarily to serve as a control.
The hash metric calculates match distance via a SHA1 cryptographic hash of tags t and u [9]. First, we concatenate t and u into a double-width bitstring v such that
Then, we use the OpenSSL library to generate a std::string digest of v. We then apply std::hash to map this digest to a std::size_t, \(v'\). Finally, we perform a floating point division to compute the matching distance as \(d(t, u) = v' / {\hat{V}}\) where \({\hat{V}}\) denotes the maximum representable std::size_t value.
Note that this metric is not commutative. As noted above, however, tag-matching systems inherently distinguish queries and operands. So, an ordering within each pair of tags processed in a tag-matching system will be well-defined. We use the convention of ordering the operand tag after the query tag when concatenating the tags’ bit representations.
Take as an example the two eight bit tags \(t = \langle 1, 0, 0, 1, 0, 0, 1, 1 \rangle \) and \(u = \langle 0, 0, 1, 0, 0, 0, 1, 1 \rangle \). The intricacy of the SHA1 algorithm precludes working through an example in full detail, but these two tags will have an arbitrary match distance — suppose it as 0.24. Changing any bit in either of the tags t or u will completely scramble match distance. For example, with the first bit of t flipped match distance might instead be computed as 0.89. Flipping the second bit instead could yield a totally different result.
2.2 Hamming metric
The Hamming metric computes match distance as the fraction of positions between tags t and u with mismatching bits. Formally, for n-bit bitstring tags,
As an example, consider eight bit tags \(t = \langle 1, 0, 0, 1, 0, 0, 1, 1 \rangle \) and \(u = \langle 0, 0, 1, 0, 0, 0, 1, 1 \rangle \). Sites 1, 4, 5, 6, 7, and 8 match. Sites 0, 2, and 3 mismatch. Because 3 sites mismatch, the Hamming metric would compute match distance as \(3 / 8 = 0.375\).
This metric is based on [21], originally after [11].
2.3 Streak metric
The streak metric computes match distance between bitstring tags t and u as a ratio of lengths of contiguously matching and mismatching substrings within those tags.
Formally, we can compute the greatest contiguously-matching length of n-long bitstrings t and u as,
Likewise, the greatest contiguously-mismatching length can be computed as,
As proposed in [7], the streak metric computes distance between n-bit bitstring tags t and u as,
where p(k) approximates the probability of a contiguous match at least k bits long between two randomly sampled bitstring tags.
[7] derives
However, this formula is subtly flawed. For instance, the probability of a 0-bit match according to this formula would be computed as \(p'(0) = (n - 0 + 1)/2^0 = n + 1\). This is clearly impossible—it would imply \(p'(0)> 1 \forall n > 0\).
Although correct probabilities can be calculated via dynamic programming, \(p'\) provides a useful approximation. For computational efficiency and consistency with the existing literature, we use the math proposed in [7] but clamp edge cases between 0.0 and 1.0. Additionally, because Downing uses the match closeness convention where a perfect match is scored 1.0, we subtract from 1 to convert to match distance. This yields the corrected streak metric d used in this work,
Downing motivates the streak metric by analogy to the biochemistry of enzyme-ligand binding. In motivating the metric, Downing reports mutational walk experiments that show it to exhibit greater robustness compared to integer and Hamming metrics. However, it is not demonstrated in an evolving system. To our knowledge, no further work on this metric has been published. (Although, through personal communication, we learned of some unpublished work applying the metric in a neuroevolution system.)
As an example, consider the eight bit tags \(t = \langle 1, 0, 0, 1, 0, 0, 1, 1 \rangle \) and \(u = \langle 0, 0, 1, 0, 0, 0, 1, 1 \rangle \). Sites 1, 4, 5, 6, 7, and 8 match. Sites 0, 2, and 3 mismatch. For these two tags, 5 conscecutive bits match: sites 4 through 8. Likewise, the longest streak of mismatching bits is 2 bits long: sites 2 and 3. Plugging these values into Downing’s formulas gives a match distance of
2.4 Integer metric
The integer metric computes match distance between tags t and u by counting upwards from t until u is reached. If necessary, the counting process wraps around at \(2^n\).
To accomplish this, the integer metric must interpret bitstring tags t and u as unsigned integers. We use a standard representation,
Formally, the integer metric computes distance between n-bit bitstring tags as,
Inclusion of this metric is motivated by [35], who used positive integers between 0 and 100 to name referents. Queries matched to the referent that had the next-larger value, wrapping around from 100 back to 0.
Like the hash metric, this metric is not commutative. We adopt the convention of using the query tag as t and the operand tag as u.
As an example, consider eight bit tags \(t = \langle 1, 0, 0, 1, 0, 0, 1, 1 \rangle \) and \(u = \langle 0, 0, 1, 0, 0, 0, 1, 1 \rangle \). The tag t encodes the integer value 147. The tag u encodes the integer value 35. Computing the difference between these integers mod 256 yields 144. This is equivalent to starting at 147 then counting up by 109 to reach 256, wrapping around to 0, then counting up a further 35. Dividing by 256 to normalize gives the match distance 0.5625.
2.5 Bidirectional integer metric
The bidirectional integer metric computes match distance between tags t and u by counting from t to u. The count from t to u may ascend or descend, whichever option is shorter. If necessary, the count wraps around at 0 and \(2^n\).
The bidirectional integer metric interprets bitstring tags t and u as unsigned integers using the same mapping, f, as the integer metric.
Formally, the bidirectional integer metric computes distance between n-bit bitstring tags as,
We included this metric to contrast with the integer metric. In particular, we wished to shed light on any consequences of its asymmetry and discontinuity. In figure axes and legends with tight space constraint, we refer to this metric as “Integer (bi).”
As an example, consider again the eight bit tags \(t = \langle 1, 0, 0, 1, 0, 0, 1, 1 \rangle \) and \(u = \langle 0, 0, 1, 0, 0, 0, 1, 1 \rangle \). As before, the tag t encodes the integer value 147 and the tag u encodes the integer value 35. We already found that counting from t to u mod 256 takes 144 steps. Counting from u to t mod 256 takes only 112 steps (i.e., \(147 - 35\)). So, we pick the shorter route and normalize by 256 to give the match distance 0.4375.
2.6 Match distance uniformification
Distance distributions of metrics before and after uniformification. A sample of 5,000 distances between randomly-generated tag pairs was used for each plot. a emphasizes the discretization artifacts in the Hamming and streak metrics present before and remaining after uniformification. (E.g., for 32-bit tags under the Hamming metric only 33 distinguishable match distance values are possible.) b masks distribution granularity to more intuitively depict the corrected metrics’ approximation of a uniform distribution. Supplementary Figure 14 visualizes the cumulative distribution of all 5,000 sampled distances for each metric before and after uniformification
For consistency of implementation and interpretation, all metrics output tag-matching distances between 0.0 (a “perfect” match) and 1.0 (a “worst” match)Footnote 2.
However, the distribution of tag-match distances within this range varies substantially between metrics. For example, the probability of a match distance \(<1/32\) between two randomly-sampled bitstring tags is 1/32 under the hash metric but \(1/2^{32}\) under the Hamming metric.
In order to ensure an intuitive interpretation of match distances that was consistent across all tag-matching metrics, we normalized metrics’ match distances so that the distances between pairs of randomly generated tags would follow a uniform distribution between 0.0 and 1.0. We call this process “uniformification.” In this discussion, we refer to match distance before uniformification as “raw.”
For example, two tags with a 0.01 uniformified match distance would be better-matched than 99% of randomly-generated tag pairs. Additionally, in situations where raw match distance plays a mechanistic role (for example, probabilistic matching or threshold-based cutoffs), this transformation ensures consistency across metrics.
We performed this uniformification independently for each tag-matching metric using the following Monte Carlo approximation method.
-
1.
We sampled 10,000 pairs of randomly-generated tags.
-
2.
We calculated raw match distance between each pair of generated tags using the chosen tag-matching metric.
-
3.
We organized these 10,000 sampled raw match distances into a list in ascending order.
-
4.
To ensure coverage of the entire [0.0, 1.0] interval of valid tag match scores, we bookended the sorted list of raw match distances with 0.0 and 1.0.
-
5.
We associated each list entry with its percentile ranking within the list.
-
(a)
i.e., the best-matching 0.0 match distance was associated with the percentile ranking 0.0,
-
(b)
the median match distance was associated with the percentile ranking 0.5, and
-
(c)
the worst-matching 1.0 match distance was associated with the percentile ranking 1.0.
-
(a)
-
6.
For subsequent tag match distance calculations during the experiment, we performed a lookup on this list.
-
If a single exactly-identical raw match distance existed in the list, we returned its percentile ranking as the uniformified match distance.
-
If two or more exactly-identical raw match distances existed in the list, we returned the mean percentile ranking of these entries as the uniformified match distance.
-
If no exactly-identical raw match distance existed in the list, we linearly interpolated between the next-largest and next-smallest list entries’ percentile rankings.
-
Figure 1 compares the distribution of match distances between randomly-sampled tags before and after this uniformification process across tag-matching metrics.
Error in the Monte Carlo approximation of the percentile for any raw match score is distributed binomially. With 10,000 samples, absolute error at the 50th percentile (0.5 match distance) can be bounded below 0.01 match distance units with 95% confidence and below 0.012 match distance units with 99% confidence. Absolute error at the 1st percentile (0.01 match distance) can be bounded below 0.0017 match distance units with 95% confidence and below 0.0024 match distance units with 99% confidence. (With five independent uniformification processes, 99% confidence per metric translates to 95% confidence over all metrics under Bonferroni correction.)
All work reported here employed match distance uniformification. A single match distance lookup table was used across all experiments with each metric. Note that match distance uniformification has no effect in experiments where tag-matching derives exclusively from relative ordering with no absolute match distance effect (i.e., evolutionary experiments with the graph-matching and changing-signal tasks introduced later on).
3 Geometric analyses
In this section, we consider the geometry that tag matching metrics impose over bitstring tag space. These geometries may affect the patterns of connectivity between tagged components that tend to arise, or are even possible at all.
As an illustrative example of potential geometric constraint, consider the bitstring tags \(t = \langle 0, 0, \ldots , 0 \rangle \) and \(u = \langle 1, 1, \ldots , 1 \rangle \) under the Hamming metric. No third tag v could simultaneously exhibit a tag match distance \(<0.5\) to both tags. Stated more generally, geometric constraint within a metric may enable tag pairs such that no single third tag can simultaneously exhibit a close affinity to both.
As another example of potential geometric constraint, consider the bitstring tags \(t = \langle 0, 0, \ldots , 0 \rangle \) and \(u = \langle 0, 0, \ldots , 1 , 0 \rangle \) under the bidirectional integer metric. (Here, the tag t would correspond to the integer 0 and the tag u would correspond to the integer 2.) No third tag v could simultaneously exhibit a match distance \(>0.9\) to t and \(<0.1\) to u. Stated more generally, geometric constraint may also enable tag pairs such that any third tag must either match both closely or match neither closely.
Geometric constraint varies by metric. For example, under the hash metric no pair of tags exists with either aspect of geometric constraint described above—how well a third tag v matches to t and how well it matches to u is always entirely independent.
Geometric constraint cannot be circumvented by mutation operator design. In both the Hamming metric and bidirectional integer metric examples above, the nonexistence of any tag v satisfying the given match distance criteria holds no matter how mutation is performed. The mutation operator only affects how tags in a genome move through bitstring space between generations and not how they match to other tags at a particular generation.
Geometric constraint seems likely to profoundly influence evolution in tag-matching systems. However, it is not obvious how to predict a priori how these implications ultimately play out. Geometric constraint might prove useful to facilitate modularity by allowing subsets of tag space to correspond to associated functionality [12]. However, it may also hinder generation of variation.
To study geometric constraint, we begin by comparing distributions of two statistics measuring constraint across our five tag-matching metrics: similarity constraint and dissimilarity constraint. Similarity constraint, presented in Sect. 3.1 quantifies the question, “If two tags both match closely to a third tag, will they necessarily match closely with each other?” In contrast, dissimilarity constraint in Sect. 3.2 quantifies the question, “If a certain tag matches a second tag closely and a third tag very poorly, will the second and third tag tend to match very poorly?” Finally, detour difference in Sect. 3.3 broadens beyond strong-match and strong-mismatch contexts to quantify the question, “How does a randomly-chosen waypoint affect distance between a pre-existing start and end tag?”
3.1 Similarity constraint
Similarity constraint of tag-matching metrics. a Summarizes the sampling process used to measure similarity constraint. b, c Compare distributions of similarity constraint across metrics. Supplementary Figure 15 visualizes the cumulative distribution of all sampled dissimilarity constraint values for each metric
To characterize similarity constraint for a metric m, we randomly sampled 5,000 target tags. Then, for each target tag R we randomly sampled tags until we found two secondarily-sampled tags \(S_1\) and \(S_2\) that were within a 0.01 match distance radius from the target. That is, we chose \(S_1\) and \(S_2\) such that \(m(R, S_1) < 0.01\) and \(m(R, S_2) < 0.01\). Finally, we computed the match distance \(d = m(S_1, S_2)\) between the pair of secondarily-sampled tags. Figure 2a summarizes this process.
Figure 2b provides our estimate of the similarity constraint statistic for each metric, with error bars representing a 95% confidence interval. Figure 2c shows the distribution of the similarity constraint statistic values among the 5,000 replicate samples in greater detail.
In a Euclidean space, similarity constraint corresponds to the average distance between points uniformly sampled from inside a ball (e.g., in two dimensions a circle, in three dimensions a sphere, etc.). In Euclidean space, this average distance increases with dimensionality. For reference, in a one-dimensional Euclidean space similarity constraint would measure approximately 0.0067. In a two dimensional Euclidean space, it would measure approximately 0.0091. In 32 dimensions, it would measure 0.0137 [8]. So, in some sense, this similarity constraint metric can be interpreted as an indirect measure of dimensionality. However, as we’ll see in Sect. 3.3, the Hamming, hash, and streak metric impose a decidedly non-Euclidean geometry.
3.1.1 Hash metric
The hash metric exhibits a very loose similarity constraint of 0.5083 in the mean case. Secondarily-sampled match distances are uniformly distributed between 0 and 1. This is unsurprising: given any particular set of operands, a well-behaved hash function should yield a uniform distribution of hash results. As expected, the hash metric exhibits no geometric structure.
3.1.2 Hamming metric
The Hamming metric exhibits a tighter range of sampled similarity constraint values. We estimated mean similarity constraint as 0.1627. As shown in Fig. 2c, many secondarily-sampled tag pairs are biased towards low match distances. However, secondarily-sampled tag pairs that break this constraint are also not uncommon. Among our 5,000 trials, we observed distances between secondarily-sampled tags as high as 0.7499.
Why is our estimate of the Hamming metric similarity constraint so much higher than the expected value of 0.0137 in a 32-dimensional Euclidean space? This phenomenon appears to be due to the uniformification process we applied to map raw match distances to a uniform distribution. We also calculated this statistic for the raw Hamming metric without uniformification, increasing the radius of our sampling ball to 0.25. (Only the exact target 32-bit tag itself falls within a sampling radius of 0.01.) The a priori expected distance between sampled points within a 32-dimensional ball with radius 0.25 is 0.3415. Our estimate of similarity constraint for the raw Hamming metric falls nearly in line with expectation at 0.3312.
3.1.3 Streak metric
The streak metric exhibited the second-loosest similarity constraint statistic with a mean value sampled at 0.2813. For this metric, we observed distances between secondarily-sampled tags as high as 0.9993. The streak metric retains some geometric constraint in the mean case, but allows for outliers that strongly break similarity constraint.
3.1.4 Integer metric
The integer metric exhibits loose similarity constraint in the mean case. We estimated this value as 0.5092. However, this looser mean similarity constraint appears to be an artifact of averaging between two very tight constraints: a tight constraint to 0 in one case and a tight constraint to 1 in the other. Figure 2c confirms that all sampled match distances fall under one of these cases. Because of the asymmetrical definition of the integer metric, half of pairs of similar scalar values will be in ascending order (resulting in a match distance close to 0) and half will be in descending order (resulting in wraparound search and a match distance close to 1). The integer metric appears to allow for a pair of tags closely related to a third tag either very strongly match or very weakly match, but permits no intermediate outcomes.
3.1.5 Bidirectional integer metric
For the bidirectional integer metric, we measured the similarity constraint statistic as 0.0068. This falls in line with expectation: this metric is essentially identical to a one-dimensional Euclidean space. As shown in Fig. 2c, the secondarily-sampled match distances are entirely bounded by the diameter of 0.02. This metric not only exhibits tight similarity constraint in the mean case, but also permits no outlying similarity constraint outcomes.
3.2 Dissimilarity constraint
To characterize dissimilarity constraint for each metric m, we randomly sampled 5,000 target tags. Then, for each target tag R we randomly sampled tags until we found a secondarily-sampled tag \(S_1\) that was within a 0.01 match distance radius from R and a secondarily-sampled tag \(S_2\) that was outside a 0.99 match distance radius from the R. That is, we chose \(S_1\) and \(S_2\) such that \(m(R, S_1) < 0.01\) and \(m(R, S_2) > 0.99\). Finally, we computed the match distance between \(S_2\) and \(S_1\), \(d = m(S_2, S_1)\). Figure 3a summarizes this process.
Dissimilarity constraint of tag-matching metrics. a summarizes the sampling process used to measure similarity constraint. Figure c and b compare distributions of similarity constraint across metrics. Supplementary Figure 16 visualizes the cumulative distribution of all sampled dissimilarity constraint values for each metric
Figure 2b provides our estimate of the dissimilarity constraint statistic for each metric, with error bars representing a 95% confidence interval. Figure 2c shows the distribution of the dissimilarity constraint statistic values among the 5,000 replicate samples in greater detail.
These results tell a similar story to our similarity constraint findings.
3.2.1 Hash metric
The hash metric exhibited no geometric structure—\(S_1\) and \(S_2\) were uniformly likely to exhibit any match distance between 0 and 1.
3.2.2 Hamming metric
The Hamming metric exhibited stronger geometric structure in the mean case than the streak metric. Mean secondarily-sampled distance was 0.8248. The Hamming metric also exhibited less extreme tail-end outcomes than the streak metric. We observed match distances between the secondarily-sampled tags only as low as 0.2355.
3.2.3 Streak metric
The streak metric exhibited some geometric structure in the mean case. We observed a mean secondarily-sampled distance 0.7127, significantly greater than the mean distance of 0.5 expected between arbitrarily-sampled tags.
However, outcomes that strongly broke geometric constraints also occurred. We observed distances between secondarily-sampled tags as low as 0.0002.
3.2.4 Integer metric
Again, the unidirectional integer metric exhibited a quirky result due to its noncommutative nature. The mean distance between secondarily-sampled tags was 0.0100. That is, instead of observing poor matches as we would expect, secondarily-sampled tags were much closer together than expected under arbitrary sampling. As shown in Fig. 2c, all secondarily-sampled distances observed with this metric were extremely small. So, although in the opposite way from what we would expect, match distances were still tightly constrained.
The mechanism behind this result stems from the metric’s asymmetrical nature. Under this metric, if you sample a tag that is close to a target it will be numerically slightly larger than the target. Likewise, if you sample a tag that is very far from a target, it will be numerically slightly smaller than the target (due to wraparound). Then, explaining this counterintuitive result, the distance from the slightly smaller to the slightly larger tag will be small.
3.2.5 Bidirectional integer metric
The bidirectional integer metric was highly constrained in both the mean and tail-end cases. The smallest distance between secondarily-sampled tags observed was 0.9802.
3.3 Detour difference
Similarity constraint and dissimilarity constraint quantify the geometric constraint imposed under preexisting strong matches and strong mismatches, respectively. To complement these measures, we set out to characterize the regularity, in a loose sense, of each space more broadly. This led us to our “detour difference” measure, which quantifies tag matching spaces’ respect for the triangle inequality.
Intuitively, detour difference is a measure of how adding a randomly-chosen waypoint affects total distance between pre-existing start and end points. Under the triangle inequality, the direct route is always shortest. So, if the triangle inequality is respected, detour difference should always be non-negative.
To measure detour difference, we uniformly sampled 5,000 triplets of tags A, B, and C. Then, for each metric m we calculated the \(m(A, B) + m(B, C) - m(A, C)\). Figure 4a provides a schematic of this process.
Figure 4b plots the distribution of the detour difference statistic for each metric. The Hamming, hash, and streak metrics show evidence of “shortcuts” that violate the triangle inequality.Footnote 3 For the Hamming, streak, and hash metrics the most extreme shortening detours observed were -0.76, -0.91, and -0.94, respectively. The integer and bidirectional integer metrics only exhibited shortening detours up to -0.02, which were due to minor stochastic imperfections of the uniformification process. As would be expected given their Euclidean basis, shortening detours were otherwise nonexistent for the integer metrics.
Surprisingly, given divergent results from the similarity and dissimilarity constraint measures, the distributions of detour difference for these three metrics appear similar. This suggests that geometric differences between these metrics are specially accentuated in contexts of preexisting strong matching and mismatching constraint.
4 Variational analysis
In Sect. 3 we investigated how existing tag-match relationships to a common tag influenced the distribution of match distances. This section, in contrast, focuses on how individual mutations and cumulative sequences of mutations affect a single tag-matching relationship.
In this work, we use a bit flip mutation operator. This approach enables straightforward, direct comparison between metrics. However, as a result, these analyses do not explore alternate, more semantic approaches to mutation—especially pertinent for integer tagging schemes where a tag’s integer value can be directly manipulated, for example by adding or subtracting a normally-distributed amount.Footnote 4 Any extrapolation of these results to systems with different mutation operators should be made with careful consideration, and may merit additional experimentation or analysis.
In Sect. 4.1, we report two single-step mutational analyses: one that examines the local mutational neighborhoods of loosely-affiliated tag pairs and a second that examines the local mutational neighborhoods of tightly-affiliated tag pairs. In Sect. 4.2, we perform mutational walk analysis to survey the broader mutational landscape.
4.1 Single-step mutations
Distributions of mutation effects on match distance for loosely matched (pre-mutation match distance \(> 0.5\)) and tightly matched (pre-mutation match distance \(< 0.01\)) tag pairs. Note that match closeness change (rather than mast distance change) is plotted so that better-matching mutational outcomes fall to the right and worse-matching mutational outcomes fall to the left. Error bars are 95% confidence intervals calculated using the Wilson score method with continuity correction [24]. Supplementary Figure 18 shows the cumulative distribution of all sampled match distance changes for each metric
We performed single-step mutational analyses to characterize the local mutational neighborhoods induced by each tag-matching metric.
To measure the effect of mutation on loosely-affiliated tag pairs, for each metric m we
-
randomly sampled a target tag R,
-
randomly sampled candidate tags until we found a second tag S with a match distance \(m(R, S) > 0.5\),
-
recorded match distance between R and S, \(d = m(R, S)\),
-
applied a one-bit mutation to the secondary tag S, yielding a mutated variant \(S'\),
-
measured the match distance between R and \(S'\), \(d' = m(R, S')\),
-
and then calculated change in match distance under mutation \(p = d' - d\).
We repeated this procedure to generate 5,000 samples.
To suit the instinct that positive change should correspond to an increase in match quality and negative change should correspond to a decrease in match quality, we pose all further discussion in terms of match cloesness change rather than match distance change.
The top panel of Fig. 5 visualizes the distribution of match closeness change under mutation of loosely-affiliated tags. Negative match closeness change—falling on left side of x-axes in Fig. 5—corresponds to a decrease in match quality. Conversly, positive match closeness change falls on the right side of x-axes in Fig. 5 and denotes an increase in match quality.
We used a similar procedure to measure the distribution of mutational perturbations on tightly-matched tag pairs, except that we uniformly sampled until we found a second tag S with match distance \(< 0.01\). The bottom panel of Fig. 5 visualizes the distribution of match distance change under mutation of tightly-affiliated tags. This distribution reflects the effects of one-step mutations on tags with pre-existing affinity.
4.1.1 Hash metric
The hash metric exhibits the thickest tails of mutational magnitude of all metrics. Extreme-effect one-step mutations are plentiful under this metric. Interestingly, compared to other metrics, the hash metric exhibits a greater fraction of mutations that reduce affinity between tightly-affiliated tags and a greater fraction of mutations that increase affinity between loosely-affiliated tags. This result can be attributed to the hash metric’s lack of geometric structure. Because all one-step mutations uniformly sample a new match distance, 99.5% of one-step mutations on tightly-affiliated tags will result in a looser coupling. Similarly, approximately 75% of one-step mutations on loosely-affiliated tags will result in a tighter coupling.
4.1.2 Hamming metric
The Hamming metric consistently exhibits small-magnitude match-closeness changes under mutation. High-magnitude one-step mutations do not occur under this metric. (Without normalizing match distance to a uniform distribution for randomly-sampled tags, all Hamming metric mutations would be of exactly the same magnitude, either increasing or decreasing the count of matching bits by 1.)
4.1.3 Streak metric
Unlike all other metrics, the streak metric frequently yields perfectly neutral outcomes under mutation. With loose affinity, 56% of all mutations were perfectly neutral. With tight affinity, 45% of all mutations were perfectly neutral. These perfectly-neutral mutations presumably affect regions of the bitstring involved in neither the longest-matching streak nor the longest-mismatching streak. The streak metric exhibits a thicker tail of mutational magnitude for mutations that couple loosely-affiliated tags than the integer metrics. In addition, the most extreme mutational outcomes that couple loosely-affiliated tags appear to be of a comparable magnitude to those under the integer metrics and the Hamming metric. Mechanistically, this might be due to mutations that disrupt longest-mismatching streaks. However, one-step mutations that decouple tightly-affiliated tags do not appear as potent. This might be because achieving a very poor match requires both increasing longest-mismatching streak length and decreasing longest-matching streak length.
4.1.4 Integer and bidirectional integer metrics
For both tightly- and loosely-affiliated tag pairs under the integer and bidirectional integer metrics, most mutations caused very small changes in match closeness. These mutations toggle less-significant bits of the tag’s integer representation. However, under these metrics, a small fraction of mutations affecting more-significant bits of the integer representation have a much stronger effect. Single-step mutations occasionally occurred that conspicuously couple loosely-affiliated tag pairs or conspicuously decouple tightly-affiliated tag pairs. For instance, under the integer metric 3.6% of mutations increased loosely-affiliated match closeness by at least 0.25 units and and 10.6% decreased tightly-affiliated match closeness by at least 0.25 units. Under the bidirectional integer metric, these percentages were 3.3% and 3.9%, respectively. Notably, the unidirectional integer metric exhibits more frequent strong decoupling mutations than the bidirectional integer metric, presumably due to its non-commutative quirks.
4.2 Mutational walks
Match distance along 5,000 sampled mutational walks from initially identical tags. Note that, unlike other metrics where tags always exhibit 0.0 self-match distance (i.e., at mutational step 0), the hash metric exhibits arbitrary self-match distance by construction (see Sect. 2.1)
We performed multi-step mutational analyses to characterize the broader mutational landscapes induced by each tag-matching metric. To conduct these mutational walks, for each metric m we
-
randomly generated a starting tag \(t^{(0)}\),
-
then sequentially applied randomly-chosen one-step bit flip mutations to that tag until a mutational saturation threshold (yielding a sequence of tags \(t^{(1)}, t^{(2)}, ...\)),
-
while recording match distance from the original starting tag \(m(t^{(0)}, t^{(i)})\) at each step i along the walk.
Back mutation was allowed in these experiments. We performed 65 step mutational walks, which allowed us to cover one binary order of magnitude past one expected mutation per site at 32 steps. We analyzed 1,000 replicate mutational walks for each metric, which was sufficient to distinguish metrics with bootstrapped 95% confidence intervals
Figure 6 shows how match distance increases along mutational walks for each tag-matching metric.
4.2.1 Hash metric
Due to the hash metric’s lack of geometric structure, bitwise equivalent tags do not exhibit low match distance. So, as expected, throughout the entire mutational walk this metric maintains a constant mean match distance of 0.5.
4.2.2 Hamming metric
The Hamming metric’s match distance diffuses upward slowest. The Hamming metric’s mutational walk is significantly slower to diverge than the streak metric’s at 16 and 32 steps (non-overlapping 95% CI). It is significantly slower to diverge than the integer metrics and the hash metric between steps 1 and 32, as well (non-overlapping 95% CI).
4.2.3 Streak metric
The streak metric diffuses away from zero match distance second-slowest, trailed only by the Hamming metric.
Interestingly, this result contradicts Downing’s presentation of the streak metric in [7], in which he suggests that the streak metric exhibits greater robustness because its match distance diverges more slowly under a mutational walk. This discrepancy presumably arises due to our uniformification to ensure a uniform distribution of raw match scores between 0 and 1 (Sect. 2.6). We believe that our result under uniformification is more representative because match distance corresponds to the probability that arbitrary tags would match more strongly by chance — which directly relates to how effectively a operand tag competes to be the “best” match for a query.
To compare mutational landscapes between the streak and integer metrics under more realistic circumstances (i.e., where tags do not begin exactly perfectly-matched), we performed a secondary mutational walk experiment. This experiment was conducted exactly as before, except instead of starting with exactly-identical tags it started with a pair of tags that was randomly sampled for match distance \(<0.01\).
As shown in Supplementary Figure 28a, this experiment confirmed greater robustness of the Hamming metric under mutation. The streak metric’s match distance was significantly greater than the Hamming metric between mutational steps 2 and 16 (non-overlapping 95% CI). Our result remained consistent when replicating the experiment with 64-bit tags (Supplementary Figure 29a).
4.2.4 Integer metric and bidirectional integer metric
The binary representation of the integer and bidirectional integer metrics (Sect. 2.4) induces a long-tailed distribution of mutational effect sizes. Under this distribution, value changes of each binary order of magnitude are equally likely (corresponding to toggling the bit at each position). Occasional large-effect mutations provide plausible explanation for the bidirectional integer metric’s rapid increase in match distance under mutation relative to the Hamming and streak metrics. The integer metric experiences even more rapid dilation of match distance under mutation. Under this metric, half of first mutational steps cause a wraparound effect, immediately spiking the average match distance to 0.5. Supplementary Figure 27 shows match distance variance decreasing as the integer metric mutational walk proceeds away from match distances biased to 0 or 1.
5 Evolutionary analysis
Sections 3 and 4 reported how each tag-matching metric induced constraints on tag match affinities and the distribution of mutational outcomes. We now move on to investigate whether—and how—these geometric and variational properties affect evolution of tag-mediated connectivity in evolutionary scenarios.
We begin with a toy problem, presented in Sect. 5.1, which allowed us to systematically vary the level of network constraint selected for. That is, these experiments compared scenarios where individual tags needed to ensure simultaneously tight affinity with several other tags (more constrained) and where individual tags only needed to ensure tight affinity with one other tag (less constrained). In this toy problem, we define a target connection topology between tagged queries and operands then select for sets of tags that exhibit high-affinity pairings between connected topology elements.
In order to investigate potential consequences of tag-matching metrics in a more complex, applied setting, we also evolved full-fledged SignalGP programs that use tag matching to mediate module activation.
The SignalGP genetic programming representation employs tag-based referencing to facilitate event-driven program execution [19]. In SignalGP, programs are segmented into modules (functions) that may be automatically triggered by exogenously- or endogenously-generated signals. Tags specify the relationship between signals and signal-handlers (program modules), triggering the module with the closest matching tag to run its linear sequence of instructions.
The SignalGP instruction set, in addition to including traditional GP operations, allows programs to generate arbitrarily-tagged internal signals and broadcast arbitrarily-tagged external signals. SignalGP also supports genetic regulation with promoter and repressor instructions that, when executed, allow programs to adjust how well subsequent signals match with a target function (specified with tag-based referencing) [17]. See [19] for a more detailed description of SignalGP.
To ensure a broad survey of tag-matching functionality, we performed experiments with a complementary pair of SignalGP problems:
-
the Changing-signal Task (Section 5.2), which is known to select for sparse tag interactions (i.e., low constraint), and
-
the Directional-signal Task (Section 5.3), which is known to select for more dense tag interactions (i.e., high constraint).
These experiments used bit flip mutation operators across all metrics. Note again that generalization to systems with alternate mutation operators (i.e., especially with respect to integer-based tags) should be made carefully.
5.1 Graph-matching task
The graph-matching task provides a barebones experimental system to isolate tag matching evolutionary dynamics. Genomes in this domain consisted solely of a set of 32 tags, partitioned evenly as 16 query tags and 16 operand tags. Operand tags serve as available tag-matching targets and query tags are used as tag-matching lookups. The goal of the graph-matching task is to establish of an arbitrary, fixed pattern of tag-matching connectivity between query tags and operands.
Example target graph layouts used in 32-vertex graph-matching evolutionary experiments. Blue dots represent tagged vertices. Black lines represent selected-for tight affinity relationships. Layouts differ in total number of selected-for affinities (“mean degree”) and whether selected-for affinities were evenly or randomly distributed between vertices (“structure”)
Trajectories of adaptive evolution for each tag-matching metric on the 32-vertex graph-matching task. Maximum fitness is the best fitness value for any individual within a population. Reported results use each metric’s best-performing per-bit mutation rate. (See Supplementary Figure 21 for survey of how mutation rate affects adaptive evolution under each metric.) Note logarithmic x-axes. Shaded area represents bootstrapped 95% confidence intervals across 50 replicate observations
Each experiment with this task selected for a distinct, randomly-determined pattern of connectivity between query tags and operand tags. Formally, we used randomly-generated bipartite graphs to specify this target connectivity. Each bitstring tag in the genome corresponded directly to a vertex in the target graph; one partition of the bipartite graph contained all query tags and the other contained all operand tags. Figure 8 shows example target graph layouts.
To evaluate the fitness of a genome, we harvested its operand tags and placed them into a tag-matching data structure. This data structure allowed us to rank operand tags in terms of match distance from each particular query tag. In order to compare tag-encoded connectivity against the target graph, we had to decide whether or not each operand was “connected to” by a query tag. For each query, we considered only the top n best-matching operands as “connected to.” This rank cutoff n was determined independently for each query tag. We used the degree of a query’s corresponding vertex as its rank cutoff n. So, for example, if a vertex had three outgoing edges then we would consider the three best-matching operands of the corresponding tag as “connected to.”
For the purposes of fitness evaluation, we considered all “connected to” matches equivalent. So, with respect to a particular query tag, we took fitness as the fraction of the “connected to” operands that shared an edge with the query in the target bipartite graph. Along these lines, we took overall fitness as the fraction of established connections that correctly corresponded to edges in the target graph. Figure 7 summarizes the evaluation process for the graph-matching task.
We controlled the degree of tag-matching constraint imposed by the target graph by manipulating:
-
1.
mean degree—the number of edges between queries and operands, and
-
2.
structure—whether edges were assigned evenly such that all vertices had identical degree (regular structure) or were assigned at random, likely causing some vertices to have high degree (irregular structure).
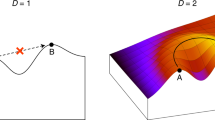
We tested target graphs with mean degree 1 and 2 and both regular and irregular construction.
Irregular, degree 2 graphs imposed the most tag-matching constraint. High-degree vertices in these graphs were strongly constrained by many simultaneous connection criteria. Figure 8b shows an example irregular, degree 2 graph.
Regular, degree 1 graphs imposed the least tag-matching constraint. Every vertex participates in exactly one connection. Figure 8c shows an example regular, degree 1 graph.
For each target graph configuration, we surveyed metrics’ performance over ten per-bit mutation rates ranging from 0.75 expected bit mutations per genome to 16.0 expected bit mutations per genome. For each combination of metric and target graph configuration, we report results from the most favorable mutation rate (as defined by sum population-maximum fitness across evolutionary generations).Footnote 5 Table 2 provides the best-performing mutation rate for each metric across target graph configurations. Best-performing mutation rates varied greatly, ranging from as low as 0.75 expected mutations per genome for the hash metric to as high as 8.0 for the integer metrics. The Hamming and streak metrics had intermediate best-performing mutation rates between 1.5 and 4.0 expected mutations per genome.
We ran 100 replicate 512-generation evolutionary runs for each mutation rate/target graph/tag-matching metric combination. Populations were initialized with genomes comprising randomly-generated tags. These runs had a well-mixed population of size 500 and used tournament selection with tournament size 7. The population was initialized with randomly-generated genomes. Figure 9 plots population-maximum fitness over the course of these evolutionary runs. We performed the same evolutionary experiment with larger 64-vertex target graphs and observed qualitatively similar results (Supplementary Figures 22 and 23).
5.1.1 Hash metric
Surprisingly, the hash metric enables faster adaptive evolution than all other metrics on the least-constrained target graph (Fig. 9; non-overlapping 95% CI). On more-constrained target graphs with mean degree 2, the hash metric’s advantage in rapid adaptive evolution disappears. In fact, on the most-constrained target graph (irregular structure with mean degree 2) the hash metric yields significantly lower-quality solutions at the end of evolutionary runs than the streak and Hamming metrics (Fig. 9; non-overlapping 95% CI).
5.1.2 Hamming and streak metrics
The streak metric facilitates slightly faster adaptive evolution than the Hamming metric, especially on mean degree 2 regularly configured target graphs (Fig. 9; non-overlapping 95% CI).
5.1.3 Integer metrics
The integer and bidirectional integer metrics successfully match the least-constrained target graph (regular structure with mean degree 1) but yield lower-quality solutions than other metrics on more constrained target graphs (Fig. 9; non-overlapping 95% CI).
5.1.4 Identical initialization experiment
Trajectories of adaptive evolution for each tag-matching metric on the 32-vertex graph-matching task with identically-initialized initial genomes. Maximum fitness is the best fitness value for any individual within a population. Reported results use each metric’s best-performing per-bit mutation rate. (See Supplementary Figure 23 for survey of how mutation rate affects adaptive evolution under each metric.) Note logarithmic x-axes. Shaded area represents bootstrapped 95% confidence intervals across 100 replicate observations
In a follow-up experiment to diagnose metrics’ capabilities to generate variation, we initialized populations with an all-identical set of initial tags (instead of random initial tags). Figure 10 shows trajectories of adaptive evolution across tag-matching metrics and target graph configurations.
Surprisingly, integer metrics exhibited the fastest initial adaptive evolution—even outperforming the volatile hash metric. However, this effect dissipated after about 10 generations and results at subsequent generations were generally the same as before.
5.2 Changing-signal task
The changing-signal task requires SignalGP programs to express a specific response to each of K environmental signals. Environmental signals correspond to a unique tagged event. Programs express a response by executing one of K response instructions. Successful programs can “hardcode” each response to the appropriate environmental signal by ensuring that each tagged environmental signal best matches the function containing its correct response. Thus, in this experiment SignalGP module tags are minimally constrained—each needs to only match with a single environmental signal.
During evaluation, we afford programs 64 virtual CPU cycles to express the appropriate response after receiving a signal. Once a program expresses a response or the allotted time expires, we reset the program’s virtual hardware (resetting all executing threads and thread-local memory), and the environment produces the next signal. Evaluation continues until the program correctly responds to each of the K environmental signals or until the program expresses an incorrect response. During each evaluation, programs experience environmental signals in a random order; thus, the correct order of responses will vary and cannot be hardcoded.
For each tag-matching metric, we evolved 200 replicate populations (each with a unique random number seed) of 500 asexually reproducing programs in an eight-signal environment for 100 generations. We identified the most performant per-bit tag mutation rates (from a range of possible mutation rates) for each metric on the changing-signal task:
-
0.01 for the Hamming and streak metrics,
-
0.002 for the hash metric, and
-
0.02 for the integer and bidirectional integer metrics.
Aside from tag mutation rate, the overall configuration used for each metric was identical.
We limited tag variation in offspring to tag mutation (bit flips) by initializing populations with a common ancestor program in which all tags were identical and by disallowing mutations that would insert instructions with random tags. Supplemental Section C gives the full configuration details for this experiment, including a guide for replication.
Figure 11 gives the number of replicates that produced a successful SignalGP program (i.e., capable of achieving maximum fitness) for each tag-matching metric on the changing-signal task. We compared the number of successful replicates across metrics using a pairwise Fisher’s exact test with a Holm correction for multiple comparisons.
5.2.1 Hash metric
The hash metric significantly outperformed both integer metrics (\(p < 4\times 10^{-10}\)).
We suspect that the hash metric performed well because it maximizes generation of phenotypic variation (i.e., signal-function relationships). Even a single bit flip in a tag is likely to completely re-order which other tags it best matches with. The capacity to quickly generate large amounts of phenotypic variation allows evolution to explore large swaths of the fitness landscape from generation to generation, which is particularly useful in this low-constraint problem. However, as evidenced by better performance of the Hamming and streak metrics, this capacity to generate phenotypic variation trades off with tag-matching robustness—under this metric, a single bit mutation may also scramble established relationships with other tags.
5.2.2 Hamming and streak metrics
The Hamming and streak metrics performed significantly better than all other metrics (\(p < 5\times 10^{-11}\)); however, there was no significant difference in performance between the Hamming and streak metrics. To assess whether the streak metric produced solutions in fewer generations than the Hamming metric, we ran 200 new replicates of each condition until 100 replicates produced a solution and recorded the number of generations that elapsed (Supplementary Figure 19). We found no difference in generations elapsed between the Hamming and streak metrics.
5.2.3 Integer metrics
Among surveyed tag-match metrics, the integer metrics performed worst. We observed no adaptive difference between the integer and bidirectional integer metrics.
5.3 Directional-signal task
As in the changing-signal task, the directional-signal task requires that programs respond to a sequence of environmental cues. In the directional-signal task, however, the correct response to signal depends on the history of previously experienced signals. In the directional-signal task, there are two possible environmental signals—a “forward signal” and a “backward signal” (each with a distinct tag)—and a cycle of four possible responses. If a program receives the forward-signal, it should express the next response in the cycle. Conversely, a program should express the previous response in the cycle if it receives the backward signal. For example, if response three is currently required, a subsequent forward signal indicates that response four is required next, while a backward signal would instead indicate that response two is required next. Because the appropriate response to both the backward and forward signals change over time, successful programs must regulate which functions these signals trigger (rather than hardcode each response to a particular signal).
SignalGP module tags are more constrained than in the changing-signal task, potentially needing to match to queries by genetic regulation instructions in addition to several tagged events (e.g., environmental signals or internally-generated signals) depending on internal regulatory state. Indeed, in other work, we have observed that the directional-signal task yields significantly more interconnected regulatory networks than the changing-signal task [18].
We evaluated programs on all possible four-signal sequences of forward and backward signals (sixteen total). For each program, we evaluated each sequence of signals independently, and a program’s fitness was taken as its aggregate performance. Otherwise, evaluation on a single sequence of signals mirrors that of the changing-signal task.
We used an identical experimental design for the directional-signal task as in the changing-signal task. However, we evolved programs for 5,000 generations (instead of 100) and re-parameterized each metric’s tag mutation rate:
-
0.001 for the Hamming and hash metrics,
-
0.002 for the integer and streak metrics, and
-
0.0001 for the bidirectional integer metric.
Full configuration details for this experiment, including a guide for replication, appears in Supplemental Section C.
Figure 12a gives the number of replicates that produced a successful SignalGP program for each tag-matching metric on the directional-signal task.
5.3.1 Hamming and streak metrics
Again, the Hamming and streak metrics performed significantly better than all other metrics (Fisher’s exact with a Holm correction for multiple comparisons, \(p < 0.0008\)). We observed no significant difference in solution count between the Hamming and streak metrics, however.
As in the changing-signal task, we assessed whether the streak metric produced solutions in fewer generations than the Hamming metric, running 200 new replicates of each condition until 100 replicates produced a solution and recorded the number of generations that elapsed (Fig. 12b). Among this subset of replicates, we found significantly faster generations-to-solution under the streak metric compared to the Hamming metric (Wilcoxon rank-sum test, \(p < 0.0016\)).
5.3.2 Integer and hash metrics
As in the changing-signal task, we observed no difference in success between the integer and bidirectional integer metrics on the directional-signal task. Again, the hash metric outperformed both the integer metrics (\(p < 3\times 10^{-5}\)).
6 Discussion
Evolutionary experiments in Sect. 5 showed that choice of tag-matching metric significantly shaped adaptive evolution. In all experiments, adaptive evolution progressed more slowly under some metrics and more quickly under others. In most experiments, final solution quality was also affected by tag-matching metric choice.
We found that network constraint (the number of tags a query or operand needs to simultaneously establish affinity with) influenced the relative performance of tag-matching metrics. In target-matching evolutionary experiments, we found that the hash metric enabled rapid adaptive evolution toward targets with low network constraint. Under high network constraint, however, the hash metric yielded poor-quality solutions. The integer metrics also yielded poor-quality solutions for target graphs with network constraint. In some more-constrained cases, the streak metric enabled more rapid adaptive evolution than the Hamming metric.
Geometrical analyses of how tag-matching metrics constrain patterns of connectivity between tags helps to explain the interplay between metrics and problem network constraint. When problems require tag-matching configurations involving more than a single query-operand tag pair (e.g., a particular query tag matching closely to two different operand tags), these geometric constraints can make certain configurations unlikely or even impossible. The bidirectional integer metric exhibited the tightest geometrical constraint in our analyses. The unidirectional integer metric also exhibited tight geometrical constraint, but quirks of its non-commutative construction allowed that constraint to split across perfect- and worst-matching extremes. Hamming and streak metrics exhibited looser geometric constraint, with the streak metric allowing for edge cases that very strongly break constraints. Finally, the hash metric exhibited no geometrical constraint because it had no geometrical structure.
Geometrical constraint explains with poor performance of the integer metrics on problems with high network constraint. However, it is surprising that the geometrically unconstrained hash metric performed best on problems with low network constraint and worst on problems with high network constraint—the opposite of what may otherwise be expected. Analysis of metrics’ mutational properties provides an explanation.
Analysis of the effect of bitwise mutation on match distance score revealed significant differences between metrics. Under the Hamming metric, all mutations have small effects on match distance score. In contrast, under the integer metrics, rare mutations can have strong effects on match distance score. The streak metric also exhibited strong-effect mutations, particularly with respect to coupling loosely-affiliated tags. The hash metric exhibited the fattest tails of mutational magnitude, with strong-effect mutations occurring frequently. Interestingly, the hash metric also exhibited sign-outcome frequencies that differed from the other metrics: mutations that decoupled tightly-matching tags and mutations that coupled loosely-matching tags were more frequent compared to other metrics.
The Hamming metric exhibited the greatest robustness to mutation along mutational walks, followed by the streak metric. The integer metrics, in particular the unidirectional integer metric, exhibited less robustness. The hash metric, where all one-step mutations scramble match distance, exhibited the least robustness.
The hash metric’s volatility under mutation may explain how it facilitates rapid adaptive evolution in unconstrained problem domains: this rapid evolution may be due to the hash metric’s ability to rapidly generate variation. Likewise, the hash metric’s poor performance on high-constraint problem domains may also be explained by mutational volatility. If tags are selected for to simultaneously satisfy two matching requirements (e.g., to match closely to two other tags), it makes sense that adaptive evolution would be stymied if any change to the focal tag necessarily scrambled behavior on both requirements. This would prevent satisfaction of a single tag-matching requirement from being used as a stepping stone towards satisfaction of both.
Results from evolution experiments with a full-fledged genetic programming system suggest that additional mechanisms may influence tag-matching metric performance.
In SignalGP experiments, we found that the Hamming and streak metrics yielded successful solutions the most frequently. On the directional-signal task, which tends to require denser (more constrained) interaction networks, we found evidence that the streak metric enabled more rapid adaptive evolution than the Hamming metric. The hash metric had the next-best performance in SignalGP experiments, yielding more solutions than the integer metrics, which both performed comparably poorly.
Although the hash metric performed best in low-constraint target-matching experiments, it was outperformed in low-constraint SignalGP experiments by the streak and Hamming metrics. Likewise, the integer metrics were outperformed by the streak and Hamming metrics in low-constraint SignalGP experiments even though they had not been in the low-constraint target-matching experiments.
The mechanisms driving degradation of hash and integer metric performance in the GP experiments remain unclear. The degradation may be due to better streak and Hash support for duplication and differentiation processes along SignalGP lineages, in which instruction and module count can grow over time. However, altering the graph-matching task to emphasize generation of variation and tag differentiation by disallowing initial tag variation did not reproduce degraded hash and integer metric performance. Another possible causal difference could be fitness landscape ruggedness. Within the target-matching experiments, the fitness benefit of a particular query-operand match does not depend on other query-operand matches In the SignalGP experiments, where modules can form arbitrary interweaving call chains, this is likely not the case. Further experiments are needed to understand the evolutionary dynamics of non-trivial tag-matching systems and the interplay of those dynamics with tag-matching metrics.
Relative to the other metrics, the streak metric appears to offer intermediate variational and geometric properties. Figure 13 depicts a schematic summary of this observation. It exhibits some, but not strict, geometric constraint. Many mutations are neutral or near-neutral (like the integer and Hamming metrics) but a fat tail of extreme-effect mutations also occur (like the hash metric). The streak metric exhibits robustness under mutational walks that falls between the Hamming and integer metrics. These mechanistic observations offer a potential explanation for the streak metric’s strong performance facilitating adaptive evolution under high-constraint conditions. However, whether these mechanistic explanations are sufficiently complete—especially with respect to the streak metric’s outperformance of the Hamming metric under high-constraint conditions—is unclear.
6.1 Practical recommendations
Our results highlight the dependence of tag-matching metrics’ performance on problem domain. So, consideration of properties of the problem domain at hand should drive the decision of which tag-matching metric to use in a particular system.
A major practical advantage of integer-based metrics is the possibility for log time lookup of operands (i.e., via binary search). However, the integer metrics performed competitively only within a particular class of problem domains. In all high-constraint problem domains, the integer metrics performed poorly. Among low-constraint problem domains, the integer metrics only performed well on the toy graph-matching task—they did not perform well on the low-constraint GP changing signals task. At present, it is unclear what problem domain property stymied the integer metrics in the low-constraint GP changing signals task—the potential for the operand set to grow over time (e.g., duplication and divergence), fitness landscape ruggedness (e.g., epistasis), or something else.
However, within the low-constraint target-matching problem domain where integer metrics performed well (i.e., a fixed-size problem with a smooth fitness landscape), the hash metric actually performed slightly better. So, under these conditions, the hash metric may be preferable when log time lookup is not critical.
Besides the low-constraint target-matching problem domain, the Hamming and streak metrics performed significantly better than the integer metrics. On the low-constraint target-matching problem, both did evolve full solutions, (although the Hamming metric was slightly slower than the streak and integer metrics). Of particular note, the Hamming and streak metrics performed best in our GP tests. So, both metrics appear to be a robst choice across most contexts.
Choosing between the two will likely depend on implementation considerations: the streak metric facilitated faster adaptive evolution in some experiments, but is more computationally expensive to calculate than the Hamming metric. Future work should investigate whether a streamlined version of the streak metric—for example, ignoring mismatching streak length and only considering matching streak length—suffices to capture its desirable properties.
A critical qualification on these recommendations must be noted: they assume the bitwise mutation used in this work. Although in experiments reported in Supplementary Section B we did not find that a simple Gaussian mutation operator improved the performance of integer metrics on the graph-matching task, future work should more thoroughly explore the extent to which our findings generalize under alternate mutation operators.
7 Conclusion
Better understanding the mechanistic properties and functional implications of tag-matching criteria will enable more effective incorporation of tag matching in evolutionary systems. Within genetic programming, tuning tag-matching criteria could facilitate faster evolution of higher-fitness solutions. In this domain, tag matching approaches have been highlighted, particularly, for their potential to enable dynamic, modular reconfiguration of evolved programs at runtime [17, 35]. Likewise, within artificial life, tuning tag-matching criteria could enable more novelty and complexity within evolving systems. There has been interest, in particular, in the potential for tag-based referencing to facilitate inter-species interactions in digital ecologies [6].
Our analyses suggests a key role of network constraint in the interaction between a tag-matching scheme and problem domain. Applications where queries much match tightly with multiple operands require high-dimensional tag-matching criteria.
The surprisingly strong performance of the hash metric on low constraint toy problems highlights how tag-matching criteria can facilitate generation of phenotypic variation.
Important open questions remain with respect to tag-matching criteria. In particular, the relationships between tag-matching criteria and specificity, modularity, robustness, and the process of duplication and divergence should be explored. Evolvability or information-theoretical analyses may prove fruitful in this regard [36]. How to systematically design new tag-matching metrics with desirable evolutionary properties also remains an open problem. Mutation operator design should also be considered. We also need algorithms capable of performing fast look ups under high-dimensional or irregular tag-matching metrics, ideally achieving sublinear time complexity on large sets of referents.
Tag-like mechanisms play a central role mediating interaction and function across the spectrum of biological scale [14]. By shining light on previously-unexplored mechanistic and evolutionary properties of tagging systems, we hope that insight into artificial tag models can translate into a more nuanced appreciation—and algorithmic mimicry—of natural systems.
Availability of data and material
The code used to perform and analyze our experiments, our figures, and data from our experiments is available via the Open Science Framework at https://osf.io/gw5mc/ [10]. Data and figures are also organized alongside related manuscript source files at https://github.com/mmore500/tag-olympics-writeup.
Notes
Many other tag representations are possible, including higher-cardinality alphabets [13] and floating point values. Although for tractability and consistency we do not explore them in this initial work, we do include some commentary in Supplementary Section A on how behavior of alternate representations might differ from the bitstring tags used in this work.
The uniformified score for the Hamming metric’s median matching case falls into the slightly leftern 31st bin instead of the central 32nd bin. This true median value for the Hamming metric was approximated to a match distance of 0.4901. This outcome falls within the outer extreme, but still plausible, error of the Monte Carlo approximation, at \(p=0.024\) under a double-tailed exact binomial test. Given the five uniformifications performed (one for each metric), this result is less surprising still. See main text for detailed discussion of expected error under the Monte Carlo approximation method.
The raw Hamming metric does respect the triangle inequality, so presumably this result is due to the uniformification.
Although not universal, use of bit flip mutation operators with integer representations is not entirely uncommon. See e.g., [7].
Additionally, in a supplementary experiment we found that the bitwise mutational operator outperforms a simple Gaussian mutation operator on the 32-vertex graph-matching task discussed in Sect. 5.1. This simple operator applies Gaussian noise to the integer value of all tags in a genome every generation; addition of an additional “mutation probability” parameter may yield better results. Supplementary Section B provides details on this experiment. This finding further motivates our limitation of scope to bitwise mutations in this initial work.
Supplementary Figure 21 shows the rate each metric’s rate of adaptive evolution across surveyed mutation rates for each target graph configurations. All treatments’ optimal mutation rates fall within the range of mutation rates surveyed, except for the hash metric on the regular target graphs. In this case, peak performance was observed on the lowest sampled mutation rate.
References
L. Altenberg et al., The evolution of evolvability in genetic programming. Adv. Genet. Program. 3, 47–74 (1994)
R. J. Bagley, J. D. Farmer, Spontaneous emergence of a metabolism. Technical report, Los Alamos National Lab., NM (USA) (1990)
W. Banzhaf, Artificial regulatory networks and genetic programming, in Genetic Programming Theory and Practice. ed. by R. Riolo, B. Worzel (Springer, Cham, 2003), pp.43–61
R.J. de Boer, A.S. Perelson, Size and connectivity as emergent properties of a developing immune network. J. Theor. Biol. 149(3), 381–424 (1991)
P. Dittrich, J. Ziegler, W. Banzhaf, Artificial chemistries-a review. Artif. Life 7(3), 225–275 (2001)
E. Dolson, C. Ofria, Digital evolution for ecology research: a review. submitted (2021)
K.L. Downing, Intelligence Emerging: Adaptivity and Search in Evolving Neural Systems (MIT Press, London, 2015)
S.R. Dunbar, The average distance between points in geometric figures. Coll. Math. J. 28(3), 187–197 (1997)
D. Eastlake, P. Jones, Us secure hash algorithm 1 (sha1) (2001)
E.D. Foster, A. Deardorff, Open science framework (OSF). J. Med. Libr. Assoc. JMLA 105(2), 203 (2017)
R.W. Hamming, Error detecting and error correcting codes. Bell Syst. Tech. J. 29(2), 147–160 (1950)
J.H. Holland, Concerning the emergence of tag-mediated lookahead in classifier systems. Phys. D 42(1–3), 188–201 (1990)
J.H. Holland, Adaptation in Natural and Artificial Systems: An Introductory Analysis with Applications to Biology, Control, and Artificial Intelligence (MIT press, London, 1992)
J.H. Holland, Signals and Boundaries: Building Blocks for Complex Adaptive Systems (MIT Press, London, 2012)
M.C. Jones, Simple boundary correction for Kernel density estimation. Stat. Comput. 3(3), 135–146 (1993)
J.R. Koza, Scalable learning in genetic programming using automatic function definition, in Advances in Genetic Programming. ed. by K.E. Kinnear Jr. (MIT Press, Cambridge, MA, USA, 1994), pp.99–117
A. Lalejini, M. A. Moreno, C. Ofria, Tag-based regulation of modules in genetic programming improves context-dependent problem solving. Genet. Program. Evol. Mach., pp. 1–31 (2021a)
A. Lalejini, M.A. Moreno, C. Ofria, Tag-based regulation of modules in genetic programming improves context-dependent problem solving. Genet. Progr. Evol. Mach. 22(3), 325–355 (2021)
A. Lalejini, C. Ofria, Evolving event-driven programs with signalgp. In: Proceedings of the genetic and evolutionary computation conference, pp. 1135–1142 (2018)
A. Lalejini, C. Ofria, Tag-accessed memory for genetic programming. In: Proceedings of the genetic and evolutionary computation conference companion on - GECCO ’19, pp. 346–347, Prague, Czech Republic. ACM Press (2019a)
A. Lalejini, C. Ofria, What else is in an evolved name? exploring evolvable specificity with signalgp, in Genetic Programming Theory and Practice XVI. ed. by W. Banzhaf (Springer, Cham, 2019), pp.103–121
E. B. Lewis, A gene complex controlling segmentation in drosophila. In: Genes, Development and Cancer, pp. 205–217. Springer (1978)
M.A. Lones, A.M. Tyrrell, Modelling biological evolvability: implicit context and variation filtering in enzyme genetic programming. Biosystems 76(1–3), 229–238 (2004)
R.G. Newcombe, Two-sided confidence intervals for the single proportion: comparison of seven methods. Stat. Med. 17(8), 857–872 (1998)
C. Ofria, E. Dolson, A. Lalejini, J. Fenton, M. A. Moreno, S. Jorgensen, R. Miller, J. Stredwick, L. Zaman, J. Schossau, L. Gillespie, N. C. G, A. Vostinar, Empirical (2019)
C. Ofria, C.O. Wilke, Avida: a software platform for research in computational evolutionary biology. Artif. Life 10(2), 191–229 (2004)
S. Ohno, Evolution by Gene Duplication (Springer, Berlin, 2013)
T.S. Ray, An approach to the synthesis of life. Artif. life II 11, 371–408 (1991)
J. Reisinger, R. Miikkulainen, Acquiring evolvability through adaptive representations. In: Proceedings of the 9th annual conference on Genetic and evolutionary computation, pp. 1045–1052 (2007)
R.L. Riolo, M.D. Cohen, R. Axelrod, Evolution of cooperation without reciprocity. Nature 414(6862), 441–443 (2001)
A. Scherer, A. Noest, R.J. de Boer, Activation-threshold tuning in an affinity model for the t-cell repertoire. Proc. R. Soc. Lond. B 271(1539), 609–616 (2004)
P.E. Seiden, F. Celada, A simulation of the humoral immune system, in Theoretical and Experimental Insights into Immunology. ed. by A.S. Perelson (Springer, Cham, 1992), pp.49–62
L. Spector, K. Harrington, T. Helmuth, Tag-based modularity in tree-based genetic programming. In: Proceedings of the 14th annual conference on Genetic and evolutionary computation, pp. 815–822. ACM (2012)
L. Spector, K. Harrington, B. Martin, T. Helmuth, What’s in an evolved name? the evolution of modularity via tag-based reference, in Genetic Programming Theory and Practice IX. ed. by R. Riolo (Springer, Cham, 2011), pp.1–16
L. Spector, B. Martin, K. Harrington, T. Helmuth, Tag-based modules in genetic programming. In: Proceedings of the 13th annual conference on Genetic and evolutionary computation, pp. 1419–1426. ACM (2011b)
D. Tarapore, J.-B. Mouret, Evolvability signatures of generative encodings: beyond standard performance benchmarks. Inf. Sci. 313, 43–61 (2015)
T. Taylor, M. Bedau, A. Channon, D. Ackley, W. Banzhaf, G. Beslon, E. Dolson, T. Froese, S. Hickinbotham, T. Ikegami et al., Open-ended evolution: perspectives from the OEE workshop in York. Artif. Life 22(3), 408–423 (2016)
J. Timmis, A. Hone, T. Stibor, E. Clark, Theoretical advances in artificial immune systems. Theoret. Comput. Sci. 403(1), 11–32 (2008)
Acknowledgements
Thanks to members of the DEVOLAB, in particular Nathan Rizik for help developing our tag-matching software infrastructure. This research was supported by Michigan State University through the computational resources provided by the Institute for Cyber-Enabled Research. This material is based upon work supported by the National Science Foundation Graduate Research Fellowship under Grant No. DGE-1424871. Any opinions, findings, and conclusions or recommendations expressed in this material are those of the author(s) and do not necessarily reflect the views of the National Science Foundation.
Funding
This research was supported in part by NSF Grants DEB-1655715 and DBI-0939454. This material is based upon work supported by the National Science Foundation Graduate Research Fellowship under Grant No. DGE-1424871.
Author information
Authors and Affiliations
Contributions
MAM and AL contributed to the study conception and design. Material preparation, data collection and analysis of genetic programming benchmarks were performed by AL. Other material preparation, data collection and analysis of genetic programming benchmarks were performed by MAM. The first draft of the manuscript was written by MAM and AL and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Code availability
We implemented our experimental systems using the Empirical library for scientific software development in C++, available at https://github.com/devosoft/Empirical [25]. Software written for these experiments is available at https://github.com/amlalejini/Exploring-tag-matching-metrics-in-SignalGP/tree/1.0 and https://github.com/mmore500/tag-olympics/tree/v1.1.1.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Moreno, M.A., Lalejini, A. & Ofria, C. Matchmaker, matchmaker, make me a match: geometric, variational, and evolutionary implications of criteria for tag affinity. Genet Program Evolvable Mach 24, 4 (2023). https://doi.org/10.1007/s10710-023-09448-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10710-023-09448-0