Abstract
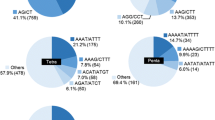
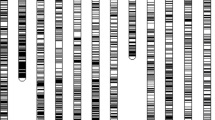
Wild rice is a valuable resource for the genetic improvement of cultivated rice (Oryza sativa L., AA genome). Molecular markers are important tools for monitoring gene introgression from wild rice into cultivated rice. In this study, Simple sequence repeat (SSR) markers were used to analyze interspecific hybrids of O. sativa–O. officinalis (CC genome), the backcrossing progenies and the parent plants. Results showed that most of the SSR primers (335 out of 396, 84.6%) developed in cultivated rice successfully amplified products from DNA samples of wild rice O. officinalis. The polymorphism ratio of SSR bands between O. sativa and O. officinalis was as high as 93.9%, indicating differences between the two species with respect to SSRs. When the SSR markers were applied in the interspecific hybrids, only a portion of SSR primers amplified O. officinalis-specific bands in the F1 hybrid (52.5%), BC1 (52.5%), and MAALs (37.0%); a number of the bands disappeared. Of the 124 SSR loci that detected officinalis-specific bands in MAAL plants, 96 (77.4%) showed synteny between the A and C-genomes, and 20 (16.1%) showed duplication in the C-genome. Sequencing analysis revealed that indels, substitution and duplication contribute to the diversity of SSR loci between the genomes of O. sativa and O. officinalis.







Similar content being viewed by others
References
Aggarwal RK, Brar DS, Khush GS (1997) Two new genomes in the Oryza complex identified on the basic of molecular divergence analysis using total genomic DNA hybridization. Mol Gen Genet 254:1–12
Aggarwal RK, Brar DS, Nandi S, Huang N, Khush GS (1999) Phylogenetic relationships among Oryza species revealed by AFLP markers. Theor Appl Genet 98:1320–1328
Brar DS, Khush GS (1997) Alien introgression in rice. Plant Mol Biol 35:35–47
Bennetzen JL (2000) Comparative sequence analysis of plant nuclear genomes: Microcolinearity and its many exceptions. Plant Cell 12:1021–1029
Bennetzen JL, Freeling M (1993) Grasses as a single genetic system: genome composition, collinearity and compatibility. Trends Genet 9:259–261
Brondani C, Rangel PHN, Borba TCO, Brondani RPV (2003) Transferability of microsatellite and sequence tagged site markers in Oryza species. Hereditas 138:187–192
Chen X, Cho YG, McCouch SR (2002) Sequence divergence of rice microsatellites in Oryza and other plant species. Mol Genet Genomics 268:331–343
Feldman M, Liu B, Segal G, Abbo S, Levy AA, Vega JM (1997) Rapid elimination of low-copy DNA sequences in polyploid wheat: a possible mechanism for differentiation of homoeologous chromosomes. Genetics 147:1381–1387
Finch RA (1983) Tissue-specific elimination of alternative whole parental genomes in one barley hybrid. Chromosoma 88:386–393
Fridman E, Carrari F, Liu YS, Fernie AR, Zamir D (2004) Zooming in on a quantitative trait for tomato yield using interspecific introgressions. Science 305:1786–1789
Fridman E, Pleban T, Zamir D (2000) A recombination hotspot delimits a wild species QTL for tomato sugar content to 484-bp within an invertase gene. Proc Natl Acad Sci USA 97:4718–4723
Fu SX, Zhan Y, Zhi HJ, Gai JY, Yu DY (2006) Mapping of SMV resistance gene Rsc-7 by SSR markers in soybean. Genetica 128:63–69
Ge S, Sang T, Lu BR, Hong DY (1999) Phylogeny of rice genomes with emphasis on origins of allotetraploid species. Proc Natl Acad Sci USA. 96:14400–14405
Gernand D, Rutten T, Varshney A, Rubtsova M, Prodanovic S, Bruß C, Kumlehn J, Matzk F, Houben A (2005) Uniparental chromosome elimination at mitosis and interphase in wheat and pearl millet crosses involves micronucleus formation, progressive heterochromatinization, and DNA fragmentation. Plant Cell 17(9):2431–2438
Grimaldi MC, Crouau-Roy B (1997) Microsatellite allelic homoplasy due to variable flanking sequences. J Mol Evol 44:336–340
Gupta PK, Varshney RK (2000) The development and use of microsatellite markers for genetic analysis and plant breeding with emphasis on bread wheat. Euphytica 113:163–185
Hai L, Wagner C, Friedt W (2007) Quantitative structure analysis of genetic diversity among spring bread wheats (Triticum aestivum L.) from different geographical regions. Genetica 130:213–225
He RF, Wang YY, Shi ZY, Ren X, Zhu LL, Weng QM, He GC (2003) Construction of genomic library of wild rice and Agrobacterium-mediated transformation of large insert DNA linked to BPH resistance locus. Gene 321:113–121
Hernandez P, Laurie DA, Martin A, Snape JW (2002) Utility of barley and wheat simple sequence repeat (SSR) markers for genetic analysis of Hordeum chilense and tritordeum. Theor Appl Genet 104:735–739
Huang Z, He GC, Shu LH, Li XH, Zhang QF (2001) Identification and mapping of two brown planthopper genes in rice. Theor Appl Genet 102:929–934
Jena KK, Khush GS (1990) Introgression of genes from Oryza officinalis Well ex Watt to cultivated rice, O. sativa L Theor Appl Genet 80:737–745
Jin HJ, Tan GX, Brar DS, Tang M, Li G, Zhu LL, He GC (2006) Molecular and cytogenetic characterization of an Oryza officinalis–O. sativa chromosome 4 addition line and its progenies. Plant Mol Biol 62:769–777
Kobayashi N, Ikeda R, Vaughan DA, Shigenaga S (1991) Resistance to Tungro in some wild relatives of rice. Rice Res Newsel 16:13
Kuleung C, Baenziger PS, Dweikat I (2004) Transferability of SSR markers among wheat, rye, and triticale. Theor Appl Genet 108(6):1147–1150
Lagercrantz U, Putterill J, Coupland G, Lydiate D (1996) Comparative mapping in Arabidopsis and Brassica, fine scale genome collinearity and congruence of genes controlling flowering time. Plant J 9:13–20
Lan WZ, Qin Q, Li G, He GC (2006) Comparative analysis of A, B, C and D genomes in the genus Oryza with Cot-1 DNA of C-genome. Chin Sci Bull 51(14):1710–1720
Linde-Laursen I, Bothmer R (1999) Orderly arrangement of the chromosomes within barley genomes of chromosome-eliminating Hordeum lechleri x barley hybrids. Genome 42:225–236
Liu L, Lafitte R, Guan D (2004) Wild Oryza species as potential sources of drought-adaptive traits. Euphytica 138:149–161
Multani DS, Khush GS, Delos Reyes BG, Brar DS (2003) Alien genes introgression and development of monosomic alien addition lines from Oryza latifolia Desv to rice, Oryza sativa L. Theor Appl Genet 107:395–405
Murray MG, Thompson WF (1980) Rapid isolation of high molecular-weight plant DNA. Nucleic Acids Res 8:4321–4325
Panaud O, Chen X, McCouch SR (1996) Development of microsatellite markers and characterization of simple sequence length polymorphism (SSLP) in rice (Oryza sativa L.). Mol Gen Genet 252:597–607
Pierantoni L, Cho KH, Shin IS, Chiodini R, Tartarini S, Dondini L, Kang SJ, Sansavini S (2004) Characterisation and transferability of apple SSRs to two European pear F1 populations. Theor Appl Genet 109:1519–1524
Riddle NC, Birchler JA (2003) Effects of reunited diverged regulatory hierarchies in allopolyploids and species hybrids. Trends in Genet 19:597–600
Shaked H, Kashkush K, Ozkan H, Feldman M, Levy AA (2001) Sequence elimination and cytosine methylation are rapid and reproducible responses of the genome to wide hybridization and allopolyploidy in wheat. Plant Cell 13:1749–1759
Tan GX, Huang Z, Weng QM, Ren X, Zhu LL, He GC (2004a) Two whitebacked planthopper resistance genes shared the same genomic intervals with brown planthopper resistance genes. Heredity 92:212–217
Tan GX, Weng QM, Ren X, Shi ZY, Zhu LL, He GC (2004b) Mapping of a new resistance gene to bacterial blight in rice line introgressed from Oryza officinalis. Acta Genetica Sinica 31:724–729
Tan GX, Jin HJ, Li G, He RF, Zhu LL, He GC (2005) Production and characterization of a complete set of individual chromosome additions from Oryza officinalis to Oryza sativa using RFLP and GISH analyses. Theor Appl Genet 111:1585–1595
Tanksley SD, McCouch SR (1997) Seed banks and molecular maps: unlocking genetic potential from the wild. Science 277:1063–1066
Uozu S, Ikehashi H, Ohmido N, Ohtsubo H, Ohtsubo E, Fukui K (1997) Repetitive sequences: cause for variation in genome size and chromosome morphology in the genus Oryza. Plant Mol Biol 35:791–799
Vaughan DA, Morishima H, Kadowaki K (2003) Diversity in the Oryza genus. Curr Opin Plant Biol 6:139–146
Wu KS, Tanksley SD (1993) Abundance, polymorphism and genetic mapping of microsatellites in rice. Mol Gen Genet 241:225–235
Zamir D (2001) Improving plant breeding with exotic genetic libraries. Nat Rev Genet 2:983–989
Acknowledgements
This study was supported by grants from the National Program of High Technology Development and the National Special Program for Research and Industrialization of Transgenic Plants of China.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Li, G., Hu, W., Qin, R. et al. Simple sequence repeat analyses of interspecific hybrids and MAALs of Oryza officinalis and Oryza sativa . Genetica 134, 169–180 (2008). https://doi.org/10.1007/s10709-007-9222-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10709-007-9222-x