Abstract
Claviceps purpurea causes ergot, a floral disease of major cereal crops, such as winter rye and winter triticale. The dark sclerotia formed by C. purpurea contain numerous toxic ergot alkaloids, which pose a major health risk for humans and animals when ingested. The countermeasures against ergot infection are currently limited to intensive tillage, seed cleaning, crop rotation or integration of optical sorting machines to remove ergot from cereals before processing. However, these practices confer only a minor protective effect and are highly energy-demanding. Thus, more effective and sustainable plant protection measures to combat ergot infections in cereals are needed. The application of Trichoderma spp. as biological control agent has already proven successful against a wide range of phytopathogenic fungi. However, there is only minor scientific evidence about its protective capacity against C. purpurea in cereal crops. In our study, we therefore investigated the antagonistic potential of several Trichoderma isolates against C. purpurea and their ability to confer protection against ergot infection. In initial in vitro tests and confrontation assays, we quantified and compared the production of iron-chelating compounds and hydrolytic enzymes as well as the antagonistic activity against C. purpurea sclerotia and mycelial cultures. Several strains showed high antifungal capacity and growth inhibitory effects towards C. purpurea. Selected Trichoderma strains were then tested for their potential to protect rye (Secale cereale) plants from ergot infection within a greenhouse trial. Rhizosphere-inoculated plants displayed a decreased disease severity compared to plants that had been sprayed with Trichoderma spores above-ground. Gene expression analysis by reverse transcription quantitative PCR also indicated the induction of systemic defense reactions. Overall, our data suggest that individual Trichoderma isolates possess a high antagonistic potential towards C. purpurea, which could be effective by direct mycoparasitism and by inducing systemic plant resistance, and therefore provide important guidance towards the development of Trichoderma treatments of cereals as biocontrol measure against ergot.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ergot, caused by the phytopathogenic ascomycete Claviceps purpurea, is a re-emerging plant disease affecting over 400 species of the Poaceae family, including economically important crops with rye most often reported (Lauber et al., 2005; Miedaner & Geiger, 2015). Ascospores and conidia of C. purpurea germinate on the stigma of flowering host plants. After infecting the ovaries, fungal mycelium differentiates into purplish-brown sclerotia replacing the host’s ovarian tissue. While ergot infection seldom causes a significant decrease in crop yield, the most serious consequence is the contamination of harvested crops with ergot sclerotia containing various alkaloid toxins (EFSA Panel on Contaminants in the Food Chain, 2012). Ergot alkaloids are secondary fungal metabolites and are defined as derivates of 4-(γ,γ dimethylallyl)tryptophan that are classified into three mayor groups as clavines, amides of lysergic acid and ergopeptines, based on their substituents (Agriopoulou, 2021; Florea et al., 2017).
By further processing the contaminated cereals, the mycotoxins enter the food and feed chain. Consumption can lead to typical clinical symptoms of ergot poisoning, such as strong vasoconstrictive and vasodilative effects, as well as severe neurological symptoms in humans and animals (Maruo et al., 2018; Merhoff & Porter, 1974). The disease caused by these alkaloids is called ergotism, also known are St. Anthony’s fire, and first reports date back to the Middle ages (Grzybowski et al., 2021). Therefore, the European Commission amended maximum levels for the content of ergot sclerotia in unprocessed rye to 500 mg/kg and for other unprocessed cereals to 200 mg/kg (Commission Regulation (EU) 2021/1399). The distribution of individual ergot alkaloids in crop samples is highly variable and does not necessarily correlate with ergot sclerotia content (Grusie et al., 2017; Kodisch et al., 2020b). Therefore, the ergot alkaloid concentration, defined by the sum of the most frequent members of ergot alkaloids, namely ergotamine, ergometrine, ergosine, ergocristine, ergocryptine, ergocornine and their epimers, in milled grain products has been newly limited to 500 µg/kg in rye and 100 µg/kg in other cereals being further reduced by 2024 to 250 µg/kg and 50 µg/kg, respectively (Commission Regulation (EU) 2021/1399).
A combination of physical and mechanical methods, based on gravity or optical separation, can be used to remove ergot sclerotia from cereals prior to processing, but these methods do not completely eliminate contamination with ergot alkaloids (Agriopoulou, 2021). Therefore, it is essential to apply countermeasures such as farming practices to reduce the incidence and severity of ergot formation.
The germination of overwintering structures of C. purpurea requires a period of cold temperatures during winter followed by a warmer period with high humidity in spring to facilitate spore production (Menzies & Turkington, 2015; Uppala et al., 2016). One of the current agricultural management strategies is therefore intensive ploughing to a depth of 5 cm to reduce the level of primary inoculum and hamper the ergot germination for following crops (Agriopoulou, 2021).
However, the fungus can survive in this dormant stage for up to three years in the soil. Crop rotation is therefore a recommended strategy to further reduce the amount of sclerotia (Agriopoulou, 2021; Menzies & Turkington, 2015). The risk of secondary infection and sclerotia formation by alternative hosts of the Poaceae family in the field margin can be reduced by application of herbicides or mowing. Preventive measures to counteract ergot infections include sowing of ergot-free seeds and avoidance of open-pollinated varieties with a long flowering period, which are more susceptible to ergot infections. In general, winter varieties are usually less susceptible than summer varieties, as the risk of secondary infections by neighboring alternative host plants is rather low (Menzies & Turkington, 2015). Another favorable approach is the cultivation of hybrid breeds with increased pollen production, which enhances the probability of fertilization and shortens the period of susceptibility (Kodisch et al., 2020a; Miedaner et al., 2022). Applying fungicides against ergot is less common and the main limitation of contact fungicides is the proper timing of the treatment, since to be functional, the compounds have to reach the plant’s ovary (McLaren, 2003).
A commercial fungicide belonging to the triazole group was shown to be effective against Claviceps africana in sorghum, but multiple applications from first anthesis to the end of flowering period are required (Ryley et al., 2003), risking the development of resistance in phytopathogenic fungi. The application of systemic antifungal compounds, which are taken up by the plant, is also restricted since the transport predominantly takes place through the xylem, limiting the amount of compounds reaching the ovary (Evans et al., 2000). However, soil-applied fungicides reduced the germination of ergot sclerotia but bears the risk to affect native beneficial microorganisms (Dung et al., 2018).
In contrast to the named countermeasures, biological control is taking advantage of beneficial organisms to control plant pests, which offers an additional strategy for a more sustainable and environmentally-friendly approach of disease management (Heydari & Pessarakli, 2010; Maruo et al., 2018; Stenberg et al., 2021; Waage & Greathead, 1988).
Trichoderma spp. (Ascomycota, Hypocreales, Hypocreaceae) are among the most commonly used fungal biocontrol agents against plant pathogens (Freeman et al., 2004; Sharma et al., 2011; Sood et al., 2020; Tyśkiewicz et al., 2022). During the versatile inter-kingdom interaction of Trichoderma spp. with host plants, both organisms are benefitting from this symbiotic relationship. While Trichoderma spp. receive photosynthates and habitat in the rhizosphere, they promote plant growth by increased nutrient availability and plant health by priming of plant defenses and alteration of root architecture (Guzmán-Guzmán et al., 2019; Naseby et al., 2000; Vinale & Sivasithamparam, 2020). The cellulolytic activity of Trichoderma spp. has been associated with the induction of local and systemic defense reactions during the root colonization process (Shoresh et al., 2010). Even though those plant growth-promoting properties are host plant specific and not present in every Trichoderma strain, it is widely common among the genus (Lorito & Woo, 2015). Further mechanisms for biological control by Trichoderma are the recognition and chemotrophic growth towards plant-pathogenic fungi, antibiosis through nutrient competition and release of cell wall degrading enzymes, and direct attack of fungal antagonists (Benítez et al., 2005; Degani & Dor, 2021; Sood et al., 2020). Mycoparasitism is a complex process which depends on a combination of various mechanisms, acting sequentially or combined (Viterbo & Horwitz, 2010).
The production of hydrolytic enzymes, such as proteases and chitinases, have been shown to have a crucial impact on their potential as biocontrol agents (Seidl et al., 2009). Several Trichoderma species have already proven to be effective against fungal phytopathogens via various synergistic mechanisms, but scientific evidence of the biocontrol ability against ergot in rye is limited.
However, Bhosale et al. (2022) identified T. viride, followed by T. hamatum and T. harzianum as effective antagonists of Claviceps fusiformis in vitro by dual plate assays. Furthermore, a commercial biocontrol preparation of T. viride showed a 50% biological efficacy in germination inhibition of ergot sclerotia from C. purpurea in vitro (Sheshegova & Shchekleina, 2020). Another approach to identify potential antagonists of ergot is the isolation from the native mycoflora of the overwintering scletoria structures. Ondřej et al. (2010) isolated fungal species from the surface of C. purpurea sclerotia and further tested them for their mycoparasitic effect on overwintering sclerotia structures in terms of germinating ability and microbial destruction. The highest effect of degradation in vitro was found by T. harzianum, but could not be verified under field conditions (Ondřej et al., 2010).
Bhuiyan et al. (2003) investigated the protective capacity of two commercial Trichoderma-based biocontrol products Trichopel and Trichoflow (Agrimm Technologies Limited, Christchurch, New Zealand) against Claviceps africana in Sorghum bicolor. Both culture filtrates significantly reduced conidial germination of C. africana macroconidia. Within a greenhouse trial, different mixtures of spores and culture filtrates of both biocontrol products were tested for their protective capacity against C. africana in planta and led to reduction of ergot severity. Summarizing, some few studies indicate biocontrol effects of Trichoderma spp. against C. purpurea vegetative growth or sclerotia germination, but were mostly performed in vitro (Bhosale et al., 2022; Ondřej et al., 2010; Sheshegova & Shchekleina, 2020).
The main purpose of this work was therefore to verify these observations in vitro and in vivo with several Trichoderma strains known for their biocontrol potential and additional isolates from agricultural land. Furthermore, the goal was to test whether any potential antagonistic effects against C. purpurea can be correlated to certain activities known to be associated with biocontrol and to evaluate if the antagonism against C. purpurea is a conserved or strain-specific property.
Materials and methods
Fungal cultures and growths conditions
For the isolation of Trichoderma, soil samples were taken from agricultural fields (Bavaria, Germany, 2022) and further processed as described previously (Mistry & Bariya, 2022). The obtained isolates as well as the following Trichoderma strains obtained from culture collection of the Chair of Phytopathology (Technical Universtiy of Munich), T. harzianum 20761 (A), T. harzianum 20770 (B), T. atroviride 20780 (C), T. atroviride 20781 (D), T. hamatum 20784 (E), T. asperellum 20866 (F) (Metz & Hausladen, 2022) and T. harzianum WM24a1 (G) (obtained from culture collection Austrian Institute of Technology GmbH) as well as T. atrobrunneum (H) (culture collection Fungal Biotechnology in Wood Science, Technical University of Munich) were routinely subcultured on ¼ potato dextrose agar (PDA) (X931.2, Carl Roth, Karlsruhe, Germany). For inoculum preparation, Trichoderma was grown on PDA for 7 days in darkness, followed by 3 days incubation with light to induce spore production. The spores were harvested by flooding the plates with sterile water and spores were carefully detached by using a spatula. The spore solution was then filtered through extra fine glass wool (Karl Hecht GmbH & Co KG, Sondheim, Germany) placed in a 50 ml tube to remove remaining mycelium debris.
The wild-type C. purpurea strain used in this experiments was isolated from sclerotium obtained of wheat cultivar (Triticum aestivum, Thuringia, Germany, harvest 2021) as described previously Faruk and Rahman (2022) and subcultured routinely on Mantle agar (Mantle & Nisbet, 1976). For inoculum production a pre-culture was prepared by inoculating 200 ml of 1% (w/v) wheat flour solution with five agar pieces containing mycelium and incubated at 25 °C for 10 days (Engelke, 2002). Rye grains were soaked in hot water for 30 min and autoclaved in 1200 ml mycocontainers (Mycogenetics, Everswinkel, Germany) and mixed with the fungal pre-culture. After two weeks of incubation, the overgrown grains were soaked in sterile water for 2 h and the spore suspension was then filtered through glass wool.
Molecular identification and phylogenetic analysis
Trichoderma isolates and C. purpurea were grown for 7 days on cellophane membranes (Natureflex 32 g/m2, 80 mm diameter, HERA Papierverarbeitung, Nidda, Germany) placed on PDA and Mantle media, respectively and approximately 20 mg mycelium was scraped with a sterile spatula and transferred into a 1.5 ml tube for genomic DNA extraction using the Animal and Fungi DNA Preparation kit (PP-208S, Jena Bioscience GmbH, Jena, Germany) according to manufacturer’s instructions. The quantity and quality of the obtained DNA was estimated with microplate reader (NanoQuant Plate, Tecan Infinite 200 Pro, Männedorf, Switzerland).
To confirm the identity of the fungal strains on molecular basis, a region of nuclear rDNA containing the ITS1-5.8S-ITS2 rRNA gene using the following primer pairs ITS 1: 5’-TCCGTAGGTGAACCTGCGG-3’ and ITS 4: 5’-TCCTCCGCTTATTGATATGC-3’ (White, 1990) was amplified.
For the verification of Trichoderma isolates the molecular identification protocol for a single Trichoderma isolate was applied following Cai & Druzhinina, 2021. Since ITS can only be used to identify Trichoderma on generic level, two additional phylogenetic markers translation elongation factor 1α (tef1) and RNA polymerase B subunit II (rpb2) were amplified using the following primer pairs EF1: 5’-ATGGGTAAGGARGACAAGAC-3’; EF2: 5’-GGARGTACCAGTSATCATGTT-3’ (O’Donnell et al., 1998) and fRPB2-5f: 5’-GAYGAYMGWGATCAYTTYGG-3’ and fRPB2-7cr: 5’ CCCATRGCTTGTYYRCCCAT-3’ (Liu et al., 1999).
PCR products were run on 1% agarose gel (1 × TAE buffer 100 ml + 5 µl Midori Green; 140 V, 500 mA, 0.6 h) and corresponding bands were cut from the gel and purified using Hi Yield® Gel/ PCR DNA Fragment Extraction Kit (Süd-Laborbedarf GmbH, Gauting, Germany). DNA sequencing of amplified regions was performed in both directions using the Mix2Seq kit (Eurofins Genomics, Ebersberg, Germany) and consensus sequence was created by alignment of forward and reverse sequence using Benchling (Benchling, 2022, San Francisco, USA).
Phylogenetic analysis
Phylogenetic analysis of the obtained soil isolates of Trichoderma spp. was computed as described by Cai and Druzhinina (2021). Summarized, retrieved tef1 and rpb2 sequences were trimmed using TrichoMark 2020 and processed sequences were used for multiple sequences comparison by log-expectation using MUSCLE (Edgar, 2004). Phylogram data was computed from concatenated tef1 and rpb2 sequences using maximum-likelihood (ML) method in IQ-TREE web server (Nguyen et al., 2015; Trifinopoulos et al., 2016) with statistical bootstrapping of 1000 replicates and best-fit model TIM2e + G4 (Kalyaanamoorthy et al., 2017). Phylogenetic trees were visualized using the online tool Interactive Tree of Life (iTOL, version 5) (Letunic & Bork, 2021). Respective sequences of Laccaria bicolor KA13-0253 were included as outgroup.
In vitro screenings for biocontrol abilities
Siderophore production
The detection of fungal siderophores in liquid culture supernatants was performed according to Joshi et al. (2006). The assay is based on the formation of a blue colored complex between CAS, hexadecyltrimethylammonium (HDTMA) bromide (199,532, Sigma-Aldrich, Taufkirchen, Germany) and iron. Hydroxamate siderophores, present in the culture filtrate, scavenge the iron leading to the release of free CAS dye, visible in a color change from blue to orange, which can be determined spectrophotometrically. For the liquid CAS assay Trichoderma strains were inoculated with 106 spores/ml in 4 ml M9 Minimal Medium in 24-well-plates and incubated for one week at 30 °C, shaking at 150 rpm. The culture supernatants were obtained by filtration through glass fiber filters (\(\varnothing\) 55 mm, MN 85/70, MACHEREY–NAGEL, Düren, Germany) using a Büchner funnel. 100 µl of the culture filtrate were mixed with 50 µl CAS solution and after incubation in the dark for 2 h the absorbance was measured at 630 nm (Tecan Infinite 200 Pro, Männedorf, Switzerland). The uninoculated medium served as reference. The amount of produced Deferoxamine B equivalent was calculated from A/Aref based on a standard curve prepared with 0 to 300 µM deferoxamine mesylate and normalized with dry biomass.
Cellulase activity
The determination of the cellulolytic activity of Trichoderma was conducted by a plate assay as described by Teather and Wood (1982). Summarized, the synthetic substrate Carboxymethyl cellulose (CMC) is stained with the direct dye Congo red. The hydrolysis of β-1,4-glycosidic linkages by cellulases, released into the medium, leads to the formation of a clearance halo around the fungal colony. Trichoderma was cultivated on solid CMC for 96 h at 25 °C followed by incubation at 45–50 °C for 18 h. Afterwards the plates were stained by adding 10 ml of a 0.2% Congo red solution (0322.1, ROTH, Karlsruhe, Germany) and incubated for 15–20 min. Staining solution was then removed by rinsing the plates with water followed by destaining with a 1 M NaCl solution for 15–20 min. Pictures were taken and the halo was measured using ImageJ V1.53e software (Wayne Rasby, National Institute of Health, USA, http://imageJ.nih.gov/ij). The enzymatic index (EI) served as a quantitative measure of cellulase production. An EI of 1 signifies no cellulase activity and was therefore set to 0. Enzymatic indices were calculated using the following formula:
Proteolytic activity
To evaluate proteolytic activity, Trichoderma was inoculated on skim milk agar, previously described by Alnahdi (2012) and after incubation at 25 °C for 4 days the plates were photographed at the end. The clear zone halo, formed by hydrolyzed milk proteins, was then quantified using ImageJ V1.53e software based on pictures taken after incubation.
Chitinase activity
For the evaluation of total chitinolytic activity Trichoderma was inoculated with 106 spores/ml in 4 ml liquid chitinase detection medium as described by Agrawal and Kotasthane (2012). The cultures were incubated at 30 °C and 150 rpm for one week and culture supernatants were obtained by filtration and used for the enzymatic assay. The reduction of 3,5-dinitrosalicylic acid to 3-amino-5-nitrosalicylic acid by the reducing sugar N-acetyl-β-D-glucosamine (NAGA) induces a shift in the absorbance at 540 nm, which serves as a quantitative measure for the NAGA concentration in the medium, a product of fungal chitinase activity (Miller, 1959). The amount of released free NAGA was determined spectrophotometrically by measuring the absorbance at 540 nm (Tecan Infinite 200 Pro, Männedorf, Switzerland) and correlating obtained values with generated standard curve of known concentrations of NAGA (A8625, Sigma-Aldrich, Taufkirchen, Germany).
Antagonistic activity
For in vitro antagonism assays in dual culture C. purpurea was inoculated on ¼ PDA in non-split and bi-compartment plates (ø 9 cm, Sarstedt, Nümbrecht, Germany) and incubated for five days at 25 °C. Trichoderma strains were inoculated with spores on the opposite site of the plate in triplicates. Pictures were taken (Nikon, Tokyo, Japan) directly after inoculation with Trichoderma as well as 4 and 6 days afterwards, corresponding to pre-contact and direct contact, respectively. Plates inoculated only with C. purpurea served as a control. Colony area (cm2) was measured using ImageJ V1.53e software. The growth inhibition by Trichoderma for air contact, medium contact and direct contact stage was then calculated as follows (Raut et al., 2014):
Where \(\Delta \mathrm{A}\) describes the difference in the colony area of C. purpurea between the evaluated time point and the previous time point. A decrease in colony area of C. purpurea between two time points, is expressed by values above 100%.
Inhibition of germinating sclerotia
It is already known that some Trichoderma species are able to colonize and degrade dormant sclerotia of plant pathogenic fungi (Elad et al., 1984). To evaluate the ability of Trichoderma strains to hamper sclerotia germination of C. purpurea, a sclerotia germination inhibition assay was conducted. Ergot sclerotia were surface sterilized by soaking them for 5 min in 2% sodium hypochlorite, washed with sterile water for 2–3 times and placed on autoclaved wet potting substrate (TS1, Klasmann-Deilmann, Geeste, Germany). After a vernalization period of 7 weeks at 4 °C with high humidity, the plates were inoculated with 2 ml of a spore solution of Trichoderma containing 106 spores/ml and the negative control was mock inoculated with sterile water. Afterwards, the plates were incubated for at least 3 weeks at 21 °C, 70% humidity and 6 h/ 18 h dark/ light cycle until the negative control started to germinate. Pictures were taken with a dissecting microscope (MDG41, Leica Microsystems, Wetzlar, Germany) and the number of germinated sclerotia per plate was assessed (Kaur et al., 2016).
In planta application
Greenhouse experiment
The Harzianum section revealed the highest number of predicted antimicrobial peptides, identified by comparative genomic screening (Rush et al., 2021). The strain T. harzianum WM24a1, belonging to the Harzianum clade, has already shown to be a strong volatile organic compound (VOC) emitter (Guo et al., 2019) and was therefore chosen for the greenhouse experiment. Furthermore, T. atrobrunneum was chosen as a second Trichoderma spp. to be tested in planta based on the registration as an active ingredient of commercial biocontrol agents under the European Union regulation of the T. atrobrunneum isolate ITEM 908 (Fanelli et al., 2018). For the in planta application, four seeds of Secale cereale (ARANTES, KWS Lochow GmbH, Bergen, Germany) were planted in 3 l pots containing C700 growth substrate and fertilized through drip irrigation. One subset of 7-week-old plants was inoculated with Trichoderma by adding 10 ml containing 106 spores/ml to each plant rhizosphere one week before infection with C. purpurea. The other subset was phyllosphere-treated with Trichoderma spore solution (106 spores/ml) using a chromatography sprayer (H451.1, Carl Roth, Karlsruhe, Germany) until run off. In order to increase air humidity, the plants were covered with plastic bags after spraying. Phyllosphere treatment was applied twice, about 3 to 5 days and one day before infection with C. purpurea.
For the in planta evaluation of Trichoderma as a potential biocontrol agent a phenotypic evaluation of ergot infection as well as an expression analysis was conducted. For the phenotypic evaluation, the plants were infected through spraying until run off, imitating the natural infection process, and intensity was monitored in terms of honeydew production and formation of ergot sclerotia (Menzies, 2004).
For the expression analysis one half of florets of blooming ears with ripe stigma, but not fully developed anthers, were directly inoculated between the palea and lemma with 5 µl conidial suspension of C. purpurea containing 106 spores/ml (in total 15–20 florets per ear) (Tenberge et al., 1996). Only the plants treated with T. harzianum WM24a1 (G) were further investigated for RT-qPCR. In order to avoid cross contamination and to achieve an increased humidity, all ears were covered with plastic bags for at least five hours or overnight.
Expression analysis
RNA extraction and RT-qPCR
One to two ears per pot were sampled 24 h, three days and five days after infection with C. purpurea. Plant material stored at -80 °C was finely ground in liquid nitrogen and 100 mg was used for RNA extraction using Direct-zol™ RNA Miniprep Plus Kit (R2073, Zymo Research Europe GmbH, Freiburg, Germany) combined with TRIzol™ reagent (Invitrogen, Carlsbad, USA) following manufacturer’s instruction, but expanding the DNase I treatment to 30 min at 37 °C. The extracted RNA was tested for genomic DNA contamination by gel electrophoresis on a 2% agarose gel running at 135 V for 0.6 h and quantified with microplate reader (NanoQuant Plate, Tecan Infinite 200 Pro, Männedorf, Switzerland) and stored at -80 °C. For cDNA synthesis, 1 µg RNA per reaction was reverse transcribed using the FastGene® Scriptase Basic cDNA Kit (LS-62, NIPPON Genetics Europe, Düren, Germany) and cDNA was stored at -20 °C until further use. RT-qPCR was conducted by using the 2 × qPCRBIO SyGreen Mix Separate-ROX kit (PB20.11, PCR Biosystems Ltd, London, United Kingdom) using a total reaction volume of 10 µl, including 5 µl SyGreen, 10 µM forward and reverse primer each, nuclease free water and 0.5 µl cDNA as template. The qPCR amplification program included 2 min initial denaturation at 95 °C, 45 cycles of amplification (5 s at 95 °C, 25 s at the primer-specific annealing temperature (Ta) and a melting curve ramping from 64.5 °C to 95 °C with an increasing temperature of 0.5 °C/s. The amplification of RNase L inhibitor-like protein from rye (Scrli NCBI: BE495958) (Giménez et al., 2011) and glyceraldehyde-3-phosphate dehydrogenase from C. purpurea (Cpgpd1; NCBI: X73282.1) (Oeser et al., 2017) served as an internal plant and fungal reference gene to normalize the amount of total RNA present in each reaction (Table 1). The gene expression levels were calculated using the 2−∆∆CT-method according to Livak and Schmittgen (2001). All samples were analyzed with three technical replicates and the amplicon lengths of all test genes as well as the respective annealing temperatures are summarized in Table 1.
Statistical analysis
Statistical data analysis was performed with Origin Pro (version 2021, OriginLab Corporation, Northampton, USA). The data was tested for normal distribution using Shapiro–Wilk test, followed by Levene’s test for variance homogeneity. One-way ANOVA was performed combined with Tukey’s test for pairwise comparison of means or Fisher’s least significant difference test.
Results
Isolation and phylogenetic analysis
In the present study altogether 12 Trichoderma strains, belonging to sections with reported biocontrol activities (Fanelli et al., 2018), were included in the in vitro screenings. T. harzianum 20761 (A), T. harzianum 20770 (B), T. atroviride 20780 (C), T. atroviride 20781 (D), T. hamatum 20784 (E), T. asperellum 20866 (F) were already tested for their biocontrol abilities in vitro, in vivo and in the field against Alternaria solani in potato and have shown to be effective (Metz & Hausladen, 2022). The available strains T. harzianum WM24a1 (G) and T. atrobrunneum (H) were also included. Furthermore, to test Trichoderma spp. that are naturally present on arable land, soil samples were collected for isolation of additional Trichoderma strains. By selective cultivation on Rose Bengal agar, a total of five Trichoderma-like fungal strains (designated I—L) were isolated from the soil samples and identified at the molecular level as T. velutinum S1 (I), T. velutinum S2 (J), T. harzianum (K) and T. velutinum S3 (L) (Fig. 1).
Phylogenetic analysis of concatenated tef1 and rpb2 sequences verified a close relationship of the soil isolate L with T. harzianum WM24a1 and the reference strain T. harzianum CBS 226.95 (Fig. 2). Furthermore, the soil isolates T. velutinum S1-S3 clustered together with the reference strains T. velutinum C.P.K. 2978 and T. velutinum DAOM 230013 (Fig. 2).
Phylogenetic tree of the soil isolates Trichoderma spp. based on concatenated sequences of translation elongation factor 1α (tef1) and RNA polymerase B subunit II (rpb2) of the genus Trichoderma. Processed sequences were aligned using Muscle 3.8.1 and maximum-likelihood phylogeny was performed with IQ-TREE web server with bootstrap values of 1000 and visualization using iTOL. Metadata values are Shimodaira-Hasegawa approximate likelihood ratio test (SH-aLRT) (≥ 70) and ultrafast bootstrap support. The strains included in this study are in bold and Laccaria bicolor KA 13–0253 was included as outgroup
In vitro screenings
Within the scope of laboratory experiments, 12 Trichoderma strains, including the four Trichoderma spp. rhizosphere isolates, were evaluated in vitro for their biocontrol and plant-growth promoting potential. To this end, the strains were screened for the release of fungal siderophores as well as the secretion of lytic enzymes, such as chitinases, proteases and cellulases. The antagonistic potential against C. purpurea was evaluated in dual plate confrontation assays with different degrees of mycelium contact and during germination of C. purpurea sclerotia.
Siderophore production
The production and release of fungal hydroxamate siderophores has been suggested as an important mechanism of Trichoderma BCAs for outcompeting other fungi in the rhizosphere (Benítez et al., 2005) and the mobilization of iron has shown to contribute to plant growth and nutrition (Altomare et al., 1999). Even though those plant growth promoting properties are host plant specific and not present in every Trichoderma spp. strain, it is widely common among the genus (Lorito & Woo, 2015). The release of siderophores was evaluated by liquid CAS assay, to test for the capacity to mobilize iron. The highest siderophore secretion was detected for T. harzianum 20770 (B), followed by T. atroviride 20781 (D), T. harzianum (K) and T. harzianum WM24a1 (G) (Fig. 3). T. atrobrunneum (H) showed the lowest chelating capacity.
Liquid chrome azurol S (CAS) assay for quantification of siderophore production. Trichoderma spp. were cultivated in liquid M9 medium for 7 days at 30 °C. Culture supernatants were incubated with CAS solution for 2 h and the absorbance measured at 630 nm. Uninoculated medium was used as reference. A/Aref-values were correlated to a standard curve prepared with deferoxamine mesylate and normalized with dry weight. Trichoderma harzianum 20761 (A), Trichoderma harzianum 20770 (B), Trichoderma atroviride 20780 (C), Trichoderma atroviride 20781 (D), Trichoderma hamatum 20784 (E), Trichoderma asperellum 20866 (F), Trichoderma harzianum WM24a1 (G), Trichoderma atrobrunneum (H), Trichoderma velutinum S1 (I), Trichoderma velutinum S2 (J), Trichoderma harzianum (K), Trichoderma velutinum S3 (L). Different letters indicate significant differences with p < 0.05, n = 9
Cellulase production
Cellulase activity is a prerequisite of Trichoderma spp. for the colonization of plant roots and has been shown to be associated with the induction of local and systemic defense reactions in plants (Saravanakumar et al., 2018; Shoresh et al., 2010). We utilized a Congo red-based plate assay to study the secretion of cellulases in the 12 Trichoderma strains. The formation of clearance halos around the colonies served therby as quantitative measure. While the two T. atroviride strains (20781 D and 20780 C) showed the highest EI (Fig. 4), the T. velutinum strains (particularly J and L) showed the lowest cellulolytic activity of all strains during the test conditions.
Carboxymethylcellulose (CMC) and Congo red assay for the detection of cellulase activity. a Trichoderma spp. cultivated on CMC agar for 96 h at room temperature, followed by incubation at 45–50 °C for 18 h. Plates were stained with 0.2% Congo red solution. Representative photos of Trichoderma harzianum 20761, Trichoderma harzianum 20770, Trichoderma atroviride 20780 and Trichoderma atroviride 20781 after staining process. b Quantitative evaluation of cellulolytic activity. Enzymatic indices were estimated from the colony and halo diameters. Trichoderma harzianum 20761 (A), Trichoderma harzianum 20770 (B), Trichoderma atroviride 20780 (C), Trichoderma atroviride 20781 (D), Trichoderma hamatum 20784 (E), Trichoderma asperellum 20866 (F), Trichoderma harzianum WM24a1 (G), Trichoderma atrobrunneum (H), Trichoderma velutinum S1 (I), Trichoderma velutinum S2 (J), Trichoderma harzianum (K), Trichoderma velutinum S3 (L). Different letters indicate significant differences with p < 0.05, SD, n = 3
Proteolytic activity
Proteolytic activity of Trichoderma is another indicator for its antagonistic and antifungal potential. For example, it is already known that aspartyl and serine proteases are involved in mycoparasitism mechanisms of important biocontrol species such as T. harzianum and T. atroviride (Atanasova et al., 2013; Benítez et al., 2005; Kubicek et al., 2011). We therefore tested the secretion of proteases in all strains by way of a skim milk plate assay. The tested Trichoderma spp. strains showed significant differences in their activity during the experimental conditions. T. harzianum WM24a1 (G) as well as the soil isolate T. velutinum S3 (L) showed the strongest proteolytic activity during the protease plate test (Fig. 5). In contrast, T. asperellum 20866 (F) was the only strain not showing any formation of a clear zone indicating no proteolytic activity under these conditions.
Skim milk plate assay for the detection of proteases. Trichoderma spp. were cultivated on skim milk agar for 4 days at room temperature. The formation of a clear zone halo was quantified. Trichoderma harzianum 20761 (A), Trichoderma harzianum 20770 (B), Trichoderma atroviride 20780 (C), Trichoderma atroviride 20781 (D), Trichoderma hamatum 20784 (E), Trichoderma asperellum 20866 (F), Trichoderma harzianum WM24a1 (G), Trichoderma atrobrunneum (H), Trichoderma velutinum S1 (I), Trichoderma velutinum S2 (J), Trichoderma harzianum (K), Trichoderma velutinum S3 (L). Different letters show significant differences with p < 0.05, SE, n = 3
Chitinase activity
Chitinases are an indispensable tool of Trichoderma spp. to mycoparasitize phytopathogenic fungi and they act as elicitors of the plant’s immune response (Kubicek et al., 2001; Nawrocka & Małolepsza, 2013). Chitinases catalyze the hydrolytic breakdown of β-1,4-glycosidic linkages between the NAGA residues of chitin. The in vitro analysis of total chitinolytic activity of Trichoderma spp. was based on the quantification of free NAGA, released from a defined concentration of colloidal chitin by the activity of fungal chitinases present in the culture supernatants. The soil isolate T. harzianum (K), T. harzianum 20761 (A), T. atroviride 20780 (C), and T. atroviride 20781 (D) were found to show the highest chitinolytic activity (Fig. 6). Under the given experimental conditions, the lowest chitinolytic activity was detected for T. atrobrunneum (H) and the soil isolate T. velutinum S3 (L).
Enzymatic assays for the quantification of chitinolytic activity by Trichoderma spp.. Strains were grown in liquid chitinase detection medium supplemented with 4.5 g/l colloidal chitin for 7 days at 30 °C. Supernatant was harvested by filtration and 1 ml mixed with 0.3 ml of 1 M sodium acetate buffer (pH 4.6) and 0.2 ml of colloidal chitin (0.5% w/v). After incubation at 40 °C for 20 h the mixture was centrifuged at 8.000 rpm for 5 min and 0.75 ml of the obtained supernatant were then mixed with 0.25 ml of a 1%-solution of 3,5-dinitrosalicylic acid. Reduction was quantified by absorbance measurements at 540 nm. The concentration of released NAGA is given per mg dry biomass. Trichoderma harzianum 20761 (A), Trichoderma harzianum 20770 (B), Trichoderma atroviride 20780 (C), Trichoderma atroviride 20781 (D), Trichoderma hamatum 20784 (E), Trichoderma asperellum 20866 (F), Trichoderma harzianum WM24a1 (G), Trichoderma atrobrunneum (H), Trichoderma velutinum S1 (I), Trichoderma velutinum S2 (J), Trichoderma harzianum (K), Trichoderma velutinum S3 (L). The graph shows the mean values ± SE of three biological replicates. Different letters show significant differences with p < 0.05
In vitro antagonistic activity against C. purpurea
The ability of Trichoderma spp. to restrict C. purpurea growth by combined mechanisms of secreted antifungal compounds, antibiosis and direct hyphal contact is a strong indicator for the antagonistic capacity and biocontrol abilities and can be evaluated with dual confrontation assays. Furthermore, by using two-compartment petri-dishes, preventing diffusion of metabolites and direct hyphal contact, the impact of volatile organic compounds can be explored. In our study, dual-plate confrontation assays showed considerable antagonistic activity of Trichoderma spp. against C. purpurea (Fig. 7). Interestingly, at 4 days after Trichoderma inoculation, the inhibitory effect on the Claviceps colonies was generally substantially higher with only air contact compared to media contact conditions, indicating a strong influence of volatile compounds. Here, T. harzianum 20761 (A) and T. harzianum 20770 (B) showed the weakest inhibitory effect. The soil isolate T. harzianum (K) could not be assessed for air contact conditions, since the mycelium grew over the separation after 3 days. After six days of incubation, allowing direct contact in this experimental setup, the inhibition strongly increased in all cases. Some of the tested strains even provoked C. purpurea to withdraw, expressed by growth inhibition values of more than 100%, as e.g. for T. asperellum 20866 (F) and T. harzianum 20770 (B) with 132,58% and 117,72%, respectively. The weakest inhibitory effect during direct contact was assessed for the soil isolate T. velutinum S3 (L) with < 45%.
Dual plate confrontation assay between Trichoderma spp. and Claviceps purpurea. Both fungi were inoculated on the opposite sides of one- or two-compartment ¼-PDA plates and incubated at room temperature. For air contact and media contact conditions, the increase in colony area (cm.2) of Claviceps purpurea from the time point of inoculation with Trichoderma spp. until 4 days post inoculation was assessed and compared to the negative control, where Claviceps purpurea was grown alone. For direct contact between Trichoderma spp. and the plant-pathogen, the change in colony area was assessed between day 4 and 6 after inoculation with the fungal antagonist, when both fungal partners where already in hyphal contact. a Representative pictures for the four contact stages of Trichoderma harzianum WM24a1 corresponding to 4 dpi, for air contact and medium contact, 6 dpi for the direct contact phase. b Quantitative evaluation of the growth inhibition of Claviceps purpurea by the respective Trichoderma spp. at the air, medium and direct contact stage. The size of the ergot colonies was measured with ImageJ. Trichoderma harzianum 20761 (A), Trichoderma harzianum 20770 (B), Trichoderma atroviride 20780 (C), Trichoderma atroviride 20781 (D), Trichoderma hamatum 20784 (E), Trichoderma asperellum 20866 (F), Trichoderma harzianum WM24a1 (G), Trichoderma atrobrunneum (H), Trichoderma velutinum S1 (I), Trichoderma velutinum S2 (J), Trichoderma harzianum (K), Trichoderma velutinum S3 (L). Significances were determined for each contact stage separately, p < 0.05, SE, n = 3
Inhibitory effect on sclerotia germination
The application of Trichoderma spp. to agricultural fields bears the potential to not only directly antagonize C. purpurea mycelium, but also to reduce sclerotia germination in the soil, which would help to minimize the risk of a following infection cycle. Some Trichoderma spp. have already shown potential to colonize and degrade resistant structures such as sclerotia of plant pathogenic fungi, leading to decreased germination (Elad et al., 1984; Ondřej et al., 2010).
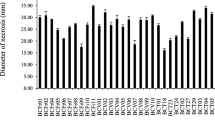
The ergot sclerotia used in this experimental setup were collected from a winter wheat cultivar from the harvest season 2020/2021 (Thuringia, Germany). The soil isolates T. velutinum S1 (I), T. velutinum S2 (J), T. harzianum (K) and T. velutinum S3 (L) could not be included in this assay, since ergot sclerotia require a cold phase of 7 weeks for germination induction, which was performed prior to their isolation. The antagonistic capacity of the tested strains was assessed by a qualitative rating score from 0 to 6, adapted from the evaluation of antagonistic activity of Trichoderma against several plant-pathogens previously described by Yu et al. (2022). Particularly T. atroviride 20781 (D), T. asperellum 20866 (F) and T. harzianum WM24a1 (G) revealed a high inhibitory effect on the formation of stroma of the germinating sclerotia (Fig. 8). T. asperellum 20866 showed an intensive mycelium growth on the surface of ergot sclerotia as well as a strong sporulation, while T. hamatum 20784 (E) did not show any visible impact.
Germination assay of ergot sclerotia inoculated with Trichoderma spp. a Close-up images of ergot sclerotia on TS 1 fully overgrown by sporulating 1: T. asperellum 20866 (F) sporulating on ergot sclerotia, 2: sclerotia covered with fine mycelium and 3: stroma formed by a germinating sclerotium of control group without Trichoderma. b Qualitative rating of the antagonistic activity of Trichoderma spp. against ergot sclerotia according to Yu et al. (2022). A score of 0 means no impact of the respective Trichoderma strain; 2 indicates growth and mycelium formation on the ergot sclerotia; 4 means sporulation on the ergot surface and 6 implies an inhibitory effect on germination of the ergot sclerotia (given by < 30% germinating sclerotia). Trichoderma harzianum 20761 (A), Trichoderma harzianum 20770 (B), Trichoderma atroviride 20780 (C), Trichoderma atroviride 20781 (D), Trichoderma hamatum 20784 (E), Trichoderma asperellum 20866 (F), Trichoderma harzianum WM24a1 (G), Trichoderma atrobrunneum (H)
Summarized in vitro Trichoderma spp. screenings
For each conducted in vitro screening assay, the isolates were grouped according to their performance from 0 to 3, referring to 0 = no activity, 1 = low activity, 2 = medium activity and 3 = high activity (Fig. 9). The thresholds for each group were set individually for each assay. T. atroviride 20781 (D) showed the overall highest total score of 18, followed by T. atroviride 20780 (C) with a score of 17, and T. harzianum 20770 (B), T. hamatum 20784 (E) and T. harzianum WM24a1 (G), all with a total score of 15 (Fig. 9). The soil isolates T. velutinum S1(I) and S3 (L) showed the lowest total score with a value of 9 and 8, respectively.
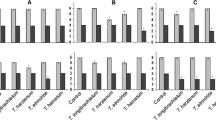
Summarized results of the in vitro screening of Trichoderma spp. a The thresholds for each group were set for each assay individually as follows: siderophore production (µM deferoxamine B equiv.): low: < 0.5, medium: 0.5–1.0, high: > 1.0; cellulase activity (EI): low: < 0.025, medium: 0.025–0.075, high: > 0.075; protease activity (clear zone, cm.2): low: < 15, medium: 15–30, high: > 30; chitinase activity (NAGA released, µg/ml): low: < 10, medium 10–17, high: > 17; growth inhibition (%): air contact (AC): low: < 30, medium: 30–35, high: > 35, medium contact (MC): low: < 10, medium: 10 15, high: > 15, direct contact (DC): low: < 55, medium: 55–85, high: > 85. red = not determined b Total score for each Trichoderma strain. Trichoderma harzianum 20761 (A), Trichoderma harzianum 20770 (B), Trichoderma atroviride 20780 (C), Trichoderma atroviride 20781 (D), Trichoderma hamatum 20784 (E), Trichoderma asperellum 20866 (F), Trichoderma harzianum WM24a1 (G), Trichoderma atrobrunneum (H), Trichoderma velutinum S1 (I), Trichoderma velutinum S2 (J), Trichoderma harzianum (K), Trichoderma velutinum S3 (L)
In planta application
Phenotypic infection assay
For the in planta application, two Trichoderma strains from the Harzianum section were chosen based on the total score of the in vitro screenings: T. harzianum WM24a1 (G) as a top-scoring strain and T. atrobrunneum (H) as a represenative of the medium-performing strains (Fig. 9). Two different treatments were included to test for their protective capacity in terms of reduced signs of ergot infection. Spores of both strains were applied either foliar by spraying the ears untill run-off, or by direct incoluation of the rhizosphere prior to infection with C. purpurea.
Honeydew production was tested, as it is mainly responsible for secondary infection cycles of ergot after successful infection of C. purpurea by containing fungal conidia which are further spread by rainsplashes or insects (Tudzynski & Scheffer, 2004). Furthermore, exclusively the sphacelial stage in the ergot development is characterized by production of honeydew, which stops with the formation of sclerotia. The initiation, duration as well as the amount of honeydew produced can therefore be used as indicators for a proper development of the ergot fungus and a balanced host–pathogen interaction (Mower & Hancock, 1975). Timing and amount of honeydew formation by the infected ears was assessed in a daily manner starting with the appearance of honeydew on the first ear (Fig. 10). Furthermore, the number of sclerotia produced per infected ear was counted.
No striking differences could be detected between both applied strains. The high variability in disease development did not allow a statistic evaluation, but overall, a reduced infection severity, in terms of appearance of honeydew and intensity of honeydew production, could be observed for plants that had been rhizosphere-treated compared to phyllosphere-treatment (Table 2). Furthermore, the number of produced sclerotia per infected ear showed a tendency to be decreased in rhizosphere-treated plants compared to phyllosphere-treated ones.
Expression analysis
To detect relative changes within the transcription of pathogenesis-related (PR) genes in the ears of rye, a RT-qPCR analysis was conducted for the plant defense genes xylinh and chit (in rye) as well as for the pathogenicity factor cell in C. purpurea. Samples for RNA extraction were only collected from plants treated with T. harzianum WM24a1 (G), since we expected an increased biocontrol ability based on the in vitro screenings. The expression of Scxylinh was found to be significantly increased in all ergot-infected plants and across all time points (Fig. 11 a-d). Those that were additionally phyllosphere-treated with T. harzianum WM24a1 spores showed the strongest induction compared to the non-treated controls after 24 h and after 5 d.
Relative expression level of xylanase inhibitor (Scxylinh) 24 h (a), 3 d (b) and 5 d (c) and relative expression of glucan endo-1,3-beta-glucosidase 8 (Scchit) 24 h (d), 3 d (e) and 5 d (f) post infection with Claviceps purpurea. Plants were phyllosphere treated with Trichoderma harzianum WM24a1 by spraying and rhizosphere treated by direct inoculation of the growth substrate with a conidial suspension containing 106 spores/ml. For infection flowers were directly inoculated with a conidial suspension of Claviceps purpurea (10.6 conidia/ml) using a micropipette. The expression of the test genes was determined by RT-qPCR analysis and the \({2}^{{-\Delta \Delta C}_{t}}\) method. The rye gene RNase L inhibitor-like protein (Scrli) served as internal reference plant gene, and the untreated and uninfected plants were used as control (grey). Relative expression of the ergot gene 1,4-β-cellobiosidase (Cpcell) in rye ears at 5 dpi (g) and the ergot gene glyceraldehyde-3-phosphate dehydrogenase (Cpgpd1) served as reference gene and the plants only infected with Claviceps purpurea were used as control. Different letters show significant differences with p < 0.05, SE, n = 3
The temporal changes in the expression of the rye gene chit (Fig. 11 d-f) revealed less dynamics than in the analysis of xylinh. After 24 h, significant differences could only be detected between phyllosphere-treated and rhizosphere-treated groups but without ergot infection. Three days after the infection with C. purpurea the chit transcript level of the rhizosphere-treated and ergot-infected plants was significantly increased compared to all other test groups. However, this effect could not be detected anymore after five days of infection, where no significant differences between all groups were observed.
Due to the limited amount of C. purpurea mRNA within the harvested plant biomass of the early time points, the expression analysis for the ergot gene cell was only performed for the 5 day samples after infection. At this time point, the expression of cell was increased in both Trichoderma-treated groups vs. the control plants without any biocontrol treatment (Fig. 11 g). The transcript level of Cpcell was most strongly increased in the previously phyllosphere-treated plants with an average fold-change of 2.5 compared to the rhizosphere-treated plants which showed an average fold-change of 1.4.
Discussion
Biological control of plant diseases caused by fungal pathogens is a promising alternative to chemical fungicides (Thambugala et al., 2020). As an opportunistic and avirulent plant symbiont, Trichoderma is by far the most extensively studied biocontrol agent against a broad spectrum of aerial and soil-born fungal pathogens but antagonism is highly species- and host-specific (Alfiky & Weisskopf, 2021; Nur & Noor, 2020). Therefore, to identify Trichoderma strains which are effective as biocontrol agents against specific pathogens, a broad variety of species should be evaluated individually (Rush et al., 2021). The isolation of Trichoderma spp. from rhizosphere soil, followed by their characterization and screening for biocontrol activities, such as production of hydrolytic enzymes and siderophores, is a quite common practice and has already proven successful in the identification of some promising biocontrol strains (Chen et al., 2021; Rai et al., 2016; Yu et al., 2022). Although in vitro screenings always bear the risk to accidentally exclude strains by not properly inducing the tested mycoparasitism-related gene clusters under laboratory conditions (Fanelli et al., 2018; Hermosa et al., 2004; Li et al., 2021), they are nevertheless the first choice when several different Trichoderma isolates have to be rated for their characteristic biocontrol abilities to identify the strains with the highest potential for follow-up experiments (Bedine Boat et al., 2020; Imran et al., 2022; Yassin et al., 2022).
During our in vitro screenings of cellulolytic, proteolytic, and total chitinolytic activity, significant differences could be detected between the tested Trichoderma strains under the given test conditions. However, strains belonging to the Harzianum section, such as T. harzianum 20761 (A), T. harzianum WM24a1 (G), and the soil isolate T. harzianum (K) displayed highest protease activities and showed a strong inhibitory effect on sclerotia germination. T. atroviride 20780 (C) and T. atroviride 20781 (D), belonging to the Atroviride section, performed also well in protease and cellulase activity plate assays, and T. atroviride 20781 (D) showed a strong inhibitory effect on sclerotia germination. Besides those enzymatic in vitro screenings testing for biocontrol properties, we aimed to evaluate the inhibitory effect of the 12 strains towards the specific plant-pathogen C. purpurea using direct confrontation assays with different degrees of contact.
In dual plate confrontation assays, all tested Trichoderma strains displayed the lowest inhibitory effect during media contact and the highest at direct contact. The increased inhibitory effect on C. purpurea during air contact compared to media contact is indicating that VOCs are involved in the interaction and have an impact in the process of antagonism of Trichoderma (Bae et al., 2016; Rajani et al., 2021). Trichoderma spp. produce VOCs that inhibit growth of other fungi, promote plant growth, and induce plant defense responses (Bruce et al., 2000; Phoka et al., 2020; Ruangwong et al., 2021; Salwan et al., 2019; Wonglom et al., 2020). Furthermore, Guo et al. (2019) showed by multivariate analysis that VOC profiles were highly species-specific among the three tested Trichoderma strains T. velutinum GL1561, T. hamatum QL15d1 and T. harzianum WM24a1 and that the latter strongly emitted the most diverse compound patterns. During fungus-fungus interaction VOC emissions and perceptions are comprising a complex host–pathogen cross-talk and can lead to a host-specific mycoparasitism initiation in Trichoderma spp. (Sharma et al., 2017; Zeilinger & Omann, 2007).
In summary, our findings showed that all Trichoderma spp. significantly suppressed mycelium growth of C. purpurea during in vitro conditions. During direct contact, the strains T. harzianum 20770 (B), T. atroviride 20781(D), T. harzianum WM24a1 (G) and T. asperellum 20866 (F) were not only able to inhibit C. purpurea growth, but even led to an active reduction of the C. purpurea colony size. The same strains, except T. harzianum 20770 (B), had a similarly strong inhibitory effect on the germination of ergot sclerotia in vitro. Smith et al. (2013) tested 21 Trichoderma isolates and T. atroviride Th002 and T. harzianum Th203 showed strong mycoparasitic activity against the plant-pathogen Sclerotinia sclerotiorum and prevented sclerotia germination of more than 70%. Haddad et al. (2017) also evaluated 120 Trichoderma spp. regarding their potential to inhibit sclerotia germination of S. sclerotiorum in vitro and identified 22 strains, including two T. atroviride strains, that caused 100% inhibition of sclerotia germination, which implies that large-scale screenings are worthwhile to identify efficient Trichoderma isolates against specific fungal pathogens.
The expression of genes connected with carbohydrate metabolism is associated with pathogenicity in plant pathogenic fungi such as Aspergillus flavus, Botrytis cinerea, and Alternaria sp. (Annis & Goodwin, 1997; Have et al., 1998; Shieh et al., 1997). As a plant-pathogenic fungus, C. purpurea is dependent on various carbohydrate-active enzymes (CAZymes), such as the investigated putative cellulase, to penetrate the plant cell walls in the rye ovary (Oeser et al., 2017; Schardl et al., 2013). Expression analysis revealed that this putative cellulase gene is induced during the first days of infection in rye (Müller et al., 1997). Furthermore, Oeser et al. (2002) showed that mutants lacking two polygalacturonase genes nearly lost their complete pathogenicity towards rye.
Trichoderma-induced resistance is not only achieved when applied directly to rhizosphere, but also after foliar application to the plants surface (Korolev et al., 2008; Perazzolli et al., 2008). Furthermore, foliar application of T. harzianum WM24a1 onto the ears surface is taking advantage of the in vitro observed antagonistic activity towards C. purpurea in planta. Therefore, the increased expression of Cpcell in the phyllosphere-treated ears could be influenced by the antagonist being present, leading to nutrient competition. However, since we obtained data for only one late time point and little is known about the direct interaction of Trichoderma and C. purpurea in planta, further studies need to be conducted to explore the observed phenomena. Xylanases from C. purpurea were found to be upregulated in planta during all infection stages (Giesbert et al., 1998). Xylanase inhibitors, expressed by the plant, inhibit the hemicellulose degradation of those secreted fungal xylanases (Tundo et al., 2022). Expression analysis revealed significantly increased mRNA levels of the xylanase inhibitor xylinh in rye plants that were infected with C. purpurea, irrespective of the Trichoderma treatment. In contrast, plants that were only treated with Trichoderma did not show an induced expression of xylinh. The expression of chit in rye was enhanced in rhizosphere-inoculated plants, but the effect was not detectable later than 3 days after infection with C. purpurea. Assigned to three families of PR proteins, PR3, PR8 and PR11, chitinolytic enzymes represent a major part of the plant defense system effective against a wide range of phytopathogenic fungi (Wang et al., 2018). In the control plants only infected with C. purpurea, no significant differences could be detected compared to the not-infected and not-treated plants, which is contradictory to the observations of an increased expression of chitinases during the first days of infection by Oeser et al. (2017).
However, this could be indicative for a strategy of host immune suppression by C. purpurea through sequestering chitin oligosaccharides (Jonge et al., 2010), which remains to be elucidated. Nontheless, the increased expression of chit in rhizosphere-inoculated plants is supporting the results of the phenotypic evaluation of the infection severity. In contrast to the control plants, the rhizosphere-treated plants showed a delayed and reduced honeydew production as well as a decrease in sclerotia formation. This effect could be explained by an induced systemic resistance in rye stimulated by the interaction with Trichoderma in the rhizosphere as it was already seen in wheat (Esmail et al., 2022; Jaroszuk-Ściseł et al., 2019). Furthermore, Pociecha and Dziurka (2015) were able to show that rhizosphere inoculation of winter rye with T. viride and T. harzianum increased resistance to the snow mold Microdochium nivale when applied after cold acclimation treatment, by a decreased carbohydrate content in the shoot and increased carbohydrate content in the roots.
A combination of different Trichoderma species, as it is the case in the commercial product Trichopel, has been previously shown to promote growth of plants in agriculture and forestry more effectively compared to single strain application (Chen et al., 2021; Chirino-Valle et al., 2016). Similarly, two mixed Trichoderma isolates showed synergistic effects and enhanced disease control of Fusarium solani in tomato plants compared to separate application (Awad-Allah et al., 2022). This could therefore also be considered for biocontrol applications against ergot. PR genes play an important role in the process of plant disease resistance and there is ample evidence that Trichoderma can induce plant defense responses, but it remains to be elucidated which biocontrol treatment and which time point of application will lead to an effective defense response in rye and subsequently to an increased resistance against ergot in the field.
Conclusion
The objective of this work was to evaluate the biocontrol ability of Trichoderma spp. against C. purpurea in vitro and the potential to protect against ergot infection of rye in planta. Taken together, a clear antagonistic activity against C. purpurea could be verified in several of the tested Trichoderma strains in vitro, and a trend for protection against ergot could be seen for rhizosphere-treated rye in the greenhouse trial. Based on the in vitro screenings and their robust ability to inhibit the growth of C. purpurea, we have identified T. harzianum 20770 (B), T. atroviride 20780 (C), T. atroviride 20781 (D), T. hamatum 20784 (E), and T. harzianum WM24a1 (G) as strong candidates for biocontrol agents against ergot. Our results therefore indicate that the application of suitable Trichoderma isolates could provide an alternative and sustainable disease management strategy for ergot infection. In the future, a mixture of several candidate strains should be evaluated regarding their effect on disease severity. Additionally, the efficacy of Trichoderma isolates in managing ergot in rye, as well as inhibiting sclerotia germination within the soil, should be further investigated and explored under field conditions.
Data Availability
All data generated or analysed during this study are included in this published article and its supplementary information files or can be obtained from the corr. author upon reasonable request.
References
Agrawal, T., & Kotasthane, A. S. (2012). Chitinolytic assay of indigenous Trichoderma isolates collected from different geographical locations of Chhattisgarh in Central India. In SpringerPlus, 1(1), 73. https://doi.org/10.1186/2193-1801-1-73
Agriopoulou, S. (2021). Ergot Alkaloids Mycotoxins in Cereals and Cereal-Derived Food Products: Ergot Alkaloids Mycotoxins in Cereals and Cereal-Derived Food Products: Characteristics, Toxicity, Prevalence, and Control Strategies. In Agronomy 11(5). https://doi.org/10.3390/agronomy11050931
Alfiky, A., Weisskopf, L. (2021). Deciphering Trichoderma-Plant-Pathogen Interactions for Better Development of Biocontrol Applications. In Journal of Fungi 7(1). https://doi.org/10.3390/jof7010061
Alnahdi, H. S. (2012). Isolation and screening of extracellular proteases produced by new Isolated Bacillus sp. In Journal of Applied Pharmaceutical Science, 2(9). https://doi.org/10.7324/JAPS.2012.2915
Altomare, C., Norvell, W. A., Björkman, T., & Harman, G. E. (1999). Solubilization of Phosphates and Micronutrients by the Plant-Growth-Promoting and Biocontrol Fungus Trichoderma harzianum Rifai 1295–22. In Applied and Environmental Microbiology, 65(7), 2926–2933. https://doi.org/10.1128/AEM.65.7.2926-2933.1999
Annis, S. L., & Goodwin, P. H. (1997). Recent advances in the molecular genetics of plant cell wall-degrading enzymes produced by plant pathogenic fungi. In European Journal of Plant Pathology, 103(1), 1–14. https://doi.org/10.1023/A:1008656013255
Atanasova, L., Le Crom, S., Gruber, S., Coulpier, F., Seidl-Seiboth, V., Kubicek, C. P., & Druzhinina, I. S. (2013). Comparative transcriptomics reveals different strategies of Trichoderma mycoparasitism. In BMC Genomics, 14, 121. https://doi.org/10.1186/1471-2164-14-121
Awad-Allah, E. F. A., Shams, A. H. M., Helaly, A. A., Ragheb, E. I. M. (2022). Effective Applications of Trichoderma spp. as Biofertilizers and Biocontrol Agents Mitigate Tomato Fusarium Wilt Disease. In Agriculture 12(11), 1950. https://doi.org/10.3390/agriculture12111950
Bae, S.-J., Mohanta, T. K., Chung, J. Y., Ryu, M., Park, G., Shim, S., et al. (2016). Trichoderma metabolites as biological control agents against Phytophthora pathogens. In Biological Control, 92, 128–138. https://doi.org/10.1016/j.biocontrol.2015.10.005
Bedine Boat, M. A., Sameza, M. L., Iacomi, B., Tchameni, S. N., Boyom, F. F. (2020). Screening, identification and evaluation of Trichoderma spp. for biocontrol potential of common bean damping-off pathogens. In Biocontrol Science and Technology 30(3), 228–242. https://doi.org/10.1080/09583157.2019.1700909
Benchling. Biology Software. (2022). Available online at https://benchling.com. Accessed 4 Jul 2022
Benítez, T., Rincón, A., Limón, M. C., & Codón, A. (2005). Biocontrol mechanism of Trichoderma strains. In International Microbiology: THe Official Journal of the Spanish Society for Microbiology, 7, 249–260.
Bhosale, G. V., Bharti, D. S., Bhosale, G. B., Jagtap, G. P., & Banne, S. N. (2022). In vitro evaluation of bio control agents against Claviceps fusiformis, causing Ergot of Bajra. Pharma Innovation, 11(1), 410–412.
Bhuiyan, S. A., Ryley, M. J., Galea, V. J., & Tay, D. (2003). Evaluation of potential biocontrol agents against Claviceps africana in vitro and in vivo. In Plant Pathology, 52(1), 60–67. https://doi.org/10.1046/j.1365-3059.2003.00799.x
Bruce, A., Wheatley, R. E., Humphris, S. N., Hackett, C. A., & Florence, M. E. J. (2000). Production of Volatile Organic Compounds by Trichoderma in Media Containing Different Amino Acids and Their Effect on Selected Wood Decay Fungi. In Holzforschung, 54(5), 481–486. https://doi.org/10.1515/HF.2000.081
Cai, F., & Druzhinina, I. S. (2021). In honor of John Bissett: Authoritative guidelines on molecular identification of Trichoderma. In Fungal Diversity, 107(1), 1–69. https://doi.org/10.1007/s13225-020-00464-4
Chen, D., Hou, Q., Jia, L., Sun, K. (2021). Combined Use of Two Trichoderma Strains to Promote Growth of Pakchoi (Brassica chinensis L.). In Agronomy 11(4). https://doi.org/10.3390/agronomy11040726
Chirino-Valle, I., Kandula, D., Littlejohn, C., Hill, R., Walker, M., Shields, M., et al. (2016). Potential of the beneficial fungus Trichoderma to enhance ecosystem-service provision in the biofuel grass Miscanthus x giganteus in agriculture. In Scientific Reports, 6(1), 25109. https://doi.org/10.1038/srep25109
Commission Regulation (EU) 2021/1399 of 24 August 2021 amending Regulation (EC) No 1881/2006 as regards maximum levels of ergot sclerotia and ergot alkaloids in certain foodstuffs (Text with EEA relevance) (2021). Official Journal, 301, 1–5. Available online at http://data.europa.eu/eli/reg/2021/1399/oj. Accessed 17 Nov 2022
Degani, O., Dor, S. (2021). Trichoderma Biological Control to Protect Sensitive Maize Hybrids against Late Wilt Disease in the Field. In Journal of Fungi 7(4). https://doi.org/10.3390/jof7040315
Dung, J. K. S., Kaur, N., Walenta, D. L., Alderman, S. C., Frost, K. E., & Hamm, P. B. (2018). Reducing Claviceps purpurea sclerotia germination with soil-applied fungicides. In Crop Protection, 106, 146–149. https://doi.org/10.1016/j.cropro.2017.12.023
Edgar, R. C. (2004). MUSCLE: A multiple sequence alignment method with reduced time and space complexity. In BMC Bioinformatics, 5(1), 113. https://doi.org/10.1186/1471-2105-5-113
EFSA Panel on Contaminants in the Food Chain (2012): Scientific Opinion on Ergot alkaloids in food and feed. In EFSA Journal 10(7). https://doi.org/10.2903/j.efsa.2012.2798
Elad, Y., Barak, R., & Chet, I. (1984). Parasitism of sclerotia of Sclerotium rolfsii by Trichoderma harzianum. In Soil Biology and Biochemistry, 16(4), 381–386. https://doi.org/10.1016/0038-0717(84)90037-3
Engelke, T. (2002). Ansätze für eine integrierte Bekämpfung des Mutterkorns (Claviceps purpurea [Fr.] Tul.). Dissertation. University Göttingen. Cuvillier Verlag, Göttingen.
Esmail, S. M., Omar, G. E., Mourad, A. I. (2022). In-depth understanding of the genetic control of stripe rust resistance (Puccinia striiformis f. sp. tritici) induced in wheat (Triticum aestivum L.) by Trichoderma asperellum T34. In Plant Disease. https://doi.org/10.1094/PDIS-07-22-1593-RE
Evans, V. J., Jenkyn, J. F., Gladders, P., Mantle, P. G. (2000). Fungicides for the control of ergot in cereal crops. In: Proceedings brighton crop protection conference - pests and diseases, pp. 511–514
Fanelli, F., Liuzzi, V. C., Logrieco, A. F., & Altomare, C. (2018). Genomic characterization of Trichoderma atrobrunneum (T. harzianum species complex) ITEM 908: insight into the genetic endowment of a multi-target biocontrol strain. In BMC Genomics, 19(1), 662. https://doi.org/10.1186/s12864-018-5049-3
Faruk, M. I., & Rahman, M. M. E. (2022). Collection, isolation and characterization of Sclerotinia sclerotiorum, an emerging fungal pathogen causing white mold disease. In Journal of Plant Science Phytopathology, 6, 43–51.
Florea, S., Panaccione, D. G., & Schardl, C. L. (2017). Ergot Alkaloids of the Family Clavicipitaceae. In Phytopathology, 107(5), 504–518. https://doi.org/10.1094/PHYTO-12-16-0435-RVW
Freeman, S., Minz, D., Kolesnik, I., Barbul, O., Zveibil, A., Maymon, M., et al. (2004). Trichoderma Biocontrol of Colletotrichum acutatum and Botrytis cinerea and Survival in Strawberry. In European Journal of Plant Pathology, 110(4), 361–370. https://doi.org/10.1023/B:EJPP.0000021057.93305.d9
Giesbert, S., Lepping, H. B., Tenberge, K. B., & Tudzynski, P. (1998). The Xylanolytic System of Claviceps purpurea: Cytological Evidence for Secretion of Xylanases in Infected Rye Tissue and Molecular Characterization of Two Xylanase Genes. In Phytopathology, 88(10), 1020–1030. https://doi.org/10.1094/PHYTO.1998.88.10.1020
Giménez, M. J., Pistón, F., & Atienza, S. G. (2011). Identification of suitable reference genes for normalization of qPCR data in comparative transcriptomics analyses in the Triticeae. In Planta, 233(1), 163–173. https://doi.org/10.1007/s00425-010-1290-y
Grusie, T., Cowan, V., Singh, J., McKinnon, J., Blakley, B. (2017). Correlation and variability between weighing, counting and analytical methods to determine ergot (Claviceps purpurea) contamination of grain. In World Mycotoxin Journal 10(3). https://doi.org/10.3920/WMJ2016.2174
Grzybowski, A., Pawlikowska-Łagód, K., & Polak, A. (2021). Ergotism and Saint Anthony’s fire. In Clinics in Dermatology, 39(6), 1088–1094. https://doi.org/10.1016/j.clindermatol.2021.07.009
Guo, Y., Ghirardo, A., Weber, B., Schnitzler, J. P., Benz, J. P., & Rosenkranz, M. (2019). Trichoderma Species Differ in Their Volatile Profiles and in Antagonism Toward Ectomycorrhiza Laccaria bicolor. In Frontiers in Microbiology, 10, 891. https://doi.org/10.3389/fmicb.2019.00891
Guzmán-Guzmán, P., Porras-Troncoso, M. D., Olmedo-Monfil, V., & Herrera-Estrella, A. (2019). Trichoderma Species: Versatile Plant Symbionts. In Phytopathology, 109(1), 6–16. https://doi.org/10.1094/PHYTO-07-18-0218-RVW
Haddad, P. E., Leite, L. G., Lucon, C. M. M., & Harakava, R. (2017). Selection of Trichoderma spp. strains for the control of Sclerotinia sclerotiorum in soybean. In Pesquisa Agropecuária Brasileira, 52(12), 1140–1148. https://doi.org/10.1590/s0100-204x2017001200002
Have, A., Mulder, W., Visser, J., & van Kan, J. A. (1998). The endopolygalacturonase gene Bcpg1 is required for full virulence of Botrytis cinerea. In Molecular Plant-Microbe Interactions, 11(10), 1009–1016. https://doi.org/10.1094/MPMI.1998.11.10.1009
Hermosa, M. R., Keck, E., Chamorro, I., Rubio, B., Sanz, L., Vizcaíno, J. A., et al. (2004). Genetic diversity shown in Trichoderma biocontrol isolates. In Mycological Research, 108(8), 897–906. https://doi.org/10.1017/s0953756204000358
Heydari, A., & Pessarakli, M. (2010). A Review on Biological Control of Fungal Plant Pathogens Using Microbial Antagonists. In Journal of Biological Sciences, 10(4), 273–290. https://doi.org/10.3923/jbs.2010.273.290
Imran, M., Abo-Elyousr, K. A. M., Mousa, M. A., & Saad, M. M. (2022). Screening and biocontrol evaluation of indigenous native Trichoderma spp. against early blight disease and their field assessment to alleviate natural infection. In Egyptian Journal of Biological Pest Control, 32(1), 40. https://doi.org/10.1186/s41938-022-00544-4
Jaroszuk-Ściseł, J., Tyśkiewicz, R., Nowak, A., Ozimek, E., Majewska, M., Hanaka, A. et al. (2019). Phytohormones (Auxin, Gibberellin) and ACC Deaminase In Vitro Synthesized by the Mycoparasitic Trichoderma DEMTkZ3A0 Strain and Changes in the Level of Auxin and Plant Resistance Markers in Wheat Seedlings Inoculated with this Strain Conidia. In International Journal of Molecular Sciences 20(19). https://doi.org/10.3390/ijms20194923
Jonge, R., van Esse, H. P., Kombrink, A., Shinya, T., Desaki, Y., Bours, R., et al. (2010). Conserved fungal LysM effector Ecp6 prevents chitin-triggered immunity in plants. In Science, 329(5994), 953–955. https://doi.org/10.1126/science.1190859
Joshi, F., Archana, G., & Desai, A. (2006). Siderophore Cross-Utilization Amongst Rhizospheric Bacteria and the Role of Their Differential Affinities for Fe3+ on Growth Stimulation Under Iron-Limited Conditions. In Current Microbiology, 53(2), 141–147. https://doi.org/10.1007/s00284-005-0400-8
Kalyaanamoorthy, S., Minh, B. Q., Wong, T. K. F., von Haeseler, A., & Jermiin, L. S. (2017). ModelFinder: fast model selection for accurate phylogenetic estimates. In Nature Methods, 14(6), 587–589. https://doi.org/10.1038/nmeth.4285
Kaur, N., Dung, J. K. S., Walenta, D. L., & Frost, K. E. (2016). Prospects for ergot disease management with biocontrol products. Oregon State University.
Kodisch, A., Wilde, P., Schmiedchen, B., Fromme, F.-J., Rodemann, B., Tratwal, A., et al. (2020b). Ergot infection in winter rye hybrids shows differential contribution of male and female genotypes and environment. In Euphytica, 216(4), 65. https://doi.org/10.1007/s10681-020-02600-2
Kodisch, A., Oberforster, M., Raditschnig, A., Rodemann, B., Tratwal, A., Danielewicz, J. et al. (2020b). Covariation of Ergot Severity and Alkaloid Content Measured by HPLC and One ELISA Method in Inoculated Winter Rye across Three Isolates and Three European Countries. In Toxins 12(11). https://doi.org/10.3390/toxins12110676
Korolev, N., Rav David, D., & Elad, Y. (2008). The role of phytohormones in basal resistance and Trichoderma-induced systemic resistance to Botrytis cinerea in Arabidopsis thaliana. In BioControl, 53(4), 667–683. https://doi.org/10.1007/s10526-007-9103-3
Kubicek, C. P., Herrera-Estrella, A., Seidl-Seiboth, V., Martinez, D. A., Druzhinina, I. S., Thon, M., et al. (2011). Comparative genome sequence analysis underscores mycoparasitism as the ancestral life style of Trichoderma. In Genome Biology, 12(4), R40. https://doi.org/10.1186/gb-2011-12-4-r40
Kubicek, C. P., Mach, R. L., Peterbauer, C. K., Lorito, M. (2001). Trichoderma: From genes to biocontrol. In Journal of Plant Pathology 83, 11–23. Available online at http://www.jstor.org/stable/41998018
Lauber, U., Schnaufer, R., Gredziak, M., & Kiesswetter, Y. (2005). Analysis of rye grains and rye meals for ergot alkaloids. In Mycotoxin Research, 21(4), 258–262. https://doi.org/10.1007/BF02957588
Letunic, I., & Bork, P. (2021). Interactive Tree Of Life (iTOL) v5: An online tool for phylogenetic tree display and annotation. In Nucleic Acids Research, 49(W1), W293–W296. https://doi.org/10.1093/nar/gkab301
Li, W. C., Lin, T. C., Chen, C. L., Liu, H. C., Lin, H. N., Chao, J. L., et al. (2021). Complete Genome Sequences and Genome-Wide Characterization of Trichoderma Biocontrol Agents Provide New Insights into their Evolution and Variation in Genome Organization, Sexual Development, and Fungal-Plant Interactions. In Microbiology Spectrum, 9(3), e0066321. https://doi.org/10.1128/Spectrum.00663-21
Liu, Y. J., Whelen, S., & Hall, B. D. (1999). Phylogenetic relationships among ascomycetes: Evidence from an RNA polymerse II subunit. In Molecular Biology and Evolution, 16(12), 1799–1808. https://doi.org/10.1093/oxfordjournals.molbev.a026092
Livak, K. J., & Schmittgen, T. D. (2001). Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2−ΔΔCT Method. In Methods, 25(4), 402–408. https://doi.org/10.1006/meth.2001.1262
Lorito, M., & Woo, S. L. (2015). Trichoderma: A Multi-Purpose Tool for Integrated Pest Management. In B. Lugtenberg (Ed.), Principles of Plant-Microbe Interactions: Microbes for Sustainable Agriculture (pp. 345–353). Springer International Publishing.
Mantle, P. G., Nisbet, L. J. (1976). Differentiation of Claviceps purpurea in Axenic Culture. Microbiology, 93.2, 321–334.
Maruo, V. M., Bracarense, A. P., Metayer, J. P., Vilarino, M., Oswald, I. P., Pinton, P. (2018). Ergot Alkaloids at Doses Close to EU Regulatory Limits Induce Alterations of the Liver and Intestine. In Toxins 10(5). Available online at doi: https://doi.org/10.3390/toxins10050183
McLaren, N. W. (2003). Efficacy of fungicides in the control of ergot (Claviceps africana) in sorghum (Sorghum bicolor) hybrid seed production. In South African Journal of Plant and Soil, 20(3), 154–156. https://doi.org/10.1080/02571862.2003.10634926
Menzies, J. G. (2004). The reactions of Canadian spring wheat genotypes to inoculation with Claviceps purpurea, the causal agent of ergot. In Canadian Journal of Plant Science, 84, 625–629. https://doi.org/10.4141/P03-086
Menzies, J. G., & Turkington, T. K. (2015). An overview of the ergot (Claviceps purpurea) issue in western Canada: challenges and solutions. In Canadian Journal of Plant Pathology, 37(1), 40–51. https://doi.org/10.1080/07060661.2014.986527
Merhoff, G. C., & Porter, J. M. (1974). Ergot intoxication: Historical review and description of unusual clinical manifestations. In Annals of Surgery, 180(5), 773–779. https://doi.org/10.1097/00000658-197411000-00011
Metz, N., & Hausladen, H. (2022). Trichoderma spp. as potential biological control agent against Alternaria solani in potato. In Biological Control, 166, 104820. https://doi.org/10.1016/j.biocontrol.2021.104820
Miedaner, T., Korzun, V., Wilde, P. (2022). Effective Pollen-Fertility Restoration Is the Basis of Hybrid Rye Production and Ergot Mitigation. In Plants 11(9). https://doi.org/10.3390/plants11091115
Miedaner, T., & Geiger, H. H. (2015). Biology, genetics, and management of ergot (Claviceps spp.) in rye, sorghum, and pearl millet. In Toxins, 7(3), 659–678. https://doi.org/10.3390/toxins7030659
Miller, G. L. (1959). Use of dinitrosalicylic acid reagent for determination of reducing sugar. Analytical Chemistry, 31, 426–428. https://doi.org/10.1021/ac60147a030
Mistry, H., Bariya, H. (2022). Isolation and Identification of Trichoderma spp. from Different Agricultural Samples. In Natarajan Amaresan, Prittesh Patel, Dhruti Amin (Eds.): Practical Handbook on Agricultural Microbiology. New York, NY: Springer US (Springer Protocols Handbooks), pp. 131–144
Mower, R. L., & Hancock, J. G. (1975). Mechanism of honeydew formation by Claviceps species. In Canadian Journal of Botany, 53(23), 2826–2834. https://doi.org/10.1139/b75-310
Müller, U., Tenberge, K. B., Oeser, B., & Tudzynski, P. (1997). Cel1, probably encoding a cellobiohydrolase lacking the substrate binding domain, is expressed in the initial infection phase of Claviceps purpurea on Secale cereale. In Molecular Plant-Microbe Interactions, 10(2), 268–279. https://doi.org/10.1094/MPMI.1997.10.2.268
Naseby, D. C., Pascual, J. A., & Lynch, J. M. (2000). Effect of biocontrol strains of Trichoderma on plant growth, Pythium ultimum polulations, soil microbial communities and soil enzyme activities. In Journal of Applied Microbiology, 88(1), 161–169. https://doi.org/10.1046/j.1365-2672.2000.00939.x
Nawrocka, J., & Małolepsza, U. (2013). Diversity in plant systemic resistance induced by Trichoderma. In Biological Control, 67(2), 149–156. https://doi.org/10.1016/j.biocontrol.2013.07.005
Nguyen, L.-T., Schmidt, H. A., Von Haeseler, A., & Minh, B. Q. (2015). IQ-TREE: A Fast and Effective Stochastic Algorithm for Estimating Maximum-Likelihood Phylogenies. In Molecular Biology and Evolution, 32(1), 268–274. https://doi.org/10.1093/molbev/msu300
Nur, A. Z., & Noor, A. B. (2020). Biological functions of Trichoderma spp. for agriculture applications. In Annals of Agricultural Sciences, 65(2), 168–178. https://doi.org/10.1016/j.aoas.2020.09.003
O’Donnell, K., Kistler, H. C., Cigelnik, E., & Ploetz, R. C. (1998). Multiple evolutionary origins of the fungus causing Panama disease of banana: Concordant evidence from nuclear and mitochondrial gene genealogies. In Proceedings of the National Academy of Sciences, 95(5), 2044–2049. https://doi.org/10.1073/pnas.95.5.2044
Oeser, B., Heidrich, P. M., Müller, U., Tudzynski, P., & Tenberge, K. B. (2002). Polygalacturonase is a pathogenicity factor in the Claviceps purpurea/rye interaction. In Fungal Genetics and Biology, 36(3), 176–186. https://doi.org/10.1016/s1087-1845(02)00020-8
Oeser, B., Kind, S., Schurack, S., Schmutzer, T., Tudzynski, P., & Hinsch, J. (2017). Cross-talk of the biotrophic pathogen Claviceps purpurea and its host Secale cereale. In BMC Genomics, 18(1), 273. https://doi.org/10.1186/s12864-017-3619-4
Ondřej, M., Cagaš, B., & Ondráčková, E. (2010). Effect of the mycoflora of ergot (Claviceps purpurea) sclerotia on their viability. In Plant Protect. Sci., 46(2), 66–71. https://doi.org/10.17221/48/2009-PPS
Perazzolli, M., Dagostin, S., Ferrari, A., Elad, Y., & Pertot, I. (2008). Induction of systemic resistance against Plasmopara viticola in grapevine by Trichoderma harzianum T39 and benzothiadiazole. In Biological Control, 47(2), 228–234. https://doi.org/10.1016/j.biocontrol.2008.08.008
Phoka, N., Suwannarach, N., Lumyong, S., Ito, S., Matsui, K., Arikit, S., Sunpapao, A. (2020). Role of Volatiles from the Endophytic Fungus Trichoderma asperelloides PSU-P1 in Biocontrol Potential and in Promoting the Plant Growth of Arabidopsis thaliana. In Journal of Fungi 6(4). https://doi.org/10.3390/jof6040341
Pociecha, E., & Dziurka, M. (2015). Trichoderma interferes with cold acclimation by lowering soluble sugars accumulation resulting in reduced pink snow mould (Microdochium nivale) resistance of winter rye. In Environmental and Experimental Botany, 109, 193–200. https://doi.org/10.1016/j.envexpbot.2014.07.009
Rai S., Kashyap P. L., Kumar S., Srivastava A. K., Ramteke P. W. (2016). Identification, characterization and phylogenetic analysis of antifungal Trichoderma from tomato rhizosphere. SpringerPlus 9, 5(1):1939. https://doi.org/10.1186/s40064-016-3657-4
Rajani, P., Rajasekaran, C., Vasanthakumari, M. M., Olsson, S. B., Ravikanth, G., & Uma Shaanker, R. (2021). Inhibition of plant pathogenic fungi by endophytic Trichoderma spp. through mycoparasitism and volatile organic compounds. In Microbiological Research, 242, 126595. https://doi.org/10.1016/j.micres.2020.126595
Raut, I., Badea-Doni, M., Calin, M., Oancea, F., Vasilescu, G., Sesan, T. E., & Jecu, L. (2014). Effect of volatile and non-volatile metabolites from Trichoderma spp. against important phytopathogens. In Revista De Chimie, 65(11), 1285–1288.
Ruangwong, O.-U., Wonglom, P., Suwannarach, N., Kumla, J., Thaochan, N., Chomnunti, P. et al. (2021). Volatile Organic Compound from Trichoderma asperelloides TSU1: Impact on Plant Pathogenic Fungi. In Journal of Fungi 7(3). https://doi.org/10.3390/jof7030187
Rush, T. A., Shrestha, H. K., Gopalakrishnan Meena, M., Spangler, M. K., Ellis., J. C., Labbé, J. L., Abraham, P. E. (2021). Bioprospecting Trichoderma: A Systematic Roadmap to Screen Genomes and Natural Products for Biocontrol Applications. In Frontiers Fungal Biology 2, Article 716511, p. 41. https://doi.org/10.3389/ffunb.2021.716511
Ryley, M., Bhuiyan, S., Herde, D., & Gordan, B. (2003). Efficacy, timing and method of application of fungicides for management of sorghum ergot caused by Claviceps africana. In Australasian Plant Pathology, 32(3), 329–338. https://doi.org/10.1071/AP03034
Salwan, R., Rialch, N., Sharma, V. (2019). Bioactive Volatile Metabolites of Trichoderma: An overview. In Singh, Harikesh Bahadur and Keswani, Chetan and Reddy, M. S. and Sansinenea, Estibaliz and García-Estrada, Carlos (Ed.): Secondary Metabolites of Plant Growth Promoting Rhizomicroorganisms: Discovery and Applications. Singapore: Springer Singapore, pp. 87–111
Saravanakumar, K., Dou, K., Lu, Z., Wang, X., Li, Y., & Chen, J. (2018). Enhanced biocontrol activity of cellulase from Trichoderma harzianum against Fusarium graminearum through activation of defense-related genes in maize. In Physiological and Molecular Plant Pathology, 103, 130–136. https://doi.org/10.1016/j.pmpp.2018.05.004
Schardl, C. L., Young, C. A., Hesse, U., Amyotte, S. G., Andreeva, K., Calie, P. J., et al. (2013). Plant-symbiotic fungi as chemical engineers: multi-genome analysis of the clavicipitaceae reveals dynamics of alkaloid loci. In PLoS Genetics, 9(2), e1003323. https://doi.org/10.1371/journal.pgen.1003323
Seidl, V., Song, L., Lindquist, E., Gruber, S., Koptchinskiy, A., Zeilinger, S., et al. (2009). Transcriptomic response of the mycoparasitic fungus Trichoderma atroviride to the presence of a fungal prey. In BMC Genomics, 10(1), 567. https://doi.org/10.1186/1471-2164-10-567
Sharma, P., Kumar, V., Ramesh, R., Saravanan, K., Deep, S., Sharma, M., et al. (2011). Biocontrol genes from Trichoderma species: A review. In African Journal of Biotechnology, 10(86), 19898–19907.
Sharma, V., Salwan, R., Sharma, P. N., & Kanwar, S. S. (2017). Elucidation of biocontrol mechanisms of Trichoderma harzianum against different plant fungal pathogens: Universal yet host specific response. In International Journal of Biological Macromolecules, 95, 72–79. https://doi.org/10.1016/j.ijbiomac.2016.11.042
Sheshegova, T. K., & Shchekleina, L. M. (2020). Control of Ergot (Claviceps purpurea (Fr) Tul.) with New Pesticides. In Russian Agricultural Sciences, 46(5), 472–475. https://doi.org/10.3103/S1068367420050183
Shieh, M. T., Brown, R. L., Whitehead, M. P., Cary, J. W., Cotty, P. J., Cleveland, T. E., & Dean, R. A. (1997). Molecular genetic evidence for the involvement of a specific polygalacturonase, P2c, in the invasion and spread of Aspergillus flavus in cotton bolls. In Applied and Environmental Microbiology, 63(9), 3548–3552. https://doi.org/10.1128/aem.63.9.3548-3552.1997
Shoresh, M., Harman, G. E., & Mastouri, F. (2010). Induced Systemic Resistance and Plant Responses to Fungal Biocontrol Agents. In Annual Review of Phytopathology, 48(1), 21–43. https://doi.org/10.1146/annurev-phyto-073009-114450
Smith, A., Beltrán, C. A., Kusunoki, M., Cotes, A. M., Motohashi, K., Kondo, T., & Deguchi, M. (2013). Diversity of soil-dwelling Trichoderma in Colombia and their potential as biocontrol agents against the phytopathogenic fungus Sclerotinia sclerotiorum (Lib.) de Bary. In Journal of General Plant Pathology, 79(1), 74–85. https://doi.org/10.1007/s10327-012-0419-1
Sood, M., Kapoor, D. K. V., Sheteiwy, M. S., Ramakrishnan, M., Landi, M., Arani, F., & Sharma, A. (2020). Trichoderma: The “Secrets” of a Multitalented Biocontrol Agent. In Plants, 9(6), 729.
Stenberg, J. A., Sundh, I., Becher, P. G., Björkman, C., Dubey, M., Egan, P. A., et al. (2021). When is it biological control? A framework of definitions, mechanisms, and classifications. In Journal of Pest Science, 94(3), 665–676. https://doi.org/10.1007/s10340-021-01354-7
Teather, R. M., & Wood, P. J. (1982). Use of Congo red-polysaccharide interactions in enumeration and characterization of cellulolytic bacteria from the bovine rumen. In Applied and Environmental Microbiology, 43(4), 777–780. https://doi.org/10.1128/aem.43.4.777-780.1982
Tenberge, K. B., Homann, V. V., Oeser, B., & Tudzynski, P. (1996). Structure and expression of two polygalacturonase genes of Claviceps purpurea oriented in tandem and cytological evidence for pectinolytic enzyme activity during infection of rye. In Phytopathology, 86, 1084–1097.
Thambugala, K. M., Daranagama, D. A., Phillips, A. J. L., Kannangara, S. D., & Promputtha, I. (2020). Fungi vs. Fungi in Biocontrol: An Overview of Fungal Antagonists Applied Against Fungal Plant Pathogens. In Frontiers in Cellular and Infection Microbiology, 10, 604923. https://doi.org/10.3389/fcimb.2020.604923
Trifinopoulos, J., Nguyen, L. T., von Haeseler, A., & Minh, B. Q. (2016). W-IQ-TREE: a fast online phylogenetic tool for maximum likelihood analysis. In Nucleic Acids Res, 44(W1), W232–W235. https://doi.org/10.1093/nar/gkw256
Tudzynski, P., & Scheffer, J. (2004). Claviceps purpurea: Molecular aspects of a unique pathogenic lifestyle. In Molecular Plant Pathology, 5(5), 377–388. https://doi.org/10.1111/j.1364-3703.2004.00237.x
Tundo, S., Mandalà, G., Sella, L., Favaron, F., Bedre, R., Kalunke, R. M. (2022). Xylanase Inhibitors: Defense Players in Plant Immunity with Implications in Agro-Industrial Processing. In International Journal of Molecular Sciences 23(23). https://doi.org/10.3390/ijms232314994
Tyśkiewicz, R., Nowak, A., Ozimek, E., & Jaroszuk-Ściseł, J. (2022). Trichoderma: The Current Status of Its Application in Agriculture for the Biocontrol of Fungal Phytopathogens and Stimulation of Plant Growth. In International Journal of Molecular Sciences, 23(4), 2329. https://doi.org/10.3390/ijms23042329
Uppala, S. S., Wu, B. M., & Alderman, S. C. (2016). Effects of Temperature and Duration of Preconditioning Cold Treatment on Sclerotial Germination of Claviceps purpurea. In Plant Disease, 100(10), 2080–2086. https://doi.org/10.1094/PDIS-02-16-0215-RE
Vinale, F., & Sivasithamparam, K. (2020). Beneficial effects of Trichoderma secondary metabolites on crops. In Phytotherapy Research, 34(11), 2835–2842. https://doi.org/10.1002/ptr.6728
Viterbo, A., & Horwitz, B. A. (2010). Mycoparasitism. In K. A. Borkovich & D. J. Ebbole (Eds.), Cellular and molecular biology of filamentous fungi (pp. 676–693). ASM Press.
Waage, J. K., Greathead, D. J. (1988). Biological control: challenges and opportunities. Philosophical Transactions of the Royal Society of London. B, Biological Sciences (318). https://doi.org/10.1098/rstb.1988.0001
Wang, X., Bi, W., Gao, J., Yu, X., Wang, H., & Liu, D. (2018). Systemic acquired resistance, NPR1, and pathogenesis-related genes in wheat and barley. In Journal of Integrative Agriculture, 17(11), 2468–2477. https://doi.org/10.1016/S2095-3119(17)61852-5
White T. J., Bruns T., Lee S., Taylor J. (1990). Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: M. A. Innis, D. H. Gelfand, J Sninsky, T. J. White (Eds.), PCR protocols (pp. 315–322). Academic Press, San Diego. https://doi.org/10.1016/B978-0-12-372180-8.50042-1
Wonglom, P., Ito, S., & Sunpapao, A. (2020). Volatile organic compounds emitted from endophytic fungus Trichoderma asperellum T1 mediate antifungal activity, defense response and promote plant growth in lettuce (Lactuca sativa). In Fungal Ecology, 43, 100867. https://doi.org/10.1016/j.funeco.2019.100867
Yassin, M. T., Mostafa, A. A. F., & Al-Askar, A. A. (2022). In vitro antagonistic activity of Trichoderma spp. against fungal pathogens causing black point disease of wheat. In Journal of Taibah University for Science, 16(1), 57–65. https://doi.org/10.1080/16583655.2022.2029327
Yu, Z., Liu, Z., Zhang, Y., & Wang, Z. (2022). The disease resistance potential of Trichoderma asperellum T-Pa2 isolated from Phellodendron amurense rhizosphere soil. In Journal of Forestry Research, 33(1), 321–331. https://doi.org/10.1007/s11676-021-01332-w
Zeilinger, S., Omann, M. (2007). Trichoderma Biocontrol: Signal Transduction Pathways Involved in Host Sensing and Mycoparasitism. In Gene Regulation and Systems Biology 1, GRSB.S397. https://doi.org/10.4137/GRSB.S397
Acknowledgements
We thank Prof. Dr. Ralph Hückelhoven and Dr. Nicole Metz (Chair of Phytopathology, TUM) for providing fungal strains, as well as Felix Hohenender (Chair of Phytopathology, TUM) for experimental advice. We are thankful to Dr. Feng Cai (International Comission on Trichoderma Taxanomy) for help with the molecular identification of the soil isolates. We also want to thank Petra Arnold, Sabrina Klasen, Manfred Reppke and Yuxin Zhang (Fungal Biotechnology in Wood Science) for their assistance in the greenhouse. Further, we want to thank KWS LOCHOW GmbH for providing ARANTES seeds and Dr. Berta Killermann (Bavarian State Research Center for Agriculture, LfL) for providing ergot sclerotia.
Funding
Open Access funding enabled and organized by Projekt DEAL. The work was partly supported by Deutsche Forschungsgemeinschaft (DFG project BE 6069/4–1 to PhB).
Author information
Authors and Affiliations
Contributions
Conceptualization: Pia Stange, Tanja Karl; Methodology: Pia Stange; Experimentation: Sophia Seidl, Pia Stange, J. Philipp Benz; Formal analysis and investigation: Sophia Seidl, Pia Stange; Writing – original draft preparation: Pia Stange; Writing – review and editing: Sophia Seidl, Tanja Karl, J. Philipp Benz; Supervision: Pia Stange, Tanja Karl; Project administration: J. Philipp Benz.
Corresponding author
Ethics declarations
Competing interests
The authors have no competing interests to declare that are relevant to the content of this article.
Additional information
This manuscript is part of the special issue ‘Biocontrol using beneficial fungi and bacteria’
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Stange, P., Seidl, S., Karl, T. et al. Evaluation of Trichoderma isolates as biocontrol measure against Claviceps purpurea. Eur J Plant Pathol 167, 651–675 (2023). https://doi.org/10.1007/s10658-023-02716-w
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-023-02716-w