Abstract
Natural monuments are IUCN Category III protected areas that play an important role in biodiversity conservation as they provide species refuge and allow species migration. Despite their status, natural monuments are often confined to cultural and fragmented landscapes due to anthropogenic land-use demands. In this population genomic study, we surveyed 11 populations of the endemic shrub Abeliophyllum distichum Nakai (Oleaceae), including five natural monument habitats, covering its range-wide distribution in South Korea. Using 2,254 SNPs as markers, our results showed a mean expected heterozygosity (He) of 0.319, with populations in the central distribution showing significantly higher He than those at the periphery. There was no significant heterozygote deficiency and inbreeding among studied populations overall (FIS = −0.098), except for a single natural monument population (GS-NM147). Population structure and differentiation was moderate to high (FST = 0.196), while recent gene flow between populations appeared weak, which can be attributed to the fragmented distribution and the outcrossing mating system of the heterostylous plant. Based on these findings, we provide suggestions for the population conservation and management of this endangered species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
On the Korean Peninsula, several natural populations of charismatic plants and animals are designated as natural monuments (NMs) (Cultural Heritage Administration 2006; Korea Protected Areas Forum 2009). The International Union for Conservation of Nature (IUCN) officially classifies NMs as a Category III protected area, which can be “a landform, sea mount, submarine cavern, geological features such as a cave or even a living feature such as an ancient grove” (Dudley 2008). Despite their conservation status, NMs are often confined to places that are otherwise cultural or fragmented (Dudley 2008). This is the case of the populations of the Korean endemic Abeliophyllum distichum Nakai (Oleaceae), a sole member in its genus and is more commonly called white forsythia.
As its common name suggests, A. distichum is similar to Forsythia species but is differentiated by its white flowers (versus yellow), as well as its flattened, broadly winged fruits (versus capsular) (Fig. 1). Its close relationship with Forsythia is well supported by molecular phylogenetic studies (Kim 1999; Kim et al. 2000; Wallander and Albert 2000; Lee et al. 2007; Kim and Kim 2011), and cytological and embryological data (Sax and Abbe 1932; Taylor 1945; Ghimire et al. 2020). Like some Forsythia species, the shrub is distylous (or heterostylous), having both ‘pin’ (long-styled) and ‘thrum’ (short-styled) plants, a floral morphological condition that enhances pollen-mediated gene flow (Ishihama et al. 2003). Early reports of the low fruit sets found in populations of A. distichum, however, could be attributed to the unequal ratio of pin/thrum morphs (Ryu et al. 1976; Kim and Maunder 1998).
The species and their habitats are considered one of the country’s “cultural properties”, which are defined as those that carry great historic, artistic, academic, and scenic values (Korea Protected Areas Forum 2009). Currently, there are five officially designated NMs for the species: NM 147, NM 220, NM 221, NM 364, NM 370a, and 370b (Cultural Heritage Administration 2006). In at least one locality (i.e. Goesan; see Fig. 2a in Materials and Methods) where the species naturally occurs, a yearly spring festival is even held in honor of the symbolic white flower. The deciduous shrub flowers in late March to April before the emergence of leaves, and is visited by various insect pollinators (Chung 1999; Kang et al. 2000). The most recent IUCN Red List category assessment for the species is ‘endangered’ due to its severely fragmented population distribution (Son et al. 2016).
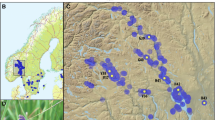
Abeliophyllum distichum a map of distribution in South Korea, showing the sampled sites (black circle) in their localities, including (insets in) Jincheon (a1), Goesan (a2), Yeongdong (a3), and Buan (a4); b results of STRUCTURE analysis, showing the optimal number of clusters (K = 2, K = 3, and K = 5) inferred from 152 individual genotypes. Each bar represents a single individual and its proportion of assignment to each of the sampled sites separated by solid black lines
Despite the protected status of many threatened species, the increasing human demand from land-use change has resulted in the isolation of species populations and their habitats (Young et al. 1996; Henle et al. 2004; Aguilar et al. 2008). Anthropogenic disturbance, in addition to the intrusion of non-native plants and animals, can cause natural populations to decline (Gemmill et al. 1998; Gonzales and Hamrick 2005). In the case of A. distichum, the pressure for infrastructure development in the early 1990s resulted in a drastic action of translocating (therefore overturning the status of) a NM in the town of Buan (see Fig. 2a in Materials and Methods). About two decades before that, the first-ever A. distichum habitat declared a NM (NM 14) in the locality of Jincheon was removed from its status due to habitat destruction (Kim and Maunder 1998).
Many reviews of population genetics studies with relation to conservation have shown that threatened or rare species are often associated with higher vulnerability to fragmentation, leading to the loss of connectivity and reduction of genetic diversity, and thus at increased risk of extinction (Ellstrand and Elam 1993; Cruzan 2001; Wallace 2002; Willi et al. 2007; Newman et al. 2013; De Vriendt et al. 2017; Chung et al. 2020). As such, it is crucial to estimate the levels of within-population genetic variation and among-population differentiation of a given species as well as its genetic structure for effective genetic management of threatened and fragmented populations (Frankham 2010; De Vriendt et al. 2017; Chung et al. 2020; DeWoody et al. 2021; García-Dorado and Caballero 2021).
In this paper, we conducted a range-wide population genomics study on A. distichum by sampling from 11 known populations, including five NMs. We estimated population genetic parameters using single nucleotide polymorphisms (SNPs) as markers through the genotyping-by-sequencing (GBS) approach. Our main objectives were as follow: (i) to determine the levels of genetic variation within populations and the degree of genetic differentiation among populations; (ii) to determine population structure and to infer significant management units for conservation; (iii) to evaluate the potential of population connectivity via recent/contemporary migration; and (iv) to provide information for the appropriate conservation and management of the study species.
Materials and methods
Study species distribution and sampling
Natural populations of A. distichum are restricted to the central provinces of South Korea, with some reports of occurrence in North Korea (Kim and Maunder 1998). The sampled sites (11 populations) represent the known geographic range of the species in the country (Fig. 2a). Some of these sites are recently discovered populations, two of which with pending application/consideration for the NM status, while others are either disturbed remnants of wild populations or those which have been subjected to previous conservation management measures (e.g. restoration) (Table 1). At each location, we collected small amounts of leaf or flower bud tissue from a maximum of 20 individuals, observing enough distance to avoid collecting potential clones. Samples were preserved in plastic bags with silica gel desiccant until DNA extraction. A voucher specimen from each population was collected and deposited in the Herbarium of Hallym University (HHU). Appropriate permits were obtained from each administrative locality before collecting plant samples and voucher specimens, particularly those from NM habitats.
DNA extractions, library preparation, and sequencing
Total genomic DNA from each sample was extracted from 20 mg of dried leaves/flower buds using a DNeasy Plant Mini Kit according to manufacturer protocol (Qiagen, Germany). The quality of extracted DNA was evaluated and quantified with a Qubit Fluorometer (Invitrogen, USA) and visualized on a 1% agarose gel. A total of 15 individuals with the highest DNA quality were selected and genotyped for each population.
Samples that passed quality and quantity filters (12–15 per population; see Table 2) were subsequently sent to Seeders Inc. (Daejeon, Korea) for GBS library construction (Elshire et al. 2011). In this approach, samples were digested with methylation-sensitive ApeKI restriction enzyme and barcoded with sequence adapters before pooling into sets of ninety-six individuals. Each library was amplified by polymerase chain reaction (PCR) and sequenced in a single lane on an Illumina HiSeq X (Illumina, USA) using 150 bp paired-end sequencing runs.
Sequence analyses, bioinformatics, and SNP identification
Given that no reference genome for A. distichum was available, a draft genome was assembled de novo to facilitate read mapping and SNP identification. The draft sequence was pre-processed using DynamicTrim (phred score ≥ 20) and LengthSort (short read length ≥ 25 bp) programs in the software package SolexaQA v.1.13 (Cox et al. 2010). The reads produced were assembled (k-mer = 59) using SOAPdenovo2 (Ver. 2.04) (Luo et al. 2012).
GBS sequencing data were demultiplexed using Cutadapt v.1.8.3 (Martin 2011), and quality-trimmed using DynamicTrim (phred score ≥ 20) and LengthSort (short read length ≥ 25 bp) in the software SolexaQA v.1.13 (Cox et al. 2010). Cleaned reads were mapped against the de novo assembled reference draft genome using the program BWA 0.6.1-r104 (Li and Durbin 2009), with the following options: seed length (–l) = 30, maximum difference in the seed (–k) = 1, number of threads (–t) = 20, mismatch penalty (–M) = 6, gap open penalty (–O) = 15, and gap extension penalty (–E) = 8, while all the rest were set to default. Using a custom script, sequence alignment map (SAM) file format was merged and sorted into a binary alignment map (BAM) with SAMtools v.0.1.16 (Li et al. 2009) to detect raw SNPs (In/Del) and extract consensus sequences. The following options were set: minimum mapping quality for SNPs (–Q) = 30, minimum mapping quality for gaps (–q) = 15, minimum read depth (–d) = 3, maximum read depth (–D) = 257, min indel score for nearby SNP filtering (–G) = 30, SNP within INT bp around a gap to be filtered (–w) = 15, and window size for filtering dense SNPs (–W) = 30.
The final matrix was generated following these SNPs filtering criteria: (i) biallelicity, (ii) a missing rate of < 30%, and (iii) a minor allele frequency (MAF) of > 5%. Hardy–Weinberg (H-W) equilibrium (p ≤ 0.01) and FST (> 0.1) (Weir and Cockerham 1984) for each SNP locus were tested using the pegas package (Paradis 2010) in R (R Core Team 2020), while linkage disequilibrium (LD) (r2 = 0.8) was computed using the software Plink v1.07 (Purcell et al. 2007). All the above bioinformatics and quality control filtering were conducted in Seeders Inc. (Daejeon, Korea). Overall, 12–15 samples per population passed the quality control (see Table 2 in Results) and were used for succeeding analyses. The online program PGDSpider version 2.1.1.5 (Lischer and Excoffier 2012) was used in converting different file formats for all downstream analyses below.
Analysis of genetic variation
To determine population genetic variation and inbreeding, we used GenAlEx 6.5 (Peakall and Smouse 2006, 2012) for the calculation of the following summary statistics: number of alleles (A), number of effective alleles (Ae), H-W expected heterozygosity (He), and observed heterozygosity (Ho). The significant difference between He and Ho for each population was determined using the t-test after testing the Bartlett test for the homogeneity of variances as implemented in the adegenet package (Jombart and Ahmed 2011) in R (R Core Team 2020). Also in R, the significant difference in He among and between populations was computed using the one-way analysis of variance (ANOVA), and the post hoc Tukey’s HSD test using the package dplyr (Wickham et al. 2022). The inbreeding coefficient (FIS) for each population was calculated using Arlequin 3.5 (Excoffier and Lischer 2010) with 20,000 permutations to determine whether values were significantly different than zero.
Analysis of genetic structure and differentiation
To evaluate genetic structuring, we assigned individuals to genetic clusters based on the Bayesian assignment analysis implemented in the program STRUCTURE version 2.3.4 (Pritchard et al. 2000). Population clustering with the admixture and correlated allele frequencies models were selected with 50,000 burn-in steps followed by 100,000 runs of Markov Chain Monte Carlo (MCMC). We ran 10 replicates at each value of K after setting the prior for the most likely number of K from 1 to 10, assuming that the samples from the single restored habitat (i.e. JC2) had been sourced from one of the ten other sites sampled. The optimal number of genetic groups was determined by calculating the Mean LnP(K) (Pritchard et al. 2000) and ΔK (Evanno et al. 2005) as implemented in Structure Harvester (Earl and vonHoldt 2012). The results of population clustering were aligned and visualized using the CLUMPAK webserver (Kopelman et al. 2015).
To test if genetic groups were simply due to the presence or absence of isolation by distance (IBD), we estimated the correlation between the matrix of pairwise Slatkin’s linearized FST (FST/(1 – FST)) and that of the log-transformed geographic distances (Rousset 1997) in a paired Mantel test with 9,999 permutations for significance testing using GenAlEx 6.5 (Peakall and Smouse 2006, 2012). In this test, we excluded the genetic and geographic data from the artificially restored JC2 to avoid bias.
We further examined the different levels of genetic differentiation by computing the F-statistics (FST, FIS, FIT), and the analysis of molecular variance (AMOVA) (Excoffier et al. 1992) with 20,000 permutations for significance testing using the program Arlequin 3.5 (Excoffier and Lischer 2010). Pairwise FST values using the same program were computed with 9,999 permutations for significance testing after Bonferroni adjustments. A principal coordinate analysis (PCoA) was performed to investigate the pattern of genotype clustering for all 152 individuals via covariance matrix with data standardization as implemented in GenAlEx 6.5 (Peakall and Smouse 2006, 2012).
Estimation of gene flow and effective population size
Recent/contemporary migration among the studied populations was inferred using the Bayesian framework in the program BayesAss 3.0 (Wilson and Rannala 2003; Mussmann et al. 2019). In this method, recent migration (m) in a population is inferred based on the per generation proportion of individuals that were migrants from another population (Rannala 2007). Mixing parameters for the delta values of migration rate (–m), allelic frequency (–a), and inbreeding (–f) were initially altered during test runs to achieve the best acceptance values (between 0.2 and 0.6) as recommended by Rannala (2007). Initial runs were set at 3,000,000 MCMC iterations, a burn-in of 1,000,000 generations, and a parameter value sampling frequency of 1,000. After optimum mixing parameter values were determined (–m = 0.18, –a = 0.55 and –f = 0.05), we completed 10 final runs with a different seed number each to validate consistency (Rannala 2007). Final runs consisted of 30,000,000 MCMC iterations, a burn-in of 3,000,000, and a sampling frequency of 1,000. Convergence models were visualized using the Tracer program (Rambaut et al. 2018), and as suggested (Faubet et al. 2007; Meirmans 2014), we selected the best-fit model as indicated by the lowest value of Bayesian deviance using the R script found in Meirmans (2014). Confidence intervals (CIs) were computed following Rannala (2007), and values of m whose 95% CIs did not include zero were reported as significant. Due to the limitations of the program, only 500 randomly selected loci with the least number of missing data, as identified in Arlequin 3.5 (Excoffier and Lischer 2010), were used. To avoid further bias, we also excluded the artificially restored population JC2 from the analysis.
To complement the test above, the contemporary effective population size (Ne) for each site was inferred using NeEstimator 2.0.1 (Do et al. 2014). Ne estimates were calculated from the bias-corrected method based on LD (Waples 2006), following the random mating model, with a minimum allele frequency cutoff of 0.05, and 95% CIs by jackknifing over individuals (Jones et al. 2016). The same number of loci used for the computation of contemporary gene flow was employed for this analysis.
Results
Levels of genetic variation
Initial quality check of nearly 6 million reads after Illumina sequencing resulted in the total SNP matrix of 528,871 loci. Initial filtering (i.e. biallecity, individual missing, and MAF) resulted to the identification of ca. 27,000 SNPs. Further filtering steps (e.g. H-W equilibrium, LD, and FST) produced 2,254 final sets of SNP loci in 152 genotyped individuals across 11 sampled locations, with an average missing data of 2.2%.
Our results revealed that overall, there were relatively small differences in within-population genetic variation between sites. Table 2 shows that the mean A for all individuals was 1.838, ranging from 1.551 to 1.991. The mean He for the species was 0.319, slightly lower than the average Ho of 0.357. The population with the lowest He was that of BA-NM370b (He = 0.263), while the highest was that of GS-NM147 (He = 0.372) (Table 2). Among all sites sampled, only GS-NM147 recorded significantly lower Ho after the t-test (p < 0.001), suggesting that the population is not in H-W equilibrium. The indication of heterozygote deficiency for this population is supported by a highly significant inbreeding coefficient (FIS = 0.131, p < 0.01). Except for this population, genetic analyses showed no other locations recording significant heterozygote deficiency, with most sampled sites even recording a slightly negative FIS.
The one-way ANOVA predicted that there was a significant difference in He between at least two populations (F (10, 2,243) = 127.10, p < 0.001). The post hoc Tukey’s HSD test indicated that significant differences in He were generally between populations in the central and peripheral distributions. In Table S1, for example, He of upper central population GS-NM147 very highly significantly differed from that of the southernmost population BA-NM370b (Tukey’s HSD = 0.109, adj. p < 0.001), and that of the northernmost population YJ (Tukey’s HSD = −0.108, adj. p < 0.001), respectively. Even when populations from the upper central range were compared with those in the lower central range (e.g. GS-NM147 vs. YD-NM364), a very highly significant difference in He (Tukey’s HSD = −0.050, adj. p < 0.001) was still observed. These differences in He are summarized in Fig. 3, but see Table S1 for the complete list of Tukey’s values between all population pairs.
Genetic structure and differentiation
Results from AMOVA (Table 3) revealed that only 19.64% of the total genetic variation was due to among-population differentiation and that the majority of differentiation was within individuals (88.22%). F-statistics from the analysis also showed that our focal species was moderately to highly structured and significantly differed from zero after 20,000 permutations (FST = 0.196, p < 0.001). Although it was shown that there was a significant total individual expected reduction in heterozygosity (FIT = 0.118, p < 0.001), the slightly negative total inbreeding coefficient (FIS = −0.098) was not shown to significantly differ from zero (p > 0.05), indicating little or no inbreeding in the species.
Pairwise FST analyses between sites showed values that ranged from 0.02 to 0.31 (Fig. S2). A significant difference in the genetic distance even after Bonferroni adjustment was recorded for all population pairs, except for the sampled populations in/between the locality of Goesan and Jincheon: GS-NM147 vs. GS-NM221 (0.03), and GS-NM147 vs. JC1 (0.03). No significant difference in pairwise FST was also shown between the artificially restored population JC2 vs. GS-NM221 (0.02), and JC2 vs. GS-NM147 (0.02). On the other hand, the highest pairwise FST estimates were found in BA-NM370b vs. YJ (0.31), and BA-NM370b vs. AD (0.29), coinciding with the large geographic distances between the southernmost range in Buan, and the northernmost and easternmost distributions in Yeoju and Andong, respectively (see Fig. 2a). The Mantel test for IBD among all sampled sites, however, revealed no significant correlation between the linearized pairwise FST and the log-transformed geographical distance (rxy = −0.107, R2 = 0.011, p > 0.05).
As visualized in Fig. S1a, ΔK estimates revealed that K = 3 was the most likely number of genetic clusters, besting the other two optimal genetic groupings K = 2 and K = 5. This inference was supported by the distribution of the mean log likelihood probability of data LnP(D), with the biggest increase at K = 2 and K = 3, and gradually being reduced after K = 5 (Fig. S1b). Based on these findings, it can be inferred that the two southernmost NM populations BA-NM370a and BA-NM370b were first to diverge (K = 2) and remained genetically distinct as K increased (K = 3 and K = 5) (Fig. 2b). At K = 3, the northernmost population YJ began to split off from the group, while the rest remained clustered, consisting of individuals admixed with mostly ‘blue’ genetic assignment. At K = 5, after the easternmost population AD split off, this ‘blue’ cluster was shown to be composed of subpopulations of individuals from sites in the upper central range locality of Goesan (GS-NM147 and GS-NM221) and Jincheon (JC1 and JC2), while the lower central populations in Yeongdong (YD and YD-NM364) and Okcheon (OC) began to be identified with their ‘green’ genetic assignment, although the latter (i.e. OC) could be also more closely associated with the members of the ‘blue’ cluster in the upper central range. Overall, this genetic structuring appeared to reflect the geographic locations of the sampled sites, with the spatially close populations tending to group together.
PCoA results (Fig. 4) showed that individuals in the same populations (or genetic groups) were clustered together, reflecting similar patterns of genetic structure as determined by STRUCTURE. In coordinate 1 (6.83%), genotypes from the southern populations in Buan (BA-NM370a and BA-NM370b) were shown to the far left of the coordinate and clustered very distinctly from the rest of the group. In coordinate 2 (5.45%), a very close genotype clustering was shown for those at the upper central distribution (along coordinate 2), with almost undiscernible clustering for individuals from sampled sites in Jincheon (JC1 and JC2) and Goesan (GS-NM147 and GS-NM221). Slightly clustered above them are lower central populations in Yeongdong (YD and YD-NM364) and Okcheon (OC). The eastern and northern populations AD and YJ, respectively, were shown plotted farther (below coordinate 2) from members of the central group.
Gene flow and effective population size
BayesAss analyses among population pairs (after excluding the artificially restored population JC2) showed weak signs of recent migration, with values of m ranging from 0.013 to 0.111. Here, only the highest estimates (above 0.015) are presented in Table 4 (but see complete migration values in Table S2). The strongest contemporary migration was symmetrically shown between two Goesan NM populations: from GS-NM221 to GS-NM147 (m = 0.111), and from GS-NM147 to GS-NM221 (m = 0.109), indicating that ca. 11% of the individuals in these populations were recent migrants from each other. Symmetric migration was also observed from GS-NM147 to the wild population remnant JC1 (m = 0.092), and from JC1 to GS-NM147 (m = 0.069), while a weaker source-sink asymmetric gene flow pattern was shown from YD to YD-NM364 (m = 0.027). All CI values were shown to overlap zero, suggesting insignificant or absence of recent/contemporary migration among all population pairs.
Analysis of contemporary Ne using the LD method also showed generally low estimates, ranging from 1.2 to 22.7 for all populations (Table S3). All the sampled sites revealed values that were lower than the suggested 50 (Jamieson and Allendorf 2012; Franklin et al. 2014), with six populations even showing Ne lower than 10, including four sampled NM habitats (i.e. YD-NM364, GS-NM221, BA-NM370a, and BA-NM370b). Ne for all sites analyzed was supported by not zero-overlapping but narrow CIs.
Discussion
Levels of genetic variation
Our data showed that although He was relatively similar among sampled sites, it was greater for populations with the occurrence at the central range and lower at the periphery. In general, the findings were supported by Tukey’s HSD test, which highlighted the statistically significant difference in He when populations in the central range were compared with those at the marginal distribution. These observations appear to be the signatures of the central-marginal/center-periphery hypothesis (Kennedy et al. 2020). This model proposes that the main core populations are genetically more diverse (“abundant center”) than the populations at the marginal range (Brussard 1984; Lawton 1993; Sagarin and Gaines 2002; Vucetich and Waite 2003; reviewed in Eckert et al. 2008). The findings can be based on the assumption that both rates of gene flow and Ne are higher at the central range (Eckert et al. 2008), where geographic distances between populations are shorter than those at the periphery. In our study, the sampled sites at the upper central range, specifically those in the locality of Goesan (GS-NM147 and GS-NM221) and Jincheon (JC1 and JC2), may represent current proximate population networks, relatively close to the slightly less genetically diverse populations at the lower central range in the locality of Yeongdong (YD and YD-MN364) and Okcheon (OC). In the past, these sites might have been once the core of a large metapopulation that split into smaller fragmented subpopulations observed today. On the other hand, peripheral populations at the south edge (BA-NM370a and BA-NM370b), marginal north (YJ), and east (AD) exhibited lower He, probably due to repeated founder events (Sagarin and Gaines 2002; Petit et al. 2003; Eckert et al. 2008). In addition, the southernmost populations occurring at the lowest-latitude (i.e. BA-NM370a and BA-NM370b) revealed relatively reduced within-population genetic variation, which is one of the characteristic features of rear-edge populations (Hampe and Petit 2005).
Overall, no sign of within-population inbreeding (except for GS-NM147) was found in this study, with most sites recording FIS that was close to or even below zero (also, note the total species FIS of −0.098 as indicated in AMOVA). These findings can be attributed to the outcrossing mating preference of the species, maintained by the distylous flowers. In pin flowers, the stigmas are placed above the anthers, while in thrum, the stigmas are positioned below; thus the exchange of pollen is promoted among different morphs. While some population genetic studies on other distylous species showed even more strongly negative inbreeding coefficient, as in Pulmonaria officinalis (FIS = −0.30) (Meeus et al. 2012), most studies we found indicated higher positive estimates (ranging from FIS = 0.10–0.43) (Van Rossum and Triest 2003; Van Rossum et al. 2004; Bresowar and McGlaughlin 2015; Shao et al. 2019). This is because in the latter cases, factors such as population size, morph bias, and natural selection were shown to play a more important role in the patterns of genetic diversity rather than self-incompatibility and mating preference among heterostylous species.
Along with this morphological feature, most distylous species include a self- and intramorph-incompatibility system, producing viable offsprings only from intermorph crosses (pin × thrum or vice versa) (Barrett and Shore 2008; Cohen 2010, 2019). An early study on A. distichum heterostyly-incompatibility revealed a very low fruit set of 1.3% under artificial intramorph pollination (i.e. pin × pin or thrum × thrum), while intermorph pollination showed 30.8% (Ryu et al. 1976). In natural settings, however, intermorph crosses were shown to produce much lower fruit sets of 10.4% for the heterostylous species Primula sieboldii E.Morren, primarily due to severe pollinator limitation (Washitani et al. 1994). The seed germination success rate for our focal species, however, could be even lower due to mechanical and chemical inhibitors (Yoo and Kim 2001; Ghimire et al. 2015).
Although a considerable variance of allozyme-based genetic variation among plant species can be attributed to their series of life-history and ecological traits (e.g. geographic range, reproductive biology, and breeding system (Hamrick and Godt 1989), mean values of genetic parameters can differ according to the markers employed in analyses. Since A. distichum is the only species in its genus, we compared our results with related studies that used a similar method of genome reduction with restriction enzymes. Our comparison showed that the mean He (0.319) fit the range (0.180–0.400) of similarly threatened woody endemics (Hasbún et al. 2016; Cheng et al. 2020; Hartvig et al. 2020). When compared with widespread perennial angiosperms (Borrell et al. 2018; Gomes Viana et al. 2018; Takach et al. 2019), our results revealed much higher He (0.319 vs. 0.087–0.167), indicating that sometimes genetic diversity for rare plants can be equivalent or greater than those of more common species, even those among/between congeneric taxa (e.g. Gitzendanner and Soltis 2000; Ellis et al. 2006; Edwards et al. 2014).
Genetic structure and differentiation
Analyses of population differentiation revealed a wide range of genetic differentiation among population pairs, but on average, it can be classified as moderate to high (FST = 0.196) (Wright 1978; Hartl and Clark 1997). The findings can be directly attributed to the lack of connectivity among sites, specifically between the central and peripheral range populations, mainly due to their large spatial separation. Pairwise FST estimates showed that more than 90% of the total population pairs significantly differed genetically, except those in close geographic proximity.
The admixed genetic assignment in STRUCTURE showed that most individuals originated within their sampled range. Population structure inference was able to define the number of genetic clusters for the species and is useful in identifying significant ‘management units’ for effective conservation strategies. Our findings are in concordance with an earlier allozyme study by Chung (1999), which revealed that his sampled population in Buan was the most genetically distinct. In this range-wide investigation, our most optimal population structure inference was able to determine additional conservation units not covered during the previous research: the recently discovered northernmost population YJ (at K = 3) and the easternmost population AD (at K = 5). Range edge or marginal populations, such as above, are often given conservation priority because of their lesser genetic diversity and their occurrence in less favorable habitats (Lawton 1993; Vucetich and Waite 2003; Hampe and Petit 2005).
Our STRUCTURE (and pairwise FST) results also support what seemed to be the possible population source(s) of materials during the restoration of JC2. The tests did not reveal that sourcing came from its nearest neighbor in Jincheon (i.e. the wild population remnant JC1) but rather suggested that the individuals transplanted to the latter may have been sourced from either of or both the two NM populations in nearby Goesan (GS-NM147 and/or GS-NM221). Regardless, this shows that the selection of individuals for the restoration followed the geographic or regional admixture provenancing (Bucharova et al. 2019), by relying on the closest regional source.
The genetic differentiation in our focal species may also be attributed to its mating system, which is often regarded to have the strongest influence in genetic structure, affecting both within-population genetic diversity and among-population genetic differentiation (Hamrick and Godt 1989). Since A. distichum does not appear to possess any mechanisms that facilitate self-pollination, the no significant inbreeding, and statistically significant among-population genetic differentiation, as shown in AMOVA and F-statistics, are most likely a consequence of its outcrossing breeding system and its long life cycle (Nybom and Bartish 2000). In comparison, the FST estimate for A. distichum (0.196) was shown to fall between those of other heterostylous taxa (Van Rossum and Triest 2003; Reisch et al. 2005; Meeus et al. 2012), which ranged from 0.1 to 0.2. On the other hand, allozyme-based studies on closely related Forsythia species revealed a range of 0.144–0.214 (Chung et al. 2013), while earlier A. distichum studies using the same marker type showed a GST of 0.141 (Chung 1999) and 0.272 (Kang et al. 2000). Caution, however, should be observed since the different values reported above can be due to the use of various markers with varying polymorphism, levels of dominance, and genome coverage (Ouborg et al. 1999; Vignal et al. 2002; Garcia et al. 2004). When compared with similar studies that employed SNP loci, genetic differentiation for our focal species was shown closest to that of another self-incompatible, woody endemic Coffea mauritiana Lam. (0.168) (Garot et al. 2019). We also do not reject the possibility that population differentiation and structuring may have been caused by the differences in regional environmental conditions and other potential evolutionary factors not described here (e.g. genetic drift).
Gene flow and effective population size
Our recent gene flow estimation showed that the highest observed values (although up to only 11%) were between population pairs with the closest geographic distance (i.e. those in the central range), becoming weaker toward more distant populations. For instance, population pairs with the highest m, such as the Goesan populations GS-NM147 and GS-NM221 are only separated by ca. 6 km distance, while GS-NM147 and JC1 in the same central range, are ca. 40 km away. Regardless, these distances may still appear to be beyond the natural dispersal ability of the winged fruit of the endemic shrub. For instance, seed dispersal experiments in the invasive tree Ailanthus altissima (Mill.) Swingle (Simaroubaceae) revealed that samaras could only reach about 100 m distance even from the edges of open fields (Landenberger et al. 2007). Populations of A. distichum growing under forest canopies (e.g. AD, OC), in particular, may encounter more difficulties in its seed dispersal over longer distances, although less likely challenging for its pollen vectors, provided the occurrence of other intermediate subpopulations.
We argue that the overall low levels of m are a result of natural geographic barriers or human-modified landscapes (e.g. road infrastructure and agriculture) that may have been obstacles to between-population gene flow and dispersal way beyond the estimated time inferred in the analysis. Since the program BayesAss measures recent migration that is estimated to have taken over the last one to three generations (Howes et al. 2009; Huey et al. 2013; Samarasin et al. 2017), we estimate that contemporary migration events may have occurred in a period of about 20 years, given our generation time estimate of six to ten years for the species, as observed in its closest Forsythia relatives (Schrader 2020). We believe that anthropogenic landscape modifications that created population fragments for the species may have at least been in existence since the past half a century when the country started its industrialization. Hence, the weak or even absence of contemporary migration found in this study strongly suggests that the current levels of structure are rather a consequence of historical migration events, if not of the long-standing biological feature of the species. We hope that a more detailed study on the A. distichum demographic history can shed light on the historical gene flow and the source-sink relationship among its fragmented populations.
NeEstimator analyses similarly showed low estimates of Ne and revealed that values were much lower than the numbers generally suggested in the 50/500 rule (Jamieson and Allendorf 2012; Franklin et al. 2014). Accordingly, an Ne of 50 is required to avoid short-term inbreeding depression, while an Ne of 500 is recommended to maintain a long-term evolutionary potential (Lehmkuhl 1984; Jamieson and Allendorf 2012). Ne estimates below these values, along with an increased level of inbreeding, are warning signs that a particular population is potentially at increased risk of genetic erosion and should therefore be subject to conservation intervention.
Another factor in the reduction of Ne can be attributed to the skewed pin/thrum morph ratios, resulting in the insufficient availability of compatible mates, thereby decreasing the chance of effective pollination between two compatible morphs, or the Allee effect (Stephens and Sutherland 1999; Stephens et al. 1999). On the contrary, in populations consisting of both pin and thrum plants, the percent fruit set increases as the ratio of pins to thrums approached unity as found in the herb Houstonia caerulea L. (Rubiaceae) (Wyatt and Hellwig 1979). Although the unequal floral morph occurrence is not uncommon and is well described among heterostylous angiosperms, even among those with fragmented distributions (e.g. Van Rossum and Triest 2006; Hodgins and Barrett 2008; Meeus et al. 2012), plant populations with unequal morph ratios, pollen limitation, and disrupted pollinator service are expected to show low levels of genetic diversity and high levels of inbreeding for the most fertile morph (Barrett 1989; Barrett and Husband 1990; Wilcock and Neiland 2002; Meeus et al. 2012). Our findings can be beneficial for the management of GS-NM147, which was found to have a significant excess of homozygotes (FIS = 0.131) and was previously assessed to have a severely biased morph ratio, with two times more pin morphs than thrum (Kim and Han 1998; Chung 1999).
Results of our Ne (1.2–22.7) computation in NeEstimator, however, should be taken with caution, as the very low estimates using the LD method indicate that some pairs of loci are closely linked (Table S3). Our data may also contain less information than assumed, attributable to the low sample sizes (England et al. 2006; Waples 2006), resulting in the downward bias of Ne as indicated by the narrow CIs. Future studies are recommended including more than 30 samples per population to achieve more reliable estimation (Waples 2006).
Implications for conservation
In this study, we evaluated the levels and degree of genetic variation within and among populations, the levels of inbreeding, and the degree of connectedness among populations of A. distichum for the ultimate goal of providing useful information for the effective conservation planning of this endangered shrub. Here, we present an additional data set of more than 2,000 SNP loci for the genetic analysis of this NM species, which was last evaluated ca. 20 years ago (e.g. Chung 1999; Kang et al. 2000). By generating range-wide data, we aimed to help identify NM habitats that are potentially at increased risk of negative genetic consequences and contribute to the further assessment of its fragmented populations, including those which were discovered after the last genetic survey. For instance, the significant excess of homozygosity estimated for the NM habitat GS-NM147 suggests that the current status of this population is a result of mating between close relatives, and could benefit from genetic management by pollen transfer from other (nearby) populations. Also, since A. distichum is a relatively long-lived woody species, the moderate to high levels of He observed in other populations could possibly be a transitory state and may not be sustained if no management actions are undertaken. Therefore, we suggest that numbers of individuals, especially in the disturbed wild remnant population JC1 and in the restored site JC2 be maintained, if not increased, to promote pollen movement.
Given the results of genetic differentiation between (and among) populations, we do not recommend sourcing individuals from any populations at the marginal range (e.g. Yeoju, Buan, or Andong) to those in the central distribution (e.g. Goesan or Jincheon), and vice versa as a conservation strategy. We suggest that future restoration efforts prefer seeds and seedlings from within the same geographic range (i.e. within the central range or the same marginal range) to avoid mating between close relatives and thus, retain the regional genetic structure. Due to the outcrossing mating system of the species, we also discourage transfers of individuals from different range margin populations (e.g. between the northernmost and the southernmost populations) to avoid outbreeding depression. Moreover, protection for rear edge populations (i.e. BA-NM370a and BA-NM370b) should be prioritized because the prolonged isolation and successful persistence of such groups suggest their evolutionary potential that can be useful for the long-term conservation of genetic diversity and phylogenetic history of the species (Hampe and Petit 2005).
Climate change could also exacerbate the challenges in the conservation of A. distichum due to the fragmentation and small Ne estimates of its populations. Under sustained warming temperatures, population genetic measures would be an essential element in creating comprehensive management plans to conserve endangered species (Schierenbeck 2017; Smith and Kay 2018). We assume that A. distichum will just have to adapt to higher temperatures under the current climate scenario, but at the same time, we do not exclude the possibility that due to postglacial recolonization, there are undiscovered natural populations that continue to migrate toward higher latitudes (e.g. those between South and North Korea). However, migrating uphill for the species may be restricted because many of its populations are situated in habitat islands surrounded and limited by human-modified landscapes. Many recently documented habitats are also occurring along slopes, some of which are bordered by streams (e.g. AD) or roadways (e.g. OC and YD). Thus, the species mode of dispersal of its samara or clonal reproduction via its air layering stems could only land to a less suitable substrate or environment. The simplest and most practical solution, therefore, is to clear the surrounding areas and expand the existing territory, particularly for the fenced NM habitats.
In countries that implement the NM system, the obstacles to the conservation of endangered taxa may not be that difficult to overcome because such species and their habitats can be a source of national pride and heritage. A greater challenge, however, as in the case of NM species conservation in Korea, can be the (poor) cooperation among different institutions and conservation groups. Due to the overlapping demarcation of responsibilities of at least three government ministries in charge of managing these national treasures, Kim (2006) pointed out the lack of communication and collaboration in the designation, restoration, ex situ conservation, and overall protection of endangered NM species habitats. This problem could extend to independent researchers, particularly the difficulty of retrieving information and technical documents related to the past restoration and augmentation works (e.g. in the artificially restored JC2). We encourage a closer partnership among these organizations, including constant communication with the managers of private lands where some NMs are located (e.g. GS-NM147 and GS-NM221). As a future study, we also recommend continuous population demographic and genetic monitoring, particularly the adaptation of the species to its natural and human-modified environment.
Data availability
The de novo assembled draft genome can be accessed at NCBI under BioProject PRJNA807751.
References
Aguilar R, Quesada M, Ashworth L, Herrerias-Diego Y, Lobo J (2008) Genetic consequences of habitat fragmentation in plant populations: susceptible signals in plant traits and methodological approaches. Mol Ecol 17:5177–5188
Barrett SCH (1989) Mating system evolution and speciation in heterostylous plants. In: Otte D, Endler J (eds) Speciation and its Consequences. Sinauer Associates, Massachusetts, pp 257–283
Barrett SCH, Husband BC (1990) Variation in outcrossing rates in Eichhornia paniculata: the role of demographic and reproductive factors. Plant Species Biol 5:41–55
Barrett SCH, Shore JS (2008) New insights on heterostyly: comparative biology, ecology and genetics. In: Franklin-Tong VE (ed) Self-incompatibility in flowering plants – evolution, diversity, and mechanisms. Springer, Berlin, Heidelberg, pp 3–32
Borrell JS, Wang N, Nichols RA, Buggs RJ (2018) Genetic diversity maintained among fragmented populations of a tree undergoing range contraction. Heredity 121:304–318
Bresowar GE, McGlaughlin ME (2015) Morphological and genetic discrepancies in populations of Oreocarya paradoxa and O. revealii: the impact of edaphic selection on recent diversification in the Colorado Plateau. Am J Bot 102:1647–1658
Brussard PF (1984) Geographic patterns and environmental gradients: the central-marginal model in Drosophila revisited. Annu Rev Ecol Syst 15:25–64
Bucharova A, Bossdorf O, Hölzel N, Kollmann J, Prasse R, Durka W (2019) Mix and match: regional admixture provenancing strikes a balance among different seed-sourcing strategies for ecological restoration. Conserv Genet 20:7–17
Cheng J, Kao H, Dong S (2020) Population genetic structure and gene flow of rare and endangered Tetraena mongolica Maxim. revealed by reduced representation sequencing. BMC Plant Biol 20:1–13
Chung MG (1999) Allozyme diversity in the endangered shrub Abeliophyllum distichum (Oleaceae): a monotypic Korean genus. Int J Plant Sci 160:553–559
Chung MY, Chung JM, López-Pujol J, Park SJ, Chung MG (2013) Genetic diversity in three species of Forsythia (Oleaceae) endemic to Korea: implications for population history, taxonomy, and conservation. Biochem Syst Ecol 47:80–92
Chung MY, Son S, Herrando-Moraira S, Tang CQ, Maki M, Kim YD et al (2020) Incorporating differences between genetic diversity of trees and herbaceous plants in conservation strategies. Conserv Biol 34:1142–1151
Cohen JI (2010) “A case to which no parallel exists”: the influence of Darwin’s different forms of flowers. Am J Bot 97:701–716
Cohen JI (2019) The conservation genomics of the endangered distylous gypsophile Oreocarya crassipes (Boraginaceae). Conserv Genet 20:1315–1328
Cox MP, Peterson DA, Biggs PJ (2010) SolexaQA: at-a-glance quality assessment of Illumina second-generation sequencing data. BMC Bioinformatics 11:1–6
Cruzan MB (2001) Population size and fragmentation thresholds for the maintenance of genetic diversity in the herbaceous endemic Scutellaria montana (Lamiaceae). Evolution 55:1569–1580
Cultural Heritage Administration (2006) Explore Heritage, Heritage Search. Retrieved from http://english.cha.go.kr/chaen/search/selectGeneralSearchList.do?mn=EN_02_02 (Accessed 02 May 2021)
De Vriendt L, Lemay MA, Jean M, Renaut S, Pellerin S, Joly S et al (2017) Population isolation shapes plant genetics, phenotype and germination in naturally patchy ecosystems. J Plant Ecol 10:649–659
DeWoody JA, Harder AM, Mathur S, Willoughby JR (2021) The long-standing significance of genetic diversity in conservation. Mol Ecol 30:4147–4154
Do C, Waples RS, Peel D, Macbeth GM, Tillett BJ, Ovenden JR (2014) NeEstimator v2: re-implementation of software for the estimation of contemporary effective population size (Ne) from genetic data. Mol Ecol Resour 14:209–214
Dudley N (2008) Guidelines for applying protected area management categories. IUCN, Switzerland
Earl DA, vonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361
Eckert CG, Samis KE, Lougheed SC (2008) Genetic variation across species’ geographical ranges: the central–marginal hypothesis and beyond. Mol Ecol 17:1170–1188
Edwards CE, Lindsay DL, Bailey P, Lance RF (2014) Patterns of genetic diversity in the rare Erigeron lemmoni and comparison with its more widespread congener, Erigeron arisolius (Asteraceae). Conserv Genet 15:419–428
Ellis JR, Pashley CH, Burke JM, McCauley DE (2006) High genetic diversity in a rare and endangered sunflower as compared to a common congener. Mol Ecol 15:2345–2355
Ellstrand NC, Elam DR (1993) Population genetic consequences of small population size: implications for plant conservation. Annu Rev Ecol Syst 24:217–242
Elshire RJ, Glaubitz JC, Sun Q, Poland JA, Kawamoto K, Buckler ES et al (2011) A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS ONE 6:e19379
England PR, Cornuet JM, Berthier P, Tallmon DA, Luikart G (2006) Estimating effective population size from linkage disequilibrium: severe bias in small samples. Conserv Genet 7:303–308
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Excoffier L, Lischer HE (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131:479–491
Faubet P, Waples RS, Gaggiotti OE (2007) Evaluating the performance of a multilocus Bayesian method for the estimation of migration rates. Mol Ecol 16:1149–1166
Frankham R (2010) Where are we in conservation genetics and where do we need to go? Conserv Genet 11:661–663
Franklin IR, Allendorf FW, Jamieson IG (2014) The 50/500 rule is still valid–reply to Frankham et al. Biol Conserv 176:284–285
Garcia AA, Benchimol LL, Barbosa AM, Geraldi IO, Souza CL Jr, Souza APD (2004) Comparison of RAPD, RFLP, AFLP and SSR markers for diversity studies in tropical maize inbred lines. Genet Mol Biol 27:579–588
García-Dorado A, Caballero A (2021) Neutral genetic diversity as a useful tool for conservation biology. Conserv Genet 22:541–545
Garot E, Joët T, Combes MC, Lashermes P (2019) Genetic diversity and population divergences of an indigenous tree (Coffea mauritiana) in Reunion Island: role of climatic and geographical factors. Heredity 122:833–847
Gemmill CE, Ranker TA, Ragone D, Perlman SP, Wood KR (1998) Conservation genetics of the endangered endemic Hawaiian genus Brighamia (Campanulaceae). Am J Bot 85:528–539
Ghimire B, Jeong MJ, Choi GE, Lee H, Lee KM, Lee CH et al (2015) Embryo, seed coat and pericarp development in Abeliophyllum distichum Nakai (Oleaceae): a rare and endemic plant of Korea. Korean J Plant Res 28:350–356
Ghimire B, Yum D, Kim JH, Jeong MJ (2020) The embryological insight into the relationship between Forsythia and Abeliophyllum (Forsythieae, Oleaceae). J Plant Res 133:611–623
Gitzendanner MA, Soltis PS (2000) Patterns of genetic variation in rare and widespread plant congeners. Am J Bot 87:783–792
Gomes Viana JP, Bohrer Monteiro Siqueira MV, Araujo FL, Grando C, Sanae Sujii P, Silvestre EDA et al (2018) Genomic diversity is similar between Atlantic forest restorations and natural remnants for the native tree Casearia sylvestris Sw. PLoS ONE 13:e0192165
Gonzales E, Hamrick JL (2005) Distribution of genetic diversity among disjunct populations of the rare forest understory herb, Trillium reliquum. Heredity 95:306–314
Hampe A, Petit RJ (2005) Conserving biodiversity under climate change: the rear edge matters. Ecol Lett 8:461–467
Hamrick JL, Godt MJW (1989) Allozyme diversity in plant species. In: Brown AHD, Clegg MT, Kahler AL, Weir BS (eds) Plant population genetics, breeding and genetic resources. Sinauer, Sunderland, MA, pp 43–63
Hartl DL, Clark AG (1997) Principles of population genetics, 3rd edn. Sinauer Associates Inc., Sunderland, MA
Hartvig I, So T, Changtragoon S, Tran HT, Bouamanivong S, Ogden R et al (2020) Conservation genetics of the critically endangered Siamese rosewood (Dalbergia cochinchinensis): recommendations for management and sustainable use. Conserv Genet 21:677–692
Hasbún R, González J, Iturra C, Fuentes G, Alarcón D, Ruiz E (2016) Using genome-wide SNP discovery and genotyping to reveal the main source of population differentiation in Nothofagus dombeyi (Mirb.) Oerst. in Chile. Int J Genomics 3654093:1–11
Henle K, Davies KF, Kleyer M, Margules C, Settele J (2004) Predictors of species sensitivity to fragmentation. Biodivers Conserv 13:207–251
Hodgins KA, Barrett SC (2008) Asymmetrical mating patterns and the evolution of biased style-morph ratios in a tristylous daffodil. Genet Res 90:3–15
Howes BJ, Brown JW, Gibbs HL, Herman TB, Mockford SW, Prior KA et al (2009) Directional gene flow patterns in disjunct populations of the black ratsnake (Pantheropis obsoletus) and the Blanding’s turtle (Emydoidea blandingii). Conserv Genet 10:407–417
Huey JA, Espinoza T, Hughes JM (2013) Natural and anthropogenic drivers of genetic structure and low genetic variation in the endangered freshwater cod, Maccullochella mariensis. Conserv Genet 14:997–1008
Ishihama F, Nakano C, Ueno S, Ajima M, Tsumura Y, Washitani I (2003) Seed set and gene flow patterns in an experimental population of an endangered heterostylous herb with controlled local opposite-morph density. Funct Ecol 17:680–689
Jamieson IG, Allendorf FW (2012) How does the 50/500 rule apply to MVPs? Trends Ecol Evol 27:578–584
Jombart T, Ahmed I (2011) adegenet 1.3-1: new tools for the analysis of genome-wide SNP data. Bioinformatics 27:3070–3071
Jones AT, Ovenden JR, Wang YG (2016) Improved confidence intervals for the linkage disequilibrium method for estimating effective population size. Heredity 117:217–223
Kang UC, Chang CS, Kim YS (2000) Genetic structure and conservation considerations of rare endemic Abeliophyllum distichum Nakai (Oleaceae) in Korea. J Plant Res 113:127–138
Kennedy JP, Preziosi RF, Rowntree JK, Feller IC (2020) Is the central-marginal hypothesis a general rule? Evidence from three distributions of an expanding mangrove species, Avicennia germinans (L.) L. Mol Ecol 29:704–719
Kim KJ (1999) Molecular phylogeny of Forsythia (Oleaceae) based on chloroplast DNA variation. Plant Syst Evol 218:113–123
Kim YS (2006) Conservation of plant diversity in Korea. Landsc Ecol Eng 2:163–170
Kim DK, Kim JH (2008) A new natural habitat of Abeliophyllum distichum Nakai. Korean J Pl Taxon 38:573–582 (in Korean with English abstract)
Kim DK, Kim JH (2011) Molecular phylogeny of tribe Forsythieae (Oleaceae) based on nuclear ribosomal DNA internal transcribed spacers and plastid DNA trnL-F and matK gene sequences. J Plant Res 124:339–347
Kim YS, Maunder M (1998) Plants in Peril, 24: Abeliophyllum distichum. Curtis’s Bot Mag 15:141–146
Kim KJ, Lee HL, Kim YD (2000) Phylogenetic position of Abeliophyllum (Oleaceae) based on nuclear ITS sequence data. Korean J Pl Taxon 30:235–250 (in Korean with English abstract)
Kim TH, Han MJ (1998) The reproductive system of the Korean endemic taxon, Abeliophyllum distichum Nakai, a distylous species (Oleaceae). Undergraduate thesis, Kyung Hee University, Seoul, Korea (in Korean with English abstract)
Kopelman NM, Mayzel J, Jakobsson M, Rosenberg NA, Mayrose I (2015) CLUMPAK: a program for identifying clustering modes and packaging population structure inferences across K. Mol Ecol Resour 15:1179–1191
Korea Protected Areas Forum (2009). Natural monuments and nature reserves of Korea. Retrieved from http://www.paforum.or.kr/area/area.html?cate_idx=19 (Accessed 06 Jan 2021)
Landenberger RE, Kota NL, McGraw JB (2007) Seed dispersal of the non-native invasive tree Ailanthus altissima into contrasting environments. Plant Ecol 192:55–70
Lawton JH (1993) Range, population abundance and conservation. Trends Ecol Evol 8:409–413
Lee HL, Jansen RK, Chumley TW, Kim KJ (2007) Gene relocations within chloroplast genomes of Jasminum and Menodora (Oleaceae) are due to multiple, overlapping inversions. Mol Biol Evol 24:1161–1180
Lee HY, Kim TG, Oh CH (2014) Recently augmented natural habitat of Abeliophyllum distichum Nakai in Yeoju-si, Gyunggi-do, Korea. Korean J Environ Ecol 28:62–70 (in Korean with English abstract)
Lehmkuhl JF (1984) Determining size and dispersion of minimum viable populations for land management planning and species conservation. Environ Manage 8:167–176
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25:1754–1760
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N et al (2009) The sequence alignment/map format and SAMtools. Bioinformatics 25:2078–2079
Lischer HE, Excoffier L (2012) PGDSpider: an automated data conversion tool for connecting population genetics and genomics programs. Bioinformatics 28:298–299
Luo R, Liu B, Xie Y, Li Z, Huang W, Yuan J et al (2012) SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. Gigascience 1:1–6
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. Embnet J 17:10–12
Meeus S, Honnay O, Brys R, Jacquemyn H (2012) Biased morph ratios and skewed mating success contribute to loss of genetic diversity in the distylous Pulmonaria officinalis. Ann Bot 109:227–235
Meirmans PG (2014) Nonconvergence in Bayesian estimation of migration rates. Mol Ecol Resour 14:726–733
Mussmann SM, Douglas MR, Chafin TK, Douglas ME (2019) BA3-SNPs: contemporary migration reconfigured in BayesAss for next-generation sequence data. Methods Ecol Evol 10:1808–1813
Newman BJ, Ladd P, Brundrett M, Dixon KW (2013) Effects of habitat fragmentation on plant reproductive success and population viability at the landscape and habitat scale. Biol Conserv 159:16–23
Nybom H, Bartish IV (2000) Effects of life history traits and sampling strategies on genetic diversity estimates obtained with RAPD markers in plants. Perspect Plant Ecol Evol Syst 3:93–114
Ouborg NJ, Piquot Y, Van Groenendael JM (1999) Population genetics, molecular markers and the study of dispersal in plants. J Ecol 87:551–568
Paradis E (2010) pegas: an R package for population genetics with an integrated–modular approach. Bioinformatics 26:419–420
Peakall R, Smouse PE (2006) GenAlEx 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research—an update. Bioinformatics 28:2537–2539
Petit RJ, Aguinagalde I, de Beaulieu JL, Bittkau C, Brewer S, Cheddadi R et al (2003) Glacial refugia: hotspots but not melting pots of genetic diversity. Science 300:1563–1565
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MAR, Bender D et al (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81:559–575
R Core Team (2020) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria
Rambaut A, Drummond AJ, Xie D, Baele G, Suchard MA (2018) Posterior summarization in Bayesian phylogenetics using Tracer 1.7. Syst Biol 67:901–904
Rannala B (2007) BayesAss edition 3.0 user's manual. University of California Davis (Last updated 28 Sept 2015). Retrieved from http://www.rannala.org/?page_xml:id=245 (Accessed 06 January 2021)
Reisch C, Anke A, Röhl M (2005) Molecular variation within and between ten populations of Primula farinosa (Primulaceae) along an altitudinal gradient in the northern Alps. Basic Appl Ecol 6:35–45
Rousset F (1997) Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics 145:1219–1228
Ryu TY, Yeam DY, Kim YJ, Kim SJ (1976) Studies on heterostyly incompatibility of Abeliophyllum distichum. Seoul Nat Univ Coll Agri Bull 1:113–120
Sagarin RD, Gaines SD (2002) The “abundant centre” distribution: to what extent is it a biogeographical rule? Ecol Lett 5:137–147
Samarasin P, Shuter BJ, Wright SI, Rodd FH (2017) The problem of estimating recent genetic connectivity in a changing world. Conserv Biol 31:126–135
Sax K, Abbe EC (1932) Chromosome numbers and the anatomy of the secondary xylem in the Oleaceae. J Arnold Arbor 13:37–48
Schierenbeck KA (2017) Population-level genetic variation and climate change in a biodiversity hotspot. Ann Bot 119:215–228
Schrader H (2020) How many years does the forsythia tree live and begin to bear fruit? ThePlantAide. Retrieved from https://www.theplantaide.com/articles/66923.html (Accessed 10 June 2021)
Shao JW, Wang HF, Fang SP, Conti E, Chen YJ, Zhu HM (2019) Intraspecific variation of self-incompatibility in the distylous plant Primula merrilliana. AoB Plants 11:1–10
Smith B, Kay KM (2018) Differing patterns of genetic diversity and inbreeding in two rare serpentine Monardellas in the northern Sierra Nevada. Madroño 65:10–21
Son SW, Kim YS, Kim H (2016) Abeliophyllum distichum. IUCN Red List Threatened Species. https://doi.org/10.2305/IUCN.UK.2016-1.RLTS.T13188339A13189399.en
Stephens PA, Sutherland WJ (1999) Consequences of the Allee effect for behaviour, ecology and conservation. Trends Ecol Evol 14:401–405
Stephens PA, Sutherland WJ, Freckleton RP (1999) What is the Allee effect? Oikos 87:185–190
Taylor H (1945) Cyto-taxonomy and phylogeny of the Oleaceae. Brittonia 5:337–367
Van Rossum F, Triest L (2003) Spatial genetic structure and reproductive success in fragmented and continuous populations of Primula vulgaris. Folia Geobot 38:239–254
Van Rossum F, Triest L (2006) Within-population genetic variation in the distylous Primula veris: does floral morph anisoplethy matter in fragmented habitats? Perspect Plant Ecol Evol Syst 7:263–273
Van Rossum F, Campos De Sousa S, Triest L (2004) Genetic consequences of habitat fragmentation in an agricultural landscape on the common Primula veris, and comparison with its rare congener, P.vulgaris. Conserv Genet 5:231–245
Vignal A, Milan D, SanCristobal M, Eggen A (2002) A review on SNP and other types of molecular markers and their use in animal genetics. Genet Sel Evol 34:275–305
von Takach DB, Jack C, Borevitz J, Lindenmayer DB, Banks SC (2019) Pervasive admixture between eucalypt species has consequences for conservation and assisted migration. Evol Appl 12:845–860
Vucetich JA, Waite TA (2003) Spatial patterns of demography and genetic processes across the species’ range: null hypotheses for landscape conservation genetics. Conserv Genet 4:639–645
Wallace LE (2002) Examining the effects of fragmentation on genetic variation in Platanthera leucophaea (Orchidaceae): inferences from allozyme and random amplified polymorphic DNA markers. Plant Species Biol 17:37–49
Wallander E, Albert VA (2000) Phylogeny and classification of Oleaceae based on rps16 and trnA-F sequence data. Am J Bot 87:1827–1841
Waples RS (2006) A bias correction for estimates of effective population size based on linkage disequilibrium at unlinked gene loci. Conserv Genet 7:167–184
Washitani I, Osawa R, Namai H, Niwa M (1994) Patterns of female fertility in heterostylous Primula sieboldii under severe pollinator limitation. J Ecol 82:571–579
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Wickham H, François R, Henry L, Müller K (2022). dplyr: A grammar of data manipulation. Retrieved from https://dplyr.tidyverse.org, https://github.com/tidyverse/dplyr
Wilcock C, Neiland R (2002) Pollination failure in plants: why it happens and when it matters. Trends Plant Sci 7:270–277
Willi Y, Kleunen MV, Dietrich S, Fischer M (2007) Genetic rescue persists beyond first-generation outbreeding in small populations of a rare plant. P Roy Soc B-Biol Sci 274:2357–2364
Wilson GA, Rannala B (2003) Bayesian inference of recent migration rates using multilocus genotypes. Genetics 163:1177–1191
Wright S (1978) Evolution and the genetics of populations: variability within and among natural populations, vol 4. The University of Chicago Press, Chicago
Wyatt RW, Hellwig RL (1979) Factors determining fruit set in heterostylous bluets, Houstonia caerulea (Rubiaceae). Syst Bot 4:103–114
Yoo YK, Kim KS (2001) Germination inhibitors in the seed coat of white forsythia (Abeliophyllum distichum Nakai). Hortic Environ Biotechnol 42:6–10
Young A, Boyle T, Brown T (1996) The population genetic consequences of habitat fragmentation for plants. Trends Ecol Evol 11:413–418
Acknowledgements
We would like to thank Seong-Hyun Cho and Dae-Hyun Kang for their assistance during fieldwork. We also thank the two anonymous reviewers and the handling editor for their valuable comments. This study was supported by grants from the National Institute of Biological Resources (NIBR) of the Republic of Korea (NIBR201905201) and the Hallym University Research Fund (H202104520001).
Funding
The authors were supported by grants from the National Institute of Biological Resources (NIBR) of the Republic of Korea (NIBR201905201) and the Hallym University Research Fund (H202104520001).
Author information
Authors and Affiliations
Contributions
This project was conceptualized by YDK and managed by JHL. Data collection was led by YIK and EKJ. Material preparation and labwork were organized by BYK. HGO designed the research, conducted the analyses, and wrote and revised the manuscript. MGC provided valuable inputs for the improvement of the final manuscript. The funding for this work was acquired by YDK. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lee, JH., Ong, H.G., Kim, BY. et al. Population genomics study for the conservation management of the endangered shrub Abeliophyllum distichum. Conserv Genet 23, 683–697 (2022). https://doi.org/10.1007/s10592-022-01447-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-022-01447-5