Abstract
Purpose
Phosphatidylinositide-3-kinase (PI3K) regulates proliferation and apoptosis; somatic PIK3CA-mutations may activate these processes. Aim of this study was to determine the prevalence of PIK3CA-mutations in a cohort of early stage breast cancer patients and the association to the course of disease.
Patients and methods
From an unselected cohort of 1270 breast cancer patients (PiA, Prognostic Assessment in routine application, NCT 01592825) 1123 tumours were tested for the three PIK3CA hotspot-mutations H1047R, E545K, and E542K by qPCR. Primary objectives were the prevalence of somatic PIK3CA-mutations and their association to tumour characteristics. Secondary objective was the association of PIK3CA-mutations to recurrence-free interval (RFI) and overall survival.
Results
PIK3CA-mutation rate was 26.7% (300 of 1123). PIK3CA-mutations were significantly more frequent in steroid hormone-receptor (SHR)-positive HER2-negative (31.4%), and G1 and G2 tumours (32.8%). Overall, we did not observe a significant association of PIK3CA-mutations to RFI. In SHR-positive BCs with PIK3CA-mutations, a strong trend for impaired RFI was observed (adjusted HR 1.64, 95% CI 0.958–2.807), whilst in SHR-negative BCs PIK3CA-mutations were insignificantly associated with improved RFI (adjusted HR 0.49; 95% CI 0.152–1.597). Of note, we observed a significantly detrimental prognostic impact of PIK3CA-mutations on RFI in SHR-positive, HER2-negative BCs if only aromatase inhibitors were administered as adjuvant therapy (adjusted HR 4.44, 95% CI 1.385–13.920), whilst no impact was observed in tamoxifen treated patients.
Conclusion
This cohort study speficies the overall mutation rate of PIK3CA in early breast cancer. The impact of PIK3CA-mutations on RFI and OS was heterogeneous. Our results suggest that estrogen deprivation failes to be active in case of PIK3CA-mutation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Phosphatidylinositol-3-kinase (PI3K) (gene symbol PIK3CA) intracellularly mediates different processes like promoting cell transformation, tumour initiation and proliferation, and resistance to apoptosis. Its activity is stimulated by extracellular growth factors and hormones [1]. The dysregulation of PI3K initiates activity of the serine/threonine kinase AKT in many cancer entities thereby modulating a range of downstream proteins that promote uncontrolled cellular and tumour growth [2]. Thirty years ago, the PI3K/AKT/mTOR signalling pathway was discovered to be associated with carcinogenesis and oncogenic development [3], as summarized by Arafeh and Samules [4], and to date, PIK3CA-targeted drugs are developed and validated in clinical trials [5, 6].
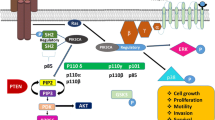
The lipid-based PI3-kinases phosphorylate the 3-hydroxyl group of phosphatidylinositol(4,5)-bisphosphate (PIP2) to phosphatidylinositol(3,4,5)-trisphosphate (PIP3) followed by the activation of AKT and downstream-signaling pathways required for cell growth and survival. PI3K activation is physiologically abrogated by the tumour suppressor phosphatase and tensin-homolog (PTEN) which converts PIP3 back to PIP2. PIP3 peptide levels depend on the competition between PI3K and PTEN. The overactivation of PI3K as well as decreased PTEN expression lead to activated and increased levels of AKT, thus pathologically promoting cell cycle progression [6].
There are three classes of PI3Ks according to their primary structures, substrate preferences and regulation: Class I (Ia, Ib), Class II and Class III. Most relevant for cellular regulation are the PI3Ks of class Ia which act as heterodimers of regulatory and catalytic subunits [7]. The catalytic subunit of the class I PI3-kinase p110α is encoded by the PIK3CA gene with a total genomic size of 86,190 base pairs in 21 exons and a final transcript of 3207 base pairs which encode a protein of 1068 amino acids. The p110α protein has five domains: an adaptor-binding-domain for linking the regulatory subunit, a Ras-binding-domain, a C2-domain for binding PIP2 and PIP3, a helical domain and a kinase domain, see https://www.uniprot.org/uniprot/P42336 [8].
Somatic mutations of the PIK3CA gene have been described in human cancers in general with a prevelance of up to 40% in primary breast cancer (http://www.sanger.ac.uk/cosmic) [9, 10]. The most frequent PIK3CA gene mutations are found in the coding sequence inducing a gain of-function of PI3K. Three hot spot non-synonymous variants represent 87% of the mutations with known clinical relevance [11] leading to amino acid substitutions: COSMIC 760 in exon 9 (17% incidence) with an E545K mutation, COSMIC 763 in exon 19 (17% incidence) affecting E545 and COSMIC 775 in exon 20 (35% incidence) altering H1047.
The three genomic aberrations are predictive for drug responsiveness, meaning that diagnostic testing can identify patients who might benefit from PI3K-targeted therapy. Recently, the PI3K inhibitor alpelisib was approved by both the FDA and EMA for patients with PIK3CA-mutated, steroid hormone receptor (SHR)-positive and HER2-negative tumours [5]. The three hotspot mutations described above can be efficiently determined by targeted sequence analysis. Nevertheless, the prognostic and predictive value of the PIK3CA mutation status as a biomarker for early breast cancer is discussed controversially for BC subgroups with respect to hormone receptor and HER2 expression [12].
In this study, we describe the prevalence of the three most common PIK3CA-mutations in subgroups of a breast cancer cohort and its association with clinical, histopathological characteristics and survival.
Material and methods
Patient and tumour characteristics
A prospective study of 1270 early breast cancer patients from five German certified breast centres (2009 to 2011) was designed in accordance with the REMARK (“Reporting Recommendations for Tumor Marker Prognostic Studies”) criteria [13] and registered as the “PiA-study” [14] (Prognostic assessment in routine application, NCT 01592825) using the following inclusion criteria: female patients, aged 18 years or older, invasive, non-metastatic BC and no secondary cancer, no limitation in tumour size, lymph node involvement, and grading or expression of estrogen receptor (ER), progesterone receptor (PgR) and human epidermal growth factor receptor 2 (HER2). Patients were diagnosed and treated (1070 with primary surgery, 200 with neoadjuvant chemotherapy, NACT) according to the annually updated German AGO Guidelines (AGO) valid at the respective times https://www.ago-online.de/leitlinien-empfehlungen/leitlinien-empfehlungen/kommission-mamma.41
In the current study, we analysed the PIK3CA gene mutation status of 1123 tumours. Median age of the patients was 60 years at time of diagnosis, with three-quarters of patients being older than 50 years, and two thirds having no lymph node involvement. Considering tumour tissue, three-quarters were well differentiated or intermediate (G1 or G2), and half of the tumours were smaller than 2 cm. The distribution of the patients' main characteristics, as well as the histopathological parameters of the analysed cohort (n = 1123), did not significantly differ from the entire PiA cohort (supplementary table S 1). The enrolment of the patients and grouping for the main analysis is shown in Fig. 1.
Endpoints and statistical analysis
We defined the prevalence of PIK3CA-mutations and the associations between PIK3CA mutation status and clinical histopathological parameters as first objective, applying binary logistic regression; recurrence free interval (RFI) and overall survival (OS) were the second objectives. RFI-related events refer to local invasive recurrence, distant recurrence, and death from breast cancer. OS included death from breast cancer, non-breast cancer causes, and unknown causes [15]. Tumour association and survival analysis were only performed for patients with one tumour. Considering the risk of multiple testing, we reduced the subgroup analyses to pre-defined, well-accepted and clinical relevant groups (e.g., tumour size, nodal status, grading, IHC-types, type of treatment etc.). The median observation time after diagnosis was 62 months (1–132). The impact of PIK3CA mutation status was calculated using Kaplan–Meier estimates; differences were described by log-rank test and multivariate analyses for selected parameters were carried out applying proportional hazard regression model and a fixed effects model (Review Manager, version 5.3). Recursive partitioning by Classification and Regression Tree (CART) analysis was performed to examine combinations of clinical and histopathological parameters to find homogenous risk groups with respect to RFI [16]. To minimize selection bias for any parameter, only significant variables were entered into the CART analysis starting with the most important prognostic parameter resulting from regression model. Patient groups were further recursively divided, considering the best split at each decision point into smaller and more homogenous groups. Unbiased parameter selection was guaranteed by following the best impact parameter (after regression model) [17].
All statistical tests were two-sided, and a p-value below 0.05 was considered to be significant. Statistical analyses were carried out using SPSS 25 (IMB, Armonk, NY, USA).
DNA extraction and hotspot mutation assay
Fresh frozen tissue (FF) (n = 813) and formalin-fixed, paraffin-embedded tumour material (FFPE) (n = 310) of the tumours were used for DNA isolation [18]. The FF-tissue was dismembrated in liquid nitrogen, and powder was used for DNA extraction using the QIAamp DNA Mini Kit [19] (Cat. No. 51304; Qiagen, Hilden, Germany). For isolation of DNA from FFPE specimens, 3–5 adjacent unstained tumour slices (5 µm) were processed using QIAamp DNA FFPE Tissue kit (Cat. No. 56404; Qiagen, Hilden, Germany). All preparations were performed according to the manufacturer’s instructions. Quality and concentration of the extracted DNA were assessed with a Tecan Infinite PRO® 200 (Tecan, Männedorf, Switzerland). A standard amount of 50 ng DNA was subjected to mutation analysis.
With respect to mutation status, we focused on the three most common hotspot mutations COSMIC C775 (H1047R), C763 (E545K) and C760 (E542K). TaqMan® Mutation Detection and reference assays were performed in duplicates (Life Technologies, Carlsbad, CA, USA) and were used for quantitative PCR (qPCR) with the StepOne Plus® Real-Time PCR System (Life Technologies) for 40 cycles with 60 °C amplification temperature.
Results
PIK3CA mutation prevalence and association with clinical and histopathological parameters
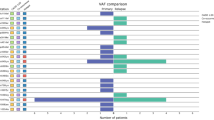
In this cohort study, 88% of the patients were tested for single nucleotide substitutions at three hotspot positions in the PIK3CA gene, and we found a mutation prevalence of 26.7% (n = 300 of 1123) considering these positions. The mutation rates for the three hot spot sites were 58% (n = 174) for COSMIC C775 (H1047R), 28% (n = 85) for C763 (E545K) and 14% (n = 43) for C760 (E542K). Co-occurrence of mutations at C775 and C763 were found in two tumours. PIK3CA-mutations were significantly more frequent in well and intermediately compared to poorly differentiated tumours (G1, OR 3.13, 95% CI 1.970–4.986; G2, OR 2.14, 95% CI 1.478–3.085). PIK3CA-mutations were significantly more often observed in steroid hormone receptor-positive than in steroid hormone receptor-negative tumours (OR 3.38, 95% CI 2.103-5.438), and in HER2-negative than in HER2-positive tumours (OR=2.25, 95% CI 1.451-3.501), respectively. We combined SHR- and HER2-status and found that the HER2-negative luminal-like IHC-type (n = 259 of 824, 31.4%) had the highest occurrence of PIK3CA-mutations (OR 4.13, 95% CI 1.753–9.712). Only 11.7% of TNBC tumours harboured a PIK3CA-mutation (15 of 128). There was no significant association with age and nodal status. The prevalences of PIK3CA-mutations in selected subgroups are reported in Table 1.
Association between PIK3CA mutation status and survival
Overall, we did not observe any significant association between presence of PIK3CA-mutations and RFI (event-free at 5 years 90.9% for mutated, 89.9% for wildtype; adjusted HR 1.19, 95% CI 0.752–1.894, Fig. 2A) and OS (alive at 5 years 88.2% for mutated, 87.2% for wildtype; adjusted HR 1.08, 95% CI 0.714–1.638, Fig. 2B), neither in univariate nor multivariate analyses (Table 2).
Survival estimates for RFI and OS stratified by detection of PIK3CA-mutations. The tables present the effective sample size for each interval (numbers at risk). A, B All patients (n = 1123), RFI (A) and OS (B). C, D Patients with SHR-positive and HER2-negative tumours, Aromatase Inhibitors (AI) treatment (n = 208), RFI (C) and OS (D). E, F Patients with SHR-negative and HER2-negative tumours (TNBC) (n = 128), RFI (E) and OS (F)
Within the cohort of patients with positive steroid hormone receptor status (irrespective of HER2 status, n = 935), 8.5% of the patients with PIK3CA-mutations experienced RFI events within 5 years of follow-up compared to 6.2% with PIK3CA-wildtype (adjusted HR 1.64, 95% CI 0.958–2.807, p = 0.071). Overall survival probability at 5 years was 88.1% and 90.5%, respectively (adjusted HR 1.37, 95% CI 0.867–2.152). We found numerically more RFI events at 5 years in patients with SHR-positive, HER2-negative and PIK3CA-mutated tumours than in patients with PIK3CA-wildtype tumours (7.9% and 6.0%, resp., Fig. S1A). More patients in this group died if their tumours were PIK3CA-mutated (11.4% and 8.5%, resp., Fig. S1B). However, the effect was not significant, neither in univariate, nor in multivariate analyses.
Of note, patients with SHR-positive HER2-negative tumours who were treated with aromatase inhibitors only (n = 208), had a significant 4.39 times higher occurrence of RFI events if they harboured a PIK3CA-mutation (n = 68) compared to those with PIK3CA-wildtype (n = 140; adjusted HR 4.39, 95% CI 1.385–13.920, p = 0.012; Fig. 2C, Table S3A) and a significantly impaired OS (adjusted HR 2.12, 95% CI 1.021–4.404, p = 0.044; Fig. 2D, Table S3B). In contrast, no association between PIK3CA mutation status and RFI or OS was observed in patients with luminal-like tumours who were treated with tamoxifen only (Fig. S1C/D, Table S3A/B).
In the hormone receptor-negative group (irrespective of HER2 status), patients with PIK3CA-mutated tumours (n = 21 of 188, 11.2%) showed numerically fewer RFI-events (3 of 21 vs 43 of 167) consistent with a higher RFI probability (84.4% and 72.9%, resp.; adjusted HR 0.49, 95% CI 0.152–1.597; Fig. S1E). In contrast, in TNBC, numerically, more patients with PIK3CA-mutations (84%) were free of RFI-events after 5 years than those with wildtype PIK3CA (71.5%; adjusted HR 0.43, 95% CI 0.103–1.822, Fig. 2E). These observations were similar after exclusion of patients without adequate (neo)adjuvant therapy. For the HER2-positive group (any SHR), we did not observe any significant impact of PIK3CA-mutations on RFI or OS (Fig. S1G/H). Interestingly, patients with SHR-negative tumours experienced a better overall survival if a PIK3CA-mutation was detected (Fig. 2F, Fig. S1F,H). The different impact of PIK3CA- mutation status on RFI in relation to steroid hormone receptor- and HER2-expression is visualized in the corresponding forest plot (Fig. S2).
To identify homogenous risk groups with regard to PIK3CA mutation status, we used a recursive partitioning procedure (Fig. 3). In node-negative, SHR-positive undifferentiated (G3) tumours, patients with a PIK3CA-mutation (n = 14) had a worse 5 year-RFI (70.5%) than those with wildtype PIK3CA (5 year RFI 96.4%, HR 11.92; 95% CI 1.724–82.461, p = 0.012). In contrast, in SHR-negative larger tumours (≥ 2 cm), patients with PIK3CA- mutations (n = 14) showed a trend to better 5 years-RFI probability (85.7%) compared to those with wildtype PIK3CA (5 years RFI 66.1%, HR 2.75; 95% CI 0.657–11.527). However, the absolute survival differences are substantial for each group and might in total be relevant for 18.9% of all patients (SHR-pos. G3 pN0, n = 91, and SHR-neg. pT2, n = 121).
Association of PIK3CA mutation status to pathologic complete response (pCR) rates
The PIK3CA mutation status was available for 120 tumour samples of TNBC patients receiving NACT, 20 of them with a PIK3CA-mutation. Due to the small numbers, only a trend was observed indicating that patients with wildtype tumours achieved pCR more often than patients with PIK3CA-mutations. No subgroup analysis was possible.
Discussion
In this study, to our knowledge we present the largest analysis of PIK3CA oncogenic mutations to date, using an unselected routine cohort of early stage breast cancer patients (n = 1123). The principal aim of our study was to evaluate the prevalence of PIK3CA-mutations and their associations with clinical and histopathological parameters and secondary the relation of a PIK3CA-mutation to clinical outcome. Since the landmark study of Samuels et al. [20], it has been known that presence of somatic PIK3CA-mutations promotes cancer progression also in breast cancer. Most previous publications have reported on heterogeneous sample sets including mixed sets of early stage and metastatic breast cancer patients from retrospective studies. Instead, our findings of a prospective well-defined homogenous cohort of early-stage breast cancer patients provide new insights to the realistic frequency of PIK3CA-mutations overall and in subgroups, as well as their association with recurrence-free interval and overall survival.
We detected an overall somatic mutation rate of 26.7% (300 of 1123 samples) when testing the three most common hot spots C775 (H1047R), C763 (E545K), and C760 (E542K) (https://www.mycancergenome.org/), which represent 87% of all mutations currently known in the PIK3CA gene [11]. Of interest, the highest frequencies (> 30%) of PIK3CA gene mutations were detected in tumours with more favourable characteristics (G1, G2, ER-positive, PgR-positive, luminal-like, HER2-negative), which is in line with most other studies and available data in the COSMIC database http://www.sanger.ac.uk/cosmic/http://www.sanger.ac.uk/cosmic/ [21]. In tumours with high risk biology (HER2-positive, TNBC) we found the lowest rate of PIK3CA-mutations (15.2% and 11.4), which is also consistent with published data [11].
These findings provoke the question why PIK3CA-mutations are more frequently detected in ER-positive disease. The current state of research postulates that PIK3CA-mutation-dependent activation of AKT phosphorylates and activates ER leading to transcriptional activity of ER in an oestradiol-independent manner and consecutively to preferential growth of ER-positive cancer [22, 23]. Thus, mutated PI3K likely promote ER-positive cancer growth and may explain the overrepresentation of PIK3CA-mutated tumours in luminal and well-differentiated breast cancer. In addition, PIK3CA-mutations are considered an early event in breast cancer development since they were detected even in small tumours as well as in non-invasive precursor lesions, like DCIS [24]. In contrast, fast growing ER-negative and undifferentiated tumours, however, may be derived from different precursor cells and independent of activating PIK3CA-mutations.
Prognostic and predictive implications
The second objective of our study was the prognostic impact of PIK3CA-mutations, and we did not find any association with recurrence free interval (RFI) or overall survival (OS) within the entire cohort of 1123 patients. We choose RFI as endpoint since we wanted to analyse the clean disease-related impact of PIK3CA-mutations. We tested an unselected and rather large cohort of early breast cancer patients, thus we assume that our data provide a realistic view, demonstrating a lack of a general impact of PIK3CA-mutations on the course of disease in breast cancer. This is in contrast to the published meta-analyses and single studies on PIK3CA-mutations that present conflicting results on its association to prognosis; studies found an association to better survival (e.g. Dumont et al., Pang et al. [25, 26]) as well as to inferior survival (Sobhani et al., Fan et al.) [12, 27]. These divergent results might presumably result from the heterogeneity of the populations that were studied with regard to sample size, subgroups, and type of treatment, so, selection bias cannot be excluded in these analyses (for review see [28]).
The published results are also inconclusive with regard to a potential predictive impact of PIK3CA-mutations. However, in our study we found a significant predictive value of PIK3CA-mutations in luminal breast cancer by observing more disease-related events in patients with PIK3CA-mutations. Most importantly, there was an association to the type of endocrine therapy: We found a significant impact of PIK3CA-mutations on the effect of adjuvant aromatase inhibitors, but no impact on the effect of adjuvant tamoxifen. This observation may be explained by PI3K-triggered estradiol-independent activation of the ER that can be observed in estradiol-deprived situations created by aromatase inhibition but might be blocked by ER-modulation through tamoxifen as postulated by Campbell and colleagues [22]. This differential therapy response has also been described for advanced BC by Ramirez-Ardila et al. [29].
It has to be acknowledged that these relations are complex and other mechanisms are involved. For example, recent findings suggest that PI3K pathway alterations might be associated with the composition of the tumour microenvironment in luminal breast cancer, including the attraction of CD8-positive T-cells [30]. Our observations are fully in line with the data of Stemke-Hale and colleagues who also did not find an association between PIK3CA-mutations and the effect of adjuvant tamoxifen [31]. However, data are again heterogeneous; some authors described resistance to tamoxifen [32, 33], whilst others found significantly improved endocrine sensitivity to tamoxifen if PIK3CA-mutations were detected [34].
The results from our observational cohort study support the finding that PIK3CA-mutations may indicate resistance to aromatase inhibitor therapy; however, prospective studies are lacking.
In patients with HER2-positive breast cancer we found no impact of PIK3CA-mutations on RFI. Similarly, a well described pooled analysis of five prospective clinical trials showed no significant impact of PIK3CA-mutation on the course of disease in patients with adequately treated HER2-positive BC, although the PIK3CA-mutated group had a significantly lower pCR rate [35]. In an uniformly treated early-stage HER2-positive Danish cohort, the PIK3CA-alterations predicted a significantly worse OS (adjusted HR 2.14), but had no significant impact on invasive disease free survival (iDFS) presumably due to the small sample size [36].
An exploratory analysis of the CLEOPATRA trial identified a subgroup of HER2-positive PIK3CA-mutated patients who were resistant to anti-HER2-therapy with trastuzumab and pertuzumab (worse OS if mutated, adjusted HR 1.48, p = 0.0025) [37]. Contradictory results may be explained by the continued activation of PI3K and an inhibitory effect on HER2 signaling [38]. Thus, in patients with HER2-positive tumours the impact of PIK3CA-mutations is not clear; at least the effects are small and not significant.
The presence of PIK3CA-mutations may have a favourable impact in early TNBC, suggested by a 2.3-times improved RFI and a 3-times improved OS, which is in line with Mosele and Takeshita, even though they worked with samples from advanced BC [39] analysing cell-free DNA [40].
To the best of our knowledge, this is the first study using CART in order to find out if combinations of variables could predict the risk of an RFI event. For nearly one fifth of the patients the gene modifications seem to have a relevant prognostic impact depending on the SHR status of the tumour. Overall, using the CART algorithms (see Fig. 3) for 121 of 1123 patients, the presence of gene alterations predict a worse prognosis in defined subgroups. Patients with PIK3CA-mutated tumours (n = 20) had lower pCR rates than wildtype tumours. This observation is similar to other studies [35].
Our real world data from a multicentre cohort adds exploratory, but valuable information, as our patients were consecutively enrolled in the daily clinical routine.
Conclusion
The real overall somatic mutation rate of PIK3CA is 26.7% when testing the three most common hot spots H1047R, E545K, and E542K in a representative cohort of patients with early breast cancer. We did not find an impact of PIK3CA-mutation on RFI and OS in general. As clinical relevant result, we demonstrated resistance of early breast cancer with somatic PIK3CA-mutation to adjuvant aromatase inhibitor therapy, suggesting tamoxifen as preferred therapy in these patients. Though only exploratory, this observation is in line with previous observations in metastatic disease. More functional studies are needed to understand the interactions and crosstalk between the activated PI3K signaling pathway and tumorigenesis.
Strength and limitations
The first intent of our study was to describe the prevalence of presumably prognostic and predictive factors including PIK3CA-mutation in the daily routine. The analysis of a prospectively collected and unselected cohort of non-metastatic breast cancer patients with a huge sample size is the strength of our PIK3CA study. Limitations may be relevant for the second endpoint of this study exploiting the prognostic and predictive impact of PIK3CA-mutations since systemic treatment was slightly heterogeneic although patients were treated in high level certified breast centres and treatment decisions were made according to national guidelines. However, it represents rather a real world situation and cannot be compared to the homogeneity of treatment that is defined by prespecified clinical trial inclusion criteria. In addition, it has to be mentioned, that during the time of enrolment the detection limit for endocrine sensitive tumours was modified on a national and international level; therefore, today we would have a higher proportion of endocrine-treated patients.
We are well aware that subgroup analyses always include the risk of type 1 error for multiple testing. We, therefore, reduced the analyses to prespecified clinical relevant and broadly accepted subgroups and used multivariate analyses to limit the risk of multiple testing.
Another limitation is that as in most published studies, we analysed only the three most frequent PIK3CA “hot spot” mutations, such that a small underrepresentation of the total number of mutations may be possible.
Data availability
The data generated in this study are available within the article and its supplementary data files. Raw data were generated and processed from the authors and are available on request to the corresponding author.
Change history
27 January 2023
Missing Open Access funding information has been added in the Funding Note.
References
Phillips J, Domingo E (2019) PIK3CA (phosphoinositide-3-kinase, catalytic, alpha polypeptide). Atlas of Genetics and Cytogenetics in Oncology and Haematology
Vivanco I, Sawyers CL (2002) The phosphatidylinositol 3-Kinase AKT pathway in human cancer. Nat Rev Cancer 2:489–501
Cantley LC (2002) The phosphoinositide 3-kinase pathway. Science 296:1655–1657
Arafeh R, Samuels Y (2019) PIK3CA in cancer: the past 30 years. Semin Cancer Biol 59:36–49
André F, Ciruelos EM, Juric D et al (2021) Alpelisib plus fulvestrant for PIK3CA-mutated, hormone receptor-positive, human epidermal growth factor receptor-2-negative advanced breast cancer: final overall survival results from SOLAR-1. Ann Oncol 32:208–217
Vitale SR, Martorana F, Stella S et al (2021) PI3K inhibition in breast cancer: identifying and overcoming different flavors of resistance. Crit Rev Oncol Hematol 162:103334
Katso R, Okkenhaug K, Ahmadi K et al (2001) Cellular function of phosphoinositide 3-kinases: implications for development, homeostasis, and cancer. Annu Rev Cell Dev Biol 17:615–675
Vanhaesebroeck B, Guillermet-Guibert J, Graupera M, Bilanges B (2010) The emerging mechanisms of isoform-specific PI3K signalling. Nat Rev Mol Cell Biol 11:329–341
Samuels Y, Velculescu VE (2004) Oncogenic mutations of PIK3CA in human cancers. Cell Cycle 3:1221–1224
Koboldt DC, Fulton R, McLellan M, Schmidt H, Kalicki-Veizer J, McMichael J, Fulton L, Dooling D, Ding L, Mardis E, Wilson R (2012) Comprehensive molecular portraits of human breast tumours. Nature 490:61–70
Martínez-Sáez O, Chic N, Pascual T et al (2020) Frequency and spectrum of PIK3CA somatic mutations in breast cancer. Breast Cancer Res 22:45
Sobhani N, Roviello G, Corona SP et al (2018) The prognostic value of PI3K mutational status in breast cancer: a meta-analysis. J Cell Biochem 119:4287–4292
McShane LM, Altman DG, Sauerbrei W et al (2005) REporting recommendations for tumour MARKer prognostic studies (REMARK). Br J Cancer 93:387–391
Hartung C, Porsch M, Stückrath K et al (2021) Identifying high-risk triple-negative breast cancer patients by molecular subtyping. Breast Care (Basel) 16:637–647
Tolaney SM, Garrett-Mayer E, White J et al (2021) Updated standardized definitions for efficacy end points (STEEP) in adjuvant breast cancer clinical trials: STEEP version 2.0. J Clin Oncol 39:2720
Breiman L, Friedman JH, Olshen RA, Stone CJ (2017) Classification and regression trees. Routledge, London
Hothorn T, Hornik K, Zeileis A (2006) Unbiased recursive partitioning: a conditional inference framework. J Comput Graph Stat 15:651–674
Bauer M, Kantelhardt EJ, Stiewe T et al (2019) Specific allelic variants of SNPs in the MDM2 and MDMX genes are associated with earlier tumor onset and progression in Caucasian breast cancer patients. Oncotarget 10:1975–1992
Bachmann HS, Jung D, Link T et al (2022) FNTB promoter polymorphisms are independent predictors of survival in patients with triple negative breast cancer. Cancers 14:468
Samuels Y, Wang Z, Bardelli A et al (2004) High frequency of mutations of the PIK3CA gene in human cancers. Science 304:554
Forbes SA, Bindal N, Bamford S et al (2011) COSMIC: mining complete cancer genomes in the catalogue of somatic mutations in cancer. Nucleic Acids Res 39:D945–D950
Campbell RA, Bhat-Nakshatri P, Patel NM et al (2001) Phosphatidylinositol 3-kinase/AKT-mediated activation of estrogen receptor alpha: a new model for anti-estrogen resistance. J Biol Chem 276:9817–9824
Dong C, Wu J, Chen Y et al (2021) Activation of PI3K/AKT/mTOR pathway causes drug resistance in breast cancer. Front Pharmacol 12:143
Dunlap J, Le C, Shukla A et al (2010) Phosphatidylinositol-3-kinase and AKT1 mutations occur early in breast carcinoma. Breast Cancer Res Treat 120:409–418
Dumont AG, Dumont SN, Trent JC (2012) The favorable impact of PIK3CA mutations on survival: an analysis of 2587 patients with breast cancer. Chin J Cancer 31:327–334
Pang B, Cheng S, Sun S-P et al (2015) Prognostic role of PIK3CA mutations and their association with hormone receptor expression in breast cancer: a meta-analysis. Sci Rep 4:1–9
Fan H, Li C, Xiang Q et al (2018) PIK3CA mutations and their response to neoadjuvant treatment in early breast cancer: a systematic review and meta-analysis. Thorac Cancer 9:571–579
Li H, Prever L, Hirsch E, Gulluni F (2021) Targeting PI3K/AKT/mTOR signaling pathway in breast cancer. Cancers 13:3517
Ramirez-Ardila DE, Helmijr JC, Look MP et al (2013) Hotspot mutations in PIK3CA associate with first-line treatment outcome for aromatase inhibitors but not for tamoxifen. Breast Cancer Res Treat 139:39–49
Sobral-Leite M, Salomon I, Opdam M et al (2019) Cancer-immune interactions in ER-positive breast cancers: PI3K pathway alterations and tumor-infiltrating lymphocytes. Breast Cancer Res 21:90
Stemke-Hale K, Gonzalez-Angulo AM, Lluch A et al (2008) An integrative genomic and proteomic analysis of PIK3CA, PTEN, and AKT mutations in breast cancer. Cancer Res 68:6084–6091
Maruyama N, Miyoshi Y, Taguchi T et al (2007) Clinicopathologic analysis of breast cancers with PIK3CA mutations in Japanese women. Clin Cancer Res 13:408–414
Beelen K, Opdam M, Severson TM et al (2014) Phosphorylated p-70S6K predicts tamoxifen resistance in postmenopausal breast cancer patients randomized between adjuvant tamoxifen versus no systemic treatment. Breast Cancer Res 16:1–9
Kalinsky K, Jacks LM, Heguy A et al (2009) PIK3CA mutation associates with improved outcome in breast cancer. Clin Cancer Res 15:5049–5059
Loibl S, Majewski I, Guarneri V et al (2016) PIK3CA mutations are associated with reduced pathological complete response rates in primary HER2-positive breast cancer: pooled analysis of 967 patients from five prospective trials investigating lapatinib and trastuzumab. Ann Oncol 27:1519–1525
Jensen JD, Knoop A, Laenkholm AV et al (2012) PIK3CA mutations, PTEN, and pHER2 expression and impact on outcome in HER2-positive early-stage breast cancer patients treated with adjuvant chemotherapy and trastuzumab. Ann Oncol 23:2034–2042
Baselga J, Cortés J, Im S-A et al (2014) Biomarker analyses in CLEOPATRA: a phase III, placebo-controlled study of pertuzumab in human epidermal growth factor receptor 2–Positive, first-line metastatic breast cancer. J Clin Oncol 32:3753–3761
Chandarlapaty S, Sawai A, Scaltriti M et al (2011) AKT inhibition relieves feedback suppression of receptor tyrosine kinase expression and activity. Cancer Cell 19:58–71
Mosele F, Stefanovska B, Lusque A et al (2020) Outcome and molecular landscape of patients with PIK3CA-mutated metastatic breast cancer. Ann Oncol 31:377–386
Takeshita T, Yamamoto Y, Yamamoto-Ibusuki M et al (2015) Prognostic role of PIK3CA mutations of cell-free DNA in early-stage triple negative breast cancer. Cancer Sci 106:1582–1589
Ditsch N, Untch M, Thill M, Müller V et al (2021) AGO Recommendations for the Diagnosis and Treatment of Patients with Early Breast Cancer: Update 2021. Breast Care 16(3) 214-227. https://doi.org/10.1159/000516419
Acknowledgements
We thank all patients participating the PiA-study and the study nurses Katrin Losse (St. Elisabeth and St. Barbara Halle), Birgit Mauder (Fürth), Andrea Meyer-Kühn (Hildesheim) for excellent documentation.
Funding
Open Access funding enabled and organized by Projekt DEAL. This work was supported by Wilhelm Roux Program of the Medical Faculty, Martin Luther University Halle-Wittenberg (FKZ 25/36), German Federal Ministry of Education and Research (e:Med FKZ: 031A429) by a grant from the Susan G. Komen Graduate Training in Disparities Research grant 16378013.
Author information
Authors and Affiliations
Contributions
EJK and MV designed the PiA study; KR, CT, and MV conceived the PIK3CA analysis; KR and MV wrote the manuscript; CH, CU, VK, TL, SP, JJ, MP, MB, FKB, EW, JB, EJK contributed substantial input to the conception and acquisition of the work; KR, KT and SK and carried out DNA mutations experiments, EJK is responsible for statistical evaluation; KR, MV, CT performed data interpretation; all authors revised the manuscript critically.
Corresponding author
Ethics declarations
Competing interest
CT reports reports support form: Martin Luther University Halle-Wittenberg, Arbeitsgemeinschaft Gynäkologische Onkologie e.V., American Diagnostica, BIOMED BMH4 - 98 - 9418, honoraria from: Amgen, AstraZeneca, Celgene, Daiichi-Sankyo, Eisai, Gilead, Lilly, MSD, NanoString, Novartis, Pfizer, Pierre Fabre, Puma, Sanofi-Aventis, Roche, Vifor, Seagen. All other authors have no relevant financial or non-financial interests to disclose.
Ethical approval
This study was performed in line with the principles of the Declaration of Helsinki. The study protocol was approved by the Ethical Board of the Medical Faculty of the Martin Luther University Halle Wittenberg.
Informed consent
All patients signed a written informed consent.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
CT reports reports support form: Martin-Luther-University Halle-Wittenberg, Arbeitsgemeinschaft Gynäkologische Onkologie e.V., Sanofi-Aventis, American Diagnostica, BIOMED BMH4 - 98 - 9418, honoraria from: Amgen, AstraZeneca, Celgene, Daiichi-Sankyo, Eisai, Gilead, Lilly, MSD, NanoString, Novartis, Pfizer, Pierre Fabre, Puma, Sanofi-Aventis, Roche, Vifor, Seagen All other authors have no relevant financial or non-financial interests to disclose.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Reinhardt, K., Stückrath, K., Hartung, C. et al. PIK3CA-mutations in breast cancer. Breast Cancer Res Treat 196, 483–493 (2022). https://doi.org/10.1007/s10549-022-06637-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-022-06637-w