Abstract
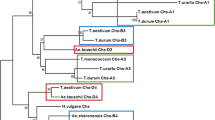
As result of a close evolutionary relationship between Triticeae B, G, and S genomes, the exchange of genetic material between them is possible and may be beneficial for broadening the genetic diversity of cultivated bread wheat. However, the extent to which regulatory networks are conserved remains poorly researched. Here, the structural organization and transcriptional activity of the B, S, and G genome copies of a gene encoding flavonoid biosynthesis enzyme chalcone-flavanone isomerase (CHI) were explored using introgression lines which differ from the wild type by carrying a non-bread wheat Chi-1 gene. Chi-S1, Chi-G1, and Chi-B1 all mapped to a comparable region of chromosomes 5S, 5G, and 5B, respectively. Nucleotide sequences of Aegilops speltoides Chi-S1 and Triticum timopheevii Chi-G1 were determined and compared with T. aestivum Chi-B1 sequences. The enzymes encoded by these three genes shared the same predicted tertiary structure and active sites. However, the replacement of Chi-B1 by Chi-S1 or Chi-G1 in a wheat background resulted in a significant decrease in the global amount of the Chi-1 transcript present in the seedling shoot indicating divergence in regulation of expression of the orthologous Chi-1 genes among Triticeae ssp.
Similar content being viewed by others
Abbreviations
- CHI:
-
chalcone-flavanone isomerase
- Chi :
-
gene encoding chalcone-flavanone isomerase
- Ka :
-
number of non-synonymous substitutions
- Ks :
-
number of synonymous substitutions
- RT-qPCR:
-
reverse transcription quantitative polymerase chain reaction
- Ubc :
-
gene encoding ubiquitin
References
Adonina, I.G.: Kharakteristika satellitnyh povtorov vidov Aegilops L. sekcii Sitopsis i ikh ispol'zovanie v kachestve molekulyarnyh markerov. [Characterization of satellite repeats of Aegilops L. Sitopsis section and their application as molecular markers.] - Dissertation, Institute of Cytology and Genetics SB RAS, Novosibirsk 2007. [In Russ.]
Adonina, I.G., Petrash, N.V., Timonova, E.M., Khristov, Y.A., Salina, E.A.: Construction and study of leaf rust-resistant common wheat lines with translocations of Aegilops speltoides Tausch. genetic material. — Russ. J. Genet. 48: 404–409, 2012.
Arnold, K., Bordoli, L., Kopp, J., Schwede, T.: The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. — Bioinformatics 22: 195–201, 2006.
Chalker-Scott, L.: Environmental significance of anthocyanins in plant stress responses. — Photochem. Photobiol. 70: 1–9, 1999.
Corpet, F.; Multiple sequence alignment with hierarchical clustering. — Nucl. Acids Res. 16: 10881–10890, 1988.
Dobrovolskaya, O., Boeuf, C., Salse, J., Pont, C., Sourdille, P., Bernard, M., Salina, E.: Microsatellite mapping of Ae. speltoides and map-based comparative analysis of the S, G, and B genomes of Triticeae species. — Theor. appl. Genet. 123: 1145–1157, 2011.
Dorofeev, V.F., Korovina, O.N. (ed.): Kul'turnaya Flora SSSR. [Flora of Cultivated Plants.] - Kolos Press, Leningrad 1979. [In Russ.]
Druka, A., Kudrna, D., Rostoks, N., Brueggeman, R., Von Wettstein, D., Kleinhofs, A.: Chalcone isomerase gene from rice (Oryza sativa) and barley (Hordeum vulgare): physical, genetic and mutation mapping. — Gene 302: 171–178, 2003.
Feldman, M.: The origin of cultivated wheat. - In: Benjean, A.P., Angus, W.J. (ed.): The Wheat Book: a History of Wheat Breeding. Pp. 3–56. Lavoisier Publishing, Paris 2001.
Goncharov, N.P.: Sravnitel'naja Genetika Pshenic i ikh Sorodichej. [Comparative genetics of wheats and their related species.] - Siberian Un-ty Press, Novosibirsk 2002. [In Russ.]
Grotewold, E. (ed.): The Science of Flavonoids. - Springer, New York 2008.
Gustafson, J.P, Sears, E.R.: An effective wheat gene manipulation system: problems and uses. - In: J. Janick (ed.): Plant Breeding Reviews. Vol. 11. Pp. 255–234. John Willey & Sons, New York, 1993.
Himi, E., Nisar, A., Noda, K.: Colour genes (R and Rc) for grain and coleoptile upregulate flavonoid biosynthesis genes in wheat. — Genome 48: 747–754, 2005.
Jez, J.M., Bowman, M.E., Dixon, R.A., Noel, J.P.: Structure and mechanism of the evolutionarily unique plant enzyme chalcone isomerase. — Natur. Struct. Biol. 7: 786–791, 2000.
Khlestkina, E.K.: The adaptive role of flavonoids: emphasis on cereals. — Cereal Res. Commun. 41: 185–198, 2013.
Khlestkina, E.K., Salina, E.A.: Genome-specific markers of tetraploid wheats and their putative diploid progenitor species. — Plant Breed. 120: 227–232, 2001.
Khlestkina, E.K., Röder, M.S., Salina, E.A.: Relationship between homoeologous regulatory and structural genes in allopolyploid genome - a case study in bread wheat. — BMC Plant Biol. 8: 88, 2008.
Khlestkina, E.K., Shoeva, O.Y.: Intron loss in the chalconeflavanone isomerase gene of rye. — Mol. Breed. 33: 953–959, 2014.
Khlestkina, E.K., Tereshchenko, O.Y., Salina, E.A.: Anthocyanin biosynthesis genes location and expression in wheat-rye hybrids. — Mol. Genet. Genomics 282: 475–485, 2009.
Khlestkina, E.K., Tereshchenko, O.Yu., Salina, E.A.: Flavonoid biosynthesis genes in wheat and wheat-alien hybrids: studies into gene regulation in plants with complex genomes. - In: Mothersill, C.E., Korogodina, V., Seymour, C.B. (ed.): Radiobiology and Environmental Security. Pp. 31–41. Springer, Dordrecht 2012.
Kilian, B., Özkan, H., Deusch, O., Effgen, S., Brandolini, A., Kohl, J., Martin, W., Salamini, F.: Independent wheat B and G genome origins in outcrossing Aegilops progenitor haplotypes. — Mol. Biol. Evol. 24: 217–227, 2007.
Kimber, G.: A reassessment of the origin of the polyploid wheats. — Genetics 78: 487–492, 1974.
Kosambi, D.D.: The estimation of map distances from recombination values. — Ann. Eugenet. 12: 172–175, 1944.
Lander, E.S., Green, P., Abrahamson, J., Barlow, A., Daly, M.J., Lincoln, S.E., Newburg, I.: MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. — Genomics 1: 174–181, 1987.
Leonova, I.N., Röder, M.S., Budashkina, E.B., Kalinina, N.P., Salina, E.A.: Molecular analysis of leaf rust resistant introgression lines obtained by crossing of hexaploid wheat Triticum aestivum with tetraploid wheat Triticum timopheevii. — Russ. J. Genet. 38: 1397–1403, 2002.
Li, W.L., Faris, J.D., Chittoor, J.M., Leach, J.E., Hulbert, S., Liu, D.J., Chen, P.D., Gill, B.S.: Genomic mapping of defense response genes in wheat. — Theor. appl. Genet. 98: 226–233, 1999.
McIntosh, R.A., Yamazaki, Y., Dubcovsky, J., Rogers, J., Morris, C., Appels, R., Xia, X.C. (ed.): Catalogue of Gene Symbols for Wheat. - IWGS, Yokohama 2013.
Mori, N., Liu, Y.-G., Tsunewaki, K.: Wheat phylogeny determined by RFLP analysis of nuclear DNA. 2. Wild tetraploid wheats. — Theor. appl. Genet. 90: 129–134, 1995.
Nei, M., Gojobori, T.: Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. — Mol. Biol. Evol. 3: 418–426, 1986.
Plaschke, J., Ganal, M.W., Röder, M.S.: Detection of genetic diversity in closely related bread wheat using microsatellite markers. — Theor. appl. Genet. 91: 1001–1007, 1995.
Schneider, A., Molnar, I., Molnar-Lang, M.: Utilization of Aegilops (goatgrass) species to widen the genetic diversity of cultivated wheat. — Euphytica 163: 1–19, 2008.
Shoeva, O.Y., Khlestkina, E.K., Berges, H., Salina, E.A.: The homoeologous genes encoding chalcone-flavanone isomerase in Triticum aestivum L.: structural characterization and expression in different parts of wheat plant. — Gene 538: 334–341, 2014.
Solovyev, V.V.: Statistical approaches in eukaryotic gene prediction. - In: Balding, D., Cannings, C., Bishop, M. (ed.): Handbook of Statistical Genetics. 3rd Ed. Pp. 97–159. Wiley-Interscience, New York 2007.
Tamura, K., Peterson, D., Peterson, N., Stecher, G., Nei, M., Kumar, S.: MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. — Mol. Biol. Evol. 28: 2731–2739, 2011.
Timonova, E.M., Leonova, I.N., Röder, M.S., Salina, E.: Marker-assisted development and characterization of a set of Triticum aestivum lines carrying different introgressions from the T. timopheevii genome. — Mol. Breed. 31: 123–136, 2013.
Author information
Authors and Affiliations
Corresponding author
Additional information
Acknowledgements: This study was partially supported by the State Budget Programme (Project No VI.53.1.5. = 0324-2015-0005), the Siberian Branch of the Russian Academy of Science (Integration project SBRAS/NAS Belarus No 22) and RFBR (Grant No 16-34-60052).We thank Ms. Galina Generalova for technical assistance and www.smartenglish.co.uk for linguistic advice in the preparation of the first manuscript draft.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Shoeva, O.Y., Dobrovolskaya, O.B., Leonova, I.N. et al. The B-, G- and S-genomic Chi genes in family Triticeae . Biol Plant 60, 279–284 (2016). https://doi.org/10.1007/s10535-016-0595-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10535-016-0595-5