Abstract
Three extremophile bacterial strains (BBCOL-009, BBCOL-014 and BBCOL-015), capable of degrading high concentrations of perchlorate at a range of pH (6.5 to 10.0), were isolated from Colombian Caribbean Coast sediments. Morphological features included Gram negative strain bacilli with sizes averaged of 1.75 × 0.95, 2.32 × 0.65 and 3.08 × 0.70 μm, respectively. The reported strains tolerate a wide range of pH (6.5 to 10.0); concentrations of NaCl (3.5 to 7.5% w/v) and KClO4− (250 to 10000 mg/L), reduction of KClO4− from 10 to 25%. LB broth with NaCl (3.5–30% w/v) and KClO4ˉ (250-10000 mg/L) were used in independent trials to evaluate susceptibility to salinity and perchlorate, respectively. Isolates increased their biomass at 7.5 % (w/v) NaCl with optimal development at 3.5 % NaCl. Subsequently, ClO4ˉ reduction was assessed using LB medium with 3.5% NaCl and 10000 mg/L ClO4ˉ. BBCOL-009, BBCOL-014 and BBCOL-015 achieved 10%, 17%, and 25% reduction of ClO4ˉ, respectively. The 16 S rRNA gene sequence grouped them as Bacillus flexus T6186-2, Bacillus marisflavi TF-11 (T), and Bacillus vietnamensis 15 − 1 (T) respectively, with < 97.5% homology. In addition, antimicrobial resistance to ertapenem, vancomycine, amoxicillin clavulanate, penicillin, and erythromycin was present in all the isolates, indicating their high adaptability to stressful environments. The isolated strains from marine sediments in Cartagena Bay, Colombia are suitable candidates to reduce perchlorate contamination in different environments. Although the primary focus of the study of perchlorate-reducing and resistant bacteria is in the ecological and agricultural realms, from an astrobiological perspective, perchlorate-resistant bacteria serve as models for astrobiological investigations.
Similar content being viewed by others
Introduction
The advancement of technology and industry has produced complex and toxic waste, leading to increased disposal into water, soil, and air, posing health and environmental risks (Gavrilescu et al. 2015). Conventional methods to tackle contamination are ineffective, expensive, and may exacerbate the issue. Hence, there’s a rising interest in biological remediation methods for their cost-effectiveness, high efficacy, and environmentally friendly (Liu et al. 2015; Azubuike et al. 2016).
Bioremediation utilizes microorganisms such as bacteria, archaea, fungi, and plants to absorb, transform, and degrade pollutants in soils, sediments, water, and air (Fenical 1993; Zolkefli et al. 2020; Belal et al. 2020). By immobilizing or altering the chemical structure of pollutants, bioremediation can lead to their partial degradation, mineralization, or transformation. Given the widespread contamination from human activities, pollutants are pervasive in various habitats, including extreme environments (Azubuike et al. 2016).
Microorganisms known as extremophiles have evolved specialized traits enabling them to thrive in harsh conditions, making them promising for biotechnological applications (Le Borgne et al. 2008). Marine microbial consortiums excel in diverse environmental variables like salinity, pH, and temperature. Bacteria from marine environments express genes for survival in saline conditions and have shown resilience in saturated matrices (Dalmaso et al. 2015). They also produce unique metabolites, enhancing their potential for bioremediation efforts amidst toxic pollutants (Bertel-Sevilla et al. 2020; Lee et al. 2021).
Some autochthonous extremophile strains generate biohydrogen as a substrate to treat organic waste to survive and use the macromolecules present in polluted marine environments as a carbon source (Oguntoyinbo 2007; Lee et al. 2009). In fact, halophytes strains are examples of extremophile microorganisms found in marine environments, with a remarkable ability to withstand harsh conditions and decontaminate the environment. Marine sediments are characterized by being the habitat of many native species and having the bioactivity of degrading xenobiotic substrates (Acevedo-Barrios et al. 2019, 2022), due to their resistance and efficient increase of biomass under extreme conditions, e.g., high salt concentrations (Cang et al. 2004; Van Ginkel et al. 2008; Srinivasan and Viraraghavan 2009). Hence, halophyte bacterial bioremediation will contribute to new alternatives for cleaning stressed environments.
The recent identification of perchlorate’s widespread occurrence in the environment, including its detection on Mars, the Earth’s moon, and in meteorites, as well as its involvement in regulating sulfidogenesis in oil reservoirs, has reignited interest in this unique ion and its microbiological implications (Carlström et al. 2016; Acevedo-Barrios and Olivero-Verbel 2021). Perchlorate, even at minute concentrations, poses significant health risks due to its toxic effects on the human thyroid gland (Shih et al. 2024). While current removal techniques like ion exchange and biological reduction show promise, their efficacy may be limited when applied individually (Shang et al. 2018; Xie et al. 2018; Fang and Naidu 2023). Integration of physico-chemical and biological processes appears essential for achieving comprehensive perchlorate decontamination (Shang et al. 2018; Xie et al. 2018). Extremophile-based bioremediation, particularly in marine environments, holds promise for perchlorate removal, as evidenced by studies isolating microorganisms capable of reducing perchlorate in diverse marine sediments (Acevedo-Barrios et al. 2023; Dong et al. 2022). For instance, in a study conducted by Acevedo-Barrios et al. (2019), marine sediment samples from different locations in the Colombian Caribbean were found to harbor various species of bacteria, such as members of the Bacillus genus, with a high potential for perchlorate reduction.
Bacillus spp. are Gram-positive bacteria widely recognized for their adaptability and resilience. They encompass over 222 species and exhibit characteristic features such as the production of spores, which enable them to endure adverse conditions (Garrity 2001; Miranda et al. 2008; Layton et al. 2011). These bacteria are commonly found in diverse environments, including deserts, marine ecosystems, and other harsh habitats, owing to their high biochemical activity and metabolic efficiency (Miranda et al. 2008; Elisashvili et al. 2019). Bacillus spp. have been isolated from various sources such as rice paddy soils (Beneduzi et al. 2008; Sriariyanun et al. 2016), pesticide-contaminated soils (Marín and Jaramillo 2015), perchlorate-contaminated soils (Acevedo-Barrios et al. 2016, 2019, 2022; Acevedo-Barrios and Olivero-Verbel 2021), hydrocarbons and their derivatives, liquid effluents from industries (Oyetibo et al. 2017; Masika et al. 2020), hypersaline mats, alkaline environments, i.e. food, sea water, or waste-water (Guadie et al. 2017; Durval et al. 2019; Mohapatra et al. 2019), and the intestinal tracks of insects and mammals (Hong et al. 2009). Furthermore, some Bacillus species have been shown to be capable of degrading some pollutants, including polycyclic aromatic hydrocarbons, long-chain alkanes (C10 to C32), and perchlorate (Feitkenhauer et al. 2003; Acevedo-Barrios et al. 2019). This versatility suggests that Bacillus spp. could serve as a basis for biotechnological methods aimed at mitigating environmental pollution.
The Bay of Cartagena, Colombia, is a tropical estuary in the Caribbean with an important biodiversity, but also historically affected by various sources of contamination (Romero-Murillo et al. 2023). This makes it an interesting area for the prospection of microorganisms for biotechnological applications. Our hypothesis lies in the reduction of perchlorate capacity in varied environmental conditions, of Bacillus genus strains isolated from the Colombian Caribbean.
Therefore, the objectives were to isolate and characterize halotolerant bacteria and evaluate their susceptibility and ability to reduce perchlorate in several pH and salinity conditions. This is the first study carried out in Bahía Cartagena, Colombian Caribbean; and it is part of our efforts to contribute to the knowledge of Colombian biodiversity with biotechnological potential. These bacteria of the genus Bacillus present suitable properties for possible biotechnological applications and constitute the basis for expanding our knowledge of salt-tolerant bacteria that can reduce perchlorate.
Materials and methods
Study area and sample collection
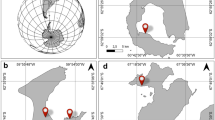
Bacterial strains of Bacillus spp. BBCOL-009, BBCOL-014 and BBCOL-015 were isolated from sediments of Cartagena Bay (10°25 × 30″N, 75°32 × 25″W), Northern Colombia (Fig. 1). Surface marine sediments were collected 100 m away from the seashore using a Van Veen grab sampler. Once gathered, the material was placed in sterile bags and transported on ice to the microbiology laboratory of the University of Cartagena, for processing.
Strains isolation and culture conditions
The isolation, purification, and conservation of the bacteria were carried out following the recommendations of Shimkets and Raffie (1990). Fluconazole (0.25 mg/ml) was used to treat pellet samples for three hours prior to inoculation into isolation media. Subsequently, the samples were incubated on NaST21CX agar. Incubating for 15–30 days at 30 °C. Media preparation was performed following the recommendations of Gaspari et al. (2005). Subsequently, individual colonies were taken and spread on Luria–Bertani (LB) agar medium for purification and preservation. Isolated colonies were plated on LB agar until pure cultures were obtained. Three bacterial strains were selected according to variations in the colonies. Later, the colonies were preserved and stored in glycerol (10% w/v) stock at − 80 °C until analysis.
Molecular identification
DNA extraction and PCR amplification of 16 S rDNA
For the DNA extraction, bacterial isolates were grown on LB medium at 30 °C overnight. The cultures underwent centrifugation at 10000 × g for 2 min, and the resulting supernatant was discarded. Genomic DNA extraction was carried out utilizing the QIAamp® DNA Mini Kit (Qiagen, CA, USA) according to the manufacturer’s instructions. To amplify the 16 S rRNA gene, the primer pairs PF (5′-AGAGTTTGATCCTGGCTCAG-3′) and 1492R (5′-ACCTTGTTACGACTT-3′) were used (Wu et al. 2005; Iizuka et al. 2006). The polymerase chain reaction (PCR) was conducted using a Veriti 96-Well Thermal Cycler (Applied Biosystems, Foster City, Ca) thermocycler. The PCR mixtures consisted of 1X AmpliTaq Gold® 360 Master Mix (Applied Biosystems), 0.4 µm of each primer and ~ 100 ng of template in a total reaction volume of 25 µL. The PCR reaction was performed with a hot start of 95 °C for 10 min, followed by 25 cycles of denaturation at 94 °C for 1 min, primer annealing at 43 °C for 1 min and primer extension at 72 °C for 1.5 min, followed by a final extension at 72 °C for 5.5 min. Amplified PCR products of the 16 S ribosomal gene were separated on 1.2% (w/v) agarose gels stained with ethidium bromide (10 mg/ml) and analyzed using a gel documentation system (IngGenius 3 System - Syngene) (Acevedo-Barrios et al. 2019; Bertel-Sevilla et al. 2020).
16 S rDNA sequencing and phylogenetic analysis
PCR products were purified with the QIAquick PCR purification kit (Qiagen, CA, USA), following the standard protocol provided by the manufacturer. Subsequently, automated DNA sequencing was performed by the National Center for Genomic Sequencing-CNSG (Medellin-Colombia) using PF and 1492R primers. The sequence reads obtained were edited and assembled using the CAP3 software (Huang and Madan 1999). The sequence was submitted to GenBank to search for similar sequences with the EzTaxon-e server (http://www.ezbiocloud.net/eztaxon; Kim et al. (2012). Sequence alignments were conducted using the CLUSTAL_W algorithm of MEGA 6 (Tamura et al. 2013), and phylogenetic analysis of all the related 16 S rRNA gene sequences was performed using MEGA 6. Distances were calculated by means of Kimura correction in a pair-wise deletion manner (Kimura 1980). Phylogenetic analyses were inferred using the Neighbour-joining (NJ) (Saitou and Nei 1987), maximum-likelihood (ML) (Felsenstein 1981), minimum-evolution (ME) and maximum-parsimony (MP) (Fitch 1971) methods in MEGA version 6.0 (Tamura et al. 2013). The tree topology was assessed by bootstrap analyses (Felsenstein 1985) based on 1000 resamplings. The sequence of Lysinibacillus tabacifolii K3514T with the GenBank accession number JQ754706 was selected as the outgroup to root the phylogenetic tree. The GenBank/EMBL/DDBJ accession numbers for the 16 S rRNA gene sequences of strains BBCOL-009, BBCOL-014 and BBCOL-015 are KU878946, KU878947 and KU878948, respectively.
Morphology
In order to isolate and describe the morphology of strains, incubation in a VY/2 medium was made, with a pH range of 7 ± 0.2 (agar 1.5%, yeast extract 0.1%, Casitone 0.3%, CaCl2·2H2O 0.1%,). Growth and morphology were observed with an optical microscope (Olympus BX41). Strains were scraped onto glass slides and stained for identification. The analyzed characteristics of the isolates included colony morphology, staining, spores if any and cell shape. Gram staining was used for microscopic description, according to Koneman et al. (2006) and Breed et al.(1957). To identify the isolated strains, scanning electron microscopy (SEM) was used (Alonso et al. 2009). Growth in LB at several pH (4.0 to 12.0 at 0.5 pH unit intervals) was determined in LB using buffer systems (Zhang et al. 2009).
Biochemical characterization
Biochemical features were identified using the BBL Crystal™ Kit according to manufacturer instructions (Ashour et al. 2011). Catalase and oxidase activities were detected by bubble production, using hydrogen peroxide solution at 3% (v/v) and the oxidation of Kovac reagent, respectively. Red metile and Voges–Proskauer tests were performed by the conventional battery test according to Winn et al. (2008) and Boone et al. (2005).
Evaluation of antimicrobial resistance
The Kirby- Bauer disc diffusion method was developed to test the antibiotic sensitivity of the studied strains (Biemer 1973; Murray et al. 1982), using as inhibition substances: ertapenem (E), vancomycine (VA), amoxicillin clavulanate (AMC), penicillin (P), erythromycin (EO) (BBLTM Sensi-DiscTMSusceptibility Test Discs – BD). Briefly, swabs of sterile cotton-tipped were used to transfer the strain to Mueller-Hinton agar plates to produce pure bacterial colonies. Antibiotic discs were put on the plate after the inoculum was dried and subsequently incubated 24–48 h, at 30°. The inhibition of bacterial growth was measured to the nearest millimeter, and their diameters as a measure of the susceptibility of the isolated strain, according to Reller et al. (2009).
Sodium chloride susceptibility assay
To test hypersaline bacterial growth, the strains were inoculated into LB broth in the presence of NaCl (3.5%, 5.0%, 7.5%, 15% and 30% w/v) by triplicates. The experiments used cell suspension with optical density (OD) = 0.6. The turbidity was recorded after 24 h incubation at 37 °C (Acevedo-Barrios et al. 2016, 2019, 2022, 2023).
Perchlorate susceptibility assay
In order to identify perchlorate susceptibility, each isolated strain was inoculated in 10 µL of LB broth with concentrations of 250, 500, 750, 1000, 2500, 5000 and 10000 mg/L of perchlorate by triplicates. After 24 h and 37 °C incubation, ClO4ˉ concentrations, cell viability and purity of strains were confirmed (Acevedo-Barrios et al. 2016, 2019, 2022, 2023).
Evaluation of perchlorate reduction by isolates
The experiments used a concentration of 10000 mg/L ClO4ˉ in LB medium with 3.5% NaCl, following the inoculation and incubation procedures of susceptibility tests. The final concentration of ClO4ˉ was measured with a selective perchlorate electrode (Thermo Fisher) (Acevedo-Barrios et al. 2016, 2019, 2022, 2023).
Results and discussion
Molecular Identification
Regarding the molecular identification of BBCOL-009, BBCOL-014 and BBCOL-015, there were sequenced of 1352, 1360 and 1361 nt of 16 S rDNA, respectively, representing an average of 90% of the Escherichia coli 16 S rRNA sequence. The phylogenetic analysis of the 16 S rRNA gene sequence revealed that the isolated bacteria belonged to the genus Bacillus spp. within the class Bacilli. The consensus tree illustrating this placement is depicted in Fig. 2. Additionally, the sequence similarities observed with closely related organisms align with the results obtained from the EzTaxon-e server analysis. Almost complete 16 S rRNA gene sequences of three isolated strains were deposited in GenBank (accession numbers: KU878946- KU878948).
Neighbour-joining tree showing the phylogenetic positions of strains BBCOL-009, BBCOL-014 and BBCOL-015 with other closely related members based on 16 S rRNA gene sequences available from the EMBL database (accession numbers are given in parentheses). The topology of the entire tree was conserved in all trees using different algorithms. The bootstrap values, expressed as percentages at the branching points, indicate the confidence level of the tree branches and were calculated based on 1000 replications. Bar 0.01 nucleotide substitutions per nucleotide position
EzTaxon-e server search analysis revealed that the strain BBCOL-009 is closely related to B. flexus T6186-2 (99.9%, 16 S rRNA gene sequence similarity), B. paraflexus RC2(T) (99.2%), B. megaterium NBRC 15308 = ATCC 14581(T) (99.1%) and other Bacilli (< 96.6%) respectively. The sequence similarities between strain BBCOL-009 and B. flexus, as determined by various clustering algorithms (99% in NJ tree, 99% in ME tree, and 99% in ML tree), in addition to the results from the EzTaxon-e server analysis, consistently indicated that B. flexus is the closest relative to strain BBCOL-009.
Strain BBCOL-014 shared highest sequence similarity with B. marisflavi TF-11(T), B. oryzaecorticis R1(T) and B. vietnamensis 15 − 1(T) with 100, 98.7 and 98.4% respectively, and nucleotide differences of 0, 14 and 21 nucleotides respectively. In the phylogenetic tree based on the neighbor-joining algorithm, strain BBCOL-014 joined the cluster comprising B. marisflavi with a bootstrap confidence value of 99% (Fig. 2), with which it shares the highest 16 S rRNA gene sequence similarity. The affiliation of strain BBCOL-014 and its closest Neighbour was also supported by the maximum-parsimony and maximum-likelihood algorithms with above 99% bootstrap values. Comparative 16 S rRNA gene sequencing revealed that strain BBCOL-015 was phylogenetically related to B. vietnamensis 15 − 1 (T) (98.4% sequence similarity), B. oryzaecorticis R1 (T) (98%), Bacillus aquimaris TF-12 (T) (98%), and B. marisflavi TF-11 (T) (97.5%). Strain BBCOL-015 demonstrated relatively low levels of 16 S rRNA gene sequence similarity compared to other species within the genus Bacillus. It formed a clade with B. vietnamensis, supported by a bootstrap value of 90% (Fig. 2).
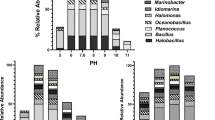
The results revealed that Bacillus spp. was the predominant group among the halotolerant bacterial communities in the sediment samples obtained from Cartagena Bay. This finding aligns with previous studies that have identified moderately halophilic bacteria belonging to the genus Bacillus in marine environments and related habitats. Examples include B. marisflavi and B. aquimaris isolated from seawater (Yoon et al. 2003), Bacillus seohaeanensis from a solar saltern (Lee et al. 2006), and B. vietnamensis from Vietnamese fish sauce (Noguchi et al. 2004).
Microscopic and biochemical characterization
The morphologic characteristics of the colonies were similar (Fig. 3), presenting a bacterial size average for 1.75 × 0.95 μm, 2.32 × 0.65 μm and 3.08 × 0.70 μm of BBCOL-009, BBCOL-014 and BBCOL-015 respectively. These dimensions are similar to the bacterial Bacillus spp. strains described by Priest et al. (1988), Noguchi et al. (2004), and Istock and Graumann (2008). In contrast, Yoon et al. (2003) described a 1500 μm Bacillus spp. length. This disparity could be explained by differences in measurement techniques, with scanning electron microscopy (SEM) being one of the most reliable (Reimer 2000).
Strains grew optimally at 37 °C ± 2, pH 7.5 ± 1 and in the absence of NaCl concentrations. All strains had a positive catalase and a negative oxidase activity, except for BBCOL-015, which had a positive oxidase. The three strains metabolized arabinose, sucrose, melibiose, rhamnose, and galactose, while the utilization of mannose, adonitol and inositol was negative. The metabolism of mannitol was positive, except for BBCOL-014. All of the strains had absence of reactions with inositol, sorbitol, indole, ONPG, ornithine, H2S production, Voges Proskauer’s, and methyl red. Each strain reduced nitrates. The biochemical characteristics of the strains were 97% compatible with Bacillus spp., sustained by the molecular results. The biochemical responses of the three strains are shown in Table 1.
Antimicrobial resistance test
The responses of the three strains to the antimicrobial test substances are shown in Table 2. All the evaluated strains presented resistance to Erythromycin. This finding suggests the presence of the erm gene (erythromycin ribosome methylation), which grants the strain the ability to produce methylase, which modifies their 23 S sRNA, as well as mef (provides resistance to 14 and 15 carbon macrolides by expulsion pump) (Davies and Davies 2010). However, this resistance has been reported mainly in grampositive cocci (Yang et al. 2024; Weisblum 1995), nonetheless, genes are acquired through the transference of transposons, probably by genetically modified microorganisms in the environment, previously exposed to antibiotics.
In addition, the strains revealed resistance to amoxicillin clavunate, vancomycine, penicillin, and ertapenem (Table 1). This behavior has been reported by Zhang et al. (2014), indicating the high capacity to activate metabolic pathways that enable the bacteria to survive in hostile conditions.
The fact that our strains were capable of resisting the aforementioned molecules indicates the property of this microorganisms to degrade pharmaceuticals and personal care products (PPCPs), considered as concerning emerging pollutants, negatively affecting aquatic ecosystems since there are no clear analyses that regulate or quantify the concentrations allowed in these environments, as well as their impact on the microbiota, flora, and fauna, is not clearly established (Liu and Wong 2013), therefore, they can be considered as potential bio-inputs in the area of biotechnology and bacterial bioremediation of emerging pollutants (e.g. other drugs like analgesics, β-blockers, preservatives, UV Filters, among others), due to structural homology. Although further studies must be done to prove the potential degradation capacity of the studied strains, the results shown in this study show the first step to consider choosing the bacterial strains, as an input for biotechnology.
Evaluation of tolerance to perchlorate, NaCl, and pH variations
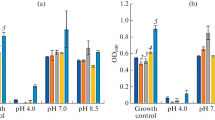
This study showed the capacity of the isolated strains BBCOL-009, BBCOL-014 and BBCOL-015 to grow at concentrations of 7.5% w/v and above of NaCl (Table 3), confirming them as moderate halotolerant species (Albuquerque et al. 2008; Zhang et al. 2009; Lei et al. 2014; Bahamdain et al. 2015). In addition to NaCl, these strains were capable to grow and tolerate KClO4− from 250 to 10000 mg/L. The NaCl and KClO4− tolerance and degrading capacity of the strains were demonstrated to be related to biofilm formation, which was present in the studied cultures. This biofilm is a physical barrier composed of extracellular polymeric substances and biomass, indicating the strains’ ability to isolate themselves from the media after being stressed by high salt concentrations via a concentration gradient (Souid et al. 2021).
Regarding pH as a condition for growth and development of the studied strains, the isolated strains showed a high capacity to resist alkaline concentrations (8 to 12 ± 0.5). This behavior suggests that Bacillus spp. are facultative alkalophilic microorganisms. Several studies conducted by Sanchez-Gonzalez et al. (2011), agree that Bacillus spp. are capable of growing and developing under alkaline conditions (Sanchez-Gonzalez et al. 2011). Due to their alkaline-resistance attribute, it is important to detect and conserve these alkalophilic microorganisms (Mandal et al. 2013; Roy and Mukherjee 2013) for their potential production of alkaline active enzymes for the treatment of alkaline sewage derivatives from the laundry detergent industry (Roy and Mukherjee 2013; Coelho et al. 2016; Lucena-Padrós and Ruiz-Barba 2016).
Evaluation of perchlorate reduction by isolates
In this study, the bacterial strains BBCOL-009, BBCOL-014 and BBCOL-015 reduced perchlorate concentrations by 10, 17 and 25%, respectively (Fig. 4). Table 4 illustrates the variety of perchlorate-reducing species.
The bacterial strains BBCOL-009, BBCOL-014, and BBCOL-015 reduced perchlorate close the values of perchlorate reduction reported for Bacillus isolated from marine sediments of the Colombian Caribbean and Antarctica (Acevedo-Barrios et al. 2019, 2022) (Table 4). The Betaproteobacteria class is the most commonly detected perchlorate-reducing bacteria. nonetheless, BBCOL-009, BBCOL-014 and BBCOL-015, of this study, offer a promising resource for the bioremediation of perchlorate-polluted environments and matrix.
A variety of perchlorate-reducing bacterial species can reduce this contaminant; however, the percentage of reduction varies according to genus and the period of exposure to the pollutant. The rates of perchlorate reduction determined in this study were comparable to those reported by Acevedo-Barrios et al. (2019), Acevedo-Barrios et al. (2022), Acevedo-Barrios et al. (2023).
Most perchlorate-reducing bacteria are anaerobic and facultative, and molecular oxygen is produced as an intermediate for microbial perchlorate reduction in a process that exudes nitrate (Acevedo-Barrios et al. 2022) and use this contaminant as an electron acceptor in their metabolic reactions. Microbial reduction of ClO4− occurs via the biochemical reaction is ClO4− → ClO3− → ClO2− → Cl− +O2). Enzymes, such as perchlorate reductase and superoxide chlorite, carry out the reduction or elimination of perchlorate (Acevedo-Barrios et al. 2019, 2022; Acevedo-Barrios and Olivero-Verbel 2021; Acevedo-Barrios et al. 2023). A reductase can reduce perchlorate to chlorate, and subsequently, to chlorite, whereas superoxide chlorite changes chlorite to chloride and molecular oxygen (Acevedo-Barrios et al. 2019). Biological reduction of perchlorate using bacteria completely degrades perchlorate ions into Cl- and O2 (Acevedo-Barrios and Olivero-Verbel 2021).
Application potential in perchlorate remediation and astrobiology
This study showed for the first time the high capacity of Colombian Caribbean Coast bacterial isolates to survive in extreme environments as well as their capacity to reduce perchlorate. These strains are potentially inputs for biotechnological, industrial, and medical processes, including the biodegradation of several xenobiotics. Lundberg et al. (2013), demonstrated the use of Bacillus spp. as an enzyme producer. In addition, the use of Bacillus spp. was reported to be a biofuel producer and biodegradator of biodiesel and other pollutants (Lundberg et al. 2013; Mukhtar et al. 2019; Acevedo-Barrios et al. 2019; Bertel-Sevilla et al. 2020). Moreover, Mukhtar et al. (2019) related the presence of halotolerant bacterial strains with lipases and hemicellulose degradation capacity, indicating the importance of these microorganisms in soil fertilization and plant metabolism.
In the context of sustainability, the utilization of bacterial-mediated processes to address perchlorate contamination is gaining interest. These bacteria have evolved mechanisms to degrade perchlorate, offering a biological potential for remediation, however, it is imperative to understand their applicability and limitations as well as the diversity of available technologies, ranging from in situ ex situ approaches and natural attenuation to controlled bioreactors at different scales (Fang and Naidu 2023). Table 5 provides a list of perchlorate bioremediation technologies.
The performance of strains BBCOL-009, BBCOL-014, and BBCOL-015 in perchlorate reduction across a wide pH range and saline conditions highlights their suitability for various remediation techniques. These bacteria exhibit resistance to saline conditions and adaptability to different pH levels (Ma et al. 2023; Adams et al. 2015). Their adaptability to broad pH ranges makes them valuable inputs for bioreactors and in situ processes, mitigating potential challenges associated with less adaptable strains (Ma et al. 2023).
Physicochemical properties, especially pH, profoundly impact bacterial growth, metabolism, and survival. Understanding the complex relationship between pH and bacterial performance is crucial in microbiology, environmental sciences, and clinical research (Saravanan et al. 2023). Bacteria capable of resisting extreme pH levels play vital roles in acidic or alkaline ecological niches, thus expanding knowledge about such environments (Harrison et al. 2013).
Operational and pilot-scale variables must be tested, as each technology, aimed at specific environmental matrices, presents inherent advantages and challenges. For example, electron donors are crucial in the perchlorate reduction process. Acetate is the most common, but the role of hydrogen, ethanol, or lactate, as well as the initial perchlorate concentration and electron donor ratio, has been investigated (Losi et al. 2002; Xie et al. 2018; Kim et al. 2020; Nozawa-Inoue et al. 2005; Gal et al. 2008; Shete et al. 2008; Nam et al. 2016).
Environmental risks, including antibiotic resistance, must be evaluated during technological scalability. While initial antibiotic resistance suggests sustained efficiency in perchlorate reduction, prolonged exposure may lead to antimicrobial resistance gene expression and microbiota alterations (Zheng et al. 2019).
The study of such extremophilic microorganisms is not limited to bioremediation; it also offers valuable insights for astrobiology, especially in Mars exploration, where Bacillus spp. could assist in perchlorate reduction, potentially addressing challenges in food production during manned missions (Oze et al. 2021; Schuerger et al. 2003; Nicholson et al. 2012).
Future studies exploring the astrobiological potential of Cartagena sectors, characterized by sandstone formations, hold promise, considering the presence of similar formations and perchlorates on Mars (Yen et al. 2017; Schieber et al. 2020; Farley et al. 2016; Martin et al. 2020). This study marks a fundamental step towards understanding extremophilic microorganisms in the region and their astrobiological and bioremediation potential.
These bacteria could be advantageous adaptive, with a broad temperature tolerance and a pH range (Ma et al. 2023; Adams et al. 2015). Our strains showed strong adaptiveness to a wide range of pH, which demonstrated them to be a highly adaptative input for using in bioreactors and in situ processes, due to potential of hydrogen could be a tedious task in less adaptative strains, decreasing efficiency. This behavior has been reported in other bacterial strains such as Exiguobacterales, Bacillales, Lactobacillales and Bacillales (Ma et al. 2023), showing, similarly to ours, biofilm formation.
Among the physicochemical properties related to bacterial performance of growth, metabolism, and survival, the pH is one of the most important. The pH of the surrounding medium can influence the solubility of essential nutrients, the efficiency of metabolic pathways, and the effectiveness of antimicrobial agents. Understanding the intricate relationship between pH and bacterial performance is essential in various fields, including microbiology, environmental science, and clinical research (Saravanan et al. 2023). The impact of pH on bacterial performance is particularly significant in extreme environments, and understanding the adaptations of bacterial strains that resist extreme pH levels is crucial, especially in the context of astrobiology. Bacteria that exhibit tolerance to extreme pH conditions play a pivotal role in ecological niches with acidic or alkaline features, such as acid mine drainage sites, alkaline soda lakes, or geothermal springs. These extremophiles have evolved unique mechanisms to thrive in environments outside the typical pH range. Studying extremophiles not only expands our knowledge of microbial diversity but also has implications for astrobiology, where the exploration of extraterrestrial environments involves considerations of extreme pH conditions. The ability of certain bacterial strains to withstand extreme pH levels is relevant to the search for life beyond Earth, as extraterrestrial environments may present challenges akin to those encountered by extremophiles on Earth. Therefore, investigations into the pH tolerance of bacterial strains hold significance not only for understanding microbial ecology on Earth but also for anticipating and exploring potential habitats in the broader scope of astro biological research (Harrison et al. 2013).
The ability to generate biofilms and adherence to surfaces made them suitable, especially for reactor-based systems or solid-phase treatments. It is a fact that in the mass production of Polydroxialkanoate (PHA), common in biofilm formation in Bacillus genus, BBCOL-009 stands out among the rest of the PHA-producers due to its capacity to tolerate extreme conditions that could be present within the bioreactor (Divyashree et al. 2009). Highlights that biofilms are shared spaces in a bacterial population.
However, pilot scale and operational variables should be tested as each technology, targeted for specific environmental matrices, comes with inherent advantages and challenges. For instance, electron donors are crucial in the perchlorate reduction process. The most common is acetate, however, the role of hydrogen, ethanol, or lactate, organic matter has been investigated (Losi et al. 2002; Xie et al. 2018; Kim et al. 2020). Also, the efficiency of reduction depends on the initial perchlorate concentration and the perchlorate electron donor ratio (Nozawa-Inoue et al. 2005; Gal et al. 2008; Shete et al. 2008) and the negative effect of nitrate has been reported (Nam et al. 2016).
Few environmental risks must be considered in the scaling of remediation technologies. An important aspect is antibiotic resistance, in which the strains showed resistance (Table 2). This condition initially suggests the capacity of the strains to sustain their efficiency in perchlorate reduction in environments containing environmental pollutants such as antibiotic’s traces, which are considered emerging pollutants. However, prolonged exposure may generate the expression of antimicrobial resistance genes and alterations in the microbiota. This should be investigated in depth (Zheng et al. 2019). The remediation species should maintain genetic stability, ensuring consistent perchlorate reduction and environmental safety over time.
All the above indicate that these microorganisms offer an opportunity to delve into other fields of study, such as astrobiology. One of the main lines of research in this transdisciplinary field (Leal et al. 2023) is the study of extremophile microorganisms that can serve as a model for understanding possible biological adaptations in other bodies of the Solar System (DasSarma et al. 2020; Thombre et al. 2020). Some studies have shown that organisms of the genus Bacillus could be promising in searching for potential past or present life on Mars, mainly due to the presence of resistance structures such as endospores (Schuerger et al. 2003; Nicholson et al. 2012).
In addition, the results provide a much more interesting perspective, such as the ability of Bacillus spp. to grow in environments contaminated with perchlorate and allow the reduction of this salt in the substrate. This is a fundamental aspect in astrobiology studies on Mars since perchlorate has been detected on the surface of the red planet, being one of the main problems for food production in future manned missions (Oze et al. 2021). However, organisms such as those reported in the present study could be a potential solution to this problem.
Although no study was carried out on the geological origin of the sediments sampled and analyzed in the present study, a future perspective to continue delving into the astrobiological potential of some Cartagena sectors is based on fine to very fine-grained sandstones’ characterizing their geological formations. Slightly feldspathic, with few lithics and parallel flat lamination in thin layers (Clavijo and Royero 2000). They are presenting the region as a potential study site of astrobiological interest, taking into account that the presence of sandstones has been identified in the Gale Crater on Mars (Yen et al. 2017; Schieber et al. 2020), as well as the presence of perchlorates (Farley et al. 2016; Martin et al. 2020). Thus, this study is presented as a first step in a chain of opportunities to continue delving deeper into the study of the extremophile microorganisms in the region and their astrobiological and bioremediation potential.
Conclusion
This is the novel study conducted in Bahia Cartagena, Colombian Caribbean, which reveals that bacteria belonging to the genus Bacillus, specifically BBCOL-009, BBCOL-014, and BBCOL-015, were extracted from marine sediment samples obtained from Cartagena Bay. It was found that these bacteria exhibit tolerance to environmental perchlorate concentrations of up to 10000 mg/L. Moreover, they are capable of reducing perchlorate concentrations by 10–25%. Furthermore, these bacteria demonstrate the ability to thrive in a wide range of pH and NaCl concentrations, showcasing their potential for remediating this contaminant through various technologies and biotechnological applications in agriculture and water sources remediation. Although the primary focus of the study of perchlorate-reducing and resistant bacteria is in the ecological and agricultural realms, from an astrobiological perspective, perchlorate-resistant bacteria serve as models for astrobiological investigations.
Data availability
No datasets were generated or analysed during the current study.
References
Acevedo Barrios RL, Hernández Rocha I, Puentes Martinez D et al (2023) Psychrobacter sp: perchlorate reducing bacteria, isolated from marine sediments from Margarita Bay, Antarctica. In: Proceedings of the 21th LACCEI International Multi-Conference for Engineering, Education and Technology (LACCEI 2023): Leadership in Education and Innovation in Engineering in the Framework of Global Transformations: Integration and Alliances for Integral Development. Latin American and Caribbean Consortium of Engineering Institutions. https://doi.org/10.18687/LACCEI2023.1.1.995
Acevedo-Barrios R, Olivero-Verbel J (2021) Perchlorate contamination: sources, effects, and technologies for Remediation. In: de Voogt P (ed) Reviews of environmental contamination and toxicology volume 256. Springer International Publishing, Cham, pp 103–120
Acevedo-Barrios R, Bertel-Sevilla A, Alonso-Molina J, Olivero-Verbel J (2016) Perchlorate tolerant bacteria from saline environments at the Caribbean region of Colombia. Toxicol Lett 259:S103. https://doi.org/10.1016/J.TOXLET.2016.07.257
Acevedo-Barrios R, Bertel-Sevilla A, Alonso-Molina J, Olivero-Verbel J (2019) Perchlorate reducing Bacteria from Hypersaline soils of the Colombian caribbean. Int J Microbiol 2019:1–13. https://doi.org/10.1155/2019/6981865
Acevedo-Barrios R, Rubiano-Labrador C, Navarro-Narvaez D et al (2022) Perchlorate-reducing bacteria from Antarctic Marine sediments. Environ Monit Assess 194:654. https://doi.org/10.1007/s10661-022-10328-w
Adams GO, Fufeyin PT, Okoro SE, Ehinomen I (2015) Bioremediation, biostimulation and bioaugmention: a review. Int J Environ Bioremediat Biodegradation 3(1):28–39. https://doi.org/10.12691/ijebb-3-1-5
Albuquerque L, Tiago I, Taborda M et al (2008) Bacillus isabeliae sp. nov., a halophilic bacterium isolated from a sea salt evaporation pond. Int J Syst Evol Microbiol 58:226–230. https://doi.org/10.1099/ijs.0.65217-0
Alonso JL, Cuesta G, Ramírez GW, Morenilla JJ, Bernácer I, Lloret RM (2009) Manual de técnicas avanzadas para la identificación y control de bacterias filamentosas. EPSAR-Generalitat Valenciana
Ashour MSED, Mansy MS, Eissa ME (2011) Microbiological environmental monitoring in pharmaceutical facility. Egypt Acad J Biol Sci G Microbiol 3(1):63–74. https://doi.org/10.21608/eajbsg.2011.16696
Azubuike CC, Chikere CB, Okpokwasili GC (2016) Bioremediation techniques–classification based on site of application: principles, advantages, limitations and prospects. World J Microbiol Biotechnol 32:180. https://doi.org/10.1007/s11274-016-2137-x
Bahamdain L, Fahmy F, Lari S, Aly M (2015) Characterization of some Bacillus strains obtained from marine habitats using different taxonomical methods. Life Sci J 12(4):58–63. http://www.lifesciencesite.com
Belal AAM, Kelany MS, Hamed MM, El-Fattah LSA (2020) Selected bacterial communities associated with macro-benthic fauna assemblages at the Timsah Lake and the Western Lagoon’s sediments, Suez Canal, Egypt. Egypt J Aquat Res 46:137–143. https://doi.org/10.1016/j.ejar.2020.02.003
Beneduzi A, Peres D, Vargas LK et al (2008) Evaluation of genetic diversity and plant growth promoting activities of nitrogen-fixing bacilli isolated from rice fields in South Brazil. Appl Soil Ecol 39:311–320. https://doi.org/10.1016/j.apsoil.2008.01.006
Bertel-Sevilla A, Cervantes-Ceballos L, Tirado-Ballestas I et al (2020) Biodegradation of biodiesel-oil by Cellulosimicrobium sp. Isolated from Colombian caribbean soils. Environ Technol 41:2337–2349. https://doi.org/10.1080/09593330.2018.1564798
Biemer JJ (1973) Antimicrobial susceptibility testing by the Kirby-Bauer disc diffusion method. Annals Clin Lab Sci 3(2):135–140
Boone DR, Castenholz RW, Garrity GM et al (2005) Bergey’s Manual® of systematic bacteriology. Springer Science & Business Media, Boston
Borden RC (2007) Concurrent bioremediation of perchlorate and 1,1,1-trichloroethane in an emulsified oil barrier. J Contam Hydrol 94:13–33. https://doi.org/10.1016/j.jconhyd.2007.06.002
Breed RS, Murray EGD, Smith NR (1957) Bergey’s Manual of. Determinative bacteriol, 7th edn. Williams Wilkins Co Baltim
Cang Y, Roberts DJ, Clifford DA (2004) Development of cultures capable of reducing perchlorate and nitrate in high salt solutions. Water Res 38:3322–3330. https://doi.org/10.1016/j.watres.2004.04.020
Carlström CI, Lucas LN, Rohde RA et al (2016) Characterization of an anaerobic marine microbial community exposed to combined fluxes of perchlorate and salinity. Appl Microbiol Biotechnol 100:9719–9732. https://doi.org/10.1007/s00253-016-7780-5
Clavijo J, Royero J (2000) Mapa Geológico Del Departamento De Bolívar. Escala 1:400000
Coelho SL, de Magalhães A, Marbach VC, Cazetta PAS ML (2016) A new alkalophilic isolate of Bacillus as a producer of cyclodextrin glycosyltransferase using cassava flour. Braz J Microbiol 47:120–128. https://doi.org/10.1016/j.bjm.2015.11.018
Dalmaso GZL, Ferreira D, Vermelho AB (2015) Marine extremophiles: a source of hydrolases for Biotechnological Applications. Mar Drugs 13:1925–1965. https://doi.org/10.3390/md13041925
DasSarma S, DasSarma P, Laye VJ, Schwieterman EW (2020) Extremophilic models for astrobiology: haloarchaeal survival strategies and pigments for remote sensing. Extremophiles 24:31–41. https://doi.org/10.1007/s00792-019-01126-3
Davies J, Davies D (2010) Origins and Evolution of Antibiotic Resistance. Microbiol Mol Biol Rev 74:417–433. https://doi.org/10.1128/mmbr.00016-10
Divyashree MS, Shamala TR, Rastogi NK (2009) Isolation of polyhydroxyalkanoate from hydrolyzed cells of Bacillus flexus using aqueous two-phase system containing polyethylene glycol and phosphate. Biotechnol Bioprocess Eng 14:482–489. https://doi.org/10.1007/s12257-008-0119-z
Dong X, Yu K, Jia X et al (2022) Perchlorate reduction kinetics and genome-resolved metagenomics identify metabolic interactions in acclimated saline lake perchlorate-reducing consortia. Water Res 227:119343. https://doi.org/10.1016/j.watres.2022.119343
Durval IJB, Resende AHM, Figueiredo MA et al (2019) Studies on biosurfactants produced using Bacillus cereus isolated from seawater with biotechnological potential for marine oil-spill bioremediation. J Surfactants Deterg 22:349–363. https://doi.org/10.1002/jsde.12218
Elisashvili V, Kachlishvili E, Chikindas ML (2019) Recent advances in the physiology of spore formation for Bacillus probiotic production. Probiotics Antimicrob Proteins 11:731–747. https://doi.org/10.1007/s12602-018-9492-x
Fang C, Naidu R (2023) A review of perchlorate contamination: analysis and remediation strategies. Chemosphere 338:139562. https://doi.org/10.1016/j.chemosphere.2023.139562
Farley KA, Martin P, Archer PD et al (2016) Light and variable 37Cl/35Cl ratios in rocks from Gale Crater, Mars: possible signature of perchlorate. Earth Planet Sci Lett 438:14–24. https://doi.org/10.1016/j.epsl.2015.12.013
Feitkenhauer H, Müller R, MAuml;rkl H (2003) Degradation of polycyclic aromatic hydrocarbons and long chain alkanes at 6070°C by Thermus and Bacillus spp. Biodegradation 14:367–372. https://doi.org/10.1023/A:1027357615649
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376. https://doi.org/10.1007/bf01734359
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791. https://doi.org/10.1111/j.1558-5646.1985.tb00420.x
Fitch WM (1971) Toward defining the course of evolution: Minimum Change for a specific Tree Topology. Syst Biol 20:406–416. https://doi.org/10.1093/sysbio/20.4.406
Flores N, Hoyos S, Venegas M et al (2020) Haloterrigena sp. Strain SGH1, a Bacterioruberin-Rich, Perchlorate-Tolerant Halophilic Archaeon isolated from Halite Microbial communities, Atacama Desert, Chile. Front Microbiol 11:324. https://doi.org/10.3389/fmicb.2020.00324
Gal H, Ronen Z, Weisbrod N et al (2008) Perchlorate biodegradation in contaminated soils and the deep unsaturated zone. Soil Biol Biochem 40:1751–1757. https://doi.org/10.1016/j.soilbio.2008.02.015
Garrity GM (2001) Bergey’s Manual® of systematic bacteriology. Springer Science & Business Media
Gaspari F, Paitan Y, Mainini M et al (2005) Myxobacteria isolated in Israel as potential source of new anti-infectives. J Appl Microbiol 98:429–439. https://doi.org/10.1111/j.1365-2672.2004.02477.x
Gavrilescu M, Demnerová K, Aamand J et al (2015) Emerging pollutants in the environment: present and future challenges in biomonitoring, ecological risks and bioremediation. New Biotechnol 32:147–156. https://doi.org/10.1016/j.nbt.2014.01.001
Guadie A, Tizazu S, Melese M et al (2017) Biodecolorization of textile azo dye using Bacillus sp. strain CH12 isolated from alkaline lake. Biotechnol Rep 15:92–100. https://doi.org/10.1016/j.btre.2017.06.007
Harrison JP, Gheeraert N, Tsigelnitskiy D, Cockell CS (2013) The limits for life under multiple extremes. Trends Microbiol 21(4):204–212. https://doi.org/10.1016/j.tim.2013.01.006
Hatzinger PB, Whittier MC, Arkins MD et al (2002) In-Situ and Ex-situ Bioremediation options for Treating Perchlorate in Groundwater. Remediat J 12:69–86. https://doi.org/10.1002/rem.10026
Hong HA, Khaneja R, Tam NMK et al (2009) Bacillus subtilis isolated from the human gastrointestinal tract. Res Microbiol 160:134–143. https://doi.org/10.1016/j.resmic.2008.11.002
Huang X, Madan A (1999) CAP3: a DNA sequence Assembly Program. Genome Res 9:868–877. https://doi.org/10.1101/gr.9.9.868
Iizuka T, Tokura M, Jojima Y et al (2006) Enrichment and phylogenetic analysis of moderately thermophilic myxobacteria from Hot Springs in Japan. Microbes Environ 21:189–199. https://doi.org/10.1264/jsme2.21.189
Istock CA, Graumann P (2008) Review of Bacillus: Cellular and Molecular Biology, Peter Graumann. Q Rev Biol 83:117–117. https://doi.org/10.1086/586951
Jacob JJ, Sumana S, Jayasri MA, Suthindhiran K (2018) Isolation, characterization and kinetics of Perchlorate reducing Magnetospirillum Species. Geomicrobiol J 35:120–126. https://doi.org/10.1080/01490451.2017.1338796
Kim OS, Cho YJ, Lee K et al (2012) Introducing EzTaxon-e: a prokaryotic 16s rRNA gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Microbiol 62:716–721. https://doi.org/10.1099/ijs.0.038075-0
Kim H-W, Hong SH, Choi H (2020) Effect of Nitrate and Perchlorate on Selenate reduction in a sequencing batch Reactor. Processes 8:344. https://doi.org/10.3390/pr8030344
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120. https://doi.org/10.1007/bf01731581
Koneman EW, Allen SD, Janda WM, Schreckenberger PC, Winn WC (2006) Diagnostico Microbiologico Texto Y Atlas a color. 6 edicion. Medica Panamericana
Layton C, Maldonado E, Monroy L et al (2011) Bacillus spp.; perspectiva de su efecto biocontrolador mediante antibiosis en cultivos afectados por fitopatógenos. Nova 9:177–187. https://doi.org/10.22490/24629448.501
Le Borgne S, Paniagua D, Vazquez-Duhalt R (2008) Biodegradation of organic pollutants by halophilic bacteria and archaea. Microb Physiol 15:74–92. https://doi.org/10.1159/000121323
Leal MA, Tovar D, Valbuena M et al (2023) The Transdisciplinary Nature of Astrobiology as a Transversal Axis of the Educational processes at the Planetarium of Bogota. Rev Mex Astron Astrofısica Ser Conf RMxAC 55:29–34. https://doi.org/10.22201/ia.14052059p.2023.55.06
Lee JC, Lim JM, Park DJ et al (2006) Bacillus seohaeanensis sp. nov., a halotolerant bacterium that contains L-lysine in its cell wall. Int J Syst Evol Microbiol 56:1893–1898. https://doi.org/10.1099/ijs.0.64237-0
Lee CW, Ng AY, Narayanan K et al (2009) Isolation and characterization of culturable bacteria from tropical coastal waters. Cienc Mar 35:153–167
Lee S-H, Hwang J-H, Kabra AN et al (2015) Perchlorate reduction from a highly concentrated aqueous solution by bacterium Rhodococcus sp. YSPW03. Environ Sci Pollut Res 22:18839–18848. https://doi.org/10.1007/s11356-015-5072-8
Lee JY, Kim H, Jeong Y, Kang C-H (2021) Lactic acid bacteria exert a hepatoprotective effect against ethanol-induced liver injury in HepG2 cells. Microorganisms 9:1844. https://doi.org/10.3390/microorganisms9091844
Lei Z, Qiu P, Ye R et al (2014) Bacillus shacheensis sp. nov., a moderately halophilic bacterium isolated from a saline-alkali soil. J Gen Appl Microbiol 60:101–105. https://doi.org/10.2323/jgam.60.101
Liu JL, Wong MH (2013) Pharmaceuticals and personal care products (PPCPs): a review on environmental contamination in China. Environ Int 59:208–224. https://doi.org/10.1016/j.envint.2013.06.012
Liu Y, Tang H, Lin Z, Xu P (2015) Mechanisms of acid tolerance in bacteria and prospects in biotechnology and bioremediation. Biotechnol Adv 33:1484–1492. https://doi.org/10.1016/j.biotechadv.2015.06.001
Losi ME, Giblin T, Hosangadi V, Frankenberger WT Jr (2002) Bioremediation of perchlorate-contaminated Groundwater using a packed Bed Biological Reactor. Bioremediat J 6:97–103. https://doi.org/10.1080/10588330208951206
Lucena-Padrós H, Ruiz-Barba JL (2016) Diversity and enumeration of halophilic and alkaliphilic bacteria in spanish-style green table-olive fermentations. Food Microbiol 53:53–62. https://doi.org/10.1016/j.fm.2015.09.006
Lundberg ME, Becker EC, Choe S (2013) MstX and a putative Potassium Channel facilitate Biofilm formation in Bacillus subtilis. PLoS ONE 8:e60993. https://doi.org/10.1371/journal.pone.0060993
Ma Y, You F, Parry D, Urban A, Huang L (2023) Adaptive growth and acidogenic fermentation performance of haloalkaliphilic bacterial communities enriched from biofilms colonising strongly alkaline and saline bauxite residue. Sci Total Environ 856:159131. https://doi.org/10.1016/j.scitotenv.2022.159131
Mandal A, Das K, Roy S et al (2013) In vivo assessment of bacteriotherapy on acetaminophen-induced uremic rats. J Nephrol 26:228–236. https://doi.org/10.5301/jn.5000129
Marín LF, Jaramillo B (2015) Aislamiento De Bacterias degradadoras de pesticidas organofosforados encontrados en suelos y en leche bovina. Rev Chil Nutr 42:179–185. https://doi.org/10.4067/S0717-75182015000200010
Martin PE, Farley KA, Douglas Archer P Jr et al (2020) Reevaluation of perchlorate in gale crater rocks suggests geologically recent perchlorate addition. J Geophys Res Planet. https://doi.org/10.1029/2019JE006156
Masika WS, Moonsamy G, Mandree P et al (2020) Biodegradation of petroleum hydrocarbon waste using consortia of Bacillus sp. Bioremediat J 25:72–79. https://doi.org/10.1080/10889868.2020.1842322
Miranda CAC, Martins OB, Clementino MM (2008) Species-level identification of Bacillus strains isolates from marine sediments by conventional biochemical, 16S rRNA gene sequencing and inter-tRNA gene sequence lengths analysis. Antonie Van Leeuwenhoek 93:297–304. https://doi.org/10.1007/s10482-007-9204-0
Mohapatra RK, Parhi PK, Pandey S et al (2019) Active and passive biosorption of pb(II)using live and dead biomass of marine bacterium Bacillus xiamenensis PbRPSD202: kinetics and isotherm studies. J Environ Manage 247:121–134. https://doi.org/10.1016/j.jenvman.2019.06.073
Mukhtar S, Mehnaz S, Mirza MS, Malik KA (2019) Isolation and characterization of bacteria associated with the rhizosphere of halophytes (Salsola Stocksii and Atriplex amnicola) for production of hydrolytic enzymes. Braz J Microbiol 50:85–97. https://doi.org/10.1007/s42770-019-00044-y
Murray PR, Zeitinger JR, Krogstad DJ (1982) Reliability of Disc Diffusion susceptibility testing. Infect Control Hosp Epidemiol 3:230–237. https://doi.org/10.1017/S0195941700056150
Nam JH, Ventura JRS, Yeom IT et al (2016) A novel perchlorate- and nitrate-reducing bacterium, Azospira sp. PMJ. Appl Microbiol Biotechnol 100:6055–6068. https://doi.org/10.1007/s00253-016-7401-3
Nicholson WL, McCoy LE, Kerney KR et al (2012) Aqueous extracts of a Mars analogue regolith that mimics the Phoenix landing site do not inhibit spore germination or growth of model spacecraft contaminants Bacillus subtilis 168 and Bacillus pumilus SAFR-032. Icarus 220:904–910. https://doi.org/10.1016/j.icarus.2012.06.033
Noguchi H, Uchino M, Shida O et al (2004) Bacillus vietnamensis sp. nov., a moderately halotolerant, aerobic, endospore-forming bacterium isolated from Vietnamese fish sauce. Int J Syst Evol Microbiol 54:2117–2120. https://doi.org/10.1099/ijs.0.02895-0
Nor SJ, Lee SH, Cho KS et al (2011) Microbial treatment of high-strength perchlorate wastewater. Bioresour Technol 102:835–841. https://doi.org/10.1016/j.biortech.2010.08.127
Nozawa-Inoue M, Scow KM, Rolston DE (2005) Reduction of perchlorate and nitrate by microbial communities in vadose soil. Appl Environ Microbiol 71:3928–3934. https://doi.org/10.1128/AEM.71.7.3928-3934.2005
Oguntoyinbo FA (2007) Monitoring of marine Bacillus diversity among the bacteria community of sea water. Afr J Biotechnol 6
Oyetibo GO, Chien M-F, Ikeda-Ohtsubo W et al (2017) Biodegradation of crude oil and phenanthrene by heavy metal resistant Bacillus subtilis isolated from a multi-polluted industrial wastewater creek. Int Biodeterior Biodegrad 120:143–151. https://doi.org/10.1016/j.ibiod.2017.02.021
Oze C, Beisel J, Dabsys E et al (2021) Perchlorate and agriculture on Mars. Soil Syst 5:37. https://doi.org/10.3390/soilsystems5030037
Priest FG, Goodfellow M, Todd C (1988) A Numerical classification of the Genus Bacillus. Microbiology 134:1847–1882. https://doi.org/10.1099/00221287-134-7-1847
Reimer L (2000) Scanning Electron Microscopy: physics of image formation and microanalysis, second edition. Meas Sci Technol. https://doi.org/10.1088/0957-0233/11/12/703
Reller LB, Weinstein M, Jorgensen JH, Ferraro MJ (2009) Antimicrobial susceptibility testing: a review of General principles and Contemporary practices. Clin Infect Dis 49:1749–1755. https://doi.org/10.1086/647952
Romero-Murillo P, Gallego JL, Leignel V (2023) Marine Pollution and advances in Biomonitoring in Cartagena Bay in the Colombian Caribbean. Toxics 11:631
Roy JK, Mukherjee AK (2013) Applications of a high maltose forming, thermo-stable α-amylase from an extremely alkalophilic Bacillus licheniformis strain AS08E in food and laundry detergent industries. Biochem Eng J 77:220–230. https://doi.org/10.1016/j.bej.2013.06.012
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Sanchez-Gonzalez M, Blanco‐Gamez A, Escalante A et al (2011) Isolation and characterization of new facultative alkaliphilic Bacillus flexus strains from maize processing waste water (nejayote). Lett Appl Microbiol 52:413–419. https://doi.org/10.1111/j.1472-765X.2011.03021.x
Saravanan A, Kumar PS, Duc PA, Rangasamy G (2023) Strategies for microbial bioremediation of environmental pollutants from industrial wastewater: a sustainable approach. Chemosphere 313:137323. https://doi.org/10.1016/j.chemosphere.2022.137323
Schieber J, Minitti ME, Sullivan R et al (2020) Engraved on the rocks—aeolian abrasion of Martian mudstone exposures and their relationship to modern wind patterns in Gale Crater, Mars. Depositional Rec 6:625–647. https://doi.org/10.1002/dep2.110
Schuerger AC, Mancinelli RL, Kern RG et al (2003) Survival of endospores of Bacillus subtilis on spacecraft surfaces under simulated martian environments:: implications for the forward contamination of Mars. Icarus 165:253–276. https://doi.org/10.1016/S0019-1035(03)00200-8
Shang Y, Wang Z, Xu X et al (2018) Bio-reduction of free and laden perchlorate by the pure and mixed perchlorate reducing bacteria: considering the pH and coexisting nitrate. Chemosphere 205:475–483. https://doi.org/10.1016/j.chemosphere.2018.04.132
Shete A, Mukhopadhyaya PN, Acharya A et al (2008) Aerobic reduction of perchlorate by bacteria isolated in Kerala, South India. J Appl Genet 49:425–431. https://doi.org/10.1007/BF03195643
Shih Y-J, Wu Z-L, Hsu C-H (2024) Perchlorate decomposition on rhodium nanoclusters supported on copper crystallite (RhxCu) electrode in an electrodialysis-assisted bipolar electro-reduction system (ED-ER). Chem Eng J 481:148477. https://doi.org/10.1016/j.cej.2023.148477
Shimkets LJ, Rafiee H (1990) CsgA, an extracellular protein essential for Myxococcus xanthus development. J Bacteriol 172:5299–5306. https://doi.org/10.1128/jb.172.9.5299-5306.1990
Shrout JD, Scheetz TE, Casavant TL, Parkin GF (2005) Isolation and characterization of autotrophic, hydrogen-utilizing, perchlorate-reducing bacteria. Appl Microbiol Biotechnol 67:261–268. https://doi.org/10.1007/s00253-004-1725-0
Souid A, Della Croce CM, Frassinetti S et al (2021) Nutraceutical potential of Leaf Hydro-Ethanolic Extract of the Edible Halophyte Crithmum maritimum L. Molecules 26:5380. https://doi.org/10.3390/molecules26175380
Sriariyanun M, Tantayotai P, Yasurin P et al (2016) Production, purification and characterization of an ionic liquid tolerant cellulase from Bacillus sp. isolated from rice paddy field soil. Electron J Biotechnol 19:23–28. https://doi.org/10.1016/j.ejbt.2015.11.002
Srinivasan A, Viraraghavan T (2009) Perchlorate: health effects and technologies for its removal from water resources. Int J Environ Res Public Health 6:1418–1442. https://doi.org/10.3390/ijerph6041418
Tamura K, Stecher G, Peterson D et al (2013) MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Mol Biol Evol 30:2725–2729. https://doi.org/10.1093/molbev/mst197
Thombre RS, Vaishampayan PA, Gomez F (2020) Chap. 7 - Applications of extremophiles in astrobiology. In: Salwan R, Sharma V (eds) Physiological and Biotechnological Aspects of Extremophiles. Academic Press, pp 89–104. https://doi.org/10.1016/B978-0-12-818322-9.00007-1
Van Ginkel SW, Ahn CH, Badruzzaman M et al (2008) Kinetics of nitrate and perchlorate reduction in ion-exchange brine using the membrane biofilm reactor (MBfR). Water Res 42:4197–4205. https://doi.org/10.1016/j.watres.2008.07.012
Wan D, Liu Y, Niu Z et al (2016) Perchlorate reduction by hydrogen autotrophic bacteria and microbial community analysis using high-throughput sequencing. Biodegradation 27:47–57
Weisblum B (1995) Erythromycin resistance by ribosome modification. Antimicrob Agents Chemother 39(3):577–585. https://doi.org/10.1128/AAC.39.3.577
William F (1993) Chemical studies of marine bacteria: developing a new resource. Chem Rev 93:1673–1683. https://doi.org/10.1021/cr00021a001
Winn WC, Allen SD, Janda WM et al (2008) Bacilos Grampositivos Aerobios facultativos. Diagnóstico microbiológico: Texto Y Atlas. Médica Panamericana, España, pp 787–789
Wu Z-H, Jiang D-M, Li P, Li Y-Z (2005) Exploring the diversity of myxobacteria in a soil niche by myxobacteria-specific primers and probes. Environ Microbiol 7:1602–1610. https://doi.org/10.1111/j.1462-2920.2005.00852.x
Xie T, Yang Q, Winkler MKH et al (2018) Perchlorate bioreduction linked to methane oxidation in a membrane biofilm reactor: performance and microbial community structure. J Hazard Mater 357:244–252. https://doi.org/10.1016/j.jhazmat.2018.06.011
Yang X, Zhang H, Chan EWC, Zhang R, Chen S (2024) Transmission of azithromycin-resistant gene, erm (T), of Gram-positive bacteria origin to Klebsiella pneumoniae. Microbiol Res. https://doi.org/10.1016/j.micres.2024.127636
Yen AS, Ming DW, Vaniman DT et al (2017) Multiple stages of aqueous alteration along fractures in mudstone and sandstone strata in Gale Crater, Mars. Earth Planet Sci Lett 471:186–198. https://doi.org/10.1016/j.epsl.2017.04.033
Yilmaz T, Yurtsever A, Sahinkaya E, Uçar D (2023) Perchlorate reduction in a thiosulfate-based denitrifying membrane bioreactor. Biochem Eng J 200:109100. https://doi.org/10.1016/j.bej.2023.109100
Yoon JH, Kim IG, Kang KH et al (2003) Bacillus marisflavi sp. nov. and Bacillus aquimaris sp. nov., isolated from sea water of a tidal flat of the Yellow Sea in Korea. Int J Syst Evol Microbiol 53:1297–1303. https://doi.org/10.1099/ijs.0.02365-0
Zhang L, Wang Y, Dai J et al (2009) H korlensis sp. nov., a moderately halotolerant bacterium isolated from a sand soil sample in China. Int J Syst Evol Microbiol 59:1787–1792. https://doi.org/10.1099/ijs.0.004879-0
Zhang F, Jiang X, Chai L et al (2014) Permanent draft genome sequence of Bacillus flexus strain T6186-2, a multidrug-resistant bacterium isolated from a deep-subsurface oil reservoir. Mar Genomics 18:135–137. https://doi.org/10.1016/j.margen.2014.09.007
Zheng X, Jiang B, Lang H et al (2019) Effects of Antibiotics on Microbial communities responsible for perchlorate degradation. Water Air Soil Pollut 230:244. https://doi.org/10.1007/s11270-019-4302-y
Zolkefli N, Ramli N, Mohamad-Zainal NSL et al (2020) Alcaligenaceae and Chromatiaceae as pollution bacterial bioindicators in palm oil mill effluent (POME) final discharge polluted rivers. Ecol Indic 111:106048. https://doi.org/10.1016/j.ecolind.2019.106048
Acknowledgements
The authors gratefully acknowledge Minciencias-Universidad de Cartagena (Grant RC-758-2011/1107-521-29360), Vice-Presidency for Research at the University of Cartagena (Research Group Support Program, 2011–2013), Cartagena, Colombia and, National Program for Doctoral Formation (MINCIENCIAS) grants 647-2014 and 727-2015.
Funding
Open Access funding provided by Colombia Consortium. This research was supported by the Doctoral Programs in Environmental Toxicology at the University of Cartagena.
Author information
Authors and Affiliations
Contributions
RAB, ABS, JLG, MAL, DT and ITB were in charge of conceptualization; investigation; visualization; writing—original draft, writing—review and editing. LCC was in charge of investigation and JOV was in charge of conceptualization, resources, and supervision.
Corresponding author
Ethics declarations
Competing interest
The authors declare no competing interests.
Ethical Approval
All authors have read, understood, and have complied as applicable with the statement on “Ethical responsibilities of Authors” as found in the Instructions for Authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Acevedo-Barrios, R., Tirado-Ballestas, I., Bertel-Sevilla, A. et al. Bioprospecting of extremophilic perchlorate-reducing bacteria: report of promising Bacillus spp. isolated from sediments of the bay of Cartagena, Colombia. Biodegradation (2024). https://doi.org/10.1007/s10532-024-10079-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10532-024-10079-0