Abstract
Humans and wildlife experience complex interactions in urban ecosystems, favoring the presence of commensal species, among which invasive species are particularly successful. Rodents are the main vertebrate group introduced to oceanic islands, where the invasion process and dispersal patterns strongly influence their evolutionary and genetic patterns. We evaluated the house mouse Mus musculus and the black rat Rattus rattus on Cozumel Island, Mexico. We assessed genetic diversity and structure, connectivity, gene flow, relatedness and bottleneck signals based on microsatellite loci. Our genetic findings suggest that introduction of individuals of different geographic sources to the island promotes high allelic diversity and the effective establishment of migrants. We identified a clear genetic structure and low connectivity for the two species, tightly linked with anthropogenic and urban features. Notably, we found that the genetic structure of the house mouse sampled within the city of San Miguel Cozumel is associated with the historical human population growth pulses accompanying the urbanization of the city. At the fine-scale genetic level, the main urban drivers of connectivity of the house mouse were both the impervious land surfaces, i.e. the urban landscape, and the informal commerce across the city (a proxy of resources availability). Chances of a secondary invasion to natural environments have been relatively low, which is crucial for the endemic taxa of the island. Nonetheless, improving urban planning to regulate future expansions of San Miguel Cozumel is of the outmost importance to prevent these invasive species to disperse further.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cities worldwide vary in age, shape, size, human population, geographic context and history, and are undergoing accelerated expansion, making them highly dynamic and unstable systems. Urban ecosystems have high heterogeneity, usually characterized by mosaics of land use, including infrastructure with different purposes (residential, commercial, industrial), urban green space (parks, gardens) and natural remnants (Johnson and Munshi-South 2017; Gaertner et al. 2017), frequently organized along gradients from the city centers to suburban surrounding landscapes. In these environments, humans and wildlife experience complex interactions, affecting the evolutionary trajectory of species (Donihue and Lambert 2015; Hulme-Beaman et al. 2016; Des Roches et al. 2021; Santangelo et al. 2022). Urbanization favors the presence of closely commensal species, organisms that have changed life history features like breeding cycles, diet, behavior, or home ranges. Such changes enabled them to thrive in urbanized environments, among which invasive species are especially successful (Hulme-Beaman et al. 2016; Gaertner et al. 2017).
Invasive (non-native) species are those that have been accidentally or intentionally transported and released outside of their natural range, have dispersed from the area of introduction and have established reproductively active populations (Blackburn et al. 2011; Gaertner et al. 2017). In recent decades, global commercial exchange and transportation have facilitated and enhanced biological invasions. The invasion dynamics, i.e. propagule acquisition, introduction, establishment, proliferation, and spread, greatly determines the abundance, structure, and impact of invasive species (Lawson et al. 2011; Estoup et al. 2016). Also, the introduction and dispersal patterns of invasive species strongly influence their genetic diversity and structure, demography, and natural selection (Duglosh and Parker 2007; Konecny et al. 2013; Suárez-Atilano et al. 2019). Indeed, the success of invasion is limited when the founding populations are small, due to decreased genetic variability, genetic drift, and inbreeding resulting from founder effects. In this scenario, the ‘new population’ becomes highly differentiated from the source population, potentially leading also to the fixation of deleterious alleles (Lambrinos 2004; Lawson et al. 2011). However, genetic erosion can be avoided by the recurrent introduction of new alleles through multiple invasion events; also, if the number of initial propagules is high and the bottleneck is short, there would likely be a rapid subsequent population expansion and increased genetic diversity. Hence, the propagule pressure (i.e. relation between the propagule size and propagule frequency) is a key factor for the success or failure of an invasion (Duglosh and Parker 2007; Suarez and Tsutsui 2008; Simberloff 2009; Suárez-Atilano et al. 2019). The availability of resources in cities facilitates the establishment and proliferation of invasive species, while road networks allow the rapid dissemination of propagules (Gaertner et al. 2017; Gortat et al. 2017). In addition, multiple social, economic and ecological factors of cities influence their distribution and abundance, with varying effects on population genetic patterns, gene flow and connectivity of animals and plants (Blackburn et al. 2011; Panty-May et al. 2016; Gaertner et al. 2017; Santangelo et al. 2022). Nonetheless, our understanding of the dynamics of invasion in urban ecosystems is still limited, particularly on islands.
Among invasive species, rodents are among the most widely introduced vertebrates. Their success is associated with their high reproductive rates, short generation times and dietary plasticity; they can consume organic matter, seeds, fruits, different invertebrate and small vertebrate species, eggs and coastal resources (Miller and Webb 2001; Caut et al. 2008; Gardner-Santana et al. 2009; Clair 2011; Shiels et al. 2013; Banks and Smith 2015). Additionally, their evolutionary history is tightly linked with their commensal ecology (Blackburn et al. 2011; Siahsarvie et al. 2012; Lucaccioni et al. 2016), to such a degree that they are considered the main group introduced to oceanic islands (Pichlmueller et al. 2020). Moreover, their global dispersal is significantly associated with human colonization, expansion and urbanization, also regulated by the intensity of commercial and cultural ties established between human populations throughout their history (Stragier et al. 2022). This is the case of the Norwegian rat (Rattus norvegicus), the black rat (Rattus rattus) and the house mouse (Mus musculus).
The black rat is considered the most damaging invasive rodent on islands (Loiseau et al. 2008). It evolved in the Indian peninsula on southern Asia, spread across Europe via the Roman trade (400 BC–100 AC), and dispersed into America and the islands of the Indian and Atlantic oceans with the explorations of the sixteenth and seventeenth centuries (Calmet et al. 2001; Aplin et al. 2011). While the house mouse originated in the Iranian Plateau and is characterized by three evolutionary lineages that diverged 100,000 years ago: M. musculus castaneus in Asia, M. musculus musculus which colonized central and eastern Europe 3000 years ago, and M. musculus domesticus that established in western Europe and arrived in America, Africa and Australia associated with European expeditions beginning in the fifteenth century (Boursot et al. 1996; Jing et al. 2014; Lippens et al. 2017). There are many genetic studies of black rats and house mice focused on diversification, phylogeographic patterns and invasion routes (e.g. Hardouin et al. 2010, 2015; Aplin et al. 2011; Lack et al. 2013; López et al. 2013; Jing et al. 2014; Gabriel et al. 2015; King 2016), both in continental regions (Bastos et al. 2011; Jones et al. 2011; Lippens et al. 2017; Combs et al. 2018) and in islands (Abdelkrim et al. 2005a, 2005b, 2010; Brouat et al. 2014; Babiker and Tautz 2015). Nonetheless, few studies have investigated their genetic and ecological dynamics associated with urban ecosystems specifically on islands (Gatto-Almeida et al. 2022).
Cozumel is the largest of the Mexican Caribbean islands and has the highest number of endemics (31 taxa), including among mammals two carnivores (Procyon pygmaeus and Nasua nelsoni) and three rodents (Oryzomys couesi cozumelae, Reithrodontomys spectabilis, and Peromyscus leucopus cozumelae; the latter potentially extinct) (Vega et al. 2007; Cuarón 2009; Fuentes-Montemayor et al. 2009; Espindola et al. 2014; Flores-Manzanero et al. 2022). Nearly 75% of the island’s surface is covered with native vegetation (Cuarón 2009). Conservation efforts have rendered two terrestrial (Refugio Estatal de Flora y Fauna Isla Cozumel and Reserva Estatal Selvas y Humedales de Cozumel) and one marine (Parque Nacional Arrecifal de Cozumel) protected areas. Despite this, a growing human population and increased cruise ships tourism have accelerated urbanization, habitat loss, and fragmentation (Trejo de la Paz 2014). Cozumel’s human population was estimated in 84,519 (census of 2020), while more than 4.5 million people (average 2018–2019) visit the island yearly (INEGI 2020). Consequently, numerous exotic species have been, and continue to be, introduced to the island (e.g. domestic cats and dogs, black rat, house mouse, boa, margay, fox), which have become feral or invasive (Cuarón 2009; Vázquez-Domínguez et al. 2012; Valenzuela-Galván et al. 2022).
Originally inhabited by native Maya, Cozumel Island was abandoned around 1570, undergoing a major population decline some decades after the Spanish conquest in 1521 (Haviland 1972). More than three hundred years later, in 1846–1847, people established again in Cozumel because of the indigenous uprising know as Caste War in the Yucatán peninsula, when rebellious Maya people escaped seeking refuge on the island (Santander and Ramos-Díaz 2011). The contemporary demographic growth of Cozumel is divided in two phases, a stationary one since the recolonization until 1950, and the tourism expansion from 1950 onwards. The latter was characterized by a constant population growth due of the intensification of human migration processes and tourism infrastructure (hotels, piers), yielding a markedly disorganized urbanization expansion (Sánchez-Crispín and Propin 2003; Palafox and Zizumbo 2009; Santander and Ramos-Díaz 2011; Trejo de la Paz 2014). Currently, Cozumel is one of the main tourist destinations in Mexico and the busiest tourist port in the Caribbean.
Hence, Cozumel is a unique study system to evaluate the evolutionary and genetic patterns of invasive rodents associated with the history of urbanization pulses and human population growth of the island. To this end, we studied two invasive rodents, the black rat and the house mouse; we assessed genetic diversity and structure, connectivity, gene flow, relatedness and contemporary and ancestral bottleneck signals using microsatellites loci. In addition, given the particularly restricted distribution we found of the house mouse within the city of San Miguel Cozumel, we evaluated the species’ colonization process and connectivity based on demography and a landscape genetics framework. We integrated environmental, anthropogenic and social variables to adequately represent the urban ecosystem. Based on Cozumel’s contemporary history of population growth and extension of tourism infrastructure, we assumed a constant introduction of individuals of both invasive species via the island seaports and, accordingly, predicted that both species will exhibit moderate genetic diversity values and weak bottleneck signals. Additionally, San Miguel Cozumel preserves some remnants of natural vegetation and, despite its small size and relatively high income from the tourist industry, has high economic heterogeneity and social inequality. Hence, we expected the two rodents populations will exhibit spatial genetic structure and low connectivity associated with urban and socioeconomic features. Yet, we predicted a differential response to urbanization for each species owing to their distinct life history and social organization, with a greater genetic differentiation in the house mouse. We show that considering the genetic history of introduction and past demographic events is key for the understanding of invasion processes of commensal rodents on islands.
Materials and methods
Study site and sampling
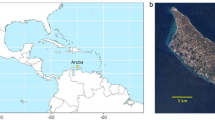
Cozumel (ca. 486 km2) is an oceanic island of coralline origin that has never been connected to the mainland (Spaw 1978; Semarnat 2016). It is located 17.5 km off the Yucatán peninsula in the Caribbean Sea (20° 16′ 18.2″–20° 35′ 32.8″ N; 86° 43′ 23.3″–87° 01′ 31.1″ W), separated from the mainland by the Cozumel Channel (400 m deep). The island has an urbanised area of approximately 45.4 km2 and can be divided based on human-influence zones as: (i) the city of San Miguel Cozumel, home to 98% of the island’s inhabitants (INEGI 2020); it includes the airport, hotel developments on the north and south of the city, and main residential and commercial infrastructure, (ii) the sanitary landfill on the east, and (iii) several scattered small settlements, e.g. several ranches and El Cedral town (Fig. 1).
Map of sampling points for the house mouse (Mus musculus; red dots), the black rat (Rattus rattus; blue) and where no individuals were captured (yellow). The urban area of San Miguel Cozumel city is shown in purple. Other human-influence zones are indicated, the sanitary landfill and El Cedral town, the maritime terminal (Muelle Fiscal) and the ferry port (Muelle Fiscal de carga), and the Puerta Maya and Punta Langosta piers
We sampled black rats and house mice with Sherman-live traps during September 2015. Sampling was site-directed, intended to capture in sites with likely probability of presence of these invasive species, thus traps were set in 12 neighborhoods within San Miguel Cozumel, plus the sanitary landfill, El Cedral town and a few small human settlements around the island (Fig. 1), baited with a mixture of rolled oats, peanut butter, and vanilla extract. Authorization to set traps was obtained at all localities, from the local authorities (piers, landfield) and private owners (food market and establishments, hotel, nearby houses). The number of traps per site and the number of houses sampled varied by neighborhood (Table S1); traps were set before sundown and collected at sunrise during 2–3 days per site, for a total trapping effort of 984 trap/nights. Trapping was performed with the corresponding scientific collecting permit from Secretaría del Medio Ambiente y Recursos Naturales (Semarnat-FAUT-0168). Use of rodenticides or permanent animal trapping techniques cannot be performed on the island due to the high risk of killing native rodent species, despite the directed sampling scheme. Hence, all animals were euthanized, following the procedures allowed by the Mexican Official Norm (Technical specifications for the protection, care and use of laboratory animals; NOM 2001). Accordingly, each captured animal was transported within the Sherman trap to an outdoor processing site where they were introduced into an anesthetic chamber with a cotton soaked with ethyl ether, the amount estimated based on the size of each individual. Death was verified by examination of each animal. The delay between collection from the trapping site and euthanasia was less than 3 h and the complete anesthesia protocol was assessed by a wildlife veterinarian. Post-mortem ear tissue samples were immediately collected and preserved in Eppendorf tubes with 96% ethanol. We also collected tail bones of run over individuals.
Microsatellite genotyping
We extracted genomic DNA from tissue samples with the Genomic DNA Tissue Miniprep kit (ZimoReserch), following the manufacturer’s instructions. For the black rat we genotyped 10 nuclear loci, eight species-specific: Rr14, Rr17, Rr21, Rr22, Rr67, Rr68, Rr107, Rr114 (Loiseau et al. 2008) and two designed for R. norvegicus: D16Rat81, D5Rat83 (Jacob et al. 1995); we chose 10 specific microsatellites for the house mouse: D5Mit149, D6Mit309, D9Mit54, D13Mit61, D15Mit98, EGG22992, PP4A02, PP7B08, PP10A02, PP10E08 (Dietrich et al. 1992; Babiker and Tautz 2015; Hardouin et al. 2015). Microsatellites were amplified by PCR reactions (see protocol details in Table S2). Fragment analysis was performed with the capillary sequencer AbiPrism3730xl Analyzer (Roy J. Carver Biotechnology Center, University of Illinois, USA) with the molecular weight marker Rox500 size standard. Fragment analysis and genotyping were done with GeneMarker v.2.6.7.
Genetic structure, diversity, relatedness and bottlenecks
To infer genetic structure, we used a Discriminant Analysis of Principal Components (DAPC) (Jombart et al. 2010), a multivariate approach that reflects the genetic variation differences between groups while minimizing variation within them. We used adegenet in R v.2.1.5 (R Core Team 2016), applying the alpha-score method to determine the proportion of successful reassignment by individual in function of the number of retained PCs (black rat: 1 PC; house mouse: 8 PCs). We also performed a spatial genetic analysis with Geneland v.4.0.4 (Guillot et al. 2005), a spatial Bayesian method based in Monte Carlo Markov chains and Poisson-Voronoi tessellations to determine genetic discontinuities; 30 independent runs with 10,000,000 interactions and 100,000 of burn-in were performed, an uncertainty coordinate value of 30 m for the black rat based on its low dispersion in urban zones (Gardner-Santana et al. 2009), while for house mice we used a 20 m value considering the species’ home range in cities (Dean et al. 2006).
The following analyses were performed for the entire population and for the different genetic clusters obtained for each species (see genetic structure Results). We screened for null alleles and stuttering in Micro-Checker v.2.2.3 (van Oosterhout et al. 2004), using 3000 randomizations and adjusting the values for multiple comparisons with a Bonferroni sequential correction. We also tested genotypes for Hardy–Weinberg equilibrium and linkage disequilibrium with the Fisher exact test with 1000 batches and 100,000 interactions in Genepop v.4.0 (Rousset 2008) and Bonferroni correction. Genetic diversity was estimated with Genalex v.6.501 (Peakall and Smouse 2012) as the average number of alleles per locus (Na), number of effective (Ae) and private alleles (Ap), observed (Ho), expected (He) and unbiased expected heterozygosity (HNEI) (corrected for small sample sizes; Nei 1974), and inbreeding coefficient FIS. We assessed relatedness by estimating the likelihood of two individuals belonging to the same group being half-siblings, full siblings, parent–offspring or unrelated, with ML-Relate (Kalinowski et al. 2006).
To evaluate genetic contemporary bottlenecks we used two approaches, a graphical method based on the distribution of allele frequencies in which recently bottlenecked populations are likely to have lost mainly the rare alleles (Luikart and Cornuet 1998). We also tested for heterozygote excess with the program Bottleneck v.5.1.26 (Cornuet and Luikart 1996; Piry et al. 1999) by comparing observed and expected heterozygosity values estimated under three mutational models: infinite alleles (IAM), stepwise mutation (SMM) and two-phase (TPM) with two variants (90% SMM, 10% IAM and 10% variance; 70% SMM, 30% and 10% variance). Models were run with 10,000 replicates and significance was calculated with a Wilcoxon test.
Differentiation and migration
Pairwise genetic differentiation between genetic clusters for both species was estimated with FST and RST with Arlequin v.3.1 (Excoffier et al. 2005). Ancestral migration was assessed with an indirect estimator of gene flow (Nm), based on the allelic differences between genetic clusters (FST), with the formula 4Nm = (1/FST). The number of migrants was estimated both by individuals and by sexes with Arlequin v.3.1. This model assumes a constant population size and symmetric migration rates (Wilson and Rannala 2003). We also estimated contemporary bidirectional migration rate using Bayesass v.3.0. (Wilson and Rannala 2003), with 5 × 106 interactions, 3 × 105 burn-in, and sample rate of 2,000. The delta value was adjusted with various trials for each species, based on which we chose values of 0.25 for the black rat and 0.20 for the house mouse. Chain convergence was evaluated with the Faubet method (Faubet et al. 2007), which calculates the Bayesian deviation based on the posterior probabilities that a certain genotype is possible under the ancestral migration rate and the posterior probability of allocation given the same migration rate (Faubet et al. 2007; Meirmans 2014).
Mus musculus colonization patterns
For our objective to evaluate the colonization patterns and connectivity for the house mouse and explore the landscape genetics of the species, we specifically assessed if the urbanization growth pulses that occurred in San Miguel Cozumel determined the observed genetic structure and genetic diversity changes in the three genetic clusters obtained (SanMi1, SanMi2, SanMi3; see Results). First, we inferred the time, expressed as number of generations, and patterns of colonization with the Approximate Bayesian computation method (ABC) (Beaumont et al. 2002), implemented in DiyABC v2.04 (Cornuet et al. 2014). We evaluated three simple scenarios built based on the history of human population presence on Cozumel, using datasets simulated by coalescent from prior distribution parameters (Bertorelle et al. 2010): (i) Scenario1, we hypothesized a sequential colonization where SanMi1 is the ancestral (introduced) Cozumel population, from which SanMi2 originated and then SanMi3 from this one; (ii) Scenario2, we tested if SanMi2 and SanMi3 originated either as two independent populations from SanMi1 or from two colonization pulses at different times; and (iii) Scenario3, we evaluated if SanMi1 and SanMi3 originated independently from the same unsampled introduced population that colonized Cozumel, while SanMi2 originated from SanMi1 (Fig. 2a). For all introduction and colonization events we assumed short genetic bottlenecks. The house mouse has a generational time of 2–3 months in captivity, but in the wild it has been observed to have on average two litters per year, depending on resource availability (Nachman and Searle 2014; Phifer-Rixey and Nachman 2015).
a Scenarios tested with DiyABC v.2.04 (Cornuet et al. 2014) for the house mouse in San Miguel Cozumel. Scenario 1: sequential colonization where SanMi1 is the ancestral (introduced) population-SanMi2-SanMi3; Scenario 2: SanMi2 and SanMi3 originated either as two independent populations from SanMi1 or from two colonization pulses at different times; Scenario3: SanMi1 and SanMi3 originated independently from the same unsampled introduced population, and SanMi2 originated from SanMi1. b Scenario model comparison performance considering the direct and logistic approaches. c Posterior predictive test of model fit corresponding to Scenario 1
We defined the priors of the divergence time between populations based on the elapsed times between the historical dates when the island was abandoned around 1570 and when people again established in 1846–1847 until 2015 and multiplied by the number of generations in one year. Hence, we defined priors accordingly for introduction of SanMi1, between 336 and 1008 generations. In 1980 Cozumel experienced a first population growth that promoted further urbanization, based on which the prior for SanMi2 was set between 70 and 210 generations; finally, another population growth wave started in 2000, so the prior for SanMi3 was between 30 and 90 generations (see below). We modeled the effective population size applying the default values (Brouat et al. 2014; Cornuet et al. 2014) and applied the generalized stepwise mutation model for microsatellite data (Estoup et al. 2012). We generated 6 × 106 simulated datasets per scenario, and scenarios were compared with the logistic and direct approaches using the following summary statistics: mean of number of alleles, genetic diversity and size variance within and between populations (Estoup et al. 2012; Suárez-Atilano et al. 2019). Confidence in the most supported scenario was evaluated by modeling 500 pods (pseudo-observed data) from the prior distribution to estimate the false allocation rate, where Type I error (i.e. probability in which the true scenario is rejected) is the proportion of pods for which the scenario does not have the highest posterior probability; and Type II error (i.e. probability of accepting a scenario when it is not true), is the proportion of pods of which the scenario with the highest posterior probability is not the scenario evaluated (Estoup et al. 2012). Based on the number of generations that coalesce supported by the best model (see Results), we estimated the dates of colonization using 4 and 6 years for the rodent’s generation time, considering that high food availability favours a high reproductive rate.
Landscape genetics and connectivity
We generated environmental and anthropogenic variables at a resolution of 30 × 30 m. For the environmental variables, we used Landsat 8 images to calculate the normalized vegetation index (NDVI) in QGis 3.8, where low NDVI values indicate impervious land surface (i.e. man-made constructions, roads; Zhang and Ye 2021). For the anthropogenic variables we used the National Housing Inventory 2015 data (INEGI 2015) to generate three indices that measure marginalization, overcrowding and commercial businesses. The marginalization index was based on the number of essential services to which the population has access (concrete floors, running water, sewer system, sanitation services and street lighting); the overcrowding index represents the percentage of housing where more than three people share a room; and the commercial businesses denote the percentage of neighborhoods that have informal sector businesses in none, some, or all of their surrounding roads.
To obtain a measure of genetic differentiation for the house mouse across its sampled distribution, we estimated the proportion of shared alleles, an individual-based genetic distance that has no biological assumptions and can be used for inbreeding populations (Shirk et al. 2017), using adegenet. To generate resistance values for our environmental and anthropogenic surfaces we followed the optimization framework developed by Peterman (Peterman et al. 2014), which uses maximum-likelihood population effects mixed models. We optimized each resistance surface with the function commuteDistance in gstudio v.1.5.2 in R (Dyer 2014), with three independent runs to verify parameters convergence. We determined the best-supported features by AICc (Akaike’s information criterion corrected for small/finite sample size; Akaike 1974). After running each surface individually, we applied a multivariate approach for which we built a set of hypotheses considering socioeconomic features most likely relevant for the distribution of the house mouse (Table 1).
Results
A total of 48 black rat individuals (25 male and 17 females; six undetermined) and 59 house mice (37 males and 17 females; five undetermined) were captured (Table S1). The microsatellite locus D5Rat83 in black rats was monomorphic and was excluded from analyses. There was no evidence of linkage disequilibrium in the black rat while in the house mouse it was only significant when the entire island was evaluated as a population (D5Mit149-D9Mit54 and D5Mit149-PP7B08) and for the genetic cluster SanMi2 (D13Mit61-EG22992). Five loci for black rats (Rr14, Rr17, Rr22, Rr68, Rr107) and four for house mice (D5Mit149, D15Mit98, PP7B08, PP10E08) deviated significantly from Hardy–Weinberg equilibrium after Bonferroni correction. Only Rr14 in black rats and eight loci in house mice (except D6Mit309 and D13Mit61) showed evidence of null alleles, though no locus was shared between genetic clusters, thus all loci were considered for further analyses. Genepop files for the black rat (Table S3) and the house mouse (Table S4) microsatellite genotypes are included in Supplementary information.
Genetic patterns
Geneland results identified three genetic clusters (K = 3) for the black rat (Fig. 3), which correspond to San Miguel Cozumel (Coz1), the sanitary landfill and El Cedral (Coz2), and the hotel development on the southwest (Coz3); in contrast, with the DAPC we found two distinctive groups, where Coz2 and Coz3 form a single cluster. All subsequent analyses for the black rat considered the three genetic clusters from Geneland. For the house mouse both Geneland and DAPC identified three clusters, all distributed within San Miguel Cozumel, where SanMi1 is located on the northeast and includes the sanitary landfill, SanMi2 in the northwest, and SanMi3 in the south (Fig. 4). Genetic diversity results showed a total of 56 alleles for the black rat (Table 2), with an average of 6.5 alleles per locus and moderate heterozygosity (0.601). Coz3 was the cluster with less genetic diversity. Regarding the house mouse, diversity was markedly higher with a total of 100 alleles (10 per locus), where SanMi3 exhibited the lowest diversity values (Table 2). All clusters for both species showed positive FIS values indicating inbreeding. Notwithstanding, relatedness results showed a high percentage of unrelated individuals (83.4% and 86.7%, black rat and house mouse respectively) (Table 2). The ancestral bottleneck test was not significant under any mutational model, while the contemporary bottleneck analysis showed rare alleles in low proportion for Coz3 and SanMi3, suggesting a bottleneck signal or a result of low sample size (Table S5, Fig. S1).
a Venn diagram for the black rat showing private and shared alleles between genetic clusters identified with Geneland v.4.0.4 (Guillot et al. 2005). b Scatter plot based on the Geneland clusters as priors and the a-score plot used to determine the number of PCs (1 in this case) and discriminant functions to retain for the DAPC. c Map of the study site showing the distribution of the three genetic clusters considering the isocline = 0.5. d Posterior probability maps belonging to each Geneland genetic cluster where yellow colors depict the high probability isoclines
a Venn diagram showing the house mouse private and shared alleles between genetic clusters identified with Geneland v.4.0.4 (Guillot et al. 2005). b Map of the study site showing the distribution of the three genetic clusters considering the isocline = 0.5. c Scatter plots based on the Geneland clusters as priors and the a-score plot used to determine the number of PCs (8 in this case) and discriminant functions to retain for the DAPC. d Posterior probability maps belonging to each Geneland genetic cluster where yellow colors depict the high probability isoclines
Ancestral and contemporary migration
Pairwise genetic differentiation between clusters was moderate to high (black rat FST = 0.107–0.309, RST = 0.086–0.52; house mouse FST = 0.094–0.184, RST = 0.122–0.274 p < 0.05) (Fig. 5a); the higher differentiation was observed between Coz2-Coz3 and SanMi1-SanMi3, respectively. Ancestral migration showed the highest exchange of individuals between Coz1-Coz2 (2.084) and SanMi1-SanMi2 (2.39) (Fig. 5b). Distinct patterns were observed for each species regarding differential migration between sexes, where black rat females disperse more and, contrastingly, house mouse males showed higher migration than females (Fig. 5a). Whereas contemporary bidirectional migration results showed extremely low migration rates in black rats where at least 95% of individuals are permanent residents; the same was observed for house mice, where only the SanMi2 showed a comparatively lower permanent resident rate (0.68), while receiving a high proportion of migrants from SanMi3 (0.30) (Fig. 5c).
a Pairwise genetic differentiation between genetic clusters based on FST (red) and RST (blue) estimators for the house mouse and the black rat. b Ancestral migration rates (Nm) based on the allelic differences between genetic clusters (FST). The thickness of the lines and arrows proportional to the intensity of migration between clusters. Color lines indicate the values for females (pink), males (blue), and global (black). c Bidirectional contemporary migration rates estimated with Bayesass v.3.0 (Wilson and Rannala 2003); proportion of non-migrant individuals by generation per genetic cluster (within the ovals) and direction of migrant individuals by generation between genetic clusters (next to each arrow) are shown
House mice colonization patterns and landscape connectivity
Model comparisons resulting from the Approximate Bayesian computation showed that Scenario1, in which the sequential colonization was tested, had the highest probability based on both the direct (P = 0.468, 95% confidence interval (CI) 0.031–0.905) and the logistic (P = 0.964, (CI) 0.929–0.994) approaches (Fig. 2b). The confidence test for the best scenario showed that type 1 error was 0.134 for the logistic approach. We performed a pre-evaluation of simulated scenarios via PCA, and results indicated that the observed data fall within the cluster of simulated data points. In addition, the goodness-of-fit test using the posterior predictions exhibited that summary statistics from the simulated datasets based on Scenario 1 closely matched the empirical data (Fig. 2c). Hence, based on the best model results and the generation times (4–6 reproductive events by year), we estimated the date of colonization events; accordingly, SanMi1 arrived at the island 676 generations ago (1846–1902), the divergence between SanMi1 and SanMi2 occurred 247 generations ago (1954–1974), and SanMi3 was founded 93 generations ago (1991–1999).
Results of model selection of the optimization resistance surfaces (environmental and anthropogenic) showed that the best-supported univariate model was geographic distance, identified as the top model 99% of the time, followed by NDVI (Table 3). The optimized NDVI surface assigned high resistance at values above zero that correspond to vegetated areas, and low resistance to bare landscape or urbanizations (Fig. S2). The linear mixed-effects models test of the composite resistance hypotheses showed that the best-supported multivariate model was Development (100% of the time; Table 4), where NDVI contributed the most to the model (88%).
Discussion
Genetic diversity, invasion models and the genetic paradox of biological invasion
As a result of the reduced number of individuals with which introduced populations are established (founder effect), populations are expected to exhibit low genetic diversity, an effect that is theoretically stronger on islands (Frankham 1999, 2005). However, in most invasive species this does not happen, which is known as the genetic paradox of biological invasion (Frankham 2005; Lawson et al. 2011; Estoup et al. 2016). In agreement with this paradox, the black rat showed moderate genetic diversity levels (Ho = 0.44, He = 0.59, Na = 6.5), with higher levels in the house mouse (Ho = 0.56, He = 0.76, Na = 10). These values are within the range found in other islands, like the black rat in Guadeloupe (He = 0.69, Na = 5), Madagascar (He = 0.72) and New Zealand (He = 0.75, Na = 5.7) (Abdelkrim et al. 2005a, 2005b, 2010; Brouat et al. 2014), as well as the house mouse on islands of New Zealand (He = 0.51–0.56), Kerguelend (He = 0.40, Na = 2.8), Madagascar (He = 0.67, Na = 9.7) and Cyprus (He = 0.77, Na = 9.7) (Hardouin et al. 2010, 2015; Pichlmueller et al. 2020), despite the comparatively small size of Cozumel island. Interestingly, continental populations, some within cities, show similar diversity values (Konecny et al. 2013; Mangombi et al. 2016; Stragier et al. 2022).
Such levels of genetic diversity can be associated with the historical and demographic processes experienced by both the source and founder populations (Estoup and Guillemaud 2010), namely the likely short ancestral bottleneck supported by our analyses, followed by rapid population growth, and coupled with multiple introduction events facilitated by the constant arrival of commercial and, more recently, cruise ships. In fact, a high number of private alleles as observed in the Cozumel populations may be the product of constant introductions of individuals of different origin, incorporating different genotypes into the genetic pool of the resident population and increasing allelic diversity (Estoup and Guillemaud 2010; Lawson et al. 2011). This incorporation of new genotypes is favored by the ‘invasive meltdown model’, which establishes that ecosystems are more easily invaded as they are modified by previously arrived invasives, increasing population density (Simberloff and Von Holle 1999; Prenter et al. 2004; Braga et al. 2018). In addition, the ‘invasive bridgehead effect’ refers to secondary invasions stemming from a particularly successful invasive population, which serves as a source of colonists for potentially new territories, increasing their probability of successful establishment (Estoup and Guillemaud 2010; Lawson et al. 2011). These scenarios apply for most human settlements, as is the case in Cozumel, where the likely continued introduction of individuals of different geographic sources into the island entails a strong propagule pressure, rendering high allelic diversity and the effective establishment of migrants. Similar examples include the house mouse in the Azores islands, which shows high allelic diversity resulting from multiple origins identified from southern Spain, Portugal, Finland and the Canary Islands (Gabriel et al. 2015).
Independent colonization events from different lineages can be another factor influencing allelic differentiation and structure, as shown by the house mouse in the Kerguelen archipelago and the Canary Islands, where the observed genetic diversity derives from the invasion of two distinct lineages (López et al. 2013; Hardouin et al. 2015). Likewise, the black rat in Madagascar comprises two genetic groups product of two independent introduction events from the same ancestral population (Brouat et al. 2014). Our results do not support an independent colonization origin for the Cozumel clusters; instead, more recent social and demographic processes may also be contributing to the currently observed genetic diversity and structure.
Genetic structure, urbanization and human population growth
As we predicted, the two rodent species exhibit a clear genetic structure. In our fine-scale genetic study, the three clusters identified for the black rat correspond with three areas of the island with different anthropogenic impact (Fig. 3), Coz1 comprises San Miguel Cozumel, Coz2 on the south of the island includes the sanitary landfill and the small El Cedral town; and Coz3 is located on the southwestern hotel zone, adjacent to the city, characterized by a combination of highly developed infrastructure surrounded by partially preserved forest remnants. These structure patterns contrast with what has been observed in other studies, for instance black rats did not show genetic differentiation in the Puketi Forest, New Zealand despite the marked habitat differentiation and significant geographic distances between samples sites (Abdelkrim et al. 2010), neither in Franceville, Gabon, a small African tropical city that has undergone a recent growth, regardless of the composite landscape including urban forest remnants and savannah (Mangombi et al. 2016).
Three genetic groups were also identified for the house mouse, tightly differentiated within San Miguel Cozumel (although SanMi1 also includes the sanitary landfill; Fig. 4). These genetic clusters are limited both by roads and high traffic avenues and by the margins of the urbanized area. There are numerous examples in which avenues or highways constitute important barriers, for example in Salvador, Brazil where gene flow of the Norway rat is limited by a main avenue (Richardson et al. 2017). Furthermore, the house mouse genetic structure is likely associated with the human population growth pulses accompanying the urbanization of San Miguel Cozumel. The spatial distribution of SanMi1 coincides with the limits of the city before 1970, when the tourism development began, and the first large-scale contemporary human migration took place. Also, the site where the sanitary landfill is currently located was initially an open-air dump, thus the constant transport of waste trucks from the city to the dump prior to 1970 (Trejo de la Paz 2014) can explain why the SanMi1 cluster extends to this area. As the urban sprawl grew, the house mouse was able to disperse and establish new populations, so that SanMi2 includes all the new neighborhoods built between 1970 and 2000. Last, SanMi3 is distributed in the neighborhoods most recently established since 2000 onwards. Outstandingly, this relationship between the species genetic structure and urban growth coincides with the ancestral migration rate; that is, gene flow between the oldest clusters (SanMi1 and SanMi2) is greater compared to that between them and the most recent cluster (SanMi3). This sequential pattern also matches the genetic diversity of each cluster. Indeed, the ABC analysis supported our Scenario1 hypothesis of sequential colonization, where SanMi1 is the ancestral Cozumel population, which harbors the highest genetic diversity; from which SanMi2 originated and, more recently, SanMi3 that has the lowest genetic diversity. Loss of genetic variability through sequential colonization routes due to successive bottlenecks is frequent, as evidenced by the house mouse invasion in Senegal (Lippens et al. 2017).
The demographic growth of Cozumel is divided in two phases, a stationary one since the recolonization until 1950, and the tourism expansion from 1950 onwards, characterized by a constant population growth due of the intensification of human migration processes (Sánchez-Crispín and Propin 2003; Palafox and Zizumbo 2009). In agreement with the spatial location of the genetic clusters, our best demographic model indicated that the initial colonization of Cozumel Island occurred in SanMi1 between 1846 and 1902. In 1846, when the Mayan farming refugees from the east of the peninsula moved to Cozumel and founded San Miguel and El Cedral. The arrival of Mayan people to the island continued until 1913, thus these historical processes could be linked to the initial introduction of both invasive rodents. The origins of SanMi2 occurred between 1954 and 1974 coinciding with the initial urbanization pulse driven by tourism. Diving and fishing tourism were promoted in 1950 but is not until 1970 that a trust is created to promote tourist and economic development, initiating the construction of the northern hotel zone coupled with a strong demographic growth (Santander and Ramos-Díaz 2011). Finally, the colonization of SanMi3 occurred between 1991 and 1999, when Cozumel joined the Caribbean cruise network (Trejo de la Paz 2014).
With this in mind and considering that Cozumel is the main port of the Mexican Caribbean, the high number of exclusive alleles in the different genetic clusters is highly relevant. Coz1 had the highest number of private alleles (17) for black rats, and SanMi1 (34) and SanMi2 (20) for house mice. A note of caution is that the low sample sizes for Coz3 and SanMi3 might underestimate the number of private alleles. Nonetheless, the clusters with highest number of private alleles occur in strategic areas of commercial exchange and tourism, supporting the hypothesis of a constant introduction of individuals to Cozumel. SanMi1, located in the oldest city square, also includes the maritime terminal (Muelle fiscal) and the cargo ferry port, the main passenger and commercial connections between the mainland and the island until the 1980s, when it started to receive cruises and two grand cruise piers were built, Puerta Maya (1997) and Punta Langosta (2000) (PRORED 2008), the latter also within SanMi1. Puerta Maya is adjacent to the southwestern hotel zone where SanMi3 and Coz3 are situated (Fig. 1). Further phylogeographic and genetic assessments are needed to more finely reconstruct the routes of invasion of these rodents into Cozumel, but we hypothesize that Coz1 and SanMi1 share a genetic ancestry related to mainland individuals, particularly from Playa del Carmen that has historically been the region with the highest interchange with the island. More recent introductions would certainly include lineages from the various Caribbean ports that constitute the main Caribbean cruise route, i.e. Belize, Roatán (Honduras), Limón (Costa Rica), Colón (Panama), Santa María (Curaçao), Montego, Falmouth, Ocho Rios (Jamaica), and Grand Cayman ports (Hernández-González and Villaseñor-Franco 2020).
The lack of ancestral and contemporary bottleneck signals in most of the genetic clusters for both species suggests a brief bottleneck after the introduction, a common pattern in species with high reproductive rate, short generation time and strong propagule pressure. In addition, the continuous introduction of individuals, from the same or different geographic origin, contributes to erasing the initial bottleneck signal and increases the global genetic diversity (Suárez-Atilano et al. 2019). The house mouse can establish rapidly, as evidenced by the experimental introduction of two founders (one female and one male) on Saddle Island, New Zealand where the system’s load capacity was reached in only six months (Nathan et al. 2015). Likewise, the Norway rat could rebound population sizes after strong bottlenecks (Richardson et al. 2019). Yet, contemporary bottleneck signals were detected in SanMi3 and Coz3. As mentioned, the colonization of SanMi3 is relatively recent, thus likely still echoing the bottleneck signal, whereas such signal in Coz3 (located along the southwestern hotel zone) can be associated with the permanent pest control carried out by hotels, causing population size reductions and decreasing genetic variation. Indeed, significant loss of genetic variation related to control strategies has been documented in Poirier Island and in Salvador, Brazil, because of an eradication plan for black rats (Abdelkrim et al. 2010) and after urban rat control campaigns (Richardson et al. 2019), respectively. However, this would require further assessment because the SanMi3 and Coz3 bottleneck signals could also be associated with their low sample size.
Ancestral and contemporary migration patterns
The dispersal capacity of a species depends, among others, on body size, morphology and physiology, as well as on biotic and abiotic factors like resource availability, habitat quality and geographic barriers. Dispersal can also vary between sexes as seen in birds and mammals, where the differential response of males and females relies on population density, competitive ability and habitat quality, among others, as a strategy to avoid inbreeding (Globert et al. 2012).
Rattus species are considered mostly philopatric, i.e. tending to return to or remain near a particular site or area, observed in females as it is more common (Gardner-Santana et al. 2009). In our study in Cozumel, the black rat showed differential ancestral migration with higher values in females, which could be associated with their social structure, where males have a strong hierarchy and control territories through which the females ‘freely’ roam; in fact, female dispersion is food-determined and shapes male dispersion (Hooker and Innes 1995). Accordingly, in the past females could have dispersed to sites with high resource availability, in agreement with the high gene flow between Coz1 and Coz2. However, contemporary gene flow showed extremely low migration between clusters. If the system is saturated (e.g. low resources, high competition) that would prevent the arrival of new migrating individuals. Only Coz3 receives some migrants from both Coz1 and Coz2, which could be explained by the population control measures implemented by hotels in this area, temporarily reducing competition for resources and allowing new migrants to arrive.
House mouse individuals tend to be nomads if population densities are low, but at high densities males are territorial (Hulme-Beaman et al. 2016). This behavior patterns might have facilitated the presence of the three genetic groups within San Miguel Cozumel, with overall low ancestral migration. Contrary to the black rat, migration in the house mouse was higher for males, concordant with their social system where dominant males control and monopolize feeding sites, shared with several females and some subordinate males (Boursot et al. 1996; Dean et al. 2006). However, because population densities are usually high, such social hierarchies do not last long and males migrate constantly. As a result, populations behave panmictically on a regional scale but are structured on a finer scale (Morgan et al. 2022). Notably, as females are philopatric and do not move freely between territories, mitochondrial DNA-based studies have revealed that populations can be resistant to secondary female invasions (Gabriel et al. 2015), also explaining the observed low migration rate in this species.
Urban drivers of connectivity in the house mouse
Connectivity analyses allow identifying key landscape elements needed to maintain genetic diversity and evolutionary processes. In commensal species they also help to elucidate dispersal and distribution patterns of invasive species (Pichlmueller et al. 2020; Stragier et al. 2022). The most significant environmental feature influencing house mice connectivity was geographic distance, while the next best supported model was vegetation (NDVI). High and continuous vegetation cover is a good predictor of connectivity in natural environments, for example in rodents like Peromyscus melanotis (Borja-Martínez et al. 2022), and in cities with non-commensal species like Peromyscus leucopus (Munshi-South 2012; Munshi-South and Nagy 2014) and the lizard Podarcis muralis (Beninde et al. 2018).
Contrastingly, as we predicted for this commensal species, we found high connectivity for the house mice along impervious land surfaces, i.e. the urban landscape, indicated by the low NDVI values recovered by the optimization surfaces. Similarly, Stragier et al. (2022) found low gene flow in this species associated with highly vegetated areas in Dakar. Additionally, Fusco et al. (2021) highlight that the variables analyzed in urban landscapes under a landscape genetics approach generally include human population density, roadways, infrastructure, impervious surfaces and industrial development, while social, cultural or political factors are rarely used. Indeed, incorporating socioeconomic variables was key in our approach. The best-ranked model in the multivariate analysis was our Development hypothesis, which considers the interaction between NDVI and Commercial businesses, further supporting that house mice connectivity is tightly linked with urbanization. The information used to build the Commercial businesses layer included all the informal commerce across the city, most of which is dedicated to street food selling. Since they produce abundant waste, we used this as a proxy of resource availability for the house mouse. Interestingly, Combs et al. (2017) show that low food waste availability decreased the genetic diversity and gene flow in another invasive rat, the Norway rat in New York.
Genetic information and invasive rodent management
The lack of biosecurity measures to prevent the arrival of invasive species to Cozumel is evidenced by our results suggesting a constant introduction of invasive rodents. This, together with the environmental and social complexities of large, inhabited islands, make rodent control (sustained reduction), rather than eradication (complete removal, feasible on smaller islands) realistic (Holmes et al. 2023).
Information regarding genetic clusters can help defining meaningful spatial units for control management (Combs et al. 2018; Gatto-Almeida et al. 2022). Indeed, we detected different genetic groups and low connectivity for each species, determined by urban and socioeconomic features. Accordingly, we located areas that could be targeted with different management strategies. Constant monitoring should be performed in high-risk locations like piers and docks (both off and on island), and in urban sites (sanitary landfill, informal commerce); routinely examining the terrestrial and maritime transport vehicles should also be implemented. Importantly, the significant association we identified of the house mouse with impervious surfaces likely limits the chances of a secondary invasion to natural environments. We highlight that in our studies of rodent population ecology and genetics on Cozumel, performed in natural vegetation and conserved areas over many years, we have not trapped black rats nor house mice (see Vega et al. 2007; Fuentes-Montemayor et al. 2009; Espindola et al. 2014). Whether these invasive rodents are indeed absent from natural areas or present in low abundance is not clear. Nonetheless, improving urban planning to regulate future expansions of San Miguel Cozumel is of the outmost importance to prevent these invasive species to disperse further on the island. Likewise, it is imperative to make authorities and people aware of the negative effects of these invasives on the native biota and of the associated high potential health risks.
Data availability
All data used in the present study is provided within the manuscript and in the Supplementary Information.
References
Abdelkrim J, Pascal M, Samadi S (2005a) Island colonization and founder effects: the invasion of the Guadeloupe islands by ships rats (Rattus rattus). Mol Ecol 14:2923–2931. https://doi.org/10.1111/j.1365-294X.2005.02604.x
Abdelkrim J, Pascal M, Samadi S (2005b) Establishing causes of eradication failure based on genetics: case study of ship eradication in Ste Anne Archipelago. Conserv Biol 21:719–230. https://doi.org/10.1111/j.1523-1739.2007.00696.x
Abdelkrim J, Byron AE, Gemell NJ (2010) Fine-scale genetic structure of mainland invasive Rattus rattus populations: implications for restauration of forested conservation areas in New Zealand. Conserv Genet 11:1953–1969. https://doi.org/10.1007/s10592-010-0085-9
Akaike H (1974) A new look at the statistical model identification. IEEE Trans Autom Control 19:716–723. https://doi.org/10.1109/TAC.1974.1100705
Aplin KP, Suzuki H, Chinen AA, Chesser RT, Ten-Have J, Donellan SC et al (2011) Multiple geographic origins of commensalism and complex dispersal history of black rats. PLoS ONE 6:11. https://doi.org/10.1371/journal.pone.0026357
Babiker H, Tautz D (2015) Molecular and phenotypic distinction of the very recently evolved insular subspecies Mus musculus helgolandicus Zimmermann 1953. BMC Evol Biol 15:160. https://doi.org/10.1186/s12862-015-0439-5
Banks PB, Smith HM (2015) The ecological impact of commensal species: black rats, Rattus rattus, at the urban-bushland interface. Wildlife Res 42:86–97. https://doi.org/10.1071/WR15048
Bastos AD, Nair D, Taylor PJ, Brettshcneider H, Kirsten F, Monsert E et al (2011) Genetic monitoring detects and overlooked cryptic species and reveals the diversity and distribution of three invasive Rattus congeners in South Africa. BMC Genet 12:26. https://doi.org/10.1186/1471-2156-12-26
Beaumont MA, Zhang W, Balding DJ (2002) Approximate Bayesian computation in populations genetics. Genetics 162:2025–2035. https://doi.org/10.1093/genetics/162.4.2025
Beninde J, Feldmeler S, Veith M, Hochkirch A (2018) Admixture of hybrid swarms of native and introduced lizards in cities is determined by the cityscape structure and invasion history. Proc R Soc B 285(1883):20180143. https://doi.org/10.1098/rspb.2018.0143
Bertorelle G, Benazzo A, Mona S (2010) ABC as a flexible framework to estimate demography over space and time: some cons, many pros. Mol Ecol 19:2609–2625. https://doi.org/10.1111/j.1365-294X.2010.04690.x
Blackburn TM, Pysek P, Bacher S, Carlton JT, Duncan RP, Jarosik V et al (2011) A proposed unified framework for biological invasions. Trends Ecol Evol 26:333–339. https://doi.org/10.1016/j.tree.2011.03.023
Borja-Martínez G, Tapia-Flores D, Shafer ABA, Vázquez-Domínguez E (2022) Highland forest’s environmental complexity drives landscape genomics and connectivity of the rodent Peromyscus melanotis. Landsc Ecol 37:1653–1671. https://doi.org/10.1007/s10980-022-01428-6
Boursot P, Din W, Anand R, Darviche D, Dod B, Von Deimling F et al (1996) Origin and radiation of the house mouse: mitochondrial DNA phylogeny. J Evol Biol 9:391–415. https://doi.org/10.1046/j.1420-9101.1996.9040391.x
Braga RR, Gómez-Aparicio L, Heger T (2018) Structuring evidence for invasional meltdown: broad support but with biases and gaps. Biol Invas 20:923–936. https://doi.org/10.1007/s10530-017-1582-2
Brouat C, Tollenaere C, Estoup A, Loiseau A, Sommer S, Soanandrasana R et al (2014) Invasion genetics of a human commensal rodent: the black rat Rattus rattus in Madagascar. Mol Ecol 23:4153–4167. https://doi.org/10.1111/mec.12848
Calmet C, Pascal M, Samadi S (2001) Is it worth eradicating the invasive pest Rattus norvegicus from Molene archipelago? Genetic structure as a decision making tool. Biodiv Conserv 10:911–928. https://doi.org/10.1023/A:1016636512734
Caut S, Angulo E, Courchamp F (2008) Dietary shift of an invasive predator: rats, seabirds and sea turtles. J Appl Ecol 45:428–437. https://doi.org/10.1111/j.1365-2664.2007.01438.x
Clair JJH (2011) The impacts of invasive rodents on island vertebrates. Biol Conserv 144:68–81. https://doi.org/10.1016/j.biocon.2010.10.006
Combs M, Puckett EE, Richardson J, Mims D, Munshi-South J (2017) Spatial populations genomics of the brown rat (Rattus norvegicus) in New York City. Mol Ecol 27:83–98. https://doi.org/10.1111/mec.14437
Combs M, Byers KA, Ghersi BM, Blum MJ, Caccone A, Costa F et al (2018) Urban rat races: spatial population genomics of brown rats (Rattus norvegicus) compared across multiple cities. Proc R Soc B 285(1880):20180245. https://doi.org/10.1098/rspb.2018.0245
Cornuet JM, Luikart G (1996) Description and power analysis of two test for detecting recent population bottlenecks from allele frequency data. Genetics 144:2001–2014
Cornuet JM, Pudio P, Veyssier J, Dehne-García A, Gautier M, Leblois R et al (2014) DIYABC v2.0: a software to make approximate Bayesian computation inferences about population history using single nucleotide polymorphism. DNA Seq Microsatellite Data Bioinform 30:1187–1189. https://doi.org/10.1093/bioinformatics/btt763
Cuarón AD (2009) Cozumel. In: Gillespie R, Clague DA (eds) Encyclopedia of Islands. University of California Press, California, pp 203–206
Dean MD, Ardlie KG, Nachman MW (2006) The frequency of multiple paternity suggests that sperm competition is common in house mouse (Mus domesticus). Mol Ecol 15:4141–4151. https://doi.org/10.1111/j.1365-294X.2006.03068.x
Des Roches S, Brans KI, Lambert MR, Rivkin LR, Savage AM, Schell CJ et al (2021) Socio-eco-evolutionary dynamics in cities. Evol Appl 14:248–267. https://doi.org/10.1111/eva.13065
Dietrich W, Katz H, Lincoln SE, Shin HS, Friedman J, Dracopoli NL, Lander E (1992) A genetic map of the mouse suitable for typing intraspecific crosses. Genetics 131:423–447. https://doi.org/10.1093/genetics/131.2.423
Donihue CM, Lambert MR (2015) Adaptative evolution in urban ecosystems. Ambio 44:194–203. https://doi.org/10.1007/s13280-014-0547-2
Duglosh KM, Parker IM (2007) Founding events in species invasions: genetic variation, adaptative evolution and the role of multiple introductions. Mol Ecol 17:431–449. https://doi.org/10.1111/j.1365-294X.2007.03538.x
Dyer RJ (2014) An R package for the spatial analysis of population genetic data. R package version 1.5.2 https://dyerlab.github.io/gstudio/
Espindola S, Gaggioti O, Cuarón AD, Vázquez-Domínguez E (2014) High genetic structure in the Cozumel Harvest mice, an endangered island endemic: conservation implications. Conserv Genet 15:393–1402
Estoup A, Guillemaud T (2010) Reconstructing routes of invasion using genetic data: Why, how and so what? Mol Ecol 19:4113–4130. https://doi.org/10.1111/j.1365-294X.2010.04773.x
Estoup A, Lombart E, Marin JM, Guillemaud T, Pudlo P, Robert CP, Cornuet JM (2012) Estimation of demo-genetic model probabilities with Approximate Bayesian Computation using linear discriminant analysis on summary statistics. Mol Ecol Res 12:846–855. https://doi.org/10.1111/j.1755-0998.2012.03153.x
Estoup A, Ravigné V, Hufbauer R, Vitalis R, Gautier M, Facon B (2016) Is there a genetic paradox of biological invasion? Ann Rev Ecol Evol Syst 47:51–72. https://doi.org/10.1146/annurev-ecolsys-121415-032116
Excoffier L, Laval G, Schneider S (2005) Arlequin (version 3.0): an integrated software package for population genetic data analysis. Evol Bioinform Online 2005:1
Faubet P, Waples RS, Gaggiotti OE (2007) Evaluating the performance of multilocus Bayesian method for the estimation of migration rates. Mol Ecol 16:1149–1166
Flores-Manzanero A, Valenzuela-Galván D, Cuarón AD, Vázquez-Domínguez E (2022) Conservation genetics of two critically endangered island dwarf carnivores. Conserv Genet 23:35–49
Frankham R (1999) Inbreeding and extinction: Island population. Conserv Biol 12:665–675. https://doi.org/10.1111/j.1523-1739.1998.96456.x
Frankham R (2005) Resolving the genetic paradox in invasive species. Heredity 94:385. https://doi.org/10.1038/sj.hdy.6800634
Fuentes-Montemayor E, Cuarón AD, Vázquez-Domínguez E, Benítez-Malvido J, Valenzuela D, Andresen E (2009) Living on the edge: roads and edge effects on small mammal populations. J Anim Ecol 78:857–865. https://doi.org/10.1111/j.1365-2656.2009.01551.x
Fusco NA, Carlen EJ, Munshi-South J (2021) Urban landscape genetics: Are biologists keeping up with the pace of urbanization? Curr Lands Ecol Rep 6:35–45. https://doi.org/10.1007/s40823-021-00062-3
Gabriel S, Matías ML, Searle JB (2015) Of mice and the age of Discovery: the complex history of colonization of the Azorean archipelago by the mouse (Mus musculus) revealed by mitochondrial DNA variation. J Evol Biol 28:130–145. https://doi.org/10.1111/jeb.12550
Gaertner M, Wilson JRU, Cadotte MW, MacIvor JS, Zenni RD, Richardson DM (2017) Non-native species in urban environments: patterns, processes, impacts and challenges. Biol Invas 19:3461–3469. https://doi.org/10.1007/s10530-017-1598-7
Gardner-Santana LC, Norris DE, Fornadel CM, Hinson ER, Klein SL, Glass GE (2009) Commensal ecology, urban landscapes, and their influence on the genetic characteristics of city-dwelling Norway rats (Rattus norvegicus). Mol Ecol 18:2766–2778. https://doi.org/10.1111/j.1365-294X.2009.04232.x
Gatto-Almeida F, de Araújo SA, Degrandi TM, Tiepolo LM, Pichlmueller F, Hass I (2022) Assessment of dispersal and population structure of Norway rats (Rattus norvegicus) in a seaport setting. Urban Ecosyst 25:535–544. https://doi.org/10.1007/s11252-021-01171-x
Globert J, Baguette M, Benton TG, Bullock JM (2012) Dispersal ecology and evolution. Oxford University Press, Oxford, UK
Gortat T, Rutkowski R, Gryczynska A, Kozakiewicz A, Kozakiewicz M (2017) The spatial genetic structure of the yellow-necked mouse in an urban environment: a recent invader vs. a closely related permanent inhabitant. Urban Ecosyst 20:581–594. https://doi.org/10.1007/s11252-016-0620-7
Guillot G, Mortier F, Estoup A (2005) GENELAND: a computer package for landscape genetics. Mol Ecol Not 5:712–715. https://doi.org/10.1111/j.1471-8286.2005.01031.x
Hardouin EA, Orth A, Teschke M, Darvish J, Tautz D, Bonhomme F (2015) Eurasian house mouse (Mus musculus L.) differentiation at microsatellite loci identifies the Iranian plateau as a phylogeographic hotspot. BMC Evol Biol 15:26. https://doi.org/10.1186/s12862-015-0306-4
Hardouin EA, Chapuis JL, Stevens MI, Van Vuuren JB, Quillfeldt P, Scavetta RJ et al (2010) House mouse colonization patterns on the sub-Antarctic Kerguelen Archipelago suggest singular primary invasions and resilience against re-invasion. BMC Evol Biol 10:325. https://doi.org/10.1186/1471-2148-10-325
Haviland WA (1972) Family size, prehistoric population estimates and the ancient Maya. Am Antiq 37(1):135–139
Hernández-González A, Villaseñor-Franco A (2020) Los cruceros turísticos en el Caribe-oeste y su articulación con la isla de Cozumel. Études Caribéennes. https://doi.org/10.4000/etudescaribeennes.20076
Holmes ND, Buxton RT, Jones HP, Méndez Sánchez F, Oppel S, Russell JC, Spatz DR, Samaniego A (2023) Chapter 15: Conservation of marine birds: biosecurity, control, and eradication of invasive species threats. In: VanderWerf E (ed) Young L. Conservation of marine birds, Academic Press, pp 403–438. https://doi.org/10.1016/B978-0-323-88539-3.00019-4
Hooker S, Innes J (1995) Ranging behaviour of forest-dwelling ship rats, Rattus rattus, and effects of poisoning with bradifacoum. New Zealand J Zool 22:291–304. https://doi.org/10.1080/03014223.1995.9518044
Hulme-Beaman A, Dobney K, Cucchi T, Searle JB (2016) An ecological and evolutionary framework for commensalism in anthropogenic environments. Trends Ecol Evol 31:633–645. https://doi.org/10.1016/j.tree.2016.05.001
Instituto Nacional de Estadística y Geografía (INEGI) (2015) Censo de Población y Vivienda Intercensal 2015. Ciudad de México, México
Instituto Nacional de Estadística y Geografía (INEGI) (2020) https://www.inegi.org.mx/app/cpv/2020/resultadosrapidos/default.html?texto=cozumel%20quintana%20roo
Jacob HJ, Brown DM, Bunker RK, Daly MJ, Dzau VJ, Goodman A et al (1995) A genetic linkage map of the laboratory rat, Rattus norvegicus. Nat Genet 9:63–69. https://doi.org/10.1038/ng0195-63
Jing M, Yu HT, Bi X, Lai YC, Jiang W, Huang L (2014) Phylogeography of Chinese house mice (Mus musculus musculus/castaneus): distribution, routes of colonization and geographical regions of hybridization. Mol Ecol 23:4387–4405. https://doi.org/10.1111/mec.12873
Johnson MTJ, Munshi-South J (2017) Evolution of life in urban environments. Science 358(6363):eaam8327. https://doi.org/10.1126/science.aam8327
Jombart T, Devillard S, Balloux F (2010) Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genet 11:94. https://doi.org/10.1186/1471-2156-11-94
Jones EP, Jensen JK, Magnussen E, Gregersen N, Hansen HS, Searle JB (2011) A molecular characterization of the charismatic Faroe house mouse. Biol J Linn Soc 102:471–482. https://doi.org/10.1111/j.1095-8312.2010.01597.x
Kalinowski S, Wagner AP, Taper ML (2006) ML-RELATED: a computer program for maximum likelihood estimation of relatedness and relationship. Mol Ecol Not 6:576–579. https://doi.org/10.1111/j.1471-8286.2006.01256.x
King CM (2016) How genetics, history and geography limit potential explanation of invasions by house mice (Mus musculus) in New Zealand. Biol Invas 18:1533–1550. https://doi.org/10.1007/s10530-016-1099-0
Konecny A, Estoup A, Duplantier JM, Bryja J, Ba K, Galan M et al (2013) Invasion genetics of the introduced black rat (Rattus rattus) in Senegal, West Africa. Mol Ecol 22:286–300. https://doi.org/10.1111/mec.12112
Lack JB, Hamilton MJ, Braun JK, Mares MA, Van Den Bussche RA (2013) Comparative phylogeography of invasive Rattus rattus and Rattus norvegicus in the U.S. reveals distinct colonization histories and dispersal. Biol Invas 15:1067–1087. https://doi.org/10.1007/s10530-012-0351-5
Lambrinos JG (2004) How interactions between ecology and evolution influence contemporary invasion dynamics. Ecology 85:2061–2070. https://doi.org/10.1890/03-8013
Lawson LJ, Estoup A, Evans DM, Thomas CE, Lombaert E, Facon B et al (2011) Ecological genetics of invasive alien species. Biocontrol 56:409–428. https://doi.org/10.1007/s10526-011-9386-2
Lippens C, Estoup A, Hima MK, Loiseau A, Tatard C, Dalecky A et al (2017) Genetic structure and invasion history of the house mouse (Mus musculus domesticus) in Senegal, West Africa: a legacy of colonial and contemporary times. Heredity 119:64–75. https://doi.org/10.1038/hdy.2017.18
Loiseau A, Rahelinirina S, Rahalinson L, Konecny A, Duplantier JM, Brouat C (2008) Isolation and characterization of microsatellites in Rattus rattus. Mol Ecol Res 8:916–918. https://doi.org/10.1111/j.1755-0998.2008.02115.x
López M, Foronda P, Feliu C, Hernández M (2013) Genetic characterization of black rat (Rattus rattus) of the Canary Islands: origin and colonization. Biol Invas 15:2367–2373. https://doi.org/10.1007/s10530-013-0466-3
Lucaccioni H, Granjon L, Dalecky A, Fossati O, Le Fur J, Duplatier JM, Handschumacher P (2016) From human geography to biological invasions: the black rat distribution in the changing southeastern of Senegal. PLoS ONE 11(9):e0163547. https://doi.org/10.1371/journal.pone.0163547
Luikart G, Cornuet JM (1998) Empirical evaluation of a test for identifying recently bottleneck populations from allele frequency data. Conserv Biol 12(1):228–237
Mangombi JB, Brouat C, Loiseau A, Banga O, Leroy EM, Bourgarel M, Duplantier JM (2016) Urban population genetics of the invasive black rats in Franceville, Gabon. J Zool 299:183–190. https://doi.org/10.1111/jzo.12334
Meirmans PG (2014) Non convergence in Bayesian estimation of migration rates. Mol Ecol Res 14:726–733. https://doi.org/10.1111/1755-0998.12216
Miller AP, Webb PI (2001) Diet of house mouse (Mus musculus) on coastal sand dunes, Otago, New Zealand. New Zealand J Zool 28:49–55. https://doi.org/10.1080/03014223.2001.9518256
Morgan AP, Hughes JJ, Didion JP, Jolley WJ, Campbell KJ, Threadgill DW et al (2022) Population structure and inbreeding in wild house mice (Mus musculus) at different geographic scales. Heredity 129:183–194. https://doi.org/10.1038/s41437-022-00551-z
Munshi-South J (2012) Urban landscape genetics: canopy cover predicts gene flow between white-footed mouse (Peromyscus leucopus) populations in New York City. Mol Ecol 21:1360–1378. https://doi.org/10.1111/j.1365-294X.2012.05476.x
Munshi-South J, Nagy C (2014) Urban park characteristics, genetic variation, and historical demography of white-footed mouse (Peromyscus leucopus) populations in New York City. PeerJ 2:e310. https://doi.org/10.7717/peerj.310
Nachman MW, Searle JB (2014) Why is the house mouse karyotype so variable? Trends Ecol Evol 10(10):397–402. https://doi.org/10.1016/s0169-5347(00)89155-7
Nathan HW, Clout NM, MacKey JW, Murphy EC, Rusell JC (2015) Experimental island invasion of house mice. Popul Ecol 57:363–371. https://doi.org/10.1007/s10144-015-0477-2
Nei M (1974) Sampling variances of heterozygosity and genetic distance. Genetics 76:379–390
Norma Oficial Mexicana NOM-062-ZOO-1999 (2001). Especificaciones técnicas para la producción, cuidado y uso de los animales de laboratorio. Diario Oficial, 22 de agosto de 2001, México, pp 107–165. https://www.gob.mx/cms/uploads/attachment/file/203498/NOM-062-ZOO-1999_220801.pdf
Palafox A, Zizumbo L (2009) Distribución territorial y turismo en Cozumel, Estado de Quintana Roo, México. Gestión Turística 11:69–88
Panty-May JA, Carvalho-Pereira TSA, Serrano S, Pedra GG, Taylor J, Pertile AC et al (2016) A two-year ecological study of Norway rats (Rattus norvegicus) in a Brazilian Urban Slum. PLoS ONE 11(3):e0152511. https://doi.org/10.1371/journal.pone.0152511
Peakall R, Smouse PE (2012) GenAlex 6.5: genetic analysis in Excel: population genetic software for teaching and research-an update. Bioinformatics 28:2537–2539. https://doi.org/10.1093/bioinformatics/bts460
Peterman WE, Connette GM, Semlitsch RD, Eggert LS (2014) Ecological resistance surfaces predict fine-scale genetic differentiation in a terrestrial woodland salamander. Mol Ecol 23:2402–2413. https://doi.org/10.1111/mec.12747
Phifer-Rixey M, Nachman MW (2015) Insights into mammalian biology from the wild house mouse Mus musculus. Elife 4:e05959. https://doi.org/10.7554/eLife.05959
Pichlmueller F, Murphy EC, MacKay JWB, Henderson J, Fewster RM, Russell JC (2020) Island invasion and reinvasion: Informing invasive species management with genetic measures of connectivity. J Appl Ecol 57:2258–2270. https://doi.org/10.1111/1365-2664.13727
Piry S, Luickart G, Cornuet JM (1999) BOTTLENECK: a computer program for detecting recent reductions in the effective population size using allele frequency data. J Heredity 90:502–503
Prenter J, MacNeil C, Dick JT, Dunn AM (2004) Roles of parasites in animal invasions. Trends Ecol Evol 19:385–390. https://doi.org/10.1016/j.tree.2004.05.002
Programa rector de desarrollo costero del estado de Quintana Roo (PRORED) (2008) Secretaría de Comunicaciones y Transportes, México
R Core Team (2016) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna https://www.r-project.org/
Richardson JL, Burak MK, Hernandez C, Shirvell JM, Mariani C, Carvalho-Pereira TSA (2017) Using fine-scale spatial genetics of Norway rats to improve control efforts and reduce leptospirosis risk in urban slum environments. Evol Appl 10:323–337. https://doi.org/10.1111/eva.12449
Richardson JL, Silveira G, Soto-Medrano I, Arietta AZ, Mariani C, Pertile AC et al (2019) Significant genetic impacts accompany an urban rat control campaign in Salvador, Brazil. Front Ecol Evol 7:115. https://doi.org/10.3389/fevo.2019.00115
Rousset F (2008) Genepop`007: a complete re-implementation of the GENEPOP software for Windows and Linux. Mol Ecol Res 8:103–106. https://doi.org/10.1111/j.1471-8286.2007.01931.x
Sánchez-Crispín A, Propin E (2003) Dependencias regionales del turismo en la Isla de Cozumel, México. Cuadernos De Turismo 11:169–180
Santangelo JS, Ness RW, Cohan B, Fitzpatrick CR, Innes SG, Koch S et al (2022) Global urban environmental change drives adaptation in white clover. Science 375(6586):1275–1281. https://doi.org/10.1126/science.abk0989
Semarnat (2016) Programa de Manejo Área de Protección de Flora y Fauna la porción norte y la franja costera oriental, terrestres y marinas de la Isla de Cozumel. Conanp, Ciudad de México, México
Shiels AB, Flores CA, Khamsing A, Krushelnycky PD, Mosher SM, Drake DR (2013) Dietary niche differentiation among three species of invasive rodents (Rattus rattus, R. exulans, Mus musculus). Biol Invas 15:1037–1048. https://doi.org/10.1007/s10530-012-0348-0
Shirk AJ, Landguth EL, Cushman SA (2017) A comparison of individual-based genetic distance metrics for landscape genetics. Mol Ecol Res 16:1308–1317. https://doi.org/10.1111/1755-0998.12684
Siahsarvie R, Auffray JC, Darvish J, Rahabi-Maham H, Yu HT, Agret S et al (2012) Patterns of morphological evolution in the mandible of the house mouse Mus musculus (Rodentia: Muridae). Biol J Linn Soc 105:635–647. https://doi.org/10.1111/j.1095-8312.2011.01821.x
Simberloff D (2009) The role of propagule pressure in biological invasions. Ann Rev Ecol Evol Syst 40:81–102. https://doi.org/10.1146/annurev.ecolsys.110308.120304
Simberloff D, Von Holle B (1999) Positive interactions pf nonindigenous species: invasional meltdown? Biol Invas 1:21–32
Spaw RH (1978) Cozumel Island: stratigraphy and depositional history of upper Pleistocene limestone. Geol Hydrogeol Northeastern Yucatan 1978:209–218
Stragier C, Piry S, Loiseau A, Mamadou K, Aliou S, Youssoupha N et al (2022) Interplay between historical and current features of the cityscape in shaping the genetic structure of the house mouse (Mus musculus domesticus) in Dakar (Senegal, West Africa). Peer Commun J 2:e11. https://doi.org/10.24072/pcjournal.85
Santander LC, Ramos-Díaz M (2011) El nacimiento de un destino turístico en el caribe mexicano. Cozumel, de una isla abandonada a puerto de cruceros. El Periplo Sustentable 21:5–30
Suarez A, Tsutsui N (2008) The evolutionary consequences of biological invasions. Mol Ecol 17:351–360. https://doi.org/10.1111/j.1365-294X.2007.03456.x
Suárez-Atilano M, Cuarón AD, Vázquez-Domínguez E (2019) Deciphering geographical affinity and reconstructing invasion scenarios of Boa imperator in the Caribbean Cozumel Island. Copeia 107:606–621. https://doi.org/10.1643/CG-18-102
Trejo de la Paz V (2014) Transformaciones espaciales de la ciudad de Cozumel, Quintana Roo, 1980–2010. Universidad Nacional Autónoma de México, Facultad de Filosofía y Letras, México
Valenzuela-Galván D, Cuarón AD, Martínez-Morales MA, Vázquez L-B, Vázquez-Domínguez E (2022) First records of margay (Leopardus wiedii) on Cozumel Island: a conservation paradox. Anim Conserv 25:1–3. https://doi.org/10.1111/acv.12748
van Oosterhout C, Hutchinson WF, Willis DPM, Shipley P (2004) Micro-checker: software for identifying and correcting genotyping error in microsatellite data. Mol Ecol Notes 4:535–538. https://doi.org/10.1111/j.1471-8286.2004.00684.x
Vázquez-Domínguez E, Suárez-Atilano M, Booth W, González-Baca C, Cuarón AD (2012) Genetic evidence of a recent successful colonization of introduced species on islands: Boa constrictor imperator on Cozumel Island. Biol Invas 14:2101–2116. https://doi.org/10.1007/s10530-012-0217-x
Vega R, Vázquez-Domínguez E, Mejía-Puente A, Cuarón AD (2007) Unexpected high levels of genetic variability and the population structure of an island endemic rodent (Oryzomys couesi cozumelae). Biol Conserv 137:210–222. https://doi.org/10.1016/j.biocon.2007.02.007
Wilson GA, Rannala B (2003) Bayesian inference of recent migration rates using multilocus genotypes. Genetics 163:1177–1191. https://doi.org/10.1093/genetics/163.3.1177
Zhang Y, Ye A (2021) Quantitatively distinguishing the impact of climate change and human activities on vegetation in mainland China with the improved residual method. Gisci Remote Sens 58:235–260. https://doi.org/10.1080/15481603.20211872244
Acknowledgements
We are grateful with all those that helped during fieldwork, O. Romero-Báez, A. Tobón-Sampedro, N. Rivas, as well as T. Garrido-Garduño for molecular advice. We deeply thank C. González-Baca, H. Ayuntamiento de Cozumel, Comisión de Agua Potable y Alcantarillado, Comisión Nacional de Áreas Naturales Protegidas, Fundación de Parques y Museos Cozumel, and other institutions, colleagues and inhabitants in Cozumel for their support. We wish to thank the editor and the anonymous reviewers for their helpful suggestions. GBM acknowledges that this paper was a part of her bachelor’s thesis in Facultad de Ciencias, Universidad Nacional Autónoma de México. This article was completed while EVD was on sabbatical at the Estación Biológica de Doñana-CSIC with financial support from Dirección General de Asuntos del Personal Académico, UNAM (DGAPA/ PASPA No. 067/2023) and CONAHCyT.
Funding
Funding for this research was partially provided by Comisión Nacional sobre el Conocimiento y Uso de la Biodiversidad (Conabio project LI028).
Author information
Authors and Affiliations
Contributions
Both authors performed the study conception and design, material preparation, field work and data collection. Data analyses were performed by Gabriela Borja-Martínez. The manuscript was written, read and approved by Gabriela Borja-Martínez and Ella Vázquez-Domínguez.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Ethical approval
Trapping was performed with the corresponding scientific collecting permit (Semarnat FAUT-0168). Animal handling was carried out in strict accordance with the recommendations allowed by the Mexican Official Norm NOM-062-ZOO-1999 (Technical specifications for the protection, care and use of laboratory animals) (NOM 2001).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Borja-Martínez, G., Vázquez-Domínguez, E. Urban colonization of invasive species on islands: Mus musculus and Rattus rattus genetics of establishment on Cozumel Island. Biol Invasions (2024). https://doi.org/10.1007/s10530-024-03343-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10530-024-03343-0