Abstract
Context
Parapatric sister species are ideal for tests of ecological interactions. Pacific (Martes caurina) and American pine (M. americana) martens are economically and culturally valuable furbearers that hybridize in the north-central Rocky Mountains. Despite preliminary evidence of biased introgression, the hybrid zone has been geographically stable for 70 years, but interspecific ecological interactions have yet to be examined in detail.
Objectives
We test whether ecological interactions may influence the outcome of hybridization in this system. To that end, we estimate the fundamental niche of each species and gauge how suitability landscapes change when the two species are in contact.
Methods
We genotyped > 400 martens from the Rocky Mountain hybrid zone to diagnose individuals to species-level and identify putative hybrids. We then built range-wide ecological niche models for each species, excluding individuals in the hybrid zone, to approximate their respective fundamental niches. Those models were projected into the hybrid zone and compared with niche models trained on individuals within the hybrid zone to assess how niche dynamics change when the species are in sympatry.
Results
The fundamental niche of each species differed significantly, while the hybrid zone was equally suitable for both. Niches of each species based on models built within the hybrid zone showed that Pacific martens utilized significantly less suitable habitat than expected based on their range-wide fundamental niche, suggesting that species interactions shape local hybridization. We detected few admixed individuals (12%), with no evidence of directional (sex or species) biases. Interstate-90 further acts as a major dispersal barrier.
Conclusions
North American martens are currently managed as a single species by some state agencies, yet significant ecological and genetic differences indicate they should be managed separately. The observed ecological displacement of Pacific martens by American pine martens may partially explain the mixed success of historical, mixed-species wildlife translocations and cautions such translocations in the future. Landscape-scale consideration of ecological dynamics, in addition to molecular compatibility, will be essential to the success of future translocations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hybridization is common among wild populations, but difficult to accommodate in zoological conservation practice (Allendorf et al. 2001; Genovart 2009; Abbott et al. 2013; Todesco et al. 2016; NASEM 2021). On one hand, hybridization can boost genetic diversity in small, inbred populations (e.g., genetic rescue, Whiteley et al. 2015) or lead to the evolution of new species (Larsen et al. 2010; Ottenburghs 2018). On the other hand, gene flow can threaten to homogenize populations or erode local adaptation, which leads to a net loss of diversity (Muhlfeld et al. 2014; Todesco et al. 2016). Red wolves (Canis rufus) offer a high profile example of how the process of hybridization, now more easily detectable with molecular data, has changed the conservation landscape (NASEM 2021). After being extirpated from the wild in the 1980s, red wolves were the subject of extensive captive breeding and reintroduction efforts. Recent molecular analyses, however, have uncovered evidence of genetic exchange between extant red wolves and coyotes (Canis latrans). Together, that information calls into question the species status and the legitimacy of on-going red wolf recovery efforts and has spurred holistic reevaluation of red wolf conservation (Reich et al. 1999; Bohling et al. 2016; vonHoldt et al. 2016, 2022; NASEM 2021).
Mammalian carnivores, specifically, face elevated conservation risks due to their higher resource demands and potential for direct conflict with humans. As such, carnivores are frequent targets of wildlife translocations, which involve the intentional introduction of related individuals from one breeding population to another often to boost local genetic diversity (Frankham 2015; Whiteley et al. 2015; Gaywood et al. 2023), thereby ‘rescuing’ a small or inbred population from the ill effects of inbreeding. Extreme inbreeding depression can manifest as physical or physiological abnormalities, as evidenced in Florida panthers (Roelke et al. 1993). Translocations of Texas pumas (P. c. stanleyana) to Florida prompted hybridization with Florida panthers and led to a doubling of the population size (USFWS 1987; Pimm et al. 2006). While panthers represent a positive conservation outcome, for most taxa the variables contributing to the outcomes of genetic mixing remain incompletely understood. Such wildlife manipulations are usually irreversible and can have different outcomes in different environments and ecological contexts (Colella et al. 2018a, 2021a). As a result, hybrid zones provide a valuable window into the complexities of interspecific interactions, critical for understanding and predicting outcomes of genetic mixing (Wolf et al. 2001).
Intrinsic genetic incompatibilities (e.g., Dobzhansky-Muller incompatibilities, reviewed by Johnson 2000) can bias the strength and directionality of introgression (Todesco et al. 2016), however, because hybrid zones often form at ecotones (Barton and Hewitt 1989; Knowles 2001; Brunsfeld et al. 2001; Swenson and Howard 2005), the extrinsic environment also importantly shapes outcomes of hybridization. An organism's environment includes abiotic features, like temperature and precipitation, and biotic interactions, including competition, predation, and parasitism. Biotic interactions are difficult to measure directly but contribute to the formation, maintenance, and movement of hybrid zones across space and time. For example, a species rarely occupies the full extent of suitable habitat; instead, their range is restricted, by a combination of accessible area and biotic interactions, into a realized niche (Hutchinson 1957). Closely related, morphologically-similar species often share ecological requirements such that their ability to coexist is limited (Darwin 1859; MacArthur 1984; Anderson et al. 2002; Manlick et al. 2017a). Consequently, congeneric zones of sympatry are usually narrow, with species segregating by habitat (Emmons 1980). Ecological niche modeling (ENM), paired with high-density genotyping across zones of sympatry, allows for tests of the geographic predictions of biotic interactions (Anderson et al. 2002; Schukman et al. 2011; Tocchio et al. 2015; Qiao et al. 2017). Understanding how niche processes (both fundamental and realized, Jiménez et al. 2019) operate at varying spatial scales can provide a framework for anticipating how species will interact. Alteration of the realized niche within a hybrid zone would suggest that interspecific interactions shape hybrid zone development, maintenance, and persistence. Further, the direction of niche change can illuminate ecological or demographic processes contributing to hybrid dynamics.
Martens are medium sized carnivores in the weasel family (Mustelidae) and economically important furbearers (Harper 2010) that have been introduced to multiple islands along the coast of Southeast Alaska, the Great Lakes region of the US, and elsewhere to increase population sizes (Paul 2009; Grauer et al. 2017; Colella et al. 2018b, c). Many historical translocations unintentionally mixed two species—Pacific marten (Martes caurina) and American pine marten (Martes americana), historically recognized as a single species—which may partially explain their mixed success (Carr and Hicks 1997; Stone et al. 2002; Dawson et al. 2017). In the 1980s, for example, American pine martens from Minnesota and Ontario Canada and Pacific martens from Colorado were introduced to Wisconsin where American pine martens are native. There remains no residual genetic evidence of Pacific martens near the introduction site, suggesting that the less abundant parental species may have been eliminated through demographic or genetic swamping or other stochastic events, including elevated mortality upon release (Grauer et al. 2017). In contrast, a translocation of 10 American pine martens from coastal Southeast Alaska to Prince of Wales (POW) Island in the mid-1930s unexpectedly led to explosive population growth. Such an outcome could be explained by a hybrid swarm, where hybrids extensively cross and backcross into parental populations, if Pacific martens were present on POW prior to translocation (Pauli et al. 2015). Different evolutionary outcomes of mixing within a single system urges expanded investigations of the role of extrinsic (e.g., abiotic, biotic) and intrinsic (e.g., genetic, physiological) factors in shaping outcomes of hybridization (Barton and Hewitt 1989; Harrison 1990; Taylor et al. 2015) to inform future management actions aimed at boosting or maintaining genetic diversity in these species or populations.
To test for evidence of ecological displacement, two species must be phylogenetically close, morphologically similar, and not broadly sympatric (Schoener 1983; Anderson et al. 2002), as is true for North American pine martens. Both marten species are hypothesized to have diverged in allopatric glacial refugia located south of the continental ice sheets in North America during the Pleistocene (~ 21 kya), with M. americana isolated in the east and M. caurina in the west (Stone et al. 2002; Dawson et al. 2017; Lynch et al. 2020; Delheimer et al. 2021). Postglacial expansion appears to have resulted in a narrow band of contact along the north-central Rocky Mountains, reaching from central British Columbia south into western Montana (Dawson et al. 2017; Lucid et al. 2020). The mixing zone was first detected in the early 1950s, based on morphology (Wright 1953), which led to M. caurina being subsumed within M. americana as a subspecies. Historical consideration of only a single marten species in North America obscured the substantial morphological and genetic differences between these taxa until recently and muddled translocation efforts (Dawson et al. 2017; Colella et al. 2018b,c, 2021b; Manlick et al. 2018; Lynch et al. 2020; Delheimer et al. 2021). Molecular investigations have since confirmed substantial differentiation between the species and the presence of admixture in the Rocky Mountain zone, although only few, early generational (e.g., F1s) hybrids have been detected (Dawson et al. 2017; Colella et al. 2018b, c, 2021b; Lucid et al. 2020). Preliminary molecular evidence of biased introgression (e.g., unidirectional movement of genetic material from M. caurina into M. americana, Colella et al. 2018c) could be explained by assortative mating, genomic incompatibilities, or demographic swamping, but it remains unclear how environmental conditions contribute to interspecific interactions.
We test for climate-associated ecological differentiation between North American marten species and examine the role of the environment in structuring the Rocky Mountain hybrid zone. We use specimen-backed occurrence records and a suite of bioclimatic variables to construct two types of ENMs for each species: (i) range-wide models that provide a null expectation of the pattern and extent of niche overlap and (ii) local models within the hybrid zone based on > 400 individuals genotyped with two (one nuclear and one mitochondrial) restriction fragment length polymorphisms (RFLPs). We compare expected and observed niche overlap to test for the influence of abiotic and potential biotic interactions in shaping realized niches within the hybrid zone. If the hybrid zone is equally suitable for both species, we expect to observe similar representation of each species and hybrids throughout the zone. In contrast, disproportionate utilization of mutually suitable areas by one species when in sympatry can highlight the strength and directionality of interspecific interactions. Less suitable environments in the hybrid zone are predicted to be disproportionately occupied by an inferior competitor or immunologically naïve species. Overall, we demonstrate how environmental variation can structure species interactions within a natural hybrid zone. Such information on how species and genes interact importantly inform expectations for species mixing in the event of intentional wildlife translocations or augmentations; thereby guiding decision making (Adavoudi and Pilot 2022). Our results exemplify the utility of long-term, voucher-backed monitoring of natural hybrid zones as an experimental model for anticipating outcomes of genetic mixing and informing wildlife management.
Materials and methods
Molecular data: DNA extraction, PCR, and RFLP profiling
Vertebrate mitochondrial genomes (mtDNA) are inherited clonally from the maternal line, whereas nuclear DNA (nuDNA) recombines maternal and paternal ancestry. In consequence, hybridization cannot be detected from mitochondrial genotypes alone, but instead requires a combination of bi-parentally (e.g., nuDNA) or differentially (e.g., mtDNA and nuDNA) inherited genetic markers (Hewitt 2001). High-throughput sequencing (e.g., whole-genome sequencing) is sensitive to introgression between closely related groups; however, those methods remain prohibitively expensive for the large sample sizes required to detect admixture at large spatial scales, with costs further increasing if hybrids are rare or backcrossed into either parental population. Instead, we used RFLPs as a cost-effective tool for genotyping many individuals with high repeatability. If hybridization is rare (e.g., < 5%, Lucid et al. 2020; Colella et al. 2021b), admixed individuals are expected to be early generational hybrids (F1s) or highly diluted backcrosses. We screened one mitochondrial RFLP nested within cytochrome b (cytb) and one nuclear RFLP within the aldolase C (aldC) gene, which gives us a 100% probability of detecting F1s and a 75% probability of detecting F2s and first-generation backcrosses, assuming random segregation of alleles. Later generation hybrids will be more difficult to detect, therefore, our methods represent a conservative, underestimation of hybridization on the landscape and could be enhanced by the addition of genomic-scale data. If an individual had mitochondrial and nuclear markers assigned to different species or if their nuclear genotype was heterozygous, they were classified as a putative hybrid. We focused sampling in western Montana due to historical evidence of regional hybridization (Wright 1953) and the availability of dense, publicly-accessible sampling accumulated through collaborations between wildlife agency biologists, citizens, and natural history museums. Importantly, these methods are tractable for basic molecular laboratories and field stations and data analysis is relatively straightforward and does not require intensive computational capacities.
Tissue subsamples (n = 409) were loaned from the Museum of Southwestern Biology (MSB) at the University of New Mexico and University of Alaska Museum of the North (UAM) (Appendix I). DNA was extracted using the QIAamp DNA Micro Kit (Qiagen Inc., Germantown, MD, USA). Approximately 830 bp of the 3′ end of cytb and the flanking region was amplified using primers MVZ14 (5′ GGTCTTCATCTYHGGYTTACAAGAC 3′; Smith and Patton 1993) and Marten37 (5′ TATATATACCCCGAAACATGGA 3′; Demboski et al. 1999). Amplifications were in 25 μl volumes, each containing 1.5 μl MgCl2, 0.5 μl 10 mM dNTPs, 0.5 μl bovine serum albumin (10 mg/mL BSA), 0.15 μl of each primer, 0.13 μl of AmpliTaq DNA polymerase, 2.5 μl 1X PCR buffer, and 1 μl of 50 ng/μl genomic DNA. Cytb was amplified under the following conditions: 1 cycle of 94 °C for 45 s, followed by 35 cycles (10 s at 94 °C, 15 s at 45 °C, 45 s at 72 °C), and 1 cycle (3 min at 72 °C). We included positive and negative controls in each reaction. PCR products were visualized on an 0.8% agarose gel stained with Gel Red Nucleic Acid Gel Stain (Biotium, San Francisco, CA, USA). Samples with a positive PCR result were processed using a restriction enzyme digestion following manufacturer's protocol to diagnose species. A silent third position transition at site 603 in the M. americana mitochondrial cytb gene (site 263 of our amplified fragment) distinguishes M. americana (C) from M. caurina (T). Nla III cuts M. caurina amplicons after 5′—…CATG…—3′, but there is no restriction enzyme cut site present in M. americana (5′—…CACG…—3′). Restriction enzyme reactions included 9.0 μl of PCR product, 1.0 μl New England Biolabs 10X buffer, 0.10 μl BSA, and 0.2 μl Nla III restriction enzyme (New England Biolabs, Ipswich, MA, USA), combined and placed in a 37 °C incubator for 3 h. We visualized banding patterns on a 0.8% TAE agarose gel (50 ml TAE, 0.4 g agarose), by combining 2 μl of PCR product with 6 μl of diluted Gel Red (400 μl water, 100 ml 6X loading dye, 100 μl GelRed Master Mix [3.5 μl 10,000X Gel Red, 1 μl 6X loading dye]) and run at 105 V. 409 cytb genotypes were recorded.
For nuclear DNA, we distinguished M. americana sequences from those belonging to M. caurina based on a silent, third position transition (site 2763 of Rattus norvegicus, Mukai et al. 1991) in exon 5 of the nuclear gene aldC (Stone and Cook 2002). Approximately 70 bp of aldC, containing this mutation, was amplified using primers Ald-1B (5′ GCTGGATGGRCTCTYRAAAC 3′) and Ald-2B (5′ CAAGTGCTGAGGGTGTGCGC 3′) (Stone and Cook 2002). An annealing temperature of 50 °C was used for PCR amplification of aldC with all other conditions identical to the mitochondrial reactions. Samples with a positive PCR result were then diagnosed to species using a restriction enzyme digestion, as detailed above, but with the HhaI enzyme (New England Biolabs), which cuts after 5′—…GCGC… —3′. Due to the small size of the amplified aldC fragment, nuclear banding patterns were visualized on a 2% TBE agarose gel (50 ml 1X TBE, 1 g agarose, 5 μl gel red), 3.5 μl of PCR product was combined with 3 μl of 2X loading dye (Thermo Fisher Scientific Inc, Waltham, MA, USA) and run at 70 V to ensure band separation. 409 nuclear aldC genotypes were recorded. A binomial test (p < 0.05; binom.test function in the rstatix v. 0.6.0 R package) was used to test for a bias in the directionality of hybridization based on the null hypothesis that hybridization is equally likely to occur in either direction (sex and species). Binomial tests are useful for examining the distribution of a single dichotomous variable by testing the difference between a sample proportion and a given proportion. In this case, we tested whether the proportion of unadmixed individuals differed significantly from the proportion of admixed individuals detected in each species and each sex.
Occurrence data
We downloaded 1,807 and 552 georeferenced collection localities for voucher-supported specimens of M. americana and M. caurina, respectively, from across their North American ranges through the Global Biodiversity Information Facility (GBIF) and Arctos (arctos.database.museum) on 30 June 2019. We removed duplicate records and sorted occurrences to species-level by comparing localities to published amplicon sequence data that defined the geographic ranges of each species (Dawson et al. 2017; Colella et al. 2018b, c). We excluded M. caurina occurrences from the temperate rainforest islands of Southeast Alaska, as those martens represent a lineage distinct from continental populations (Colella et al. 2021b) and those environments are non-analog to those in the Rocky Mountains. Occurrences near or within the hybrid zone, defined as a 150 km buffer around a minimum convex polygon of coordinates from genotyped hybrid individuals, were excluded from range-wide niche analyses.
We cleaned occurrences following established protocols (sensu Feng et al. 2019). This included omitting records with no verifiable geographic coordinates, mis-matched coordinates with locality descriptions, geographic coordinate uncertainty > 15 km (to account for spatial uncertainty), and generally erroneous records (e.g., those located in bodies of water, on continents where the species do not occur, mis-signed longitude coordinates, and records from zoos or other institutions). To mitigate model overfitting that may stem from areas of exceptionally dense sampling (e.g., Michigan’s Upper Peninsula for M. americana), we spatially thinned remaining records to a minimum distance of 50 km using spThin (Aiello-Lammens et al. 2015), as a generous estimate of dispersal distance and home range size. Home range sizes for North American martens are estimated between 4 and 20 km2 (Mech and Rogers 1977; Bull and Heater 2001; Wasserman et al. 2010), although greater dispersal distances of up to 100 km2 have been recorded (Flynn and Schumacher 1999). For occurrences in the hybrid zone, we also performed sensitivity analyses where we thinned records only to a single occurrence per species per grid cell instead of a 50 km buffer to better understand local-scale hybrid dynamics.
Environmental data
We used a subset of the 19 bioclimatic variables from WorldClim 2.0 (Fick and Hijmans 2017) at 5 arc-minute resolution (~ 10 km grid cells). We selected variables assumed a priori to have a meaningful effect on the physiological tolerances of these species. We ensured the variables were not highly correlated (Pearson correlation > 0.7), favoring seasonal means over absolute maxima/minima (e.g., Mean Temperature of Warmest Quarter over Maximum Temperature of Warmest Month). This approach is reasonable because martens are relatively good dispersers (4–20 km, Mech and Rogers 1977; 15 km, Wasserman et al. 2010) and can behaviorally and physiologically accommodate seasonal variation in situ (Buskirk 1984; Chapin et al. 1997). Our final variables included: Bio4, Temperature Seasonality; Bio10, Mean Temperature of the Warmest Quarter; Bio12, Mean Annual Precipitation; and Bio15, Precipitation Seasonality.
Range-wide “fundamental niche” estimation
We generated range wide ENMs for each species to approximate their fundamental niche (Peterson et al. 2011; Jiménez et al. 2019). Models were based on 200 M. americana and 70 M. caurina thinned occurrence records. We defined training regions for each species as cells within a 250 km buffer around respective M. americana/M. caurina occurrence points, again excluding cells within the hybrid zone. We generated ENMs based on those data, then projected the models into the hybrid zone to determine whether there were broad-scale niche differences between the species that may influence the frequency or distribution of hybridization. Understanding how hybrid zone formation and maintenance is influenced by both broad- and local-scale niche processes is key to understanding how these zones emerge and change over time.
We applied a maximum entropy (Maxent v 3.4.1) approach to fitting ENMs (Phillips et al. 2006) via the dismo (v1.4; (Hijmans et al. 2020) R package. Maxent is a suitable algorithm for our question, given a lack of reliable absence data. Maxent approaches additionally fit models of flexible complexity. We built niche models that approximated each species' range-wide fundamental niche (Peterson et al. 2011) following an approach similar to the ENMeval framework (Muscarella et al. 2014). To minimize risk of overfitting, we tuned model parameters and validated models based on established criteria for presence-background ENMS, such as Maxent; only selecting simple feature classes (linear, quadratic, and product, and quadratic only) and a sequence of regularization multipliers (0.025, 0.05, 0.10, 0.25, 0.50, 0.75, and 1.0–5.0 in steps of 0.50). This framework permits optimization of a reasonable (e.g., 30) number of realistic (i.e., simple) models that are likely to reflect the physiological tolerances of species (Phillips et al. 2017). We selected the best of these models by identifying those with the lowest AICc values and comparing any models with AICc values within two of the optimal model to the minimum AICc via fivefold cross validation.
We projected range wide models into the hybrid zone and compared suitability values for genotyped M. americana/M. caurina/hybrid individuals sampled in the hybrid zone using a Hotelling’s T2 test to determine if broad-scale niche patterns may influence local-scale hybridization patterns. Hotelling’s T2 tests for multivariate differences in the means of where Martes individuals plotted in bivariate M. americana/M. caurina suitability space (i.e., is any group expected to exist in lower or higher suitability habitat in the hybrid zone?). A significant T2 statistic would suggest that broad-scale range-wide niche processes partially shape patterns of hybridization, which would confound analyses of local-scale competitive processes. Because Hotelling’s T2 test only evaluates a multivariate mean, we used the logistic, un-thresholded Maxent suitabilities for both species despite these values being incongruent (i.e., despite suitability values of 0.5 from two separate models being unequal; Elith et al. 2010). In this context, a significant T2 statistic indicates that the environments in which each Martes group (M. americana, M. caurina, hybrids) is found are significantly different from each other and also differ from the background environment. Thus, this test permitted us to assess whether range wide niche patterns for each species were sufficiently different to cause unbalanced suitability within the contact zone. Equivalent suitability for both species within the hybrid zone would suggest that any imbalances in the direction and intensity of hybridization are driven by factors other than the sampled climatic variables (e.g., biotic interactions).
Ecological Niche modeling: realized niche within the hybrid zone
Next, we constructed ENMs based on genotyped occurrences from within the hybrid zone to quantify niche overlap and assess the role of local interspecific interactions in shaping the hybrid zone. The hybrid zone (402,149 km2) represents a small fraction of each species’ total distributional area as inferred from International Union for Conservation of Nature (IUCN) range polygons, but represents a greater component for Pacific martens (~ 5% M. americana, ~ 18% M. caurina). As above, we used Maxent to construct local models. Non-admixed, genotyped individuals from within the hybrid zone were included in analyses. Realized niches are not expected to be convex and simple as is the case for functional niches (Soberón and Peterson 2020); therefore, we allowed for more complex feature classes (linear + quadratic, linear + quadratic + hinge, and hinge-only). We again used a sequence of regularization multipliers (0.10, 0.25, 0.50, 0.75, 1.0–5.0 in steps of 0.50, 7.50, and 10). Higher regularization values paired with a more complex feature class(es) has been shown to outperform simpler models (Shcheglovitova and Anderson 2013). Because we used more complex feature classes here, we did not use AICc to determine model quality/performance. Instead, we evaluated models using fivefold cross-validation as mentioned above, selecting models with realistic (i.e., not concave-up) response curves (Peterson et al. 2011; Araújo et al. 2019) and a balance of low testing omission error and low cross-validation variability using the one-standard-error rule (Hastie et al. 2009). These choices ensured that models fit well to the data while allowing for additional complexities where statistically justified. We determined if the clustering of M. americana/M. caurina/hybrid individuals in bivariate suitability space were significantly different from each other in the Rocky Mountain hybrid zone using a Hotelling’s T2 test as before (e.g., whether a group was more likely to occur in environments of higher/lower suitability for one or both marten species?).
Results
RFLP profiling
NlaIII digestions of cytb PCR products produced two small bands for M. caurina and a single, larger (uncut) band for M. americana. Nuclear restriction enzyme digestion using HhaI also produced multiple small bands for M. caurina and a single, large uncut band for M. americana (Fig. 1). Due to the small size of the amplified aldC fragment and central position of the RFLP cut site, the two smaller bands in M. caurina appear as a single band that is half the size of the uncut M. americana fragment. Therefore, nuclear heterozygotes appear as having two bands: one longer (M. americana) and one shorter (M. caurina), that fluoresce at half of the intensity of homozygotes (e.g., fewer copies). For cytb, 264 specimens were genotyped as M. caurina and 145 as M. americana. Nuclear genotypes identified 259 M. caurina, 127 M. americana, and 23 heterozygotes. Overall, 356 martens had consistent species diagnosis across differentially inherited markers, including 116 M. americana (60 male, 53 female, 3 unknown sex) and 240 M. caurina (122 male, 112 female, 6 unknown), and 53 of 409 (~ 13%) genotyped martens exhibited some degree of discordance and were classified as a putative hybrid. Thirty of the putative hybrids exhibited mismatched mitochondrial and nuclear markers: 11 had M. americana mtDNA and M. caurina nuDNA (4 male, 6 female, 1 unknown) and 19 had the other combination (10 male, 9 female). There were 23 nuclear heterozygotes; of those, 9 had M. americana mtDNA (4 male, 5 female) and the remaining 14 had M. caurina mtDNA (8 male, 5 female, 1 unknown). Binomial tests did not identify a sex or directional bias among hybrids. There were similar numbers of admixed individuals with M. americana mitochondrial and M. caurina nuclear DNA, as the reverse combination.
ENMs: range-wide “fundamental niche” estimates
ENMs built on range wide biodiversity data for M. americana and M. caurina identified meaningful differentiation in their respective fundamental niches (Figs. 2, 3). Across North America, suitable M. americana habitat is widespread from coast to coast, whereas M. caurina models illustrate a more restricted subset of suitable habitat. Mean summer temperature was the most important variable for both M. americana (67% importance) and M. caurina (82%), with M. caurina occurring less frequently in areas with hotter summers, by about 5 °C (as determined by the temperature at half the maximum suitability at the upper end of the distribution; Fig. 2). Increased temperature seasonality was inversely correlated with suitability for both species. Only temperature variables were influential in shaping the niche of M. americana. Increased precipitation seasonality (11% importance) conferred higher suitability for M. caurina. Decreased temperature seasonality and increased annual precipitation were associated with higher suitability for M. caurina, but these variables each had < 5% importance. Of the 1807 M. americana and 552 M. caurina records, 800 and 289 records for each species, respectively, passed cleaning/duplication filters and 200 and 70, respectively, remained after spatial thinning. Within the hybrid zone, we retained 36 and 116 records for M. americana and M. caurina respectively, of which 13 and 36 remained post-spatial thinning, respectively.
Response curves for range-wide models of each species, M. caurina (blue) and M. americana (red), for the 4 predictor variables in each model: (A) Temperature Seasonality, (B) Mean Summer Temperature, (C) Mean Annual Precipitation, and (D) Precipitation Seasonality. Lines represent the weighted average based on fivefold cross-validation. Numbers indicate percent importance to the overall model for each species ± 1 standard deviation. Vertical dashed lines indicate the input range of the environmental data used to train models for each species. Martes caurina tends to prefer colder, wetter environments than M. americana
Range-wide niche models for M. americana (A) and M. caurina (B) suitability projected across North America, with higher suitability denoted in darker blues. Occurrences (non-spatially thinned) are shown as red dots. Training regions for each species are outlined with solid black lines and the Rocky Mountain hybrid zone is outlined with a dashed line. Areas in white indicate non-analog environmental conditions relative to the respective training regions
Despite differences in fundamental niches between the two species, hybrid zone suitability was generally high for both species (SI Figs. 2–4). Our Hotelling’s T2 test on the range-wide models projected into thxe hybrid zone was non-significant (p = 0.2437), indicating that the multivariate means of each species (in bivariate suitability space) were not meaningfully different from each other. This test only specifies that niche processes were non-differentiable from each other (e.g., niches projected to the hybrid zone were similarly suitable) as opposed to high suitability for one species and low suitability for the other. This suggests that range wide fundamental niche processes may only provide the pre-conditions for hybridization (i.e., hybrids are likely to occur only in areas where both parent species are present), whereas local, processes are more likely to be the primary driver.
ENMs: realized niche within the hybrid zone
ENMs based on occurrences from within the hybrid zone recovered significant results (Hotelling’s T2 tests p < 0.05 for all tests and p < 0.01 for most) supporting a role for non-fundamental-niche-based biological interactions in shaping the structure of the hybrid zone. Hybrid zone-based models for each species showed relatively tight clustering of M. americana individuals across the bivariate suitability space (particularly for cells with high M. americana suitability and moderate M. caurina suitability; Fig. 4A). In contrast, M. caurina are dispersed more broadly (Fig. 4B). Interestingly, hybrid individuals mostly occur in cells with low M. americana suitability, but moderate to high M. caurina suitability (Fig. 4C). We further found meaningful geographic and elevational separation between the species within the hybrid zone. Martes americana are detected almost exclusively north of Interstate-90 (I-90), whereas M. caurina are detected almost exclusively south of I-90 (Fig. 5). Areas in the hybrid zone south of I-90 are higher in elevation on average than areas north of I-90 (Fig. 5E). On each side of the interstate, hybrids are distributed similarly to the most common species on that side of the freeway both spatially and elevationally (Fig. 5B–D).
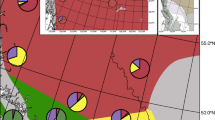
Environmental suitability space within the Rocky Mountain hybrid zone for the two Martes species and hybrids, built on occurrences (spatially thinned to a single occurrence per grid cell) and environmental variables from within the hybrid zone for each species. Black dots represent grid cells within the hybrid zone. Colored dots represent occurrences for M. americana (A), M. caurina (B), and caurina-americana hybrids (C). Point positions along the x-axis denote suitability based on the M. americana model. Point positions along the y-axis denote suitability based on the M. caurina model. Models built on occurrences within the hybrid zone identify M. americana and M. caurina as occupying different parts (i.e., grid cells) of the hybrid zone, suggesting that these species may be partitioning their niche within the zone. Hybrids occupy grid cells that both parental species also occupy
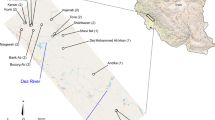
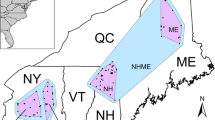
Summary of > 400 genotyped martens collected within the Rocky Mountain hybrid zone; (A) Map of genotyped individuals collected within the hybrid zone (area inside dashed line). Martes americana occurrences are shown in red, M. caurina in blue, and putative hybrids in purple. Interstate-90 (I-90) is shown as the purple path running East–West across the map, separating the hybrid zone into northern and southern regions. The elevation background layer is from MERIT DEM (Yamazaki et al. 2017); Elevation distribution of genotyped M. americana (B), caurina-americana hybrids (C), and M. caurina (D) individuals, along with their respective sample sizes and provenance within either the northern (N) or southern (S) part of the hybrid zone; (E) Density plot of elevations within the northern (dashed line) or southern (solid line) part of the hybrid zone. Crossing I-90 marks a turnover from M. americana in the north to M. caurina in the south. Martes americana individuals are rarely (~ 5%) found south of I-90, whereas M. caurina are uncommon (~ 11%) north of I-90. Hybrids are found in roughly equal proportions (~ 43% north, ~ 56% south) on either side of I-90. Within the hybrid zone, M. caurina individuals are generally found at higher elevations
Discussion
The process of hybridization conflicts with traditional definitions of species based on reproductive isolation (e.g., Biological Species Concept), which challenges the rigidity of conservation paradigms (Jackiw et al. 2015; NASEM 2021). Improved ability to detect hybridization using molecular methods, however, has forced acknowledgement of the ubiquity of hybridization in nature (Taylor and Larson 2019) and the slow accommodation of gene flow into conservation policy and practice (Draper et al. 2021). In fact, hybridization is now considered among a wildlife managers’ toolkit (Jackiw et al. 2015; Chan et al. 2019), whereby the intentional geographic reshuffling of species can be used to restore genetic diversity to small, inbred populations, increase resilience in the face of changing climate, or intentionally increase population sizes of economically valuable species (e.g., furbearers, game animals). As many mammals, and especially carnivores, are declining, they are regular candidates for genetic rescue for both conservation and commercial purposes. Visible success stories, like that of Florida panthers (Johnson et al. 2010), encourage expanded use of genetic management tools, however, variation in the evolutionary outcomes of mixing cautions indiscriminate use (e.g., Grauer et al. 2017; Gippoliti et al. 2018). As reflected by the mixed success of marten translocations (Alaska, Pauli et al. 2015; Minnesota, Grauer et al. 2017), which have often aimed to boost population sizes or fur quality, we have only a shallow understanding of the expected outcomes of interspecific interactions in this commercially valuable group (Harper 2010; Todesco et al. 2016; van Wyk et al. 2017). While much research has focused on identifying genetic incompatibilities (e.g., Maheshwari and Barbash 2011) and quantifying the fitness effects of gene flow (e.g., Fredrickson et al. 2007; Johnson et al. 2010), less attention has been paid to ecological responses to sympatry. Our results underscore the importance of both biotic and abiotic factors in shaping the evolutionary outcomes of hybridization (e.g., introgression, reinforcement, genetic swamping, etc.; Moran et al. 2021).
Historical treatment of North American martens as a single species since the 1950s has obscured ecological differences and limited our understanding of gene flow, now recognized as interspecific hybridization (Wright 1953; Carr and Hicks 1997; Dawson et al. 2017; Lucid et al. 2020; Colella et al. 2021b). To understand the role of the environment in shaping species distributions and the Rocky Mountain hybrid zone, we used ENMs to assess suitability landscapes and test for a pattern of ecological displacement in areas of sympatry. Overall, suitability landscapes in North America differed between the two species but were similar when projected into the Rocky Mountain hybrid zone. Suitable and unsuitable areas for each species were also similar in spatial extent (SI Fig. 1). When in sympatry, however, suitability patterns for each species were markedly different than expected. Spatial projections of ENMs based on genotyped individuals sampled within the hybrid zone were complementary and non-overlapping (SI Figs. 4–5), with the two species segregated in different environments (SI Fig. 6). Putative hybrids were distributed more uniformly, in cells with both high M. americana and high M. caurina suitability. That pattern is consistent with biological interactions influencing hybridization.
We found only low levels of mixing (13%) within the Rocky Mountain hybrid zone, consistent with other recent molecular investigations (Wasserman et al. 2010, 2021; Lucid et al. 2020). Detection of few hybrids is consistent with two potentially related scenarios: (1) true rarity of hybrids on the landscape (Lucid et al. 2020; Colella et al. 2021b) due to genetic or ecological factors or (2) underestimation of hybrids as an artifact of the coarse molecular resolution of RFLPs. Assortative mating or infertility after the first generation (i.e., hybrid breakdown), for example, is expected to lead to a rarity of hybrids on the landscape. Alternatively, unidirectional backcrossing would lead to many later generational hybrids with small portions of minor parental ancestry, which would be hard to detect without genome-level resolution. Because hybrid classes cannot be reliably distinguished with this set of RFLPs, we cannot distinguish between these scenarios. Unlike previous investigations (Colella et al. 2018c; Lucid et al. 2020), we did not detect a significant bias in the directionality of hybridization by either sex or species (Fig. 5). Ultimately, consideration of both the environmental and spatial contexts is essential to anticipating the evolutionary outcomes of hybridization and wildlife translocations.
Species-specific niche characterization
Range wide niche suitability models well represent the current understanding of the geographic distributions of the two species (Fig. 2). As more of a generalist, M. americana has broad suitability across much of northern North America, extending from east to west coasts across the boreal zone. Martes americana suitability also reaches into the islands along the North Pacific Coast, where the species has invaded naturally and been intentionally introduced (Stone et al. 2002; Paul 2009; Pauli et al. 2015). Their suitability also stretches further into the Southwest than they are currently distributed, showing large overlap with the current distribution of M. caurina. Suitable area for mainland M. caurina, in contrast, is highly fragmented, with peak suitability in areas of higher elevation in the Pacific Northwest. Other areas with relatively high M. caurina suitability are only connected by thin corridors, with the Great Plains forming a barrier to eastward dispersal.
Climatic variables partially explain niche differentiation between the two species. Martes caurina is restricted to wetter, cooler localities relative to M. americana. This observation is initially counter-intuitive given the more southerly distribution of M. caurina, however, it is the result of the higher elevation (mountain top sky islands) distribution of the species. Pacific marten’s affinity for higher elevations, that tend to be cooler and wetter than surrounding lowlands, is also what makes large portions of northwestern North America environmentally suitable for the species despite their current distribution. Mean summer temperature was particularly informative for both species, with peak suitability for M. caurina occurring at the lowest sampled temperatures and decreasing sharply for warmer temperatures (Fig. 2). In contrast, M. americana exhibits moderate suitability across a range of mean summer temperatures, biased in favor of warmer conditions, with suitability decreasing near the extremes of the sampled distribution. These results are consistent with previous investigations that found temperature and precipitation (Baltensperger et al. 2017; Lynch 2018) to be the most informative climatic variables for martens; however, climate may be less important than static landscape variables in predicting species responses to changing environments (Baltensperger et al. 2017). Although not explicitly modeled, our results suggest the two species will respond differently to warming, with M. caurina suitability expected to shrink and M. americana better able to accommodate such change, potentially even expanding their distribution (Baltensperger et al. 2017; Baltensperger 2018). Precipitation was highly informative for M. caurina models, but did not significantly influence M. americana, further reinforcing the resilience of M. americana in the face of environmental change.
Within the hybrid zone, the two species are delimited across an elevational gradient (Fig. 5). Martes caurina occurs at higher elevations (average 2000 m), whereas M. americana is found at lower elevations (average 1110 m). Hybrids occur across a range of elevations, but are generally found at intermediate elevations, consistent with observations in northern Idaho, where resistance to gene flow was lowest at 1500 m (Wasserman et al. 2010). We excluded elevation as a predictor from our ENMs as it is highly correlated with temperature and precipitation (Peterson 2006); however, within a narrow geographic extent, such as this hybrid zone, one can reasonably interpret the elevational structure as being aligned with the distinct ecology of each species. The higher elevation distribution of M. caurina and the association with greater precipitation and cooler temperatures leads us to hypothesize that M. caurina were historically more widespread, consistent with paleo-distributional reconstructions of other montane species (Waltari and Guralnick 2009). Since the Last Glacial Maximum (LGM, ~ 21 kya), M. caurina populations have likely become increasingly fragmented on mountaintop sky islands, as the climate has warmed (Brown 1971, 1978). Although suitable habitat for M. caurina remains connected further north, there are limited corridors connecting lower latitude sky islands to each other or to northern populations (Wasserman et al. 2021). The impact of warming on dispersal corridors and increasing habitat fragmentation should be considered when developing management plans for M. caurina.
The geographic and elevational separation of species in the hybrid zone provides insights into how the zone may have formed and persisted for 70 years. Hybrids broadly follow the spatial/elevational pattern of the dominant species on either side of I-90 (SI Figs. 4–6). Response curves between hybrid zone occurrences also differ for each parent species (SI Figs. 7–8). Martes americana is generally found north of I-90 and M. caurina generally to the south, suggesting that highways may act as an important dispersal barrier for martens. That result is consistent with prior work by Cushman et al. (2011) demonstrating that North American martens strongly avoid non-forested and open canopy habitats. One exception to that pattern occurs at Homestake Pass, where I-90 turns abruptly north–south before resuming its northwestern route through Missoula and into Idaho. At Homestake, we genotyped one M. caurina population located north and east of I-90. That population may be a residual M. caurina population “trapped” north of I-90 following highway construction (completed in 1966, Meeks 2000; Axline 2013). That population would have been disconnected from the rest of the meta-population further south, resulting in a local sink of M. caurina that may have slowly dwindled across more than five decades. The converse pattern would have similarly occurred for populations of M. americana historically located south of I-90. In this sense, the risk of swamping would be highly localized and unlikely to affect the rest of the species. However, in such a scenario, interspecific translocations are also unlikely to be effective at altering local genetic diversity because of the different environments occupied by these species when in sympatry. Last, the species divide across I-90 also suggests that the hybrid zone may have shifted south by ~ 130 km over the last 70 years. The zone was first described in the early 1950s (Wright 1953) with morphological intergradation documented at Swan River, South Fork, and Sun River Montana prior to construction of I-90. Today, those sites are located 95 to 160 km north of I-90.
Biotic interactions influence niche dynamics in sympatry
As sister species that are morphologically similar but not broadly sympatric, North American martens are well suited to tests of ecological interactions. The hybrid zone is similarly suitable for both species when inferred from ENMs built on range-wide occurrences (SI Fig. 2, 3). In contrast, ENMs built within the hybrid zone show spatial partitioning and lack of overlap, with M. americana more likely to occur in areas of high M. americana suitability and vice versa (Fig. 4). Hybrids are also more common in areas of high M. caurina suitability (Fig. 4), a pattern that is reinforced by the greater elevational overlap observed between M. caurina and putative hybrids (Fig. 5B–E). Lack of spatial overlap in suitability projections in the hybrid zone is consistent with a pattern of ecological displacement (Anderson et al. 2002). Carnivores, in particular, exhibit strong interspecific competition and niche partitioning to minimize agonistic interactions (Manlick et al. 2017a). That hypothesis is consistent with the generalist and specialist ecologies of each marten species (Martin et al. 1994; Slauson and Zielinski 2017; Manlick et al. 2019) and size differences, as mainland M. caurina are physically smaller than M. americana (Colella et al. 2018b). Size differences can affect assortative mating preferences, although we did not find evidence of a directional sex or species bias among hybrids. Therefore, it is possible that M. americana is a superior competitor relative to M. caurina, but the presence of I-90 may have altered the dynamics between these species (Craighead et al. 2001, Serveen and Shoemaker 2003), in this case by potentially lessening competition within the hybrid zone. The extent of parasite coevolution and dynamics of possible exchange between these two host species holds interesting possibilities as well (Koehler et al. 2009; Hoberg et al. 2012).
Interpretation of species interactions is complicated by both deep-time historical biogeography and more contemporary regional development. Martes caurina are hypothesized to have diverged in the American West, while M. americana persisted in the east during glacial advances (Small et al. 2003; Stone et al. 2002; Dawson et al. 2017; Lynch et al. 2020; Colella et al. 2021b). Following glacial recession, M. caurina likely tracked the historical environmental conditions of the western refugium northward in latitude and also up in elevation, leading to their fragmented contemporary distribution. Martes americana, instead, are adapted to boreal forests. Reassembly of boreal forests following the LGM may have facilitated a transition of M. americana to a more generalist lifestyle (Lynch et al. 2020). In this sense, niche partitioning in the hybrid zone may reflect longer-term biogeographic processes in addition to contemporary ecological dynamics.
Interpretation and management recommendations
Martens in North America are currently managed as a single species by some (e.g., Idaho Department of Fish and Game 2019; Montana Fish and Game 2021), but not all state agencies (e.g., Alaska Department of Fish and Game). Genetic, morphological, and now ecological differentiation indicate these two species should be managed independently (Carr and Hicks 1997; Dawson and Cook 2012; Dawson et al. 2017; Colella et al. 2021b). The fragmented contemporary distribution and thin suitability corridors of M. caurina in western North America, combined with a narrower range of suitability in areas of cooler temperatures and higher precipitation, position M. caurina at risk under future warming scenarios. In contrast, the breadth of environmental conditions suitable for M. americana suggest a greater breadth thermal tolerance and the large, connected areas of suitable boreal habitat indicate that the species may be more resilient to change. As anticipated for other taxa, I-90 serves as a barrier to dispersal for North American martens that further reinforces the North/South divide between the two species. Such anthropogenic barriers can affect local hybrid dynamics by reducing exchange that otherwise may have occurred on natural landscapes.
Martens are among the most commonly translocated carnivores in North America. Rationale for translocations has cited improvement in fur quality and increasing local population sizes, with the goal of boosting local harvests (Paul 2009; Powell et al. 2012). The two species have been repeatedly, often unintentionally, mixed as part of those management efforts (Paul 2009; Powell et al. 2012). Distinct niches and evidence of ecological displacement when in sympatry may partially explain the low success of historical translocations (Grauer et al. 2017; Manlick et al. 2017b). For genetic rescue to be a viable management option, there must be (i) evidence of low fitness in the recipient population, (ii) a closely related donor, and (iii) experimental data that support the validity of genetic rescue in the proposed context (Hedrick and Fredrickson 2010). Subspecies, which ostensibly represent distinctive units of geographic variation within a single species, are capable of interbreeding with other conspecific subspecies and represent viable donor populations for active genetic management (Hedrick and Fredrickson 2010). Mixed species introductions, however, as have occurred repeatedly for North American martens, are not advised. Introgression almost always occurs from the less numerous group into the group with greater numbers, thereby diluting the genetic signal of the less numerous group through the process of demographic genetic swamping (Currat et al. 2008). That pattern creates a genetic sink that can lead to the replacement of the minor parental species, as observed for martens in the Great Lakes region (Grauer et al. 2017) and suspected on some North Pacific Coast islands (Pauli et al. 2015). Natural hybrid zones act as a valuable empirical test for understanding and anticipating the evolutionary outcomes of genetic mixing, that can proactively inform the outcomes of intentional genetic manipulations. The accumulation of sufficient sampling across space and time is increasingly possible through collaborative programs that permanently archive hunter and trapper salvaged materials in natural history museum biorepositories. Continued sampling within this hybrid zone will be critical to determining whether ecological displacement leads to longer-term consequences for M. caurina and at what timescales.
References
Abbott R, Albach D, Ansell S, Arntzen JW, Baird SJE, Bierne N et al (2013) Hybridization and speciation. J Evol Biol 26(2):229–246. https://doi.org/10.1111/j.1420-9101.2012.02599.x
Adavoudi R, Pilot M (2022) Consequences of hybridization in Mammals: a systematic review. Genes 13(1):50. https://doi.org/10.3390/genes13010050
Aiello-Lammens ME, Boria RA, Radosavljevic A, Vilela B, Anderson RP (2015) spThin: an R package for spatial thinning of species occurrence records for use in ecological niche models. Ecography 38(5):541–545. https://doi.org/10.1111/ecog.01132
Allendorf FW, Leary RF, Spruell P, Wenburg JK (2001) The problems with hybrids: setting conservation guidelines. Trends Ecol Evol 16(11):612–622. https://doi.org/10.1016/S0169-5347(01)02290-X
Anderson RP, Peterson AT, Gómez-Laverde M (2002) Using niche-based GIS modeling to test geographic predictions of competitive exclusion and competitive release in South American pocket mice. Oikos 98(1):3–16. https://doi.org/10.1034/j.1600-0706.2002.t01-1-980116.x
Araújo MB, Anderson RP, Barbosa AM, Beale CM, Dormann CF, Early R, Garcia RA, Guisan A, Maiorano L, Naimi B, O’Hara RB, Zimmermann NE, Carsten R (2019) Standards for distribution models in biodiversity assessments. Sci Adv 5(1):eaat4858. https://doi.org/10.1126/sciadv.aat4858
Axline J (2013) A massive undertaking: construction of Montana’s interstate highways, 1956–1988. Montana Historical Soc 63(3):60
Baltensperger AP (2018) Using interactions among species, landscapes, and climate to inform ecological niche models: a case study of American Marten (Martes americana) Distribution in Alaska. In Machine Learning for Ecology and Sustainable Natural Resource Management, pp. 205–25. Springer. https://doi.org/10.1007/978-3-319-96978-7_10
Baltensperger AP, Morton JM, Huettmann F (2017) Expansion of American marten (Martes americana) distribution in response to climate and landscape change on the Kenai Peninsula, Alaska. J Mammal 98(3):703–714. https://doi.org/10.1093/jmammal/gyx011
Barton NH, Hewitt GM (1989) Adaptation, speciation and hybrid zones. Nature 341(6242):497–503. https://doi.org/10.1038/341497a0
Bohling JH (2016) Strategies to address the conservation threats posed by hybridization and genetic introgression. Biol Cons 203:321–327. https://doi.org/10.1016/j.biocon.2016.10.011
Brown JH (1971) Mammals on mountaintops: nonequilibrium insular biogeography. Am Nat 105(945):467–478. https://doi.org/10.1086/282738
Brown JH (1978) The theory of insular biogeography and the distribution of boreal birds and Mammals. Great Basin Naturalist Memoirs 2:209–227
Brunsfeld SJ, Sullivan J, Soltis DE, Soltis PS (2001) Comparative phylogeography of Northwestern North America: a synthesis. Special Publ Br Ecol Soc 14:319–340
Buskirk SW (1984) Seasonal use of resting sites by marten in South-Central Alaska. J Wildl Manag 48(3):950–953. https://doi.org/10.2307/3801445
Carr SM, Hicks SA (1997) Are there two species of marten in North America? Genetic and evolutionary relationships within Martes. In: Martes: Taxonomy, Ecology, Techniques, and Management. pp. 15–28
Chan WY, Hoffmann AA, van Oppen MJ (2019) Hybridization as a conservation management tool. Conserv Lett 12(5):e12652. https://doi.org/10.1111/conl.12652
Chapin TG, Harrison DJ, Phillips DM (1997) Seasonal habitat selection by marten in an untrapped forest preserve. J Wildl Manag 61(3):707–717. https://doi.org/10.2307/3802178
Colella JP, Frederick LM, Talbot SL, Cook JA (2018a) Extrinsically reinforced hybrid speciation within Holarctic ermine (Mustela spp.) produces an insular endemic. Divers Distrib 27(4):747–762. https://doi.org/10.1111/ddi.13234
Colella JP, Johnson EJ, Cook JA (2018b) Reconciling molecules and morphology in North American Martes. J Mammal 99(6):1323–1335. https://doi.org/10.1093/jmammal/gyy140
Colella JP, Wilson RE, Talbot SL, Cook JA (2018c) Implications of introgression for wildlife translocations: the Case of North American martens. Conserv Genet 20(2):153–166. https://doi.org/10.1007/s10592-018-1120-5
Colella JP, Lan TY, Schuster SC, Talbot SL, Cook JA, Lindqvist C (2021a) Whole-genome analysis of Mustela erminea finds that pulsed hybridization impacts evolution at high latitudes. Commun Biol 1(1):51. https://doi.org/10.1038/s42003-018-0058-y
Colella JP, Lan T, Talbot SL, Lindqvist C, Cook JA (2021b) Whole-genome resequencing reveals persistence of forest-associated mammals in late Pleistocene refugia along North America’s North Pacific Coast. J Biogeogr 48(5):1153–1169. https://doi.org/10.1111/jbi.14068
Craighead L, Craighead A, Roberts EA (2001) Bozeman Pass wildlife linkage and highway safety study. In: Irwin CL, Garrett P, McDermott KP (eds) Proceedings of the 2001 International Conference on Ecology and Transportation. Center for Transportation and the Environment, North Carolina State University, Raleigh, pp 405–422
Currat M, Ruedi M, Petit RJ, Excoffier L (2008) The hidden side of invasions: massive introgression by local genes. Evolution 62(8):1908–1920. https://doi.org/10.1111/j.1558-5646.2008.00413.x
Cushman SA, Raphael MG, Ruggiero LF, Shirk AS, Wasserman TN, O’Doherty A (2011) Limiting factors and landscape connectivity: the American marten in the Rocky Mountains. Landscape Ecol 26:1137–1149. https://doi.org/10.1007/s10980-011-9645-8
Darwin C (1859) On the Origin of Species. Published on: https://ncse.ngo/files/pub/evolution/Illustrated%20Origin--Darwin--small--credit.pdf
Dawson NG, Cook JA (2012) Behind the genes: diversification of North American martens (Martes americana and Martes caurina). In: Aubry KB, Zielinski WJ, Raphael MG, Proulx G, Buskirk SW (eds) Biology and conservation of Martens, Sables, and Fishers: a new synthesis. Cornell University Press, Ithaca, pp 23–38. https://doi.org/10.7591/9780801466076-005
Dawson NG, Colella JP, Small MP, Stone KD, Talbot SL, Cook JA (2017) Historical biogeography sets the foundation for contemporary conservation of martens (genus Martes) in northwestern North America. J Mammal 98(3):715–730. https://doi.org/10.1093/jmammal/gyx047
Delheimer MS, Moriarty KM, Slauson KM, Roddy AM, Early DA, Hamm KA (2021) Comparative reproductive ecology of two subspecies of Pacific marten (Martes caurina) in California. Northwest Sci 94(3–4):271–285. https://doi.org/10.3955/046.094.0305
Demboski JR, Stone KD, Cook JA (1999) Further perspectives on the Haida Gwaii glacial refugium. Evolution 53(6):2008–2012. https://doi.org/10.2307/2640462
Draper D, Laguna E, Marques I (2021) Demystifying negative connotations of hybridization for less biased conservation policies. Front Ecol Evol 9:637100. https://doi.org/10.3389/fevo.2021.637100
Elith J, Kearney M, Phillips S (2010) The art of modeling range-shifting species. Methods Ecol Evol Br Ecol Soc 1(4):330–342. https://doi.org/10.1111/j.2041-210X.2010.00036.x
Emmons LH (1980) Ecology and resource partitioning among nine species of African rain forest squirrels. Ecol Monogr 50(1):31–54. https://doi.org/10.2307/2937245
Feng X, Park DS, Liang Y, Pandey R, Papeş M (2019) Collinearity in ecological niche modeling: confusions and challenges. Ecol Evol 9(18):10365–10376. https://doi.org/10.1002/ece3.5555
Fick SE, Hijmans RJ (2017) WorldClim2: new 1 km spatial resolution climate surfaces for global land areas. Int J Climatol 37(12):4302–4315
Flynn RW, Schumacher TV (1999) Ecology of martens in Southeast Alaska. Available at: http://www.arlis.org/docs/vol1/D/45087730.pdf
Frankham R (2015) Genetic rescue of small inbred populations: meta-analysis reveals large and consistent benefits of gene flow. Mol Ecol 24(11):2610–2618. https://doi.org/10.1111/mec.13139
Fredrickson RJ, Siminski P, Woolf M, Hedrick PW (2007) Genetic rescue and inbreeding depression in Mexican wolves. Proc Royal Soc Lond Biol Sci 274(1623):2365–2371. https://doi.org/10.1098/rspb.2007.0785
Gaywood MJ, Ewen JG, Hollingsworth PM, Moehrenschlager A (2023) Conservation translocations. Cambridge University Press. https://doi.org/10.1017/9781108638142
Genovart M (2009) Natural hybridization and conservation. Biodivers Conserv 18(6):1435. https://doi.org/10.1007/s10531-008-9550-x
Gippoliti S, Cotterill FPD, Groves CP, Zinner D (2018) Poor taxonomy and genetic rescue are possible co-agents of silent extinction and biogeographic homogenization among ungulate mammals. Biogeographia 33:41–54. https://doi.org/10.21426/B633039045
Grauer JA, Gilbert JH, Woodford JE, Eklund D, Anderson S, Pauli JN (2017) Unexpected genetic composition of a reintroduced carnivore population. Biol Cons 215:246–253. https://doi.org/10.1016/j.biocon.2017.09.016
Harper P (2010) Furbearer Management Report of Survey-Inventory Activities 1 July 2006–30 June 2009. Alaska Department of Fish and Game, Division of Wildlife Conservation
Harrison RG (1990) Hybrid zones: windows on evolutionary processes. Oxf Surv Evol Biol 7:69–128
Hastie T, Tibshirani R, Friedman JH, Friedman JH (2009) The elements of statistical learning: data mining, inference, and prediction. Springer, New York, pp 1–758. https://doi.org/10.1007/978-0-387-84858-7
Hedrick PW, Fredrickson R (2010) Genetic rescue guidelines with examples from Mexican wolves and Florida panthers. Conserv Genet 11:615–626. https://doi.org/10.1098/rspb.2007.0785
Heppenheimer E, Brzeski KE, Wooten R, Waddell W, Rutledge LY, Chamberlain MJ et al (2018) Rediscovery of red wolf ghost alleles in a canid population along the American Gulf Coast. Genes 9(12):618. https://doi.org/10.3390/genes9120618
Hewitt GM (2001) Speciation, hybrid zones and phylogeography—or seeing genes in space and time. Mol Ecol 10(3):537–549. https://doi.org/10.1046/j.1365-294x.2001.01202.x
Hijmans RJ, Phillips S, Leathwick J, Elith J (2020) Dismo: Species Distribution Modeling. R Package (1.1-4) [Computer Software]
Hoberg E, Koehler AVA, Cook JA (2012) Complex host-parasite systems in Martes: Implications for conservation biology of endemic faunas. In: Aubry K, Zielinski WJ, Raphael MG, Proulx F, Buskirk SW (eds) Biology and conservation of Marten, Sables, and Fisher. A new synthesis. Cornell University Press, Ithaca, pp 39–57. https://doi.org/10.7591/9780801466076-006
Hutchinson GE (1957) Concluding remarks. Cold Spring Harb Symp Quant Biol 22:415–427. https://doi.org/10.1101/SQB.1957.022.01.039
Idaho Department of Fish and Game (2019) Upland Game, Furbearer, and Turkey Seasons and Rules. Boise, ID: Idaho Department of Fish and Game
Jackiw RN, Mandil G, Hager HA (2015) A framework to guide the conservation of species hybrids based on ethical and ecological considerations. Conserv Biol 29(4):1040–1051. https://doi.org/10.1111/cobi.12526
Jiménez L, Soberón J, Andrés Christen J, Soto D (2019) On the problem of modeling a fundamental niche from occurrence data. Ecol Model 397:74–93. https://doi.org/10.1016/j.ecolmodel.2019.01.020
Johnson NA (2000) Speciation: Dobzhansky-Muller incompatibilities, dominance, and gene interactions. Trends Ecol Evol 15(12):480–482. https://doi.org/10.1016/s0169-5347(00)01961-3
Johnson WE, Onorato DP, Roelke ME, Land ED, Cunningham M, Belden RC et al (2010) Genetic restoration of the Florida panther. Science 329(5999):1641–1645. https://doi.org/10.1126/science.1192891
Knowles LL (2001) Did the Pleistocene glaciations promote divergence? Tests of explicit refugial models in montane grasshoppers. Mol Ecol 10(3):691–701
Koehler AVA, Hoberg EP, Dokuchaev NE, Tranbenkova NA, Whitman JS, Nagorsen DW, Cook JA (2009) Phylogeography of a Holarctic nematode Soboliphyme baturini among mustelids: climate change, episodic colonization, and diversification in a complex host-parasite system. Biol J Linn Soc Lond 96:651–663. https://doi.org/10.1111/j.1095-8312.2008.01145.x/full
Larsen PA, Marchán-Rivadeneira MR, Baker RJ (2010) Natural hybridization generates mammalian lineage with species characteristics. Proc Natl Acad Sci USA 107(25):11447–11452. https://doi.org/10.1073/pnas.1000133107
Lucid M, Cushman S, Robinson L, Kortello A, Hausleitner D, Mowat et al (2020) Carnivore Contact: a species fracture zone delineated amongst genetically structured North American marten populations (Martes americana and Martes caurina). Front Genet 11:735. https://doi.org/10.3389/fgene.2020.00735
Lynch LM (2018) Limb skeletal morphology of North American pine martens, Martes americana and Martes caurina, correlates with biome and climate. Biol J Lin Soc 126(2):240–255. https://doi.org/10.1093/biolinnean/bly175
Lynch LM (2020) Fossil calibration of mitochondrial phylogenetic relationships of North American pine martens, Martes, suggests an older divergence of M. americana and M. caurina than previously hypothesized. J Mammal Evol 27(3):535–548. https://doi.org/10.1007/s10914-019-09476-7
MacArthur RH (1984) Geographical ecology: patterns in the distribution of species. Princeton University Press
Maheshwari S, Barbash DA (2011) The genetics of hybrid Incompatibilities. Annu Rev Genet 45:331–355. https://doi.org/10.1146/annurev-genet-110410-132514
Manlick PJ, Woodford JE, Zuckerberg B, Pauli JN (2017a) Niche compression intensifies competition between reintroduced American martens (Martes americana) and fishers (Pekania pennanti). J Mammal 98(3):690–702. https://doi.org/10.1093/jmammal/gyx030
Manlick PJ, Woodford JE, Gilbert JH, Eklund D, Pauli JN (2017b) Augmentation provides nominal genetic and demographic rescue for an endangered carnivore. Conserv Lett 10(2):178–185. https://doi.org/10.1111/conl.12257
Manlick PJ, Romanski MC, Pauli JN (2018) Dynamic colonization history in a rediscovered Isle Royale carnivore. Sci Rep 8(1):12711. https://doi.org/10.1038/s41598-018-31130-0
Manlick PJ, Petersen SM, Moriarty KM, Pauli JN (2019) Stable isotopes reveal limited Eltonian niche conservatism across carnivore populations. Funct Ecol 33(2):335–345. https://doi.org/10.1111/1365-2435.13266
Martin SK, Buskirk S, Harestad A, Raphael M, Powell R (1994) Martens, Sables and Fishers: biology and conservation. Cornell University Press, Comstock Publishing Associates, p 536
Mech LD, Rogers LL (1977) Status, distribution, and movements of marten in northeastern Minnesota. U.S. Forest Service Research Papers, NE-14–3, 1–7
Meeks H (2000) On the Road to Yellowstone: the Yellowstone Trail and American Highways, 1900–1930. Pictorial Histories Publishing, Missoula, Montana, p 127
Montana Fish, Wildlife, & Parks (MFWP) (2021) 2021 Furbearer and Trapping, Trapping and Hunting Regulations. Montana Fish, Wildlife, and Parks. https://fwp.mt.gov/binaries/content/assets/fwp/hunt/regulations/2021/2021-furbearer-final-for-web.pdf
Moran BM, Payne C, Langdon Q, Powell DL, Brandvain Y, Schumer M (2021) The genomic consequences of hybridization. Elife 10:e69016. https://doi.org/10.7554/eLife.69016
Muhlfeld CC, Kovach RP, Jones LA, Al-Chokhachy R, Boyer MC, Leary RF, Lowe WH, Luikart G, Allendorf FW (2014) Invasive hybridization in a threatened species Is accelerated by climate change. Nat Clim Chang 4(7):620–624. https://doi.org/10.1038/nclimate2252
Mukai T, Yatsuki H, Masuko S, Arai Y, Joh K, Hori K (1991) The structure of the brain-specific rat aldolase C gene and its regional expression. Biochem Biophys Res Commun 174(2):1035–1042. https://doi.org/10.1016/0006-291x(91)91523-f
Murphy SM, Adams JR, Cox JJ, Waits LP (2019) Substantial red wolf genetic ancestry persists in wild canids of southwestern Louisiana. Conserv Lett 12(12):e12621. https://doi.org/10.1111/conl.12621
Muscarella R, Galante PJ, Soley-Guardia M, Boria RA, Kass JM, Uriarte M, Anderson RP (2014) ENM Eval: an R Package for conducting spatially independent evaluations and estimating optimal model complexity for maxent ecological niche models. Methods Ecol Evol Br Ecol Soc 5(11):1198–1205
National Academies of Sciences, Engineering, and Medicine (NASEM) (2021) A research strategy to examine the taxonomy of the red wolf. The National Academies Press. Washington, D.C
Ottenburghs J (2018) Exploring the hybrid speciation continuum in birds. Ecol Evol 8(24):13027–13034. https://doi.org/10.1002/ece3.4558
Paul T (2009) Game Transplants in Alaska. No. 4. Juneau, Alaska: Alaska Department of Fish and Game, Division of Wildlife Conservation
Pauli JN, Moss WE, Manlick PJ, Fountain ED, Kirby R, Sultaire SM, Perrig PL, Mendoza JE, Pokallus JW, Heaton TH (2015) Examining the uncertain origin and management role of martens on Prince of Wales Island, Alaska. Conserv Biol 29(5):1257–1267. https://doi.org/10.1111/cobi.12491
Peterson AT (2006) Uses and requirements of ecological niche models and related distributional models. Biodiv Inf 3:59–72. https://doi.org/10.17161/bi.v3i0.29
Peterson AT, Soberón J, Pearson RG, Anderson RP, Martínez-Meyer E, Nakamura M, Araújo MB (2011) Ecological niches and geographic distributions (MPB-49). Princeton University Press
Phillips SJ, Anderson RP, Schapire RE (2006) Maximum entropy modeling of species geographic distributions. Ecol Modeling 190(3):231–259. https://doi.org/10.1016/j.ecolmodel.2005.03.026
Phillips SJ, Anderson RP, Dudík M, Schapire RE, Blair ME (2017) Opening the black box: an open-source release of Maxent. Ecography 40(7):887–893. https://doi.org/10.1111/ecog.03049
Pimm SL, Dollar L, Bass OL (2006) The genetic rescue of the Florida panther. Anim Conserv 9(2):115–122. https://doi.org/10.1111/j.1469-1795.2005.00010.x
Powell RA, Lewis JCJ, Slough BG, Brainerd SM, Jordan NR, Abramov AV, Monakhov V, Zollner PA, Murakami T (2012) Evaluating translocations of Martens, Sables, and Fishers: testing Model Predictions with Field Data. In: Aubry KB, Zielinski WJ, Raphael MG, Proulx G, Buskirk SW (eds) Biology and conservation of Martens, Sables, and Fishers: a new synthesis, vol 536. Cornell University Press, Ithaca. https://doi.org/10.7591/9780801466076-009
Qiao H, Escobar LE, Peterson AT (2017) Accessible areas in ecological niche comparisons of invasive species: recognized but still overlooked. Sci Rep 7(1):1–9. https://doi.org/10.1038/s41598-017-01313-2
Reich DE, Wayne RK, Goldstein DB (1999) Genetic evidence for a recent origin by hybridization of red wolves. Mol Ecol 8(1):139–144. https://doi.org/10.1046/j.1365-294x.1999.00514.x
Roelke ME, Martenson JS, O’Brien SJ (1993) The consequences of demographic reduction and genetic depletion in the endangered Florida panther. Curr Biol 3(6):340–350. https://doi.org/10.1016/0960-9822(93)90197-v
Schoener TW (1983) Field experiments on interspecific competition. Am Nat 122(2):240–285. https://doi.org/10.1086/284133
Schukman JM, Lire-Noriega A, Peterson AT (2011) Multiscalar ecological characterization of Say’s and Eastern phoebes and their zone of contact in the Great Plains. The Condor 113(2):372–384. https://doi.org/10.1525/cond.2011.100073
Servheen C, Shoemanker R (2003) A sampling of wildlife use in relation to structure variables for brdiges and culverts under I-90 between Alberton and St. Regis, Montana. UC Davis Road Ecology Center. Available at: https://escholarship.org/uc/item/4tt86932
Shcheglovitova M, Anderson RP (2013) Estimating optimal complexity for ecological niche models: a jackknife approach for species with small sample sizes. Ecol Modeling 269:9–17. https://doi.org/10.1016/j.ecolmodel.2013.08.011
Slauson KM, Zielinski WJ (2017) Seasonal specialization in diet of the Humboldt marten (Martes caurina humboldtensis) in California and the importance of prey size. J Mammal 98(6):1697–1708. https://doi.org/10.1093/jmammal/gyx118
Small MP, Stone KD, Cook JA (2003) American marten (Martes americana) in the Pacific Northwest: population differentiation across a landscape fragmented in time and space. Mol Ecol 12(1):89–103. https://doi.org/10.1046/j.1365-294X.2003.01720.x
Smith MF, Patton JL (1993) The diversification of South American murid rodents: evidence from mitochondrial DNA sequence data for the Akodontine tribe. Biol J Linnean Soc 50(3):149–177. https://doi.org/10.1111/j.1095-8312.1993.tb00924.x
Soberón J, Peterson AT (2020) What is the shape of the fundamental Grinnellian niche? Thyroid Res 13(1):105–115. https://doi.org/10.1007/s12080-019-0432-5
Stone KD, Cook JA (2002) Molecular evolution of Holarctic martens (genus Martes, Mammalia: Carnivora: Mustelidae). Mol Phylogenet Evol 24(2):169–179. https://doi.org/10.1016/S1055-7903(02)00229-4
Stone KD, Flynn RW, Cook JA (2002) Post-glacial colonization of Northwestern North America by the forest-associated American marten (Martes americana, Mammalia: Carnivora: Mustelidae). Mol Ecol 11(10):2049–2063. https://doi.org/10.1046/j.1365-294X.2002.01596.x
Swenson NG, Howard DJ (2005) Clustering of contact zones, hybrid zones, and phylogeographic breaks in North America. Am Nat 166(5):581–591. https://doi.org/10.1086/491688
Taylor SA, Larson EL (2019) Insights from genomes into the evolutionary importance and prevalence of hybridization in nature. Nat Ecol Evol 3(2):170–177. https://doi.org/10.1038/s41559-018-0777-y
Taylor SA, Larson EL, Harrison RG (2015) Hybrid zones: windows on climate change. Trends Ecol Evol 30(7):398–406. https://doi.org/10.1016/j.tree.2015.04.010
Tocchio LJ, Gurgel-Gonçlaves R, Escobar LE, Peterson AT (2015) Niche similarities among white-eared opossums (Mammalia: Didelphidae): is ecological niche modeling relevant to setting species limits? Zool Scripta 44(1):1–10. https://doi.org/10.1111/zsc.12082
Todesco M, Pascual MA, Owens GL, Ostevik KL, Moyers BT, Hübner S et al (2016) Hybridization and extinction. Evol Appl 9(7):892–908. https://doi.org/10.1111/eva.12367
United States Fish and Wildlife Service (USFWS) (1987) Florida panther (Felis concolor coryi) recovery plan. Prepared by Florida Panther Interagency Committee for the US Fish and Wildlife Service, Atlanta, GA
van Wyk AM, Dalton DL, Hoban S, Bruford MW, Russo I-RM, Birss C et al (2017) Quantitative evaluation of hybridization and the impact on biodiversity conservation. Ecol Evol 7(1):320–330. https://doi.org/10.1002/ece3.2595
vonHoldt BM, Cahill JA, Fan Z, Gronau I, Robinson J, Pollinger JP et al (2016) Whole-genome sequence analysis shows that two endemic species of North American wolf are admixtures of the coyote and gray wolf. Sci Adv 2(7):e1501714. https://doi.org/10.1126/sciadv.1501714
vonHoldt BM, Brzeski KE, Aardema ML, Schell CJ, Rutledge LY, Fain SR et al (2022) Persistence and expansion of cryptic endangered red wolf genomic ancestry along the American Gulf coast. Mol Ecol 31(21):5440–5454. https://doi.org/10.1111/mec.16200
Waltari E, Guralnick RP (2009) Ecological niche modeling of montane mammals in the Great Basin, North America: examining past and present connectivity of species across basins and ranges. J Biogeogr 36(1):148–161
Wasserman TN, Cushman SA, Schwartz MK, Wallin DO (2010) Spatial scaling and multi-model inference in landscape genetics: Martes americana in Northern Idaho. Landscape Ecol 25:1601–1612. https://doi.org/10.1007/s10980-010-9525-7
Wasserman TN, Cushman SA, Shirk AS, Landguth EL, Littell JS (2021) Simulating the effects of climate change on population connectivity of American marten (Martes americana) in the Northern Rocky Mountains, USA. Landscape Ecol 27:211–225. https://doi.org/10.1007/s10980-011-9653-8
Whiteley AR, Fitzpatrick SW, Funk WC, Tallmon DA (2015) Genetic rescue to the rescue. Trends Ecol Evol 30(1):42–49. https://doi.org/10.1016/j.tree.2014.10.009
Wolf DE, Takebayashi N, Reiseberg LH (2001) Predicting the risk of extinction through hybridization. Conserv Biol 15(4):1039–1053. https://doi.org/10.1046/j.1523-1739.2001.0150041039.x
Wright PL (1953) Intergradation between Martes americana and Martes caurina in Western Montana. J Mammal 34(1):74–86. https://doi.org/10.2307/1375946
Yamazaki D, Ikeshima D, Tawatari R, Yamaguchi T, O’Loughlin F, Neal JC, Sampson CC, Kanae S, Bates PD (2017) A high-accuracy map of global terrain elevations. Geophys Res Lett 44(11):5844–5853. https://doi.org/10.1002/2017gl072874
Acknowledgements
We thank faculty and staff at the University of New Mexico’s Museum of Southwestern Biology and the University of Alaska’s Museum of the North for collecting, curating, and loaning the specimens used for this project; Carrie Hicks and Kody Baumgarder for laboratory assistance; Natalie Dawson, N. Anderson, and the Montana trapping community for sample collection; A. Townsend Peterson and Marlon E. Cobos for expert guidance in niche modeling.
Funding
This work was supported by a Grant-in-Aid to J.P.C. from the American Society of Mammalogists.
Author information
Authors and Affiliations
Contributions
J.P.C. and J.A.C. conceptualized the study. K.D.S. developed and piloted the molecular methods. J.P.C. and D.M.L. generated the molecular data. J.P.C. performed molecular analyses. N.A.F. built niche models, performed analyses, and made the figures. J.P.C. wrote the first draft. D.M.L., B.J.W., and J.A.C. provided significant editorial contributions. J.A.C funded the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Colella, J.P., Freymueller, N.A., Land, D.M. et al. Ecological displacement in a Rocky Mountain hybrid zone informs management of North American martens (Martes). Landsc Ecol 39, 125 (2024). https://doi.org/10.1007/s10980-024-01915-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10980-024-01915-y