Abstract
Objectives
Major lignocellulosic inhibitory compounds found in sugarcane-based industrial hydrolysate samples were tested in laboratory and industrial yeast strains, as well as in lactic acid bacteria, in order to verify their effects on important physiological parameters.
Results
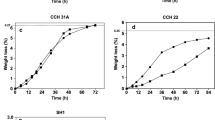
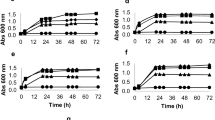
Saccharomyces cereviaise SA-1, an industrial strain, stood out as compared to the remaining strains for virtually all inhibitors investigated. This strain presented the highest growth rate and the lowest lag-phase in the presence of acetic acid, levulinic acid, p-coumaric acid, ferulic acid, and HMF, when compared to the other strains. In sugarcane-based hydrolysate fermentations, both SA-1 and CEN.PK113-7D presented similar fermentation performances. Industrial isolates of contaminating lactic acid bacteria were evaluated in the presence of an inhibitory cocktail, containing a mixture of 76.6 mM acetic acid, 1.3 mM HMF, 7.1 mM furfural, and 1.9 mM p-coumaric acid. Whilst all yeast strains were unable to grow under such conditions, bacteria had an average inhibition of roughly 50% on their growth rates.
Conclusions
Overall, industrial strain SA-1 might be a promising microbial chassis for second generation ethanol production and for future metabolic and evolutionary engineering strategies, and for strain robustness understanding.




Similar content being viewed by others
References
Adeboye PT, Bettiga M, Olsson L (2014) The chemical nature of phenolic compounds determines their toxicity and induces distinct physiological responses in Saccharomyces cerevisiae in lignocellulose hydrolysates. AMB Express 4:1–10. https://doi.org/10.1186/s13568-014-0046-7
Almeida JR, Modig T, Petersson A et al (2007) Increased tolerance and conversion of inhibitors in lignocellulosic hydrolysates by Saccharomyces cerevisiae J Chem Technol Biotechnol 82:340–349. https://doi.org/10.1002/jctb.1676
Argueso JL, Carazzolle MF, Mieczkowski PA et al (2009) Genome structure of a Saccharomyces cerevisiae strain widely used in bioethanol production. Genome Res 19:2258–2270. https://doi.org/10.1101/gr.091777.109
Auesukaree C, Damnernsawad A, Kruatrachue M et al (2009) Genome-wide identification of genes involved in tolerance to various environmental stresses in Saccharomyces cerevisiae J Appl Genet 50:301–310. https://doi.org/10.1007/BF03195688
Avanzi IR, Gracioso LH, Baltazar M, PG et al (2017) Rapid bacteria identification from environmental mining samples using MALDI-TOF MS analysis. Environ Sci Pollut Res 24:3717–3726. https://doi.org/10.1007/s11356-016-8125-8
Azhar AF, Bery MK, Colcord AR et al (1981) Factors affecting alcohol fermentation of wood acid hydrolysate. Biotechnol Bioeng 11:293–300
Basso LC, de Amorim HV, de Oliveira AJ, Lopes ML (2008) Yeast selection for fuel ethanol production in Brazil. FEMS Yeast Res 8:1155–1163. https://doi.org/10.1111/j.1567-1364.2008.00428.x
Basso TO, Gomes FS, Lopes ML et al (2014) Homo- and heterofermentative lactobacilli differently affect sugarcane-based fuel ethanol fermentation. Antonie van Leeuwenhoek Int J Gen Mol Microbiol 105:169–177. https://doi.org/10.1007/s10482-013-0063-6
Benjaphokee S, Hasegawa D, Yokota D et al (2012) Highly efficient bioethanol production by a Saccharomyces cerevisiae strain with multiple stress tolerance to high temperature, acid and ethanol. N Biotechnol 29:379–386
Bosma EF, Forster J, Nielsen AT (2017) Lactobacilli and pediococci as versatile cell factories—evaluation of strain properties and genetic tools. Biotechnol Adv 35:419–442. https://doi.org/10.1016/J.BIOTECHADV.2017.04.002
Boyer LJ, Vega JL, Klasson KT et al (1992) The effects of furfural on ethanol production by Saccharomyces cerevisiae in batch culture. Biomass Bioenergy 3:41–48. https://doi.org/10.1016/0961-9534(92)90018-L
Caspeta L, Nielsen J (2015) Thermotolerant yeast strains adapted by laboratory evolution show trade-off at ancestral temperatures and preadaptation to other stresses. MBio. https://doi.org/10.1128/mBio.00431-15
Caspeta L, Castillo T, Nielsen J (2015) Modifying yeast tolerance to inhibitory conditions of ethanol production processes. Front Bioeng Biotechnol 3:184. https://doi.org/10.3389/fbioe.2015.00184
Collograi KC, da Costa AC, Ienczak JL (2019) Effect of contamination with Lactobacillus fermentum I2 on ethanol production by Spathaspora passalidarum Appl Microbiol Biotechnol 103:5039–5050. https://doi.org/10.1007/s00253-019-09779-y
Costa VMM, Basso TOO, Angeloni LHPHP et al (2008) Production of acetic acid, ethanol and optical isomers of lactic acid by Lactobacillus strains isolated from industrial ethanol fermentations. Cienc E Agrotecnol 32:503–509
Cunha JT, Aguiar TQ, Romaní A, Oliveira C, Domingues L (2015) Contribution of PRS3, RPB4 and ZWF1 to the resistance of industrial Saccharomyces cerevisiae CCUG53310 and PE-2 strains to lignocellulosic hydrolysate-derived inhibitors. Bioresour Technol 191:7–16
De Angelis M, Gobbetti M (2011) Lactic Acid bacteria – Lactobacillus spp.: general characteristics, encyclopedia of dairy sciences, 2nd edn. pp 78–90
Della-Bianca BE, Basso TO, Stambuk BU et al (2013) What do we know about the yeast strains from the Brazilian fuel ethanol industry? Appl Microbiol Biotechnol 97:979–991. https://doi.org/10.1007/s00253-012-4631-x
Della-Bianca BE, Gombert AK (2013) Stress tolerance and growth physiology of yeast strains from the Brazilian fuel ethanol industry. Antonie van Leeuwenhoek Int J Gen Mol Microbiol 104:1083–1095. https://doi.org/10.1007/s10482-013-0030-2
Kammerer D, Claus A, Carle R, Schieber A (2004) Polyphenol screening of pomace from red and white grape varieties (Vitis vinifera L.) by HPLC-DAD-MS/MS. J Agric Food Chem. https://doi.org/10.1021/JF049613B
Klinke HB, Thomsen AB, Ahring BK (2004) Inhibition of ethanol-producing yeast and bacteria by degradation products produced during pre-treatment of biomass. Appl Microbiol Biotechnol 66:10–26. https://doi.org/10.1007/s00253-004-1642-2
Lam FH, Ghaderi A, Fink GR, Stephanopoulos G (2014) Engineering alcohol tolerance in yeast. Science. https://doi.org/10.1126/science.1257859
Larsson S, Palmqvist E, Hahn-Hägerdal B et al (1999) The generation of fermentation inhibitors during dilute acid hydrolysis of softwood. Enzyme Microb Technol 24:151–159. https://doi.org/10.1016/S0141-0229(98)00101-X
Liu ZL, Ma M, Song M (2009) Evolutionarily engineered ethanologenic yeast detoxifies lignocellulosic biomass conversion inhibitors by reprogrammed pathways. Mol Genet Genomics 282:233–244. https://doi.org/10.1007/s00438-009-0461-7
Luttik MAH, Kötter P, Salomons FA, van der Klei IJ, van Dijken JP, Pronk JT (2000) The Saccharomyces cerevisiaeICL2 gene encodes a mitochondrial 2-methylisocitrate lyase involved in propionyl-coenzyme a metabolism. J Bacteriol 182(24):7007–7013
Navarro AR (1994) Effects of furfural on ethanol fermentation by Saccharomyces cerevisiae: mathematical models. Curr Microbiol 29:87–90. https://doi.org/10.1007/BF01575753
Olsson L, Nielsen J (1997) On-line and in situ monitoring of biomass in submerged cultivations. Trends Biotechnol 15:517–522. https://doi.org/10.1016/S0167-7799(97)01136-0
Palmqvist E, Hahn-Hägerdal B (2000) Fermentation of lignocellulosic hydrolysates. I: inhibition and detoxification. Bioresour Technol 74:17
Pampulha ME, Loureiro-Dias MC (1990) Activity of glycolytic enzymes of Saccharomyces cerevisiae in the presence of acetic acid. Appl Microbiol Biotechnol 34:375–380. https://doi.org/10.1007/BF00170063
Reis VR, Bassi APG, Cerri BC et al (2018) Effects of feedstock and co-culture of Lactobacillus fermentum and wild Saccharomyces cerevisiae strain during fuel ethanol fermentation by the industrial yeast strain PE-2. AMB Express 8:23. https://doi.org/10.1186/s13568-018-0556-9
Russell JB (1992) Another explanation for the toxicity of fermentation acids at low pH: anion accumulation versus uncoupling. J Appl Bacteriol 73:363–370. https://doi.org/10.1111/j.1365-2672.1992.tb04990.x
Skerker JM, Leon D, Price MN et al (2013) Dissecting a complex chemical stress: chemogenomic profiling of plant hydrolysates. Mol Syst Biol 9:674. https://doi.org/10.1038/msb.2013.30
Soni ML, Meehrotra RN, Kapoor RC (1982) Dissociation constant and some thermodynamic parameters of leavulinic acid. Indian J Chem 21:1960197
van den Broek M, Bolat I, Nijkamp JF et al (2015) Chromosomal copy number variation in Saccharomyces pastorianus is evidence for extensive genome dynamics in industrial lager brewing strains. Appl Environ Microbiol 81:6253–6267. https://doi.org/10.1128/AEM.01263-15
van der Pol EC, Bakker RR, Baets P, Eggink G (2014) By-products resulting from lignocellulose pretreatment and their inhibitory effect on fermentations for (bio)chemicals and fuels. Appl Microbiol Biotechnol 98:9579–9593. https://doi.org/10.1007/s00253-014-6158-9
van Dijken JP, Bauer J, Brambilla L et al (2000) An interlaboratory comparison of physiological and genetic properties of four Saccharomyces cerevisiae strains. Enzyme Microb Technol 26:706–714
van Maris AJA, Abbott DA, Bellissimi E et al (2006) Alcoholic fermentation of carbon sources in biomass hydrolysates by Saccharomyces cerevisiae: current status. Antonie Van Leeuwenhoek Int J Gen Mol Microbiol 90:391–418. https://doi.org/10.1007/S10482-006-9085-7
Wahlbom CF, Hahn-Hägerdal B (2002) Furfural, 5-hydroxymethyl furfural, and acetoin act as external electron acceptors during anaerobic fermentation of xylose in recombinant Saccharomyces cerevisiae Biotechnol Bioeng 78:172–178. https://doi.org/10.1002/bit.10188
Zetty-Arenas AM, Alves RF, Portela CAF et al (2019) Towards enhanced n-butanol production from sugarcane bagasse hemicellulosic hydrolysate: Strain screening, and the effects of sugar concentration and butanol tolerance. Biomass Bioenergy 126:190–198. https://doi.org/10.1016/j.biombioe.2019.05.011
Acknowledgements
We would like to express our gratitude to Dr. Maria Anita Mendes and Dr. Claudio Augusto Oller do Nascimento for equipment and infrastructure, as well as for performing the identification of the bacterial isolates. We are very grateful to the excellent technical assistance performed by Thamiris Giacon. P.C. and D.P.P would like to thank the scholarship provided by CAPES and FAPESP, respectively. I.S.V and L.R.C. would like to thank the scholarship provided by CNPq.
Funding
This work was supported by Fundação de Apoio à Pesquisa do Estado de São Paulo (Grant Nos. 2015/50612-8; 2018/01759-4; 2018/17172-2) and also by Coordenação de Aperfeiçoamento de Pessoal de Nível Superior - Brasil (CAPES) - Finance Code 001.
Author information
Authors and Affiliations
Contributions
Conceptualization: [PC, DPP, TOB]; Methodology: [PC, BLVC, IVS]; Formal analysis and investigation: [PC, DPP, ATCA, LRC, TOB]; Writing—original draft preparation: [PC, DPP, TOB]; Writing—review and editing: [DPP, BLCV, TOB]; Funding acquisition: [TOB]; Resources: [TOB]; Supervision: [BLVC, TOB].
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Cola, P., Procópio, D.P., Alves, A.T.C. et al. Differential effects of major inhibitory compounds from sugarcane-based lignocellulosic hydrolysates on the physiology of yeast strains and lactic acid bacteria. Biotechnol Lett 42, 571–582 (2020). https://doi.org/10.1007/s10529-020-02803-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-020-02803-6