Abstract
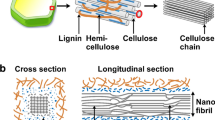
The cellulosome complex has evolved to degrade plant cell walls and, as such, combines tenacious binding to cellulose with diverse catalytic activities against amorphous and crystalline cellulose. Cellulolytic microorganisms provide an extensive selection of domains; those with affinity for cellulose, cohesins and their dockerin binding partners that define cellulosome stoichiometry and architecture, and a range of catalytic activities against carbohydrates. These robust domains provide the building blocks for molecular design. This review examines how protein modules derived from the cellulosome have been incorporated into chimaeric proteins to provide biosynthetic tools for research and industry. These applications include affinity tags for protein purification, and non-chemical methods for immobilisation and presentation of recombinant protein domains on cellulosic substrates. Cellulosomal architecture provides a paradigm for design of enzymatic complexes that synergistically combine multiple catalytic subunits to achieve higher specific activity than would be obtained using free enzymes. Multimeric enzymatic complexes may have industrial applications of relevance for an emerging carbon economy. Biocatalysis will lead to more efficient utilisation of renewable carbon-fixing energy sources with the added benefits of reducing chemical waste streams and reliance on petroleum.


Similar content being viewed by others
References
Adams JJ, Webb BA, Spencer HL et al (2005) Structural characterization of type II dockerin module from the cellulosome of Clostridium thermocellum: calcium-induced effects on conformation and target recognition. Biochemistry 44:2173–2182
Alber O, Noach I, Lamed R et al (2008) Preliminary X-ray characterization of a novel type of anchoring cohesin from the cellulosome of Ruminococcus flavefaciens. Acta Crystallogr Sect F Struct Biol Cryst Commun 64:77–80
Azriel-Rosenfeld R, Valensi M, Benhar I (2004) A human synthetic combinatorial library of arrayable dingle-chain antibodies based on shuffling in vivo formed CDRs into general framework regions. J Mol Biol 335:177–192
Barak Y, Handelsman T, Nakar D et al (2005) Matching fusion protein systems for affinity analysis of two interacting families of proteins: the cohesin–dockerin interaction. J Mol Recognit 18:491–501
Baratieri M, Baggio P, Fiori L et al (2008) Biomass as an energy source: thermodynamic constraints on the performance of the conversion process. Bioresour Technol 99:7063–7073
Bayer EA, Morag E, Lamed R (1994) The cellulosome—a treasure-trove for biotechnology. Trends Biotechnol 12:379–386
Bayer EA, Belaich J-P, Shoham Y et al (2004) The cellulosomes: multienzyme machines for degradation of plant cell wall polysaccharides. Annu Rev Microbiol 58:521–554
Bayer EA, Lamed R, Himmel ME (2007) The potential of cellulases and cellulosomes for cellulosic waste management. Curr Opin Biotechnol 18:237–245
Blouzard J-C, Bourgeois C, de Philip P et al (2007) Enzyme diversity of the cellulolytic system produced by Clostridium cellulolyticum explored by two-dimensional analysis: identification of seven genes encoding new dockerin-containing proteins. J Bacteriol 189:2300–2309
Bolam DN, Ciruela A, McQueen-Mason S et al (1998) Pseudomonas cellulose-binding domains mediate their effects by increasing enzyme substrate proximity. Biochem J 331:775–781
Boraston AB, Creagh AL, Alam MM et al (2001) Binding specificity and thermodynamics of a family 9 carbohydrate-binding module from Thermotoga maritima xylanase 10A. Biochemistry 40:6240–6247
Boraston AB, Nurizzo D, Notenboom V et al (2002) Differential oligosaccharide recognition by evolutionarily-related beta-1, 4 and beta-1, 3 glucan-binding modules. J Mol Biol 319:1143–1156
Boraston AB, Bolam DN, Gilbert HJ et al (2004) Carbohydrate-binding modules: fine-tuning polysaccharide recognition. Biochem J 382:769–781
Bruce ER (2008) Opportunities for renewable bioenergy using microorganisms. Biotechnol Bioeng 100:203–212
Carrard G, Koivula A, Soderlund H et al (2000) Cellulose-binding domains promote hydrolysis of different sites on crystalline cellulose. Proc Natl Acad Sci USA 97:10342–10347
Carvalho AL, Dias FM, Prates JA et al (2003) Cellulosome assembly revealed by the crystal structure of the cohesin–dockerin complex. Proc Natl Acad Sci USA 100:13809–13814
Carvalho AL, Dias FMV, Nagy T et al (2007) Evidence for a dual binding mode of dockerin modules to cohesins. Proc Natl Acad Sci USA 104:3089–3094
Cha J, Matsuoka S, Chan H et al (2007) Effect of multiple copies of cohesins on cellulase and hemicellulase activities of Clostridium cellulovorans mini-cellulosomes. J Microbiol Biotechnol 17:1782–1788
Cho H-Y, Yukawa H, Inui M et al (2004) Production of Minicellulosomes from Clostridium cellulovorans in Bacillus subtilis WB800. Appl Environ Microbiol 70:5704–5707
Craig SJ, Foong FC, Nordon RE (2006) Engineered proteins containing the cohesin and dockerin domains from Clostridium thermocellum provides a reversible, high affinity interaction for biotechnology applications. J Biotechnol 121:165–173
Craig SJ, Shu A, Xu Y et al (2007) Chimeric protein for selective cell attachment onto cellulosic substrates. Protein Eng Des Sel 20:235–241
Ding SY, Rincon MT, Lamed R et al (2001) Cellulosomal scaffoldin-like proteins from Ruminococcus flavefaciens. J Bacteriol 183:1945–1953
EU (2006) Directive 2003/30/EC of the European Parliament and of the Council of 8 May 2003 on the promotion of the use of biofuels and other renewable fuels for transport Institution. City Document
Fierobe H-P, Bayer EA, Tardif C et al (2002) Degradation of cellulose substrates by cellulosome chimeras. Substrate targeting versus proximity of enzyme components. J Biol Chem 277:49621–49630
Fierobe H-P, Mingardon F, Mechaly A et al (2005) Action of designer cellulosomes on homogeneous versus complex substrates: controlled incorporation of three distinct enzymes into a defined trifunctional scaffoldin. J Biol Chem 280:16325–16334
Fujita Y, Ito J, Ueda M et al (2004) Synergistic saccharification, and direct fermentation to ethanol, of amorphous cellulose by use of an engineered yeast strain codisplaying three types of cellulolytic enzyme. Appl Environ Microbiol 70:1207–1212
Galbe M, Zacchi G (2007) Pretreatment of lignocellulosic materials for efficient bioethanol production. Adv Biochem Eng Biotechnol 108:41–65
Ha J-S, Lee Y-M, Choi S-L et al (2008) Thermostable beta-glycosidase-CBD fusion protein for biochemical analysis of cotton scouring efficiency. J Microbiol Biotechnol 18:443–448
Hahn-Hagerdal B, Wahlbom CF, Gardonyi M et al (2001) Metabolic engineering of Saccharomyces cerevisiae for xylose utilization. Adv Biochem Eng Biotechnol 73:53–84
Haimovitz R, Barak Y, Morag E et al (2008) Cohesin–dockerin microarray: diverse specificities between two complementary families of interacting protein modules. Proteomics 8:968–979
Hammel M, Fierobe H-P, Czjzek M et al (2005) Structural basis of cellulosome efficiency explored by small angle X-ray scattering. J Biol Chem 280:38562–38568
Harhangi HR, Freelove ACJ, Ubhayasekera W et al (2003) Cel6A, a major exoglucanase from the cellulosome of the anaerobic fungi Piromyces sp. E2 and Piromyces equi. Biochim Biophys Acta/Gene Struct Expression 1628:30–39
Harry JG (2007) Cellulosomes: microbial nanomachines that display plasticity in quaternary structure. Mol Microbiol 63:1568–1576
Henrissat B, Coutinho PM, Danchin E et al (2008) Carbohydrate-active enzymes database. Retrieved August 2008, from http://www.cazy.org
Heyman A, Barak Y, Caspi J et al (2007) Multiple display of catalytic modules on a protein scaffold: nano-fabrication of enzyme particles. J Biotechnol 131:433–439
Hildén L, Johansson G (2004) Recent developments on cellulases and carbohydrate-binding modules with cellulose affinity. Biotechnol Lett 26:1683–1693
Hong SY, Lee JS, Cho KM et al (2006) Assembling a novel bifunctional cellulase-xylanase from Thermotoga maritima by end-to-end fusion. Biotechnol Lett 28:1857–1862
Hong J, Ye X, Zhang YHP (2007) Quantitative determination of cellulose accessibility to cellulase based on adsorption of a nonhydrolytic fusion protein containing CBM and GFP with its applications. Langmuir 23:12535–12540
Hsu S-h, Chu W-P, Lin Y-S et al (2004) The effect of an RGD-containing fusion protein CBD-RGD in promoting cellular adhesion. J Biotechnol 111:143–154
Hwang S, Ahn J, Lee S et al (2004) Evaluation of cellulose-binding domain fused to a lipase for the lipase immobilization. Biotechnol Lett 26:603–605
Jervis EJ, Haynes CA, Kilburn DG (1997) Surface diffusion of cellulases and their isolated binding domains on cellulose. J Biol Chem 272:24016–24023
Jervis EJ, Guarna MM, Doheny JG et al (2005) Dynamic localization and persistent stimulation of factor-dependent cells by a stem cell factor/cellulose binding domain fusion protein. Biotechnol Bioeng 91:314–324
Kang H-J, Uegaki K, Fukada H et al (2007) Improvement of the enzymatic activity of the hyperthermophilic cellulase from Pyrococcus horikoshii. Extremophiles 11:251–256
Katahira S, Mizuike A, Fukuda H et al (2006) Ethanol fermentation from lignocellulosic hydrolysate by a recombinant xylose- and cellooligosaccharide-assimilating yeast strain. Appl Microbiol Biotechnol 72:1136–1143
Kavoosi M, Meijer J, Kwan E et al (2004) Inexpensive one-step purification of polypeptides expressed in Escherichia coli as fusions with the family 9 carbohydrate-binding module of xylanase 10A from T. maritima. J Chromatogr B 807:87–94
Kavoosi M, Lam D, Bryan J et al (2007) Mechanically stable porous cellulose media for affinity purification of family 9 cellulose-binding module-tagged fusion proteins. J Chromatogr A 1175:187–196
Kumar GA, Sekaran G, Krishnamoorthy S (2006) Solid state fermentation of Achras zapota lignocellulose by Phanerochaete chrysosporium. Bioresour Technol 97:1521–1528
Kumar R, Singh S, Singh O (2008) Bioconversion of lignocellulosic biomass: biochemical and molecular perspectives. J Ind Microbiol 35:377–391
Kwan M, Guarna M, Boraston AB et al (2002) Self-activating factor X derivative fused to the C-terminus of a cellulose-binding module: production and properties. Biotechnol Bioeng 79:724–732
Lamed R, Setter E, Bayer EA (1983) Characterization of a cellulose-binding, cellulase-containing complex in Clostridium thermocellum. J Bacteriol 156:828–836
Lehtio J, Sugiyama J, Gustavsson M et al (2003) The binding specificity and affinity determinants of family 1 and family 3 cellulose binding modules. Proc Natl Acad Sci USA 100:484–489
Levy I, Paldi T, Shoseyov O (2004) Engineering a bifunctional starch-cellulose cross-bridge protein. Biomaterials 25:1841–1849
Linde M, Galbe M, Zacchi G (2008) Bioethanol production from non-starch carbohydrate residues in process streams from a dry-mill ethanol plant. Bioresour Technol 99:6505–6511
Madkour M, Mayer F (2003) Structural organization of the intact bacterial cellulosome as revealed by electron microscopy. Cell Biol Int 27:831–836
Malça J, Freire F (2006) Renewability and life-cycle energy efficiency of bioethanol and bio-ethyl tertiary butyl ether (bioETBE): assessing the implications of allocation. Energy 31:3362–3380
Maurice S, Dekel M, Shoseyov O et al (2003) Cellulose beads bound to cellulose binding domain-fused recombinant proteins; an adjuvant system for parenteral vaccination of fish. Vaccine 21:3200–3207
McAloon A, Taylor F, Yee W et al (2000) Determining the cost of producing ethanol from corn starch and lignocellulose feedstocks. National Renewable Energy Laboratory, U.S. Department of Energy, Denver, Colorado, pp 1–15 (NREL/TP-580-28893)
McLean BW, Bray MR, Boraston AB et al (2000) Analysis of binding of the family 2a carbohydrate-binding module from Cellulomonas fimi xylanase 10A to cellulose: specificity and identification of functionally important amino acid residues. Protein Eng 13:801–809
Merino ST, Cherry J (2007) Progress and challenges in enzyme development for biomass utilization. Adv Biochem Eng/Biotechnol 108:95–120
Mingardon F, Chanal A, Lopez-Contreras AM et al (2007a) Incorporation of fungal cellulases in bacterial minicellulosomes yields viable, synergistically acting cellulolytic complexes. Appl Environ Microbiol 73:3822–3832
Mingardon F, Chanal A, Tardif C et al (2007b) Exploration of new geometries in cellulosome-like chimeras. Appl Environ Microbiol 73:7138–7149
Mosier N, Wyman C, Dale B et al (2005) Features of promising technologies for pretreatment of lignocellulosic biomass. Bioresour Technol 96:673–686
Murashima K, Chen C-L, Kosugi A et al (2002) Heterologous production of Clostridium cellulovorans engB, using protease-deficient Bacillus subtilis, and preparation of active recombinant cellulosomes. J Bacteriol 184:76–81
Nagy T, Simpson P, Williamson MP et al (1998) All three surface tryptophans in Type IIa cellulose binding domains play a pivotal role in binding both soluble and insoluble ligands. FEBS Lett 429:312–316
Nahálka J, Gemeiner P (2006) Thermoswitched immobilization—a novel approach in reversible immobilization. J Biotechnol 123:478–482
Nishiyama Y, Sugiyama J, Chanzy H et al (2003) Crystal structure and hydrogen bonding system in cellulose; from synchrotron X-ray and neutron fiber diffraction. J Am Chem Soc 125:14300–14306
Nordon RE, Shu A, Camacho F, Milthorpe BK (2004) Hollow fiber assay for ligand-mediated cell adhesion. Cytometry 57A:39–44
Ong E, Alimonti JB, Greenwood JM et al (1995) Purification of human interleukin-2 using the cellulose-binding domain of a prokaryotic cellulase. Bioseparation 5:95–104
Otero JM, Panagiotou G, Olsson L (2007) Fueling industrial biotechnology growth with bioethanol. Adv Biochem Eng/Biotechnol 108:1–40
Pangu G, Johnston E, Petkov J et al (2007) Targeted particulate adhesion to cellulose surfaces mediated by bifunctional fusion proteins. Langmuir 23:10682–10693
Pell G, Williamson MP, Walters C et al (2003) Importance of hydrophobic and polar residues in ligand binding in the family 15 carbohydrate-binding module from Cellvibrio japonicus Xyn10C. Biochemistry 42:9316–9323
Perret S, Belaich A, Fierobe H-P et al (2004a) Towards designer cellulosomes in Clostridia: mannanase enrichment of the cellulosomes produced by Clostridium cellulolyticum. J Bacteriol 186:6544–6552
Perret S, Casalot L, Fierobe HP et al (2004b) Production of heterologous and chimeric scaffoldins by Clostridium acetobutylicum ATCC 824. J Bacteriol 186:253–257
Qi M, O’Brien JP, Yang J (2008) A recombinant triblock protein polymer with dispersant and binding properties for digital printing. Peptide Sci 90:28–36
Sabathe F, Soucaille P (2003) Characterization of the CipA scaffolding protein and in vivo production of a minicellulosome in Clostridium acetobutylicum. J Bacteriol 185:1092–1096
Sakon J, Irwin D, Wilson DB et al (1997) Structure and mechanism of endo/exocellulase E4 from Thermomonospora fusca. Nat Struct Biol 4:810–818
Schaeffer F, Matuschek M, Guglielmi G et al (2002) Duplicated dockerin subdomains of Clostridium thermocellum endoglucanase CelD bind to a cohesin domain of the scaffolding protein CipA with distinct thermodynamic parameters and a negative cooperativity. Biochemistry 41:2106–2114
Shapouri HDJ, Wang M (2001) The energy balance of corn ethanol: an update. Department of Agriculture, USA. Washington, DC, pp 1–15
Shi J, Chinn MS, Sharma-Shivappa RR (2008) Microbial pretreatment of cotton stalks by solid state cultivation of Phanerochaete chrysosporium. Bioresour Technol 99:6556–6564
Sticklen MB (2008) Plant genetic engineering for biofuel production: towards affordable cellulosic ethanol. Nat Rev Genet 9:433–443
Tardif C, Belaıch A, Fierobe HP et al (2006) Cellulosomes and cellulolysis. In: Kataeva I, Uversky VN (eds) The cellulosome. Nova Sciences Publishers, Hauppauge, NY, pp 231–260
Thormann K, Feustel L, Lorenz K et al (2002) Control of butanol formation in Clostridium acetobutylicum by transcriptional activation. J Bacteriol 184:1966–1973
Tomme P, Boraston A, McLean B et al (1998) Characterization and affinity applications of cellulose-binding domains. J Chromatogr B 715:283–296
von Ossowski I, Eaton JT, Czjzek M et al (2005) Protein disorder: conformational distribution of the flexible linker in a chimeric double cellulase. Biophys J 88:2823–2832
Wahlbom CF, van Zyl WH, Jonsson LJ et al (2003) Generation of the improved recombinant xylose-utilizing Saccharomyces cerevisiae TMB 3400 by random mutagenesis and physiological comparison with Pichia stipitis CBS 6054. FEMS Yeast Res 3:319–326
Waltz E (2008) Cellulosic ethanol booms despite unproven business models. Nat Biotechnol 26:8–9
Wang WX, Pelah D, Alergand T et al (2002) Characterization of SP1, a stress-responsive, boiling-soluble, homo-oligomeric protein from aspen. Plant Physiol 130:865–875
Warren RA (1996) Microbial hydrolysis of polysaccharides. Annu Rev Microbiol 50:183–212
Watkins K (2007) Human Development Report 2007/2008. Fighting climate change. Human solidarity in a divided world. United Nations Development Programme, New York, pp 1–384
Wingren A, Galbe M, Zacchi G (2003) Techno-economic evaluation of producing ethanol from softwood: comparison of SSF and SHF and identification of bottlenecks. Biotechnol Prog 19:1109–1117
Xu Q, Gao W, Ding SY et al (2003) The cellulosome system of Acetivibrio cellulolyticus includes a novel type of adaptor protein and a cell surface anchoring protein. J Bacteriol 185:4548–4557
Yang G, Miyamoto H, Yamane C et al (2007) Structure of regenerated cellulose films from cellulose/aqueous NaOH solution as a function of coagulation conditions. Polym J 39:34–40
Yeh M, Craig S, Lum M-G et al (2005) Effects of the PT region of EngD and HLD of CbpA on solubility, catalytic activity and purification characteristics of EngD-CBDCbpA fusions from Clostridium cellulovorans. J Biotechnol 116:233–244
Zhang P (2006) Investigation of novel quantum dots/proteins/cellulose bioconjugate using NSOM and fluorescence. J Fluoresc 16:349–353
Zverlov VV, Kellermann J, Schwarz WH (2005) Functional subgenomics of Clostridium thermocellum cellulosomal genes: identification of the major catalytic components in the extracellular complex and detection of three new enzymes. Proteomics 5:3646–3653
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Nordon, R.E., Craig, S.J. & Foong, F.C. Molecular engineering of the cellulosome complex for affinity and bioenergy applications. Biotechnol Lett 31, 465–476 (2009). https://doi.org/10.1007/s10529-008-9899-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-008-9899-7