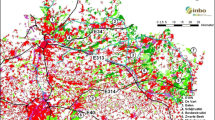
Abstract
Comparative landscape genetics studies provide insights on the impact of landscape elements on gene flow patterns of different species inhabiting the same geographic area. We investigated the population genetic structure of two sympatric ungulates, roe deer Capreolus capreolus and Northern chamois Rupicapra rupicapra, in a mountain area of the central Italian Alps (Trentino, northern Italy). A total of 122 chamois and 72 roe deer samples were genotyped by two species-specific panels of 11 polymorphic microsatellite loci and analyzed by aspatial and spatially explicit analyses. While the roe deer population resulted unstructured, a clear population structure was detected in chamois, with two main groups, one inhabiting the eastern and the other spread in the western part of the study area. Landscape genetics analysis confirmed these scenarios and revealed a different effect of landscape on gene flow. An IBD (Isolation-By-Distance) model best explained genetic variation in roe deer, while IBR (Isolation-By-Resistance) was found as the process underlying genetic variation patterns in chamois, suggesting arable lands, coniferous forests, watercourses, and main roads as potential barriers. Species distribution and landscape use might explain these results: roe deer mostly occupy valley floors relatively connected to each other, and their spatial behavior may promote gene flow across areas. On the other hand, chamois prefer higher elevations and their movements may be hindered by valleys, rivers, and road networks. This study highlights the different impacts of natural and anthropic landscape elements on gene flow in two sympatric species, resulting from their different ecological requirements.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Landscape genetic analyses integrate ecology, spatial statistics, and population genetics to develop and assess predictions about the effect of landscape features on animal movements and gene flow, and to identify environmental elements potentially acting as barriers and generating genetic discontinuities (Manel et al. 2003; Storfer et al. 2007). Landscape genetic studies could be particularly helpful to inform the management and conservation of different species/populations, including wild ungulate species, in a wide variety of situations (Segelbacher et al. 2010; Shafer et al. 2012; Portanier et al. 2018; Lecis et al. 2022). Besides a rich literature of landscape genetic studies focused on single taxa (e.g., Storfer et al. 2010; Keller et al. 2014; Richardson et al. 2016), comparative studies on distinct species inhabiting the same environment and/or geographic area have become more common (Olsen et al. 2011; Kierepka et al. 2016; Burkhart et al. 2017; Emel et al. 2019), providing interesting insights on different species’ interactions with landscape that can lead to contrasting patterns of genetic connectivity.
Landscape might similarly influence the spatial distribution of genetic diversity in closely related and geographically overlapping species (Olsen et al. 2011). However, it is not predictable how connectivity relates to site-specific conditions, and similar organisms may respond in a different way to the same landscape. For instance, three species of Ambystomatid salamanders showed different genetic structures in response to habitat fragmentation in USA (Whiteley et al. 2014), a motorway in Belgium was found to represent barrier to gene flow for red deer Cervus elaphus but not for wild boar Sus scrofa (Frantz et al. 2012), and different species–landscape interactions were described when analyzing two syntopic frog species in China (Garcia et al. 2017) and two closely related but ecologically distinct newt species in Poland (Antunes et al. 2022). Landscape effects on gene flow can thus sometimes differ strongly even among related sympatric species.
The Northern chamois (Rupicapra rupicapra, Linnaeus 1758) is an alpine ungulate inhabiting Europe and the Near East, with a typical range from 300 to 2500 m above sea level (a.s.l.) (Corlatti et al. 2022). In the past century, the species has been generally affected over most of its range by overhunting and consequent decline, although it now appears recovered and is widespread in the Alps (Buzan et al. 2013, Corlatti et al. 2022). Populations tend to be separated by low valleys, especially when these are impacted by human activities; however, they do not always represent barriers to chamois movements (Loison et al. 1999; Safner et al. 2019). Landscape genetic studies have been recently conducted on this species: Soglia et al. (2010) found three main clusters and genetic differences among chamois sampled over the Italian Alps, while Buzan et al. (2013) identified four geographically associated clusters in Slovenia and detected genetic isolation in some small populations. More recently, Leugger et al. (2022) described two main clades in the Alpine chamois population corresponding to south-western and eastern Alps, and limited connectivity in combination with large rivers or valleys acting as barriers to dispersal.
The European roe deer (Capreolus capreolus, Linnaeus 1758) is the most widespread ungulate species in Europe, inhabiting various ecosystems from the Mediterranean to Scandinavia, in a range between sea level and 2000 m a.s.l (Lorenzini et al. 2022). At the beginning of the last century, the species was almost extinct in different European regions, but has subsequently recovered almost everywhere (Apollonio et al. 2010). Among the landscape genetics studies recently conducted on this species, Zannèse et al. (2006) combined spatial and microsatellite analysis with animal morphology to define management units within the roe deer population in north-eastern Italy identifying two main regions, while Coulon et al. (2006) found that a combination of several landscape features with low permeability such as highways and rivers led to roe deer population differentiation in south-western France. Breyne et al. (2014) detected distance and infrastructure effects on Belgian roe deer populations, and Hepenstrick et al. (2012) identified fenced freeways as the main obstacles to gene flow in roe deer in Switzerland.
The two species are widespread over the Italian Alps. Chamois are represented by populations with varying densities, and in Trento province (covering around 6,200 km2), a total of 29,810 heads were estimated in 2020 (Provincia Autonoma di Trento Dipartimento Risorse Forestali—official data 2020). Roe deer inhabit most of central and northern Italy, after recovering due to natural dispersal and re-introductions (Apollonio et al. 2010; Lecis et al. 2023). In Trento province, a total of 35185 individuals were estimated in 2020 (Provincia Autonoma di Trento Dipartimento Risorse Forestali—official data 2020). These two ungulates have different ecology, land use, and altitude preferences, with chamois ranging at higher altitudes, forest, steep and rocky terrains, and roe deer usually inhabiting a wider range of habitats including woods and pastures. Italian Alps, and specifically the study area, are characterized by natural and artificial elements which can restrict species movements, such as topography, roads, valley urbanization, and fences used in agricultural practices.
Herein, we apply a comparative landscape genetic approach to two sympatric populations of Northern chamois and roe deer in an Alpine area, with the following objectives: (i) to estimate levels of genetic diversity and test for past bottlenecks in both populations, (ii) to assess the population genetic structure of the two species in the area, and (iii) to investigate spatial patterns and identify the effect of barriers or landscape connectivity on gene flow in both species.
Materials and methods
Study area and sample collection
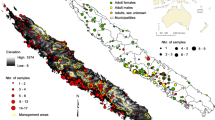
The study site is represented by the western part of Trento province, in the east-central Italian Alps. The area includes the Adamello-Presanella and Brenta massifs in the north and Mt. Cadria-Altissimo and Ledro Alps in the south (Fig. 1). The northern part of the study site is occupied by the Adamello Brenta Natural Park. Climate is transitional between semi-continental and alpine, average temperatures range between −5 and −10 °C in January and 20–25 °C in July (Chirichella et al. 2020). Elevation ranges from 65 to 3550 m a.s.l. (Presanella Massif). The study site, covering 1366 km2, is represented by a typical montane habitat, with more than 60% of the area above 1000 m a.s.l., and covered by forests (51% of the entire study area, among which 45% coniferous forests, 42% mixed forests, and 13% broad-leaved forests), shrubs (10%), meadows (8%), and rocky outcrops (21%). Two main rivers cross the area: Chiese and Sarca, and the region is characterized by a rich diversity of flora and fauna. Apart from Northern chamois and roe deer, red deer (Cervus elaphus) are abundant but seldom use the areas above the tree line (2000 m a.s.l.). European mouflon (Ovis gmelini musimon) and Alpine ibex (Capra ibex), which potentially use chamois habitat, are limited to few locations, having been introduced and reintroduced in the recent past only. Potential terrestrial predators include a stable population of brown bears (Ursus arctos), a small number of lynx (Lynx lynx), and occasionally wolves (Canis lupus; Groff et al. 2023). The golden eagle (Aquila chrysaetos; Pedrini et al. 2005) might rarely predate upon juveniles and is present in the area (Adamello Brenta Nature Park 2011). Rendena valley divides the study area in a NE-SW direction, separating western (Adamello, Presanella) from eastern massifs (Brenta, Cadria-Altissimo, Ledro). An orographic barrier might be represented by Canyon Limarò, between Cadria-Altissimo massif and Brenta Dolomites. Roads and linear infrastructures, urban settings along the valleys, fences used in agricultural practice, and human-exploited land are among the main potential landscape barriers found in this area (see Fig. S1).
The study site includes seven Northern chamois management areas (Adamello, Destra Chiese, Presanella, Brenta, Cadria-Altissimo, Misone-Casale, and Ledro) and four roe deer management districts (Chiese, Ledro, Rendena, Giudicarie), as well as a total of 90 hunting reserves. For the purpose of this study, a total of 122 chamois samples were collected between 2010 and 2015, while 72 roe deer samples were gathered during the 2011 hunting season (Fig. 1). Sampling was designed to be as representative as possible of local population distribution for both species. All samples consisted of ear tissue stored in absolute ethanol and at −20 °C until the analysis.
Microsatellite amplification and genotyping
DNA extraction was performed using GenElute Mammalian Genomic DNA Miniprep Kit (Sigma), following the manufacturer’s instructions. Roe deer samples were analyzed with a panel of 11 polymorphic autosomal microsatellites (Roe01, Roe06, Roe08, Roe09, NV16, NV21, NV24, RT1, ILSTS011, FCB304, BMC1009, Fickel and Reinsch 2000), amplified in three multiplex PCRs (multiplex 1A: Roe01, Roe08, Roe09; multiplex 2A: RT1, BMC1009; multiplex 1B: Roe06, ILSTS011), and the remaining loci in four simplex PCRs (NV16, NV21, NV24, FCB304) (Biosa et al. 2015; Lecis et al. 2023). Chamois samples were also genotyped at 11 microsatellite loci (ILSTS05, ILSTS011, BMC1009, FCB304, SRCRSP05, SRCRSP08, SRCRSP09, ETH10, ETH225, INRA05, INRA11, Arevalo et al. 2009). Three multiplex reactions were developed to amplify 9 loci (multiplex 1: ILSTS05, INRA05, SRCRSP08, SRCRSP09; multiplex 2: FCB304, SRCRSP05; multiplex 3: INRA11, ETH10, ETH225), and the remaining 2 loci (ILSTS011, BMC1009) were amplified independently. PCR conditions are reported in Supplementary Material (Tables S1 and S2).
PCR products were analyzed by BMR Genomics (Padova, Italy) using capillary electrophoresis on an ABI PRISM 3130xl Avant automatic sequencer (Applied Biosystems). Allele size was determined by analyzing electropherograms in Peak Scanner 1.0 software (Applied Biosystems).
Genetic diversity and population structure
Micro-Checker 2.2.3 (Van Oosterhout et al. 2004) was used to detect evidence of null alleles, stuttering, or large allele dropout. Microsatellite genotypes were then analyzed in GenAlex v.6.5 (Peakall and Smouse 2012) to calculate observed heterozygosity (Ho), expected heterozygosity (He), mean number of alleles per locus (k), inbreeding coefficient (Fis, 1000 permutations), and to calculate Fst indexes in both populations. We used SPAGeDi (Hardy andVekemans 2002) to compute pairwise relatedness (Rousset’s ar) in both species. Genepop v. 4 (Raymond and Rousset 1995) was implemented to assess deviations from Hardy-Weinberg equilibrium (HWE) and from linkage equilibrium (LE), using 10,000 dememorizations, 50 batches, and 10,000 iterations per batch in both tests. Sequential Bonferroni corrections for multiple comparisons were used to find significance levels. Evidence of recent effective population size reductions was investigated using the program Bottleneck V.1.2.02 (Piry et al. 1999), to test whether local C. capreolus and R. rupicapra populations showed an excess of heterozygosity, as expected after a recent bottleneck. The test was performed under a two-phase mutation model (TPM with 95% SMM, step mutation model), assessing significance by the Wilcoxon sign-rank test.
Population genetic structure was analyzed by Bayesian clustering and assignment tests implemented in Structure 2.3.4 (Pritchard et al. 2000; Falush et al. 2003) for both species, in order to identify the most likely number of genetic clusters within both datasets. Our results were based on simulations from 1 to 10 inferred populations (K = 1–10), and the following settings were applied for both species: admixture ancestry model, no population information, correlated allele frequencies, 200,000 burn-in, and 200,000 subsequent iterations of data collection. Results were summarized and the best K value was calculated using the ΔK approach developed by Evanno et al. (2005) in Structure Harvester (Earl and VonHoldt 2012) to identify the best clustering solution. We employed Pophelper (Francis 2017) to edit Structure results and produce the final plots. GenAlex was used to assess genetic differentiation across samples by principal coordinate analysis (PCoA) and to examine microgeographic patterns of genetic structure by spatial genetic autocorrelation analysis. The latter was based on matrices of pairwise inter-individual genetic distances and pairwise topographic distances estimated by the program. Up to five different distance classes of relatively even sample size were selected, and 95% confidence interval was estimated by 999 permutations.
Landscape genetics analyses
A landscape genetics approach was used to test the effect of Isolation-By-Distance (IBD) and Isolation-By-Resistance (IBR) on the genetic variation patterns of the roe deer and chamois populations in the study area. We parametrized landscape resistance by simultaneously optimizing multiple categorical and continuous surfaces through the genetic algorithm implemented in the R package ResistanceGA (Peterman 2018). Based on species ecology, we tested the effect of two categorical (land cover and main roads) and three continuous (elevation, slope, and snow cover) surfaces on chamois and roe deer gene flow. The land cover surface (500 × 500-m pixel resolution) was obtained by overlapping the Trentino digital forest map (http://www.territorio.provincia.tn.it/portal/server.pt/community/portale_geocartografico_trentino/254/portale_geocartografico_trentino/18994) over the Corine Land Cover 2018 (https://land.copernicus.eu/pan-european/corine-land-cover/clc2018). The original land cover classes at the 4th CLC level were reclassified into ten categories: urban areas, arable lands, mixed broadleaved forests, beech forests, coniferous forests, meadows and pastures, transitional vegetation, rocky areas, glaciers, water bodies. The road surface was obtained from OpenStreetMap (OpenStreetMap contributors 2023) rasterized at a 500 × 500-m resolution map where cells crossed by roads had a value equal to 1 while all other cells had a value equal to 0. Elevation and slope surfaces were derived from the Digital Elevation Model of Trentino (http://www.territorio.provincia.tn.it/portal/server.pt/community/portale_geocartografico_trentino/254/ portale_geocartografico_trentino/18994) resampled at a 500 × 500-m pixel resolution. Finally, the snow cover surface was obtained by averaging the maximum snow cover over the period 2000–2015 (Hall and Riggs 2021). The optimization process proceeds by calculating pairwise ecological distances from alternative single or composite resistance surfaces and by regressing genetic against ecological distances and fitting linear mixed-effects models with a maximum likelihood population effects parameterization (MLPEs) (Peterman 2018). Optimization is repeated until the best model is found, according to parametrization of the genetic algorithm and a chosen goodness-of-fit measure. We used the pairwise Rousset’s ar genetic distance (Rousset 2000) calculated for all pairs of individuals as dependent variable, and the ecological distance calculated using Circuitscape 5.0 implemented in Julia (Hall et al. 2021; McRae et al. 2008, 2016) as fixed effect. For categorical surfaces, the optimization process iteratively develops alternative resistance surfaces by assigning a new set of resistance values to each class. For continuous variables, alternative resistance surfaces are obtained by transforming original values through alternative functions, described by a shape and a magnitude parameter. For this study, we tested four alternative functions: monomolecular, reverse monomolecular, inverse monomolecular, inverse-reverse monomolecular (Peterman 2018). For both categorical and continuous surfaces, we set a maximum resistance value equal to 100. We compared all possible combinations of surfaces, including single and multiple, which are composed by summing up alternative single surfaces.
Models were ranked according to AICc and those with a ΔAICc < 2 were considered as equally valid. A pseudo bootstrap procedure (N = 100) was applied to assess the contribution of each surface and to evaluate model performance by reducing type I error rates (Winiarski et al. 2020). Pairwise genetic and ecological distance matrices generated from each optimized surface were subsampled (75% of the complete sample) without replacement, each time the MLPE model was refitted and the AICc values calculated (Peterman 2018). Surface contribution is defined as the frequency at which a model is the top-ranked. For each species, we ran the whole optimization procedure three times to confirm convergence and parameter estimates (Peterman 2018). We used the optimized resistance surface to develop a current density map of the study area to represent the species’ gene flow. Following the approach of Koen et al. (2014), we designed a 12-km-wide buffer around the study area, roughly 20% of its length and 30% of its width, and randomly selected 100 nodes around the buffer perimeter. Circuitscape was run on these 100 nodes, and then, the buffer was removed to obtain a current density map independent from the geographic location of the original samples. Analyses were performed using R v. 4.2.0 (R Core Team 2022) within the GALILEO 100 HPC System at CINECA (www.hpc.cineca.it/).
Results
A total of 72 roe deer samples and 122 chamois samples were successfully genotyped at 11 microsatellite loci respectively, with no evidence of genotyping error or null alleles identified by the preliminary inspection in Micro-Checker. Alleles ranged from 2 to 14 per locus in chamois and from 2 to 9 per locus in roe deer. In sampled chamois, the less polymorphic locus was ILSTS011 (2 alleles, He = 0.025) and the most polymorphic loci were INRA11 and FCB304 (14 alleles, He = 0.755 and 12 alleles, He = 0.809, respectively), while in roe deer, Roe01 resulted as the less variable locus (2 alleles, He = 0.488), and RT1 and NV21 were the most polymorphic (9 alleles, He = 0.801 and 8 alleles, He = 0.810, respectively). Tables 1 and 2 summarize genetic diversity statistics in chamois and roe deer populations. Overall, genetic diversity was higher in roe deer (He = 0.707, Ho = 0.699) than in chamois (He = 0.663, Ho = 0.586).
Departure from Hardy-Weinberg equilibrium, due to heterozygote deficiency, was highly significant at two loci (SRCRSP08, ETH225) and significant at locus BMC1009 in the chamois population (see Table 1), while deviation resulted significant at three loci (Roe01, Roe06, Roe08) in roe deer. In the latter, there was evidence of heterozygote excess only at one locus (Roe01, Table 2). No evidence of non-random association of genotypes at different loci was detected by linkage equilibrium analysis (non-significant p-values for all locus pairs after sequential Bonferroni correction). The values of Fis over all loci resulted low and not significant in roe deer (Fis = 0.0057, ns) indicating low levels of local inbreeding, but high and significant in chamois (Fis = 0.1163, p < 0.001) showing local isolation of different nuclei. There was no evidence of bottleneck effects by analyzing data of chamois and roe deer, as both analyses showed a normal L-shaped distribution of allele frequencies and no significant deviation from the expected levels of heterozygosity in Bottleneck (TPM model, Wilcoxon test not significant for both species: chamois p = 0.995; roe deer p = 0.319). Spatial correlation analysis performed on the roe deer population showed a significant positive correlation for the first distance class only (< 8 km; p = 0.004) and values not significantly different from zero at higher distances (see Supplementary Information, Fig. S2).
Bayesian analyses of population structure revealed sharply different results for chamois and roe deer populations in Trento province. When analyzing the chamois population dataset, the most likely partition was identified at K = 2 (Fig. S3), with individuals’ assignment to cluster I averaging q = 0.46, while q = 0.54 to cluster II. After mapping the chamois specimens over the sampling area, the Northwest (NW, Adamello-Presanella) appeared predominantly inhabited by animals assigned to cluster I (mean qI = 0.735), while the Southeast (SE, Cadria-Altissimo-Ledro) was occupied by individuals mostly assigned to cluster II (mean qII = 0.905). Chamois sampled in the Northeast (NE, Brenta) region appeared intermediate between the two clusters (mean qII = 0.680, Fig. 2). Instead, cluster analysis of roe deer genotypes did not evidence any structure, showing a slight likelihood decrease when increasing K, and therefore suggesting a panmictic population (Figs. S4 and 3).
Bar plots illustrating the genetic structure of chamois in Trento province, inferred by Bayesian cluster analysis in Structure. K = 2 was selected as the best clustering option and individual assignment proportion (q-value) to the two inferred clusters is represented by colors. Population codes: NW, North-West; NE, North-East; SE, South-East
Bar plots illustrating the absence of genetic structure in roe deer in Trento province, inferred by Bayesian cluster analysis in Structure (K = 2). Individual assignment proportion (q-value) to the two inferred clusters is represented by colors. The best solution is represented by K = 1. Population codes: NE, North-East; NW, North-West; SE, South-East
PCoA analyses confirmed these results, with some degree of differentiation and a partial overlap among chamois from different massifs (NW, NE, SE, Fig. 4), and complete overlap among roe deer from different areas (Fig. 5).
Landscape genetics analyses revealed a different effect of landscape composition and configuration on each species’ gene flow.
As for the chamois, the AICc-based bootstrap model selection supported IBR as the process underlying the observed genetic variation patterns. Specifically, two optimized surfaces, a single surface (land cover) and a multiple surface (land cover + main roads), were associated to the first (land cover: avg.weight = 0.52, Δavg.AICc = 0, 71% top model) and second (land cover + main roads: avg.weight = 0.26, Δavg.AICc = 1.93, 24% top model) top models, respectively. Since models were both valid (Δavg.AICc < 2), we considered the resistance values generated in the resistance map built from the multiple surface (land cover + main roads) to account for all landscape features that had a non-negligible effect in driving chamois gene flow. In the optimized resistance map, the lowest resistance to the chamois movement was offered by rocky areas (i.e., optimal habitat type for the species) followed by urban areas, glaciers, beech forests, and transitional vegetation cover. Intermediate resistance values were associated with mixed broadleaved forests and meadows and pastures, while highest values were associated with arable lands, water bodies, and coniferous forests (Table 3). In cells crossed by roads, the resistance value of each land cover class increased by about 50 units, showing that these linear infrastructures appeared to act as barriers to chamois (Table 3). Conversely, when analyzing the roe deer population, the Euclidean distance model was the top-ranked by the AICc-based bootstrap model selection (avg.weight = 0.96, Δavg.AICc = 0, 100% top model), showing that an IBD process drove the genetic variation patterns observed for this species.
A current density map was generated for chamois from the resistance surface optimized under the IBR hypothesis and is reported in Fig. 6, showing corridors for the species’ gene flow obtained from the optimized multisurface “land cover + main roads.”
Current density map obtained using Circuitscape, showing corridors for chamois gene flow obtained from the optimized multisurface “land cover + main roads.” The green-red color gradient indicates increasing current density values (threshold values of displayed classes are the quantiles of current density values distribution). Main roads are represented in grey
Discussion
Genetic and landscape genetics results obtained in this study, based on roe deer and Alpine chamois sampled over the Trento province in Northern Italy, allow us to infer several aspects of their population structure and their relationships with the same landscape elements and potential barriers. Both ungulate populations apparently retain a good amount of genetic variability, with some differences. Observed heterozygosity values in chamois appeared lower than expected, while in roe deer, expected and observed heterozygosity values almost coincided (Tables 1 and 2). These results are consistent with those obtained in previous studies on Northern chamois (Crestanello et al. 2009; Soglia et al. 2010) and on roe deer inhabiting other European areas (Coulon et al. 2006; Biosa et al. 2015; Vasiljevic et al. 2022). No bottleneck effect was detected in the two populations analyzed, consistently with the described genetic variation. Chamois in Trento province were however characterized by a high inbreeding coefficient (Fis, see Table 1), especially when compared to the value observed in roe deer (Table 2), although a large Fis variation among loci was detected in both species.
In order to maintain such levels of genetic variation, preserving gene flow within these two native wild populations would be essential. Bayesian cluster analysis performed using Structure highlighted a different scenario when applied to the two ungulate populations analyzed. We did not find any significant population structure in sampled roe deer inhabiting the study area, apparently connected by gene flow (Fig. 3). Instead, a clear population structure was detected in chamois, with two main clusters: one predominant in individuals sampled in the North-West (NW) of the study area (Adamello, Presanella) and the other found in animals inhabiting Mt Cadria in the South-East (SE) and prevalent in chamois sampled in the North-East (NE), corresponding to the Brenta massif (Fig. 2).
Landscape genetics analysis confirmed these scenarios and revealed a different effect of landscape on the two species’ gene flow. In the case of chamois, IBR was identified as the process underlying the observed genetic variation patterns, and the selected model suggested the role of arable lands, coniferous forests, watercourses, and main roads as potential barriers for this species. Indeed, chamois prefer higher elevations and their movements may be hindered by landscape elements such as anthropized valleys, large rivers, and road networks. As already mentioned, this species inhabits most of the main mountain massifs in Europe, being adapted for life in rocky terrains, which represent escape grounds from predators, and cold climates (Corlatti et al. 2022). Chamois populations are found at elevations up to 2500 m a.s.l., in a range of habitats including alpine and subalpine meadows, pastures, forests, and clearings, but also on slopes, rocky areas, and shrublands (Corlatti et al. 2022). Lack of continuity in these habitats and overhunting (Baumann et al. 2005) highly fragmented chamois populations. Overall, the Northern chamois is a habitat specialist ungulate occupying “continental archipelagos” of fragmented high altitudes rocky habitats (Buzan et al. 2013). Our results suggest that connectivity among these “islands” might be limited, therefore playing a role in generating the observed genetic structure. However, the observed pattern refers to a limited geographic scale and might change when replicating the study over a broader area.
Landscape elements, local orography, and land use, as well as the different species’ ecology, might thus explain our results. Rendena valley, that parts the study area in East-West direction, with western massifs (Adamello, Presanella) on one side and eastern massifs (Brenta, Cadria-Altissimo, Ledro) on the other side, is anthropized and crossed by Sarca river and SS239 road. The current density map (Fig. 6) shows that this valley represents a NE-SW barrier to gene flow, determining genetic isolation between chamois inhabiting the western and eastern massifs. Canyon Limarò, between Cadria-Altissimo massif and Brenta Dolomites, is also crossed by Sarca river, and apparently represents a semi-permeable barrier for the species movements, reducing genetic connectivity between NE and SE of the study area. On the eastern side of the study area, a potential corridor is present between Cadria-Altissimo and Adamello, enabling North-South movements, due to higher valley elevation and less road network development. High connectivity levels can support North-South movements within the western part (Adamello, Presanella massifs) of the study area. Finally, Campo Carlo Magno (1700 m), an alpine pass in the north, might also represent a potential corridor for chamois.
Although gene flow might also be affected by other variables (such as demography, ethology, and genetic drift), the effect of orography on the connectivity between chamois populations was also highlighted in recent studies conducted at the contact zone of two mountain ranges in Slovenia (Buzan et al. 2013) and across the Alps (Leugger et al. 2022).
As mentioned, no genetic structure was detectable in roe deer, and an IBD model was best explaining genetic variation in this species, which tends to occupy valley floors relatively connected to each other and is characterized by a spatial behavior promoting gene flow across areas, especially during the rut period (Debeffe et al. 2014). The observed pattern of genetic autocorrelation, indicating that individuals living within a range of 8 km are slightly more related than random pairs in the population, suggests that the apparent lack of structure and the IBD are generated by a majority of philopatric individuals and a minority of long-range dispersers in the population, as observed in this species (Debeffe et al. 2012). As reviewed by Lorenzini et al. (2022), roe deer is an ungulate species adapted to wooded habitat, particularly at edges with open areas providing good feeding resources for browsers. Roe deer also show behavioral plasticity, as habitat composition, availability, and distribution of resources might affect habitat selection. Main roads probably do not represent a significant barrier for roe deer, as demonstrated by the high number of road kills registered in Trento province (295 per year between 1993 and 2008, Mustoni et al. 2012), compared to a very low number (0.68 per year) registered for chamois in the same period. Previous studies on roe deer in Switzerland revealed the influence of transportation infrastructure on their genetic divergence, but no impact on genetic diversity (Kuehn et al. 2007) and no effect on gene flow of a highly frequented but unfenced railway track (Hepenstrick et al. 2012).
The different effects of landscape elements on the two ungulate species in this area were also suggested by a study promoted by Trento province, highlighting the diverse use of wildlife corridors by chamois and roe deer (Mustoni et al. 2012). Among 67 potential corridors for wild ungulates, only 7% were suitable for chamois, while 58% may be used by roe deer, providing a higher connectivity for this species’ movements. Albeit our results’ robustness would benefit from replication, they already highlight the different impacts of orography, land use, and anthropic infrastructures on gene flow in two sympatric species, as a result of their different ecological requirements. The observed population structure and restricted gene flow among chamois nuclei inhabiting the Trento province should be taken into account for future species management and conservation planning. Further research is recommended to improve landscape connectivity and assure the presence of potential corridors for wild ungulate species in the alpine region.
Data availability
Datasets generated during the current study are available from the corresponding author on request.
References
Adamello Brenta Nature Park (2011) Golden eagle (Aquila chrysaetos) monitoring in Adamello Brenta Nature Park. Technical report edited by Adamello Brenta Natural Park, Strembo, Trento, Italy
Antunes B, Figueiredo-Vázquez C, Dudek K, Liana M, Pabijan M, Zieliński P, Babik W (2022) Landscape genetics reveals contrasting patterns of connectivity in two newt species (Lissotriton montandoni and L. vulgaris). Mol Ecol. https://doi.org/10.1111/mec.16543
Apollonio M, Ciuti S, Pedrotti L, Banti P (2010). In: Apollonio M, Andersen R, Putman R (eds) European ungulates and their management in the 21st century. Cambridge University Press, Cambridge, UK
Arevalo E, Holder DA, Derr JN, Bhebhe E, Linn RA, Ruvuna F, Davis SK, Taylor JF (2009) Caprine microsatellite dinucleotide repeat polymorphisms at the SR-CRSP-1, SR-CRSP-2, SR-CRSP-3, SR-CRSP-41 and SRCRSP-5 loci. Anim Genet 25(3):202–202
Baumann M, Babotai C, Schibler J (2005) Native or naturalized? Validating alpine chamois habitat models with archaeozoological data. Ecol Appl 15:1096–1110
Biosa D, Scandura M, Tagliavini J, Luccarini S, Mattioli L, Apollonio M (2015) Patterns of genetic admixture between roe deer of different origin in central Italy. J Mammal 96(4):827–838
Breyne P, Mergeay J, Casaer J (2014) Roe deer population structure in a highly fragmented landscape. Eur J Wildl Res 60(909–917):82
Burkhart JJ, Peterman WE, Brocato ER et al (2017) The influence of breeding phenology on the genetic structure of four pond-breeding salamanders. Ecol Evol 7:4670–4681. https://doi.org/10.1002/ece3.3060
Buzan VE, Bryja J, Zemanová B, Kryštufek B (2013) Population genetics of chamois in the contact zone between the Alps and the Dinaric Mountains: uncovering the role of habitat fragmentation and past management. Conserv Genet 14:401–412
Chirichella R, Stephens PA, Mason TE, Apollonio M (2020) Contrasting effects of climate change on Alpine chamois. J Wildl Manage. https://doi.org/10.1002/jwmg.21962
Coulon A, Guillot G, Cosson JF, Angibault JMA, Aulagnier S, Cargnelutti B, Galan M, Hewison AJM (2006) Genetic structure is influenced by landscape features: empirical evidence from a roe deer population. Mol Ecol 15:1669–1679
Corlatti L, Herrero J, Ferretti F, Anderwald P, Garcia-Gonzalez R, Hammer S, Nores C, Rossi L, Lovari S (2022) Northern Chamois Rupicapra rupicapra (Linnaeus, 1758) and Southern Chamois Rupicapra pyrenaica (Bonaparte, 1845). Handbook of the European Mammals, Springer Nature
Crestanello B, Pecchioli E, Vernesi C, Mona S, Martinkova N, Janiga M, Hauffe HC (2009) Bertorelle G (2009) The genetic impact of translocations and habitat fragmentation in chamois (Rupicapra) spp. J Hered 100(6):691–708
Debeffe L, Morellet N, Cargnelutti B, Lourtet B, Bon R, Gaillard JM, Hewison AJM (2012) Condition dependent natal dispersal in a large herbivore: heavier animals show a greater propensity to disperse and travel further. J Anim Ecol 81:1327–1337
Debeffe L, Focardi S, Bonenfant C, Hewison AJM, Morellet N, Vanpé C, Heurich M, Kjellander P, Linnell JDC, Mysterud A, Pellerin M, Sustr P, Urbano F, Cagnacci F (2014) A one night stand? Reproductive excursions of female roe deer as a breeding dispersal tactic. Oecologia 176:431–443
Earl DA, VonHoldt BM (2012) Structure Harvester: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361
Emel SL, Olson DH, Knowles LL, Storfe A (2019) Comparative landscape genetics of two endemic torrent salamander species, Rhyacotriton kezeri and R. variegatus: implications for forest management and species conservation. Conserv Genet 20:801–815
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software structure: a simulation study. Mol Ecol 14:2611–2620
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567–1587
Fickel J, Reinsch A (2000) Microsatellite markers for the European roe deer (Capreolus capreolus). Mol Ecol 9:994–995
Francis RM (2017) Pophelper: an R package and web app to analyse and visualize population structure. Mol Ecol Resour 17:27–32
Frantz AC, Bertouille S, Eloy MC, Licoppe A, Chaumont F, Flamand MC (2012) Comparative landscape genetic analyses show a Belgian motorway to be a gene flow barrier for red deer (Cervus elaphus), but not wild boars (Sus scrofa). Mol Ecol 21:3445–3457
Garcia VO, Ivy C, Fu J (2017) Syntopic frogs reveal different patterns of interaction with the landscape: a comparative landscape genetic study of Pelophylax nigromaculatus and Fejervarya limnocharis from central China. Ecol Evol 7(22):9294–9306
Groff C, Angeli F, Baggia M, Bragalanti N, Zanghellini P, Zeni M (eds) (2023) 2022 Large Carnivores Report. Autonomous Province of Trento’s Wildlife Department, Trento, Italy. https://grandicarnivori.provincia.tn.it/Rapporto-Orso-e-grandi-carnivori
Hall KR, Anantharaman R, Landau VA, Clark M, Dickson BG, Jones A, Platt J, Edelman A, Shah VB (2021) CIRCUITSCAPE in Julia: empowering dynamic approaches to connectivity assessment. Land 10(3):301
Hall DK, Riggs GA (2021) MODIS/Terra Snow Cover 8-Day L3 Global 500m SIN Grid, Version 61 [Data Set]. Boulder, Colorado USA. NASA National Snow and Ice Data Center Distributed Active Archive Center. https://doi.org/10.5067/MODIS/MOD10A2.061. Accessed 17 Apr 2023
Hardy OJ, Vekemans X (2002) SPAGeDi: a versatile computer program to analyse spatial genetic structure at the individual or population levels. Mol Ecol Notes 2:618–620
Hepenstrick D, Thielb D, Holderegger R, Gugerlia F (2012) Genetic discontinuities in roe deer (Capreolus capreolus) coincide with fenced transportation infrastructure. Basic Appl Ecol 13:631–638
Keller D, Holderegger R, van Strien MJ, Bolliger J (2014) How to make landscape genetics beneficial for conservation management? Conserv Genet 16:503–512. https://doi.org/10.1007/s10592-014-0684-y
Kierepka EM, Anderson SJ, Swihart RK, Rhodes OE Jr (2016) Evaluating the infuence of life-history characteristics on genetic structure: a comparison of small mammals inhabiting complex agricultural landscapes. Ecol Evol 6:6376–6396
Koen EL, Bowman J, Sadowski C, Walpole AA (2014) Landscape connectivity for wildlife: development and validation of multispecies linkage maps. Methods Ecol Evol 5(7):626–633
Kuehn R, Hindenlang KE, Holzgang O, Senn J, Stoeckle B, Sperisen C (2007) Genetic effect of transportation infrastructure on roe deer populations (Capreolus capreolus). J Hered 98(1):13–22
Lecis R, Dondina O, Orioli V, Biosa D, Canu A, Fabbri G, Iacolina L, Cossu A, Bani L, Apollonio M (2022) Scandura M (2022) Main roads and land cover shaped the genetic structure of a Mediterranean island wild boar population. Ecol Evol 12:e8804
Lecis R, Bongi P, Russo C, Cecchi F, DelFrate M, Chirichella R, Apollonio M, Scandura M (2023) Contribution of a native roe deer lineage to the recolonisation of the northern Apennines. Hystrix, Italy. https://doi.org/10.4404/hystrix-00585-2022
Leugger F, Broquet T, Karger DN, Rioux D, Buzan E, Corlatti L, Crestanello B, Gaudin N, Hauffe HC, Roleckova B, Sprem N, Tissot N, Tissot S, Valterova R, Yannic G, Pellissier L (2022) Dispersal and habitat dynamics shape the genetic structure of the Northern chamois in the Alps. J Biogeogr 49:1848–1861
Loison A, Jullien JM, Menaut P (1999) Subpopulation structure and dispersal in two populations of chamois. J Mammal 80:620–632
Lorenzini , Hewison AJM, Gaillard JM, Garofalo L, Rossi L et al (2022) European roe deer Capreolus capreolus (Linnaeus, 1758) Hackländer K, Zachos FE (eds) Handbook of the Mammals of Europe. Springer
Manel S, Schwartz MK, Luikart G, Taberlet P (2003) Landscape genetics: combining landscape ecology and population genetics. Trends Ecol Evol 18:189–197
McRae BH, Dickson BG, Keitt TH, Shah VB (2008) Using circuit theory to model connectivity in ecology and conservation. Ecology 10:2712–2724
McRae BH, Shah VB, Edelman A (2016) Circuitscape: modeling landscape connectivity to promote conservation and human health. The Nature Conservancy, Fort Collins CO
Mustoni A, Zibordi F, Cavedon M, Armanini M (2012) Provincia Autonoma di Trento I Grandi Mammiferi in Trentino: Corridoi faunistici e investimenti stradali. Report by Adamello Brenta Nature Park
Olsen JB, Crane PA, Flannery BG, Dunmall K, Templin WD, Wenburg JK (2011) Comparative landscape genetic analysis of three Pacific salmon species from subarctic North America. Conserv Genet 12:223–241
OpenStreetMap contributors (2023) Planet Dump. Retrieved from https://osmit-estratti.wmcloud.org/
Peakall R, Smouse PE (2012) Genalex 6.5: genetic analysis in Excel. Population genetic software for teaching and research- an update. Bioinformatics 28:2537–2539
Pedrini P, Caldonazzi M, Zanghellini S (2005) Atlante degli Uccelli nidificanti e svernanti in provincia di Trento. Museo Tridentino di Scienze Naturali, Trento. Studi Trentini di Scienze Naturali. Acta Biologica 80:154–157
Peterman WE (2018) ResistanceGA: An R package for the optimization of resistance surfaces using genetic algorithms. Methods Ecol Evol 9(6):1638–1647
Piry S, Luikart G, Cornuet JM (1999) Bottleneck: a computer program for detecting recent reductions in the effective population size using allele frequency data. J Hered 90:502–503
Portanier E, Larroque J, Garel M, Marchand P, Maillard D, Bourgoin G, Devillard S (2018) Landscape genetics matches with behavioral ecology and brings new insight on the functional connectivity in Mediterranean mouflon. Landscape Ecol 33:1069–1085
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Provincia Autonoma Di Trento, Dipartimento Risorse Forestali e Montane SFF, Ufficio Faunistico (2020) Report, Analisi delle consistenze e dei prelievi di ungulati, tetraonidi e coturnice
R Core Team (2022) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/
Raymond M, Rousset F (1995) GENEPOP (version 1.4): population genetics software for exact tests and ecumenicism. J Hered 86:248–249
Richardson JL, Brady SP, Wang IJ, Spear SF (2016) Navigating the pitfalls and promise of landscape genetics. Mol Ecol 25:849–863
Rousset F (2000) Genetic differentiation between individuals. J Evol Biol 13:58–62
Safner T, Buzan E, Rezic A, Šprem N (2019) Small-scale spatial genetic structure of Alpine chamois (Rupicapra rupicapra) in Northern Dinarides. Eur J Wildl Res 65:23
Segelbacher G, Cushman SA, Epperson BK et al (2010) Applications of landscape genetics in conservation biology: concepts and challenges. Conserv Genet 11:375–385. https://doi.org/10.1007/s10592-009-0044-5
Shafer ABA, Northrup JM, White KS, Boyce MS, Cote SD, Coltman DW (2012) Habitat selection predicts genetic relatedness in an alpine ungulate. Ecology 93(6):1317–1329
Soglia D, Rossi L, Cauvin E, Citterio C, Ferroglio E, Maione S, Meneguz PG, Spalenza V, Rasero R, Sacchi P (2010) Population genetic structure of Alpine chamois (Rupicapra r. rupicapra) in the Italian Alps. Eur J Wildl Res 56:845–854
Storfer A, Murphy MA, Evans JS et al (2007) Putting the “landscape” in landscape genetics. Heredity 98:128–142
Storfer A, Murphy MA, Spear SF, Holderegger R, Waits LP (2010) Landscape genetics: where are we now? Mol Ecol 19:3496–3514
Van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P (2004) Micro-checker: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4(3):535–538
Vasiljevic N, Morf NV, Senn J, Pérez-Espona S, Mattucci F, Mucci N, Moore-Jones G, Pisano SRR, Kratzer A, Ogden R (2022) Phylogeography and population genetic structure of the European roe deer in Switzerland following recent recolonization. Ecol Evol 12(2):e8626
Whiteley AR, McGarigal K, Schwartz MK (2014) Pronounced differences in genetic structure despite overall ecological similarity for two Ambystoma salamanders in the same landscape. Conserv Genet 15:573–591
Winiarski KJ, Peterman WE, Whiteley AR, McGarigal K (2020) Multiscale resistant kernel surfaces derived from inferred gene flow: an application with vernal pool breeding salamanders. Mol Ecol Res 20(1):97–113
Zannèse A, Morellet N, Targhetta C, Coulon A, Fuser S, Hewison AJM, Ramanzin M (2006) Spatial structure of roe deer populations: towards defining management units at a landscape scale. J Appl Ecol 43:1087–1097
Acknowledgements
We are grateful to the Province of Trento and the Trento provincial hunting association (Associazione Cacciatori Trentini) for support in sampling activities. We thank M. Rocca, A. Brugnoli, and all the game wardens for their invaluable support during data collection.
Funding
Open access funding provided by Università degli Studi di Sassari within the CRUI-CARE Agreement. The research was supported by the program “Fondo di Ateneo per la Ricerca 2020” and received support from The European Union – NextGenerationEU as part of the National Biodiversity Future Center, Italian National Recovery and Resilience Plan (NRRP) Mission 4 Component 2 Investment 1.4.
Author information
Authors and Affiliations
Contributions
Planning: MA, MS, RC, RL; sample collection: RC; lab analyses: SA, AC; data elaboration: RL, MS, OD, ET; support to data elaboration: VO, LB; writing and editing: RL, OD, MS, RC; review: MA.
Corresponding author
Ethics declarations
Ethical approval
No animals were killed specifically for the study. No living animals were sampled.
Competing interests
MS is associate editor of the journal.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lecis, R., Chirichella, R., Dondina, O. et al. Same landscape, different connectivity: contrasting patterns of gene flow in two sympatric ungulates in a mountain area. Eur J Wildl Res 70, 42 (2024). https://doi.org/10.1007/s10344-024-01796-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10344-024-01796-1