Abstract
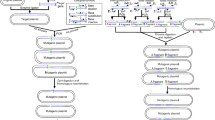
The gene integration method is an important tool to stably express desirable genes in bacteria. To avoid heavy workload and cost, we constructed a rapid and efficient method for genome modification. This method depended on a mobilizable plasmid, which contains a P tac promoter, an introduced multiple cloning site (iMCS), and rrnBT1T2 terminator. Briefly, the mobilizable plasmid pK18-MBPMT with the P tac-iMCS-rrnBT1T2 cartridge derived from pK18mobsacB was prepared to directly integrate hetero-/homologous DNA into the Corynebacterium glutamicum genome. Like our previous method, this method was based on insertional inactivation and double-crossover homologous recombination, which simultaneously achieved gene overexpression and inactivation in the genome without the use of genetic markers. Compared to the previous method, this protocol omitted the construction of a recombinant expression plasmid and clone of the target gene(s) cassette, which significantly decreased the workload, cost, and operational time. Using this method, the heterologous gene amy and the homologous gene lysC T311I were successfully integrated into the C. glutamicum genome at alaT and avtA loci, respectively. Moreover, the operation time of this method was shorter than that of the previous method, especially for repeated integration. This method, which is based on the mobilizable plasmid pK18-MBPMT, thus represents a potentially attractive protocol for the integration of genes in the course of genetic modification of C. glutamicum.





Similar content being viewed by others
Abbreviations
- AKFr :
-
Allosterically feedback-resistant aspartokinase
- AlaT:
-
Aspartate aminotransferase
- AvtA:
-
PLP-dependent aminotransferase
- iMCS:
-
Introduced multiple cloning sites
- Km:
-
Kanamycin
- DCW:
-
Dry cell weight
- μ max :
-
Maximum specific growth rate
- LB:
-
Luria-Bertani
- LBG:
-
LB + 5 g L−1 glucose
- LBS:
-
LB + 100 g L−1 sucrose
- LBK:
-
LB + 50 µg mL−1 Km
- LBHIS:
-
LB + brain heart infusion + sorbitol
References
Wieschalka S, Blombach B, Bott M, Eikmanns BJ (2013) Bio-based production of organic acids with Corynebacterium glutamicum. Microb Biotechnol 6:87–102. doi:10.1111/1751-7915.12013
Hou XH, Chen XD, Zhang Y, Qian H, Zhang WG (2012) L-Valine production with minimization of by-products’ synthesis in Corynebacterium glutamicum and Brevibacterium flavum. Amino Acids 43:2301–2311. doi:10.1007/s00726-012-1308-9
Pátek M, Nešvera J, Guyonvarch A, Reyes O, Leblon C (2003) Promoters of Corynebacterium glutamicum. J Biotechnol 104:311–323. doi:10.1016/j.jbiotec.2005.12.027
Becker J, Zelder O, Häfner S, Schröder H, Wittmann C (2011) From zero to hero-design-based systems metabolic engineering of Corynebacterium glutamicum for l-lysine production. Metab Eng 13:159–168. doi:10.1016/j.ymben.2011.01.003
van Ooyen J, Noack S, Bott M, Reth A, Eggeling L (2012) Improved l-lysine production with Corynebacterium glutamicum and systemic insight into citrate synthase flux and activity. Biotechnol Bioeng 109:2070–2081. doi:10.1002/bit.24486
Xu JZ, Han M, Zhang JL, Guo YF, Zhang WG (2014) Metabolic engineering Corynebacterium glutamicum for the l-lysine production by increasing the flux into l-lysine biosynthetic pathway. Amino Acids 46:2165–2175. doi:10.1007/s00726-014-1768-1
Xu JZ, Xia XH, Zhang JL, Guo YF, Zhang WG (2014) A method for gene amplification and simultaneous deletion in Corynebacterium glutamicum genome without any genetic markers. Plasmid 72:9–17. doi:10.1016/j.plasmid.2014.02.001
Ikeda M, Katsumata R (1998) A novel system with positive selection for the chromosomal integration of replicative plasmid DNA in Corynebacterium glutamicum. Microbiol 144:1863–1868
Hu JY, Li YY, Zhang HL, Tan YZ, Wang XY (2014) Construction of a novel expression system for use in Corynebacterium glutamicum. Plasmid 75:18–26. doi:10.1016/j.plasmid.2014.07.005
Correia A, Martin JF, Castro JM (1996) Targeted integration of foreign genes into repetitive sequences of the Brevibacterium lactofermentum genome. FEMS Microbiol Lett 142:259–264. doi:10.1111/j.1574-6968.1996.tb08440.x
Amador E, Franciso J, José M, Castro M (2007) A Brevibacterium lactofermentum 16S rRNA gene used as target site for homologous recombination. FEMS Microbiol Lett 185:199–204. doi:10.1111/j.1574-6968.2000.tb09062.x
Suzuki N, Inui M, Yukawa H (2007) Site-directed integration system using a combination of mutant lox sites for Corynebacterium glutamicum. Appl Microbiol Biotechno 77:871–878. doi:10.1007/s00253-007-1215-2
Xu DQ, Tan YZ, Huan XJ, Hu XQ, Wang XY (2010) Construction of a novel shuttle vector for use in Brevibacterium flavum, an industrial amino acid producer. J Microbiol Methods 80:86–92. doi:10.1016/j.mimet.2009.11.003
Schäfer A, Tauch A, Jäger W, Kalinowski J, Thierbach G, Pühler A (1994) Small mobilizable multi-purpose cloning vectors derived from the Escherichia coli plasmids pK18 and pK19: selection of defined deletions in the chromosome of Corynebacterium glutumicum. Gene 145:69–73. doi:10.1016/0378-1119(94)90324-7
van der Rest ME, Lange C, Molenaar D (1999) A heat shock following electroporation induces highly efficient transformation of Corynebacterium glutamicum with xenogeneic plasmid DNA. Appl Microbiol Biotechnol 52:541–545. doi:10.1007/s002530051557
Keilhauer C, Eggeling L, Sahm H (1993) Isoleucine synthesis in Corynebacterium glutamicum: molecular analysis of the ilvB-ilvN-ilvC operon. J Bacteriol 175:5595–5603
Srivastava P, Deb JK (2005) Gene expression systems in corynebacteria. Protein Expression Purif 40:221–229. doi:10.1016/j.pep.2004.06.017
Ohnishi J, Mitsuhashi S, Hayashi M, Ando S, Yokoi H, Ochiai K, Ikeda M (2002) A novel methodology employing Corynebacterium glutamicum genome information to generate a new l-lysine-producing mutant. Appl Microbiol Biotechnol 58:217–223. doi:10.1007/s00253-001-0883-6
Xu JZ, Zhang JL, Guo YF, Zhang WG (2015) Genetically modifying aspartate aminotransferase and aspartate ammonia-lyase affects metabolite accumulation in l-lysine producing strain derived from Corynebacterium glutamicum ATCC13032. J Mol Catal B Enzym 113:82–89. doi:10.1016/j.molcatb.2014.12.015
Marienhagen J, Eggeling L (2008) Metabolic function of Corynebacterium glutamicum aminotransferases AlaT and AvtA and impact on l-valine production. Appl Environ Microbiol 74:7457–7462. doi:10.1128/AEM.01025-08
Ugorčáková J, Jucovič M, Bukovská G, Timko J (1996) Construction and characterization of new corynebacterial plasmids carrying the α-amylase gene. Folia Microbiol 41:10–14. doi:10.1007/BF02816332
Tauch A, Götker S, Pühler A, Kalinowski J, Thierbach G (2002) The alanine racemase gene alr is an alternative to antibiotic resistance genes in cloning systems for industrial Corynebacterium glutamicum strains. J Biotechnol 99:79–91. doi:10.1016/S0168-1656(02)00159-1
Schäfer A, Tauch A, Droste N, Pühler A, Kalinowski J (1997) The Corynebacterium glutamicum cglIM gene encoding a 5-cytosine methyltransferase enzyme confers a specific DNA methylation pattern in an McrBC-deficient Escherichia coli strain. Gene 203:95–101. doi:10.1016/S0378-1119(97)00519-2
Vertès AA, Hatakeyama K, Inui M, Kobayshi M, Kurusu Y, Yukawa H (1993) Replacement recombination in Coryneform bacteria: high efficiency integration requirement for non-methylated plasmid DNA. Biosci Biotechnol Biochem 57:2036–2038. doi:10.1271/bbb.57.2036
Vogt M, Haas S, Klaffl S, Polen T, Eggeling L, van Ooyen J, Bott M (2014) Pushing product formation to its limit: metabolic engineering of Corynebacterium glutamicum for l-leucine overproduction. Metab Eng 22:40–52. doi:10.1016/j.ymben.2013.12.001
Kinoshita S, Udaka S, Shimono M (1957) Studies on the amino acid fermentation Part I. production of l-glutamic acid by various microorganisms. J Gen Microbiol 3:193–205
Acknowledgments
This work was financially supported by the Youth Program of Natural Science Foundation of Jiangsu Province (No. BK20150149) and the Youth Foundation of Jiangnan University (No. JUSRP115A19).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Xu, J., Zhang, J., Han, M. et al. A method for simultaneous gene overexpression and inactivation in the Corynebacterium glutamicum genome. J Ind Microbiol Biotechnol 43, 1417–1427 (2016). https://doi.org/10.1007/s10295-016-1806-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-016-1806-y