Abstract
Despite the prominence of iron–sulfur cluster (ISC) proteins in bioenergetics, intermediary metabolism, and redox regulation of cellular, mitochondrial, and nuclear processes, these proteins have been given scarce attention in Drosophila. Moreover, biosynthesis and delivery of ISCs to target proteins requires a highly regulated molecular network that spans different cellular compartments. The only Drosophila ISC biosynthetic protein studied to date is frataxin, in attempts to model Friedreich’s ataxia, a disease arising from reduced expression of the human frataxin homologue. One of several proteins involved in ISC biogenesis is heat shock protein cognate 20 (Hsc20). Here we characterize two piggyBac insertion mutants in Drosophila Hsc20 that display larval growth arrest and deficiencies in aconitase and succinate dehydrogenase activities, but not in isocitrate dehydrogenase activity; phenotypes also observed with ubiquitous frataxin RNA interference. Furthermore, a disruption of iron homeostasis in the mutant flies was evidenced by an apparent reduction in induction of intestinal ferritin with ferric iron accumulating in a subcellular pattern reminiscent of mitochondria. These phenotypes were specific to intestinal cell types that regulate ferritin expression, but were notably absent in the iron cells where ferritin is constitutively expressed and apparently translated independently of iron regulatory protein 1A. Hsc20 mutant flies represent an independent tool to disrupt ISC biogenesis in vivo without using the RNA interference machinery.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Iron–sulfur clusters (ISCs) may have participated in reactions leading to the origin of life on Earth [1, 2]. They are central to most key processes sustaining living ecosystems, including carbon and nitrogen fixation [3]. In animals, ISC proteins are involved in oxidative phosphorylation and the regulation of iron homeostasis [4, 5]. ISC proteins are also conspicuously abundant in the nucleus, but the specific functions for the ISCs in DNA replication and repair are currently under investigation [6–9]. Cells actively build and deliver ISCs to target proteins [10–13]; defects in this process lead to disease, notably Friedreich’s ataxia [14–16], which is caused by the expansion of a GAA trinucleotide repeat element in an intron of the human frataxin gene [17]. The study of related human disorders has revealed tissue-specific requirements of specific ISC biosynthesis genes [18–20].
One factor considered important for delivery of ISCs to target proteins, in particular under conditions of oxidative stress, is heat shock protein cognate 20 (Hsc20) [21, 22]. The yeast homologue of Hsc20 is known as Jac1 and was recovered from a genetic screen as a suppressor of superoxide dismutase deficiency [23]. Reduced activity of Jac1 resulted in a decrease in activity of iron/sulfur-containing mitochondrial proteins and an accumulation of iron in mitochondria [24–26]. Jac1 interacts with the iron–sulfur scaffold protein Isu1p [27] and this interaction appears to be conserved in evolution [21, 28]. Hsc20 proteins from higher animals contain a metal-binding, cysteine-rich N-terminal domain, which is important for the integrity and function of the human co-chaperone [21, 29].
ISC biosynthesis has been studied in Drosophila, following the cloning of dfh, the fly homologue of the human frataxin gene [30]. A number of laboratories have used RNA interference (RNAi) induced by the heterologous yeast Gal4/upstream activating sequence (UAS) transgenic induction system [31] to model Friedreich’s ataxia in Drosophila [32–36]. Strong ubiquitous reduction in dfh expression resulted in giant long-lived larvae that failed to initiate metamorphosis and had defects in ISC enzymes [32], whereas more moderate reductions in expression resulted in adult flies with locomotion defects and increased sensitivity to oxidative stress [33]. Scavenging of hydrogen peroxide but not of superoxide rescued many dfh phenotypes [34], including the reversal of inactivation of aconitase, whose ISC is a bona fide target for superoxide [37]. This result is consistent with a signaling role for hydrogen peroxide as recently uncovered in the fly hematopoietic and wound-healing processes [38–40]. Reduced dfh expression in the central nervous system led to defective mitochondrial axonal transport and membrane potential when RNAi was induced in neurons [35], and accumulation of lipids and lipid peroxidation when RNAi was induced in glia [36]. These studies and also observations that ubiquitous overexpression of dfh affected specifically the development of embryonic muscles [41] have highlighted the variable tissue-specific roles also for Drosophila dfh. As no animal models are available beyond dfh RNAi flies and frataxin knockout mice [42], we undertook to characterize two transgenic insertions disrupting the Drosophila Hsc20 homologue.
Materials and methods
Drosophila stocks
All Drosophila melanogaster strains were maintained on standard cornmeal/yeast/agar medium at 25 °C. Iron supplementation was in the form of ferric ammonium citrate. PBac{PB}l(3)72Do c05018 /TM6 and PBac{WH}l(3)72Do f02457 /TM6, Tb were from the Exelixis collection at Harvard Medical School. Both insertions were rebalanced to the TM3, P{GAL4-Kr.C}DC2, P{UAS-GFP.S65T}DC10, Sb 1 fluorescent balancer (GFP is green fluorescent protein) from Bloomington stock no. 5195 to identify the time of larval lethality and select homozygous mutant larvae for further analysis. The resulting stocks are abbreviated Hsc20 c5018 /TM3, Kr-GFP, Sb and Hsc20 f2457 /TM3, Kr-GFP, Sb, respectively, in the text. Actin-Gal4 used for rescue experiments and Fer1HCH G188 (Fer1HCH is ferritin-1 heavy chain homologue) used for visualization of intestinal ferritin have been described elsewhere [43].
RNA isolation and reverse transcription PCR
Total RNA was isolated from larvae using a modified Trizol method [44] and was purified using an RNeasy Mini kit (QIAGEN). The RNase-free DNase set (QIAGEN) was applied for on-column digestion of residual DNA. Total RNA of each sample was first reverse-transcribed into complementary DNA using the SuperScript III system (Invitrogen). The sequence encoding the full-length Hsc20 protein was amplified by PCR using the total match primers 5′-ATGAGTAGAGTTATTAATGGCTTTAAAAGTGCT-3′ and 5′-TCAGCTGCCCAGCAAACTTTGTTGCT-3′ (Fig. 1a). The PCR product was purified by agarose gel electrophoresis, excised, and eluted with a MinElute gel extraction kit (QIAGEN). Cloning of the PCR product was performed using a Zero Blunt TOPO PCR cloning kit (Invitrogen). For a C-terminal fusion of Hsc20 with red fluorescent protein, the sequence was amplified from the cloning vector with the primers hsc20NotF 5′-TGAATTCATGAGTAGAGTTATTAATG GCTTTAAAGT-3′ and hsc20AgeR 5′-TTACCGGTAATCCGCTGCCCAGCAAACT-3′. After subcloning in pCR2.1-TOPO, the transcript was excised with EcoRI and AgeI and inserted in the corresponding sites of the mammalian expression vector pmCherry-N1 (Clontech). All constructs used were verified by sequencing.
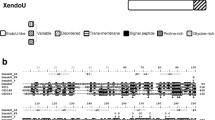
The Drosophila Hsc20 gene is broadly expressed at low levels and encodes a mitochondrial protein. a The information presented was retrieved and adapted from the FlyBase and modENCODE databases, where Hsc20 is listed as l(3)72Do or CG34246. The gene is composed of four exons and three introns. The open reading frame is shown in beige and the noncoding region of the messenger RNA in gray. The two piggyBac insertions used in this study disrupt the first exon. The gene shows ubiquitous low level of expression throughout development and into adulthood. b Confirmation of the presence and small size of introns 2 and 3 by reverse transcription PCR (RT-PCR; the location of primers used shown is in a) and localization of an Hsc20–red fluorescent protein (RFP) fusion protein in mitochondria of transfected HeLa cells
Cell culture and confocal microscopy
HeLa cells were obtained from the American Type Culture Collection and were grown in the Dulbecco’s modified Eagle’s medium with 4.5 g/l glucose, supplemented with 10 % (v/v) fetal bovine serum, 100 U/ml penicillin and 100 mg/ml streptomycin at 37 °C and 5 % CO2. Cells were seeded in microwell Lab-Tek chambered cover glass (Nunc International), and Fugene 6 (Roche) was used for transient transfections with mCherry-tagged Hsc20 according to the recommendations of the manufacturers. Immunofluorescence was imaged with a confocal microscope system (LSM 510 META; Zeiss) typically 24–36 h after transfection. For imaging red fluorescence, the 543-nm line of an He–Ne laser was used with a 488 nm/543 nm dichroic mirror, and the fluorescence was collected with a 560-nm long-pass filter.
Enzyme assays
Three third instar larvae were transferred to a 1.5-ml microcentrifuge tube and were washed with 1× phosphate-buffered saline (PBS). Subsequently they were homogenized in about 5 vol of ice-cold lysis buffer [50 mM tris(hydroxymethyl)aminomethane (Tris)/HCl at pH 8.0, containing 1 % (v/v) NP-40 (Fluka), and EDTA-free protease inhibitor (one tablet per 10 ml) (Roche)] with a disposable plastic pestle and briefly sonicated on ice. The homogenates were cleared by centrifugation (16,000g, 4 °C, 15 min). Total aconitase activity was determined by following the reduction of NADP+ at 340 nm in the subsequent isocitrate dehydrogenase reaction. Five microliters of the homogenate was added to 50 μl buffer R1 [50 mM Tris/HCl, pH 8.0, with 50 mM NaCl, 5 mM MgCl2, 0.5 mM NADP+, 0.01 U isocitrate dehydrogenase (NADP+-dependent) from porcine heart (Sigma)]. The reaction was started by the addition of 45 μL buffer R2 (buffer R1 without NADP+ and isocitrate dehydrogenase, with 2.5 mM cis-aconitate). Activity was expressed as ΔE(340 nm)/Δt and was normalized for protein concentration. Isocitrate dehydrogenase activity was determined by following the reduction of NADP+ at 340 nm as described above. Five microliters of lysate was added to 45 μl buffer R3 (buffer R1 without isocitrate dehydrogenase). The reaction was initiated by the addition of 50 μl buffer R4 (buffer R3 without NADP+, with 5 mM isocitrate).
Succinate dehydrogenase activity in situ stain
Succinate dehydrogenase activity was evaluated on the basis of the succinate-dependent iodonitrotetrazolium chloride reduction. Prior to dissection the larvae were kept for 10 min in cold 1× PBS with 25 % (v/v) glycerol. Subsequently the cuticle and fat body of the larva (totally submerged in cold buffer) were carefully removed. The dissected tissue was briefly washed in cold isotonic buffer and was transferred on a coverslip. One hundred microliters of 50 mM Tris/HCl pH 7.5 with 1 mM rotenone, 1 mg/ml antimycin, 10 mM KCN, 25 mM azide, 0.1 % digitonin, and proteinase inhibitor was added to the sample. Tissues were completely submerged in this solution for 10 min. Following preincubation, 100 μl of 50 mM Tris/HCl pH 7.5 with 4 mM iodonitrotetrazolium chloride, 0.5 mM EDTA, 15 g/l CremophorEL (Sigma), and 50 mM succinate (for negative control without succinate) was added. Incubation was at room temperature for 10 min. The staining solution was removed with a pipette and filter paper and samples were imaged unfixed in 1× PBS with 25 % (v/v) glycerol. The reduction of iodonitrotetrazolium chloride leads to red, water-insoluble formazan.
Enhanced Prussian blue stain
The dissection was performed as described in the previous section. The samples were rinsed three times with 1× PBS and subsequently fixed and stained simultaneously in 4 % paraformaldehyde (v/v) and Perls’s solution [1 % K4Fe(CN)6 and 1 % HCl] for 30 min at room temperature. Enhancement with diaminobenzidine was performed as described in [45].
Results
Drosophila Hsc20 encodes a mitochondrial protein
An alignment between Drosophila and human Hsc20 protein sequences has revealed extended homology and 28 % identity between the homologue proteins of the two species [21]. The Drosophila Hsc20 gene is currently annotated in FlyBase as l(3)72Do, following a detailed genetic analysis of the polytene chromosome region 72A-D that revealed CG34246 as an essential gene [46]. Here we refer to CG34246 or l(3)72Do as Hsc20.
Recent genomic approaches under the Model Organism Encyclopedia of DNA Elements (modENCODE) project [47], supported by the National Human Genome Research Institute, have provided a developmental transcriptome in Drosophila melanogaster [48]. This work has revealed that Hsc20 contains four exons and is expressed at low levels throughout development (Fig. 1a) and in all tissues of larvae and adults (see http://flybase.org), consistent with a housekeeping role for a putative mitochondrial protein. The second intron is predicted to be 52 bp long and the third intron 60 bp long. Primers used to amplify the open reading frame of Hsc20 by reverse transcription PCR confirmed the difference of 112 bp between the PCR on the genomic template and the reverse transcription PCR product and also that Hsc20 is expressed in flies (Fig. 1b). A construct linking red fluorescent protein to the C-terminus of Hsc20 was transfected in HeLa cells to determine the subcellular localization of the protein, which was found to accumulate in mitochondria (Fig. 1c), a result that was expected on the basis of the characterization of the human protein and the conserved mitochondrial targeting sequence in the N-terminus of Drosophila Hsc20 [21].
Drosophila Hsc20 mutants show growth arrest and reduced ISC biogenesis
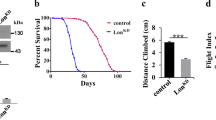
The two transgenic insertions used in this study were selected because their insertion points disrupt the first exon of Hsc20 at a position in the 5′ untranslated region (Fig. 1a). Consistent with what was shown previously for other transposons in this locus [46], both the Hsc20 c5018 allele and the Hsc20 f2457 allele are homozygous lethal and do not complement each other. The developmental time point of lethality was investigated by balancing the respective insertions to TM3, Kr-GFP, Sb, making heterozygous and homozygous genotypes recognizable through the presence or absence of fluorescence, respectively, throughout the organism’s development (Fig. 2a). The time point of lethality was identical for both alleles (shown only for the Hsc20 f2457/f2457 genotype); homozygous mutant larvae grew through the first and second instars, but halted growth after they reached the third instar (Fig. 2a).
Drosophila Hsc20 piggyBac insertion mutants arrest growth as third instar larvae. a Hsc20 f2457 /TM3, Kr-GFP, Sb (f2457/+) flies (GFP is green fluorescent protein) laid eggs on agar plates supplied with yeast; embryos and larvae were visualized with a fluorescent microscope and homozygous mutants (f2457/f2457) were identifiable from the absence of fluorescent signal. Homozygous mutant larvae failed to grow during the third instar. b Actin-Gal4-driven genetic rescue of homozygous f2457/f2457 mutants by virtue of the upstream activating sequence (UAS) elements present in the PBac{WH}l(3)72Do f02457 transgene, which permit expression of the endogenous Hsc20 gene following activation by Gal4. N depicts the total number of flies scored. The rescue is not always complete, as many flies complete metamorphosis but fail to eclose from their pupal cases. UAS elements are absent from the PBac{PB}l(3)72Do c05018 transgene, which serves as a control for this experiment. FRT flippase recombinase target
As the piggyBac element inserted in the Hsc20 f2457 allele included UAS, which can be used to induce neighboring gene expression (from exons 2-4 that encode the Hsc20 open reading frame), we generated Actin-Gal4/Cyo; Hsc20 f2457 /TM3, Kr-GFP, Sb flies and looked for rescue (Fig. 2b). Given that balancer chromosomes are homozygous lethal, a full rescue would result in 33 % Hsc20 f2457/f2457 homozygous adult flies and 67 % Hsc20 f2457/+ heterozygous adult flies. In the experiment shown we counted 249 progeny flies, of which 183 (73 %) were heterozygous and 66 (27 %) were homozygous, of which 53 (22 %) died while trying to eclose from their pupal cases (Fig. 2b, left panel) and only 13 (5 %) survived into adulthood. The Actin-Gal4/Cyo; Hsc20 c5018 /TM3, Kr-GFP, Sb genotype was also generated as a control, but no homozygous viable adults were obtained in this stock at any point. Hence, our rescue experiments show that the lethality associated with Hsc20 f2457/f2457 flies is due to a specific reduction in Hsc20 expression.
We collected third instar homozygous Hsc20 mutant larvae and their heterozygous sibling controls and quantified total aconitase and isocitrate dehydrogenase activities in lysates prepared from these. Both Hsc20 c5018/c5018 and Hsc20 f2457/f2457 larvae showed a marked decrease of aconitase activity to approximately 50 % of the activity measured in the respective heterozygous flies (Fig. 3a, left panel). To verify that the loss of activity was due to a specific loss of ISCs in aconitase enzymes, we also tested isocitrate dehydrogenase, another housekeeping enzyme that does not depend on ISCs, in the same lysates and found no difference between the genotypes (Fig. 3a, right panel). We also tested for succinate dehydrogenase activity, this time employing an activity assay in dissected tissues of the larvae. We observed high activity in the anterior midgut of the intestine in Hsc20 f2457/+ larvae, which was markedly reduced in Hsc20 f2457/f2457 larvae (Fig. 3b). These results showed that ISC proteins are specifically impaired in Hsc20 mutant larvae.
Drosophila Hsc20 mutants have reduced aconitase and succinate dehydrogenase activities but normal isocitrate dehydrogenase activity, suggesting specific defects in iron–sulfur cluster (ISC) biosynthesis. a Lysates were prepared from heterozygous Hsc20 c5018 /TM3, Kr-GFP, Sb (c/+) and Hsc20 f2457 /TM3, Kr-GFP, Sb (f/+) adult files and homozygous Hsc20 c5018/Hsc20 c5018 (c/c) and Hsc20 f2457 /Hsc20 f2457 (f/f) adult flies. Lysates were assayed for ISC-dependent aconitase activity and for ISC-independent isocitrate dehydrogenase activity and normalized against total protein. Note the differences in Hsc20 homozygous mutants for aconitase (asterisks denote p < 0.01 from the t test). b Succinate dehydrogenase activity normally present in the anterior midgut is severely reduced in intestines from Hsc20 f2457 /Hsc20 f2457 larvae. Specificity of this stain was shown by removal of the substrate for the reaction in the left panel, in which case minimal signal is obtained
Drosophila Hsc20 mutants accumulate iron in mitochondria instead of in ferritin
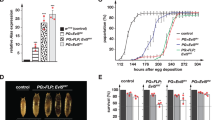
To test the effect of defective ISC biogenesis on iron homeostasis, we recombined a GFP–ferritin trap line [43, 49] with the piggyBac insertions in Hsc20. We then dissected intestines from Hsc20 f2457, Fer1HCH G188 /Hsc20 f2457 third instar larvae and from Hsc20 f2457, Fer1HCH G188 /TM3, Sb controls and observed ferritin under a fluorescence microscope. Ferritin accumulation in the iron region was unaffected between the two genotypes (Fig. 4a, middle panels); expression of ferritin in these cells was previously shown to be independent of iron [49]. In contrast, cells in the anterior and posterior midgut showed low, but readily detectable levels of ferritin accumulation in the heterozygous Hsc20 mutant larvae; these cells were devoid of ferritin in the homozygous Hsc20 mutants (Fig. 4a, top and bottom panels). Dietary iron induces ferritin in the anterior midgut of larvae [43, 49], so we tested the response of Hsc20 mutants when grown on yeast supplemented with 1 mM ferric ammonium citrate. As expected, Hsc20 f2457, Fer1HCH G188 /TM3, Sb larvae showed a clear induction of ferritin in the anterior midgut, whereas induction of ferritin in Hsc20 f2457, Fer1HCH G188 /Hsc20 f2457 intestines was compromised (Fig. 4b, top panels).
Drosophila Hsc20 mutants show punctate iron accumulation and fail to induce ferritin in the intestine. a A GFP–ferritin protein trap line [43] was crossed into Hsc20 mutants to monitor ferritin accumulation. Intestines were dissected from Hsc20 f2457, Fer1HCH G188 /TM3, Sb (f2457, G188/+) heterozygous control flies (Fer1HCH is ferritin-1 heavy chain homologue) and Hsc20 f2457, Fer1HCH G188 /Hsc20 f2457 mutants (f2457, G188/f2457). Note that induction of ferritin appears to be reduced in the anterior and posterior midgut, but is unaffected in the iron region of the middle midgut. b Feeding larvae with iron normally leads to a pronounced induction of ferritin in the anterior midgut, not seen in Hsc20 mutants, which still accumulate iron in the anterior midgut, but in a pattern more reminiscent of mitochondria (arrows)
Ferritin induction in the anterior midgut serves as a store for iron absorbed by the diet [43]. We stained the intestines dissected from Hsc20 f2457/f2457 larvae (and from the respective controls) for iron using the enhanced Prussian blue stain (Fig. 4b, middle panels). Despite the absence of ferritin induction in Hsc20 f2457/f2457 mutants, we still observed marked accumulation of iron in the anterior midgut. Examination of the stained cells at higher magnification revealed a punctate pattern, which was not seen in Hsc20 f2457/+ heterozygous control stains (Fig. 4b, bottom panels).
Discussion
Hsc20 encodes a mitochondrial protein involved in ISC biogenesis
We have characterized two piggyBac insertions in the Hsc20 gene that result in reduced activities of classic ISC enzymatic activities and have shown that Hsc20 is a nuclear-encoded mitochondrial protein. Our results are largely similar to those obtained by other investigators who studied ubiquitous RNAi phenotypes of the dfh gene [32, 33] and therefore validate our conclusion that the Hsc20 mutants described here provide the first example of Drosophila mutants in ISC biogenesis.
Iron accumulation in mitochondria is characteristic of disrupted ISC biosynthesis
We observed a punctate staining for iron in Hsc20 mutant cells localized in the anterior intestine of larvae (Fig. 4b, lower panels). Accumulation of iron in mitochondria is a characteristic of Friedreich’s ataxia patients and is also observed in animal models of the disease [50] and yeast lacking Jac1 [24, 26]. Such pathologic iron is present in aggregate form, but is clearly distinct from ferritin iron [51]. Given the evidence accumulated in studies with yeast, mice models, and human patients [42], we believe that the punctate staining we have observed specifically in Hsc20 mutants could represent similar mitochondrial iron inclusions, suggesting these may also develop in the new Drosophila model of disrupted ISC biogenesis we have described. The phenotypes of reduced ISC activities, mitochondrial iron overload, and cytosolic iron deficiency are consistent with those reported for the human and yeast Hsc20 homologues [21–29]. For these reasons we suggest that Hsc20 should be considered as a candidate disease gene in humans.
Cell-type-specific effects in ferritin regulation in Hsc20 mutants
Intestinal ferritin regulation is complex and regulates both iron absorption and iron storage [43, 52–54]. The iron cells in the middle midgut express ferritin in a constitutive manner and independent of systemic iron concentrations [49]. Cells in the very posterior of the intestine appear to be specialized in zinc homeostasis, where ferritin may also have a specialized function that is not yet understood [55]. In this study, we have noted no changes in ferritin accumulation in the middle midgut in Hsc20 mutants (Fig. 4a, middle panels), consistent with the notion that iron homeostasis in these cells is specialized. The apparent downregulation of ferritin in cells of the anterior midgut (Fig. 4a, b, upper panels) may be explained by the activation of iron regulatory protein 1A as a result of defective ISC biogenesis and hence translational repression of ferritin [56, 57], or by mitochondrial iron accumulation causing a relative cytoplasmic iron deficiency. Whichever explanation holds (or both may be true), we note that not all cell types respond in the same way on genetic inactivation of Hsc20, calling for further investigations into the in vivo roles of the ISC biosynthetic machinery. The mutants described in this study should help expand in vivo studies.
An iron-related mitochondrial to nucleus signal may be relevant to the circadian clock
Drosophila melanogaster provides an elegant system for the parallel investigation of molecular, biochemical, cellular, physiological, and behavioral biology using genetic and environmental manipulations. The Hsc20 mutants we describe offer an alternative model to validate results obtained via RNAi experiments [32–36, 58]. Most other genes known from other systems to be involved in ISC biosynthesis are conserved in Drosophila, where they were implicated in the maintenance of circadian rhythms [58], representing the first proposed function of ISCs in the regulation of animal behavior. At least one ISC-containing enzyme, dihydropyrimidine dehydrogenase, has been shown to have circadian rhythmic expression in the head of flies [59], but it is unknown which ISC protein may interact directly with the biological clock’s transcriptional feedback loops [60, 61]. The function of ISC proteins in regulating redox homeostatic mechanisms may help explain how the recently discovered free-running metabolic and redox cycles link to the cellular transcriptional and translational timekeeping networks [62, 63]. In plants, a retrograde signal from chloroplasts to nucleus signaling an iron deficiency has been suggested to affect the period length of the clock [64–66]. Such a signal could be the unknown chemical exported by the ABCB7 mitochondrial transporter [17, 67]. Owing to the lethality of Hsc20 mutants during their larval stage of development (Fig. 2a), we were unable to test whether loss of Hsc20 would disrupt the circadian clock. However, this lethality underscores the essential role of ISCs in animal growth.
Abbreviations
- Fer1HCH:
-
Ferritin-1 heavy chain homologue
- GFP:
-
Green fluorescent protein
- Hsc20:
-
Heat shock protein cognate 20
- ISC:
-
Iron–sulfur cluster
- modENCODE:
-
Model Organism Encyclopedia of DNA Elements
- PBS:
-
Phosphate-buffered saline
- RNAi:
-
RNA interference
- Tris:
-
Tris(hydroxymethyl)aminomethane
- UAS:
-
Upstream activating sequence
References
Wickramasinghe RH (1973) Space Life Sci 4:341–352
Schoepp-Cothenet B, van Lis R, Philippot P, Magalon A, Russell MJ, Nitschke W (2012) Sci Rep 2:263
Hall DO, Evans MC (1969) Nature 223:1342–1348
Crack JC, Green J, Thomson AJ, Le Brun NE (2012) Curr Opin Chem Biol 16:35–44
Tong WH, Rouault TA (2007) Biometals 20:549–564
Stehling O, Vashisht AA, Mascarenhas J, Jonsson ZO, Sharma T, Netz DJ, Pierik AJ, Wohlschlegel JA, Lill R (2012) Science 337:195–199
Gari K, Leon Ortiz AM, Borel V, Flynn H, Skehel JM, Boulton SJ (2012) Science 337:243–245
Benjdia A (2012) Curr Opin Struct Biol 22:711–720
White MF, Dillingham MS (2012) Curr Opin Struct Biol 22:94–100
Rouault TA (2012) Dis Model Mech 5:155–164
Rouault TA, Tong WH (2005) Nat Rev Mol Cell Biol 6:345–351
Shepard EM, Boyd ES, Broderick JB, Peters JW (2011) Curr Opin Chem Biol 15:319–327
Lill R, Hoffmann B, Molik S, Pierik AJ, Rietzschel N, Stehling O, Uzarska MA, Webert H, Wilbrecht C, Muhlenhoff U (2012) Biochim Biophys Acta 1823:1491–1508
Rouault TA, Tong WH (2008) Trends Genet 24:398–407
Pandolfo M, Pastore A (2009) J Neurol 256:9–17
Stemmler TL, Lesuisse E, Pain D, Dancis A (2010) J Biol Chem 285:26737–26743
Campuzano V, Montermini L, Moltò MD, Pianese L, Cossée M, Cavalcanti F, Monros E, Rodius F, Duclos F, Monticelli A, Zara F, Cañizares J, Koutnikova H, Bidichandani SI, Gellera C, Brice A, Trouillas P, De Michele G, Filla A, De Frutos R, Palau F, Patel PI, Di Donato S, Mandel JL, Cocozza S, Koenig M, Pandolfo M (1996) Science 271:1423–1427
Sanaker PS, Toompuu M, Hogan VE, He L, Tzoulis C, Chrzanowska-Lightowlers ZM, Taylor RW, Bindoff LA (2010) Biochim Biophys Acta 1802:539–544
Crooks DR, Jeong SY, Tong WH, Ghosh MC, Olivierre H, Haller RG, Rouault TA (2012) J Biol Chem 287:40119–40130
Xia H, Cao Y, Dai X, Marelja Z, Zhou D, Mo R, Al-Mahdawi S, Pook MA, Leimkuhler S, Rouault TA, Li K (2012) PLoS ONE 7:e47847
Uhrigshardt H, Singh A, Kovtunovych G, Ghosh M, Rouault TA (2010) Hum Mol Genet 19:3816–3834
Shan Y, Cortopassi G (2012) Hum Mol Genet 21:1457–1469
Strain J, Lorenz CR, Bode J, Garland S, Smolen GA, Ta DT, Vickery LE, Culotta VC (1998) J Biol Chem 273:31138–31144
Voisine C, Cheng YC, Ohlson M, Schilke B, Hoff K, Beinert H, Marszalek J, Craig EA (2001) Proc Natl Acad Sci USA 98:1483–1488
Lutz T, Westermann B, Neupert W, Herrmann JM (2001) J Mol Biol 307:815–825
Kim R, Saxena S, Gordon DM, Pain D, Dancis A (2001) J Biol Chem 276:17524–17532
Andrew AJ, Dutkiewicz R, Knieszner H, Craig EA, Marszalek J (2006) J Biol Chem 281:14580–14587
Ciesielski SJ, Schilke BA, Osipiuk J, Bigelow L, Mulligan R, Majewska J, Joachimiak A, Marszalek J, Craig EA, Dutkiewicz R (2012) J Mol Biol 417:1–12
Bitto E, Bingman CA, Bittova L, Kondrashov DA, Bannen RM, Fox BG, Markley JL, Phillips GN Jr (2008) J Biol Chem 283:30184–30192
Canizares J, Blanca JM, Navarro JA, Monros E, Palau F, Molto MD (2000) Gene 256:35–42
Martinek S, Young MW (2000) Genetics 156:1717–1725
Anderson PR, Kirby K, Hilliker AJ, Phillips JP (2005) Hum Mol Genet 14:3397–3405
Llorens JV, Navarro JA, Martinez-Sebastian MJ, Baylies MK, Schneuwly S, Botella JA, Molto MD (2007) FASEB J 21:333–344
Anderson PR, Kirby K, Orr WC, Hilliker AJ, Phillips JP (2008) Proc Natl Acad Sci USA 105:611–616
Shidara Y, Hollenbeck PJ (2010) J Neurosci 30:11369–11378
Navarro JA, Ohmann E, Sanchez D, Botella JA, Liebisch G, Molto MD, Ganfornina MD, Schmitz G, Schneuwly S (2010) Hum Mol Genet 19:2828–2840
Missirlis F, Hu J, Kirby K, Hilliker AJ, Rouault TA, Phillips JP (2003) J Biol Chem 278:47365–47369
Owusu-Ansah E, Banerjee U (2009) Nature 461:537–541
Moreira S, Stramer B, Evans I, Wood W, Martin P (2010) Curr Biol 20:464–470
Juarez MT, Patterson RA, Sandoval-Guillen E, McGinnis W (2011) PLoS Genet 7:e1002424
Navarro JA, Llorens JV, Soriano S, Botella JA, Schneuwly S, Martinez-Sebastian MJ, Molto MD (2011) PLoS ONE 6:e21017
Martelli A, Napierala M, Puccio H (2012) Dis Model Mech 5:165–176
Missirlis F, Kosmidis S, Brody T, Mavrakis M, Holmberg S, Odenwald WF, Skoulakis EM, Rouault TA (2007) Genetics 177:89–100
Bogart K, Andrews J (2006) CGB technical report 2006–2010. Indiana University, Bloomington
Tong WH, Rouault TA (2006) Cell Metab 3:199–210
Cooper MT, Kennison JA (2011) PLoS ONE 6:e23509
modENCODE Consortium (2010) Science 330:1787–1797
Graveley BR, Brooks AN, Carlson JW, Duff MO, Landolin JM, Yang L, Artieri CG, van Baren MJ, Boley N, Booth BW, Brown JB, Cherbas L, Davis CA, Dobin A, Li R, Lin W, Malone JH, Mattiuzzo NR, Miller D, Sturgill D, Tuch BB, Zaleski C, Zhang D, Blanchette M, Dudoit S, Eads B, Green RE, Hammonds A, Jiang L, Kapranov P, Langton L, Perrimon N, Sandler JE, Wan KH, Willingham A, Zhang Y, Zou Y, Andrews J, Bickel PJ, Brenner SE, Brent MR, Cherbas P, Gingeras TR, Hoskins RA, Kaufman TC, Oliver B, Celniker SE (2011) Nature 471:473–479
Mehta A, Deshpande A, Bettedi L, Missirlis F (2009) Biochimie 91:1331–1334
Puccio H, Simon D, Cossee M, Criqui-Filipe P, Tiziano F, Melki J, Hindelang C, Matyas R, Rustin P, Koenig M (2001) Nat Genet 27:181–186
Whitnall M, Rahmanto YS, Huang ML, Saletta F, Lok HC, Gutierrez L, Lazaro FJ, Fleming AJ, St Pierre TG, Mikhael MR, Ponka P, Richardson DR (2012) Proc Natl Acad Sci USA. doi:10.1073/pnas.1215349109
Georgieva T, Dunkov BC, Harizanova N, Ralchev K, Law JH (1999) Proc Natl Acad Sci USA 96:2716–2721
Tang X, Zhou B (2012) FASEB J. doi:10.1096/fj.12-213595
Kosmidis S, Botella JA, Mandilaras K, Schneuwly S, Skoulakis EM, Rouault TA, Missirlis F (2011) Neurobiol Dis 43:213–219
Gutierrez L, Sabaratnam N, Aktar R, Bettedi L, Mandilaras K, Missirlis F (2010) FEBS Lett 584:2942–2946
Lind MI, Missirlis F, Melefors O, Uhrigshardt H, Kirby K, Phillips JP, Soderhall K, Rouault TA (2006) J Biol Chem 281:18707–18714
Surdej P, Richman L, Kuhn LC (2008) Insect Biochem Mol Biol 38:891–894
Mandilaras K, Missirlis F (2012) Metallomics 4:928–936
Van Gelder RN, Bae H, Palazzolo MJ, Krasnow MA (1995) Curr Biol 5:1424–1436
Veleri S, Brandes C, Helfrich-Forster C, Hall JC, Stanewsky R (2003) Curr Biol 13:1758–1767
Nitabach MN, Taghert PH (2008) Curr Biol 18:R84–R93
Masri S, Zocchi L, Katada S, Mora E, Sassone-Corsi P (2012) Ann N Y Acad Sci 1264:103–109
Edgar RS, Green EW, Zhao Y, van Ooijen G, Olmedo M, Qin X, Xu Y, Pan M, Valekunja UK, Feeney KA, Maywood ES, Hastings MH, Baliga NS, Merrow M, Millar AJ, Johnson CH, Kyriacou CP, O’Neill JS, Reddy AB (2012) Nature 485:459–464
Salomé PA, Oliva M, Weigel D, Krämer U (2013) EMBO J 32:511–523
Hong S, Kim SA, Guerinot ML, McClung CR (2013) Plant Physiol 161:893–903
Chen YY, Wang Y, Shin LJ, Wu JF, Shanmugam V, Tsednee M, Lo JC, Chen CC, Wu SH, Yeh KC (2013) Plant Physiol [Epub ahead of print]
Metzendorf C, Wu W, Lind MI (2009) Biochem J 421:463–471
Acknowledgments
The authors acknowledge the invaluable support of Mary Lilly, who provided her laboratory fly facility that allowed this project to be conducted. This work was supported by the Intramural Program of the Eunice Kennedy Shriver National Institute of Child Health and Human Development in the USA and CONACYT project 179835 in Mexico.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Uhrigshardt, H., Rouault, T.A. & Missirlis, F. Insertion mutants in Drosophila melanogaster Hsc20 halt larval growth and lead to reduced iron–sulfur cluster enzyme activities and impaired iron homeostasis. J Biol Inorg Chem 18, 441–449 (2013). https://doi.org/10.1007/s00775-013-0988-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-013-0988-2