Abstract
Cecropins constitute an important family of insect antimicrobial peptides involved in humoral innate immune response. In comparison with the highly basic cecropins A and B, cecropins D are less cationic and more hydrophobic. Interestingly, cecropins D were described only in lepidopteran insects, e.g., the greater wax moth Galleria mellonella. In the present study, interactions of neutral cecropin D (pI 6.47) purified from hemolymph of G. mellonella with living Escherichia coli cells were investigated. Fluorescence lifetime imaging microscopy using fluorescein isothiocyanate-labeled cecropin D revealed very fast binding of the peptide to E. coli cells. Fourier transform infrared spectroscopy analyses showed that G. mellonella cecropin D interacted especially with E. coli LPS and probably other lipid components of the bacterial cell envelope and exhibited an ordering effect with regard to lipid chains. This effect is consistent with the peptide binding mechanism based upon its incorporation into the lipid phase of the cell membrane. The interaction resulted in permeabilization of the bacterial cell membrane. Upon cecropin D binding, the cells lost characteristic surface topography, which was accompanied by altered nanomechanical properties, as revealed by atomic force microscopy. The interaction of the peptide with the bacterial cells also led to intracellular damage, i.e., loss of the cell envelope multilayer structure, formation of membrane vesicles, and enlargement of periplasmic space, which eventually caused death of the bacteria. In summary, it can be concluded that amphipathic character of α-helices, exposure of small positively charged patches on their polar surfaces and hydrophobic interactions are important physicochemical characteristics related to effective binding to E. coli cells and antibacterial activity of neutral G. mellonella cecropin D.
Similar content being viewed by others
Introduction
Antimicrobial peptides (AMPs) are small, amphipathic, and mainly cationic molecules that upon interaction with microbial membranes lead to disruption of the membrane structure and function, which eventually results in microbial cell death. Several models have been proposed to explain the interactions of AMPs with microbial membranes, including the barrel-stave, carpet-like, aggregate, and toroidal pore model. Nevertheless, irrespective of the model, it is generally accepted that the initial step of the binding is mediated by electrostatic interactions between the cationic peptide and negatively charged membrane components followed by hydrophobic interactions between amphipathic domains in the peptide molecule and lipid components of the membrane (Nguyen et al. 2011; Scocchi et al. 2011; Cytryńska and Zdybicka-Barabas 2015).
Cecropins constitute a family of invertebrate AMPs, found especially in insects. They are amphipathic and mostly highly basic linear peptides composed of 31–39 amino acids, which can form two flexible α-helices bound with a hinge region in a hydrophobic environment. The distribution of hydrophilic and hydrophobic groups is conserved among all cecropins. They possess a hydrophilic N-terminal part containing many charged residues, a very hydrophobic stretch between residues 22–30, and a more hydrophilic C-terminal end. Usually, the N-terminal part of a cecropin molecule is strongly basic, whereas the C-terminal part is hydrophobic. The peptides were first characterized in Hyalophora cecropia (H. cecropia) (Lepidoptera) and classified into several subfamilies: A, B, C, D, E (Hultmark et al. 1982). Cecropins A, B, and C were then found in species belonging to other insect orders, i.e., Diptera and Coleoptera (Otvos 2000; Bulet and Stöcklin 2005; Yi et al. 2014). Interestingly, cecropins D were described only in lepidopteran insects, suggesting that this subfamily appeared relatively late in the evolution of insects (Gudmundsson et al. 1991).
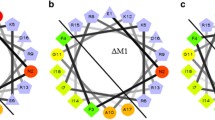
A great arsenal of diverse AMPs was reported in the greater wax moth Galleria mellonella (G. mellonella), a model insect species widely used in studies on innate immunity mechanisms, host–pathogen interactions, and pathogenicity and virulence factors of different microorganisms (Cytryńska et al. 2007; Brown et al. 2008, 2009; Vogel et al. 2011). Among G. mellonella AMPs, two cecropins have been described to date, i.e., a highly basic cecropin A (pI 10.38) (Kim et al. 2004) and a neutral cecropin D (pI 6.47) (Mak et al. 2001; Cytryńska et al. 2007). Cecropin D, purified from G. mellonella immune hemolymph, was active against selected Gram-negative and Gram-positive bacteria as well as a filamentous fungus Aspergillus niger (Cytryńska et al. 2007). More recently, anti-Salmonella enterica and anti-Proteus mirabilis activity of synthetic G. mellonella cecropin D, named Gm1, has been demonstrated (Correa et al. 2014a). The interactions of Gm1 and its positively charged analog ∆Gm1 with model lipid membranes have also been analyzed (Correa et al. 2014b). In addition, Oñate-Garzón et al. (2017a; b) investigated the antibacterial activity and binding to model lipid membranes of two cationic peptides, ∆M1 and ∆M2, with a net charge +5 and +9, respectively, derived from G. mellonella cecropin D.
To provide more data on the interactions of the native G. mellonella cecropin D with living bacterial cells, the effects of the peptide action on Escherichia coli (E. coli) cells were investigated in the present study. The alterations in bacterial cell surface components that occur upon the peptide binding were investigated in detail by Fourier transform infrared (FTIR) spectroscopy. Furthermore, fluorescently labeled G. mellonella cecropin D was used for analysis of the peptide binding to bacterial cells by fluorescence lifetime imaging microscopy (FLIM). The cell surface topography and properties were analyzed by atomic force microscopy (AFM), whereas the intracellular damage was imaged by transmission electron microscopy (TEM).
Materials and methods
Bacterial strains and culture conditions
Gram-negative bacteria Escherichia coli D31 and E. coli JM83 and Gram-positive bacterium Micrococcus luteus (M. luteus) ATCC 10,240 were grown in 2.5% Lysogeny Broth (LB; Sigma-Aldrich) at 37 and 28 °C, respectively. In the experiments, bacteria in the logarithmic phase of growth were used.
Insect immune challenge, hemolymph collection, and preparation of hemolymph methanolic extracts
The larvae of G. mellonella (Lepidoptera: Pyralidae) were reared on a natural diet—honeybee nest debris at 30 °C in the dark. For the immune challenge, the last instar larvae (250–300 mg in weight) were pierced with a needle dipped into a pellet containing live E. coli D31 and M. luteus ATCC 10,240, and the hemolymph was collected 24 h after the immunization. Acidic/methanolic extracts rich in antimicrobial peptides and proteins below 30 kDa were prepared from the hemocyte-free hemolymph as described previously (Cytryńska et al. 2007; Mak et al. 2010).
Purification and fluorescent labeling of G. mellonella cecropin D
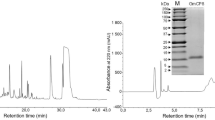
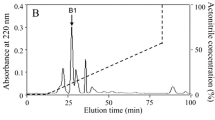
The cecropin D peptide was purified from the immune hemolymph extracts using a modified procedure described in our previous papers (Cytryńska et al. 2007; Palusińska-Szysz et al. 2012; Zdybicka-Barabas et al. 2012) and a P680 high-performance liquid chromatography (HPLC) system (Dionex). In brief, the lyophilized and lipid-free hemolymph extract was reconstituted in 0.1% (v/v) trifluoroacetic acid (TFA), filtered through a 0.45 μm filter, and subjected to HPLC using a Discovery Bio Wide Pore C18 250 mm × 4.6 mm column (Sigma-Aldrich) and two buffer sets: A—0.1% TFA (v/v), B—0.07% TFA, 80% acetonitrile (v/v). A linear 30–70% gradient of buffer B over 35 min, spectrophotometric detection at 220/280 nm, and a 1 ml/min flow rate were applied. The fraction of cecropin was collected and then purified to homogeneity using the same column as above and a linear 49–52% B gradient over 20 min. The purity of the collected final peptide was confirmed independently by analytical HPLC separation on the C18 column as above and by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS–PAGE) (Schägger and von Jagow 1987), while the identity was confirmed by determination of the N-terminal sequence on a Procise 491 automatic protein sequencer (Applied Biosystems). The concentration of the peptide was determined by amino acid analysis (Bocheńska et al. 2013).
Labeling of cecropin D with fluorescein isothiocyanate (FITC, isomer 1; Sigma-Aldrich) was performed according to a procedure described in our previous paper (Zdybicka-Barabas et al. 2014). Briefly, 4 nmol of the peptide was dissolved in 50 µl of sodium borate, pH 9.0, and mixed with 100-times molar excess of FITC dissolved in dimethylsulfoxide (DMSO). The solution was incubated for 20 h at room temperature in darkness and then the unreacted FITC was quenched by addition of 10 μl of 1 M glycine water solution. After additional 0.5-h incubation in darkness, the solution was subjected to gel filtration on a Superdex Peptide 10/300 GL column (GE Healthcare). The separation was carried out at a 0.7 ml/min flow rate in 10 mM ammonium acetate buffer, pH 5.8, containing 30% (v/v) acetonitrile. The peak containing the monosubstituted FITC-cecropin D peptide was collected and freeze-dried. Examination of the labeled peptide by SDS–PAGE revealed a single yellow band at approx. 5 kDa.
Bacterial membrane permeabilization assay
The E. coli JM83 strain bearing a pCH110 plasmid was used in the assay (Pharmacia-Amersham). The plasmid encodes constitutively synthesized cytoplasmic β-galactosidase and ampicillin resistance (Mak et al. 2007). The assay was performed using a suspension of mid-logarithmic phase cells prepared in 20 mM phosphate buffer, pH 6.8, essentially as described previously (Zdybicka-Barabas et al. 2012). Briefly, 2 μl of the suspension (5 × 105 colony forming units, CFU) was added to 23 μl of pre-incubated (37 °C, 15 min) cecropin D in the phosphate buffer (final concentrations of peptide: 0.625–20.0 μM). The samples were then incubated at 37 °C for 5–60 min. Subsequently, 20 mM HEPES/150 mM NaCl buffer, pH 7.5, and 50 mM p-nitrophenyl-β-d-galactopyranoside were added, the samples were incubated for 1.5 h at 37 °C, and absorbance was measured at 405 nm. Simultaneously, two types of the control samples were analyzed: (1) live bacteria incubated without the peptide and (2) bacteria completely killed by treatment with 5 µM cecropin B (Sigma-Aldrich). At the given time point, the perforation level of the live bacteria was subtracted from the results of the tested samples. Then, the perforation percent was calculated assuming the dead bacteria perforation level as 100%. Three independent experiments with three repetitions for each type of sample were performed.
Atomic force microscopy (AFM)
Preparation of the bacterial samples
100 μl of the suspension containing log-phase E. coli JM83 cells (OD600 = 0.2) cultured in the LB medium was incubated without (control) and in the presence of cecropin D (final concentrations 2.5 and 1 μM) for 0.5, 1, and 1.5 h at 37 °C. The bacterial suspensions were centrifuged at 8000×g for 10 min at 4 °C. The bacterial pellets were gently washed once with 20 mM phosphate buffer, pH 6.8, and next twice with non-pyrogenic water (100 μl). After final centrifugation, the bacteria were suspended in 5 μl of non-pyrogenic water and applied on the surface of freshly cleaved mica disks. Before imaging, the samples were allowed to dry at 28 °C (Zdybicka-Barabas et al. 2011, 2012).
AFM imaging and analysis
All AFM measurements in the tapping or Peak Force QNM operation modes were carried out using a NanoScope V AFM (Veeco) equipped with NanoScope 8.10 software and a piezoscanner with a maximum scan range of 150 × 150 μm (Analytical Laboratory, Faculty of Chemistry, UMCS, Lublin, Poland). A rectangular Si cantilever/tip (Veeco) with a spring constant of 20–80 N/m and resonance frequency of 300 kHz was used. The resolution of the scans obtained was 256 × 256 pixels. The height and peak force error images were obtained simultaneously. Four fields on each mica disk were imaged. The data were analyzed with WSxM 5.0 software (Nanotec). The roughness values were measured over the entire bacterial cell surface on 400 × 400 nm areas. The average surface root-mean-square (RMS) roughness of the cells was calculated from forty fields examined during two independent experiments.
For estimation of the G. mellonella cecropin D influence on the elasticity and adhesion properties of bacterial cells, Derjaguin–Muller–Toporov (DMT) modulus and adhesion forces were determined, respectively. The force measurements were performed in the Peak Force QNM operation mode using a silicon tip at the nitride lever, SCANASYST-AIR, with a spring constant of 0.4 N/m (Veeco). The data were analyzed with Nanoscope Analysis ver. 1.40 software (Veeco).
Transmission electron microscopy (TEM)
The bacteria (18 Eppendorf tubes with 100 µl suspension in LB medium; OD600 = 0.4) were incubated for 1 h at 37 °C without (control) or with cecropin D (final concentration 2.5 µM). Immediately after the incubation, the cells were directly fixed in the culture medium to avoid osmotic problems in the cells. The cells were flooded with the first fixative—2.5% (v/v) glutaraldehyde dissolved in 0.1 M phosphate-buffered saline (PBS, pH 7.4). After 30 min, the cells were gently centrifuged at 3000×g for 30 min at 4 °C. After rinsing several times with 0.1 M PBS, the cell pellets were post-fixed in a 1% (v/v) osmium tetroxide solution in 0.1 M phosphate buffer (pH 7.4) for 2 h at 4 °C. The bacterial cells were then dehydrated in a series of alcohol and acetone, and embedded in LR White resin. Ultrathin sections (65 nm) were cut with a diamond knife on a microtome RMC MT-XL (Tuscon, USA). The samples were collected on copper grids and contrasted with the use of uranyl acetate and Reynold’s liquid. The samples were observed under a LEO-Zeiss 912 AB electron microscope (Oberkohen, Germany).
Confocal laser scanning microscopy imaging (CLSM)
For CLSM imaging, the procedure was carried out essentially as described previously (Zdybicka-Barabas et al. 2014). Briefly, 60 μl of the suspension containing log-phase E. coli JM83 cells (OD600 = 0.2) was incubated with 10 µM FITC-labeled cecropin D at 37 °C for 15 min. Then, the bacterial suspension was centrifuged (2000×g, 10 min, 4 °C) and washed twice with 20 mM phosphate buffer, pH 7.4, containing 0.9% NaCl (100 μl). Next, the bacterial cells were suspended in 10 μl of 20 mM phosphate buffer, pH 7.4, and imaged using a laser scanning confocal microscope LSM 5 PASCAL (Carl Zeiss Microscopy) (excitation and emission wavelengths were 470 and 520 nm, respectively; excitation time was 600 ms).
Fluorescence lifetime imaging microscopy (FLIM)
For FLIM analysis, the log-phase E. coli JM83 cells (100 μl of suspension; OD600 = 0.2) in 20 mM phosphate buffer, pH 7.4, were incubated without (control) or with 2.5 μM FITC-labeled cecropin D at 37 °C for up to 60 min. The bacterial suspensions were centrifuged at 4000×g for 10 min at 4 °C. The cells were gently washed once with 20 mM phosphate buffer, pH 7.4 (500 μl), twice with non-pyrogenic water (300 μl), and finally they were suspended in 100 μl of non-pyrogenic water. Then, 10 μl of the resulting suspension was applied on a polylysine-coated cover slip and after 10 s carefully rinsed three times with water.
Fluorescence lifetime images were collected with a MicroTime 200 confocal system (Picoquant) coupled to an inverted microscope Olympus IX71 equipped with a 60 × water immersed objective (Olympus PlanApo NA = 1.2). For excitation purpose, a solid-state laser operating at 470 nm wavelength (set to 1.8 µW power at the sample) was used and dichroic (ZT 473 rdcxt) and band-pass 520/35 fluorescence filters (both from Analysentechnik) were applied for observation. The confocal mode of the measurements was accomplished with a pinhole 75 µm in diameter. Signal detection and processing was based on a Single Photon Avalanche Diode (τ-SPAD, Picoquant) and HydraHarp 400, respectively. The emission intensity decays were analyzed in an exponential model using SymPhoTime software v. 2.3 (Picoquant). Due to so-called color effect (Luchowski et al. 2009a, b) characteristic for detectors used in following experiments, the intensity decays I(t) for each sample were tail-fitted (taking into account response time of photodetector) with the multi-exponential model:
where τi are the characteristic decay times and αi are the pre-exponential factors. In particular, the exponential components are listed in the Results section. Average lifetime was calculated based on the intensity-weighted formula:
Fourier transform infrared (FTIR) spectroscopy
For FTIR analysis of cecropin D interactions with bacterial cells, 3 μl from 10 µl of the E. coli JM83 suspension (OD600 = 0.2) prepared in PBS, pH 7.4, supplemented with 5% D2O (v/v), immediately after cecropin D addition (final concentration 2.5 μM) was placed onto the attenuated total reflection (ATR) crystal element. For analyses of cecropin D-LPS interactions, cecropin D (10 µM) was incubated with E. coli K-235 lipopolysaccharide (LPS) (final concentration 1.5 mg/ml; Sigma-Aldrich, cat. no L-2143) in non-pyrogenic water containing 5% D2O (v/v) for 1 h at 37 °C (final volume 4 µl). The samples were deposited on the ATR crystal element by 5-min evaporation and the absorption spectra were recorded. The absorption spectra of the control cells, the cells exposed to cecropin D, cecropin D alone, as well as the LPS and PBS used were collected. The time period of 5 min was selected and standardized for all the experiments on the basis of preliminary optimization experiments where IR absorption spectra were recorded repeatedly till the moment at which no spectral changes were observed in the O–H stretching region. Such a procedure allowed to remove only the bulk water before recording of the spectra, whereas the water molecules inside cells and the water fraction tightly bound to the cells were preserved to maintain conditions as close as possible to the physiological ones.
Infrared absorption spectra were recorded using a Fourier transform infrared absorption spectrometer equipped with an attenuated total reflection set-up (ATR–FTIR). The internal reflection element (diamond prism) was used as an attenuated total reflection element. Continuous purging with argon during the measurements was applied. The spectra were recorded with a Nicolet iS50R spectrometer (Thermo Scientific). Typically, 10 scans were collected, Fourier transformed, and averaged for each measurement. Absorption spectra at a resolution of one data point every 2 cm−1 were obtained in the region between 4000 and 400 cm−1 using a clean crystal as the background. Taking into consideration that spectroscopic measurements in the phase transition region of the lipid may affect interpretation, all the spectroscopic experiments were performed at 21 °C, relatively far from the phase transition temperature of LPS (~ 35 °C) (Brandenburg and Seydel 1998; Hassan and Ilev 2014; Heinbockel et al. 2015). Spectral analysis was performed with Grams Al software (ThermoGalactic).
Other methods
The protein concentration in the hemolymph and methanolic extracts was estimated with the Bradford method using bovine serum albumin (BSA) as a standard (Bradford 1976).
To compare two means, statistical analysis was performed using Student’s t test. The data were presented as the means ± standard deviation (± S.D.). The differences were considered statistically significant at *p < 0.05, **p ≤ 0.01, ***p ≤ 0.001.
Results
Permeabilization of E. coli cell membrane
The exposure of E. coli JM83 cells to G. mellonella cecropin D at the concentration range of 2.5–20 µM led to permeabilization of the bacterial cell membrane with similar kinetics of the peptide action during the 60-min incubation time, regardless of the concentration (Fig. 1). After 60 min, cecropin D at the concentration of 2.5 and 10 µM permeabilized the E. coli cell membrane in approx. 20% and approx. 50%, respectively. The lower concentrations of the peptide used, i.e., 0.625 and 1.25 µM, were ineffective in the cell membrane perturbation during the 60-min incubation.
Level of permeabilization of E. coli cells treated with G. mellonella cecropin D. The bacteria in the logarithmic phase of growth were incubated without (negative control) or in the presence of synthetic cecropin B (5 µM; positive control) or G. mellonella cecropin D (final concentration range 0.625–20 µM) for up to 60 min. Then the absorbance proportional to the amount of released β-galactosidase was measured at 405 nm. First, the perforation level of the negative control was subtracted from all measurements and then the perforation level of the dead bacteria was considered as 100%. The values in the graph are presented as the means from three independent experiments ± S.D. At cecropin D concentrations of 2.5–20 µM, the differences between the peptide-treated and control cells at corresponding incubation times were statistically significant (p˂0.05). At cecropin D concentrations of 0.625 and 1.25 µM, the differences were not statistically significant
Effects of G. mellonella cecropin D on the cell surface and ultrastructure of E. coli
AFM imaging and analysis
Figure 2 presents AFM images of E. coli JM83 cells treated with 2.5 µM G. mellonella cecropin D for 30 and 60 min. The control cells were characterized by a rod-shape and well-visible flagella. The regular cortex bend-like structures were especially well recognized on the cell surface of the control cells incubated for 60 min. (Figure 2, bottom panel), similarly to topography of the E. coli cell surface demonstrated previously (Zdybicka-Barabas et al. 2012; Rahnamaeian et al. 2015, 2016). Although the surface topography of the control cells incubated for 30 min was less regular, the characteristic cortex bend-like structures were visible as well (Fig. 2, bottom panel). Dramatically damaged bacterial cells were found among those exposed to cecropin D for 30 min (Fig. 2, upper panel). In addition, the cell surface profiles revealed surface depressions 20–30 nm in depth and approx. 250 nm in diameter (Fig. 3a). Although no such extreme damage was detected in cells exposed to cecropin D for 60 min, these cells also lost the characteristic surface topography and were covered with irregular wrinkles (Fig. 2, bottom panel). Moreover, clearly delineated hollows (12–15 nm in depth and 180–200 nm in diameter) were detected (Fig. 3b). These results suggested that E. coli cells are the most susceptible to the 2.5 µM concentration of cecropin D during the first 30 min of exposure. On the other hand, when the cells were treated with 1 µM cecropin D for 90 min, the cell surface profiles revealed the presence of long-branched humps (20 nm in height and 200 nm in width) in addition to the hollows (20–25 nm in depth and 300 nm in diameter) (Fig. 3c). The treatment of E. coli cells with 2.5 µM cecropin D resulted in an approx. 3.5-fold increase in DMT modulus and an approx. 40% increase in the RMS roughness value in comparison with the control cells, regardless of the incubation time. On the other hand, adhesion forces of cells exposed to the peptide for 30 and 60 min decreased by 50% and increased by 30%, respectively, compared with the relevant controls. Interestingly, the adhesion forces of the control cells after the 60-min incubation decreased by 70% in comparison with the control cells incubated for 30 min, reflecting alterations in the cell surface properties occurring during growth. All the biophysical parameters illustrating surface roughness, cell elasticity, and adhesion forces are summarized in Table 1.
Effect of G. mellonella cecropin D on E. coli cell surface topography. The bacteria in the logarithmic phase of growth were incubated without (control) or with 2.5 µM G. mellonella cecropin D for 30 and 60 min. Then the cells were imaged by AFM. The three-dimensional (3D), height, and “peak force error” images are presented. The upper and bottom panels demonstrate imaged areas of 3 × 3 µm and 500 × 500 nm, respectively. The white arrows in the upper panel indicate dramatically damaged cells. The white ellipse and arrowheads in the bottom panel indicate, respectively, cortex bend-like structures on the surface of control cells and irregular wrinkles on the surface of cecropin D-treated cells
Cell surface section profiles of the E. coli cells treated with G. mellonella cecropin D. The bacteria in the logarithmic phase of growth were incubated without (control) or with 2.5 µM cecropin D for 30 min a and 60 min b or 1 µM cecropin D for 90 min c. Then the cells were imaged by AFM as described in the “Materials and Methods” section. The diagrams present section profiles corresponding to the lines marked in the upper images
Effect of cecropin D on the ultrastructure of E. coli cells
To examine the effect of cecropin D on the E. coli cell ultrastructure, the bacterial cells were observed under a transmission electron microscope. Microscopic observations of the control cells (non-treated with cecropin D) revealed the presence of a cell envelope that appeared as a wavy multilayered structure, characteristic of Gram-negative bacteria (Fig. 4). Starting from the outside, a well discernible outer membrane (OM) was found. The next layer was the intermediate layer (IL) followed by the periplasmic space (PS) that was adjacent to the cytoplasmic membrane (CM). The periplasmic space appeared as a non-electron (unstained)-dense band. The cytoplasm was also observed with slightly granular structure in which a nucleoid appearing as a fibrous material (FM) was discernible. Cytoplasmic integrity was maintained in all cells (Fig. 4a–f). Microscopic investigations of the cells treated with the examined antimicrobial peptide revealed many alterations in the structure of E. coli cells, especially the cell envelope damage and changes in the cytoplasm appearance (Fig. 4g–l). In some cells, the cytoplasm was clear with no points of disruption in the cell wall; while in other cells, it was denser with apparent loss of the multilayer structure of the cell envelope (Fig. 4g, i, j–k). Cells with zones of disruption in the cell wall and an enlarged periplasmic space were detected as well (Fig. 4j). The distortion of the cell envelope was also found in the form of vesicles of the outer membrane, which were noticed on the cell surface (Fig. 4g–h). Some swollen cells containing clear cytoplasm were also visible (Fig. 4l).
Influence of G. mellonella cecropin D on E. coli cell ultrastructure. The bacteria were incubated without (control) or in the presence of cecropin D (2.5 µM) for 60 min and then imaged by TEM. Control cells (a–f) and cecropin D-treated cells (g–l). a—many control cells showing a well-preserved cell envelope; b—enlarged view of control cells showing distinct fibrillar material (FM) and a well-preserved cell envelope (arrow); c—control cells with one cell in the longitudinal section; the outer and inner membrane in the cell envelope are prominent; d, e—enlarged view of control cells with fibrillar material, granular cytoplasm, and a well-retained multilayered cell envelope (arrow); f—a fragment of a bacterium with well visible arrangement of the cell envelope containing the outer membrane (OM), intermediate layer (IL), periplasmic space (PS), and cytoplasmic membrane (CM); g—cells cultured in the presence of cecropin D showing many different changes, e.g., alterations in the cell envelope, dense cytoplasm; h—enlarged view of a bacterium with rolls (thin arrows) on the cell wall and visible distortion of the cell envelope (thick arrows); i—bacteria with apparent loss of one or more envelope layers (arrows) and with dense cytoplasm; j—bacteria with visible different alterations in the cell envelope (arrows); k—cells showing cell envelope deteriorations (thin arrows); l—swollen cells with clear cytoplasm (thick arrows)
Analysis of interactions of G. mellonella cecropin D with E. coli cells
FLIM analysis of cecropin D binding to bacterial cells
AFM and TEM imaging revealed that the treatment with G. mellonella cecropin D caused damage to the E. coli JM83 cells. These effects resulted from the interaction of cecropin D with bacterial cells, as demonstrated by CLSM imaging using FITC-labeled cecropin D (Fig. 5).
Laser scanning confocal microscope imaging of FITC-cecropin D binding to E. coli cells. The bacteria were incubated in the presence of FITC-labeled cecropin D for 15 min and then imaged by LSM as described in the “Materials and Methods” section. The bar represents 5 µm
Further analysis of the interaction with application of FLIM provided evidence on direct peptide binding to the bacterial cell surface. Two types of E. coli cells were analyzed: the control ones and those incubated with FITC-labeled cecropin D. Examples of images are presented in Fig. 6. All fluorescence photons collected during the cell imaging were analyzed in terms of multi-exponential decay. Two fluorescence lifetime components were resolved in the case of the auto-fluorescence of the control cells characterized by the following lifetime parameters: 1.9 ns and 6.7 ns with relative amplitudes 70 and 30%, respectively. A three-component approach appeared to be necessary to describe satisfactorily the fluorescence decay properties of FITC-labeled cecropin D: 1.9 ns (58%), 3.7 ns (12%), and 6.7 ns (30%), indicating heterogeneous binding of fluorophores to the peptide. The FLIM images of the bacteria incubated with cecropin D were analyzed by fitting all the components listed above and the process of cecropin binding to the bacteria was monitored by calculating the amplitude ratio corresponding selectively to the pure peptide (3.7 ns) relative to the amplitude of the component resolved in both the bacteria and the fluorescence-labeled peptide (6.7 ns). Immediately after mixing with cecropin D, the E. coli cells were imaged and the amplitude ratio was determined as 0.46 ± 0.16; the ratio was determined as 1.29 ± 0.31 after 10-min incubation and 5.90 ± 0.28 after 60-min incubation (averaged from three cells ± S.D.). The results indicate directly the process of the cecropin D binding to the surface of the bacteria. This process can be seen from the color-coded fluorescence lifetimes of the bacterial images shown in Fig. 6.
FLIM images of E. coli cells incubated with FITC-labeled cecropin D, recorded directly after subjecting to the interaction and after 10- and 60-min incubation (as indicated). The apparent blue coloration of the bacteria accompanying the incubation with cecropin D is associated with increased relative amplitude of the short-lifetime component of FITC (3.7 ns) and is a manifestation of cecropin D binding to E. coli
FTIR study of cecropin D interaction with LPS and live E. coli cells
The effects of the cecropin D interaction with E. coli cells and with LPS were analyzed at the molecular level by FTIR spectroscopy. During the procedure for FTIR measurements, the bulk water was removed while water bound to the cells, cecropin D and LPS was manifested by the presence of intensive bands centered at 3205, 3301 and 3353 cm−1, respectively, representing the OH stretching in water molecules tightly associated with these components (Figs. 7 and 8).
FTIR analysis of the interaction of G. mellonella cecropin D with E. coli lipopolysaccharide. Infrared absorption spectra of cecropin D, LPS, LPS incubated with cecropin D; the difference spectrum is shown in the lowest panel. To calculate the difference spectrum, the spectra recorded for pure cecropin D and pure LPS were multiplied by appropriate scaling factors selected to minimize the contribution of the absorption bands at wavenumbers specific for those components: at 1204 and 1070 cm−1, respectively
FTIR analysis of E. coli cells treated with G. mellonella cecropin D. Infrared absorption spectra of cecropin D, E. coli, and E. coli exposed to cecropin D; the difference spectrum is shown in the lowest panel. To calculate the difference spectrum, the spectra recorded for pure cecropin D and for E. coli were multiplied by appropriate scaling factors selected to minimize the contribution of the absorption bands at wavenumbers specific for those components: at 1204 and 973 cm−1, respectively
Figure 7 presents the infrared absorption spectra of cecropin D, LPS, LPS modified by interaction with the peptide, and a difference spectrum. To calculate the difference spectrum [(LPS + cecropin) minus cecropin], the pure cecropin D spectrum was multiplied by the factor selected to minimize the intensity of the spectral band centered at 1204 cm−1 present exclusively in the cecropin D spectrum. Further, the LPS spectrum was subtracted from the resulting spectrum [(LPS + cecropin) minus cecropin]. To do so, the LPS spectrum was taken with the factor calculated to minimize the band centered at 1070 cm−1, particularly intensive in the spectrum of pure LPS. Such a procedure yields a spectrum that reflects selectively and exclusively spectral changes basically free of the contributions from the individual components. Some changes are observed in the difference spectrum as a result of the cecropin D and LPS interaction. A relatively broad band centered at 3382 cm−1 is observed and can be an indication of the water fraction bound via hydrogen bonds to the LPS. The spectral shift towards lower frequencies of the bands centered at 2850 and 2922 cm−1 assigned to the symmetric and antisymmetric C–H stretching in the CH2 groups of alkyl chains of lipids is observed as well. This indicates an ordering effect of cecropin D with respect to lipid chains directly involved in the interaction with the peptide. In the spectral band centered at 1649 cm−1, corresponding to both the α-helix structure of cecropin D and the stretching vibration of the carbonyl group of LPS, a considerable increase in the oscillator strength is noted. Such an effect can be interpreted in terms of the interaction between LPS and cecropin D via water bridges. It has been noted that deformation vibrations of water molecules may also contribute to the spectrum in this region.
E. coli cells exposed to cecropin D were also examined with application of the FTIR technique (Fig. 8). The broad band at 3291 cm−1 represents O–H stretching in water molecules in a bulk phase present in living cells. The results of our measurements show that cecropin D caused a decrease in the water fraction bound to the cells. In the spectral band centered at 1061 cm−1, representing C–O–P–O–C and C–O–C stretching modes of lipids, the very strong effect observed can be interpreted in terms of breakage of hydrogen bonds (a strong negative band at 1061 cm−1 accompanied by a positive band at 1203 cm−1). Such combined changes in a difference spectrum represent a hypsochromic spectral shift but this effect has to be interpreted with caution due to the fact that spectral contribution of the phosphate group of the PBS buffer may interfere with the spectra of endogenous phospholipids, despite precise calculation of the difference spectra recorded from the samples containing the same buffer. The principal band between 2800 and 3000 cm−1 represents the C–H stretching modes with the maxima of peaks at 2852 and 2922 cm−1, corresponding to the symmetric and antisymmetric stretching in the CH2 groups of alkyl chains. The spectral shift towards lower frequencies is attributed to the ordering effect with regard to the lipid fraction of the E. coli cells. It is possible that the certain ordering effect with respect to lipid acyl chains observed is directly related to incorporation of the peptide or its part into the lipid phase of the membrane.
Discussion
Cecropins constitute a family of defense peptides characterized by two α-helices bound with a flexible hinge. In contrast to the highly basic cecropins A and B, cecropins D are less basic and more hydrophobic (Hultmark et al. 1982; Sidén and Boman 1983). So far, much attention has been paid to the antimicrobial action of cecropins A and B, most probably because of their highly cationic character, effective killing of pathogens, and much broader occurrence among insect species (Bulet and Stöcklin 2005; Yi et al. 2014).
In the present study, we analyzed the anti-E. coli activity of native G. mellonella cecropin D and interactions of this peptide with live E. coli cells as well as LPS. As revealed by the cell membrane permeabilization assay, cecropin D used at the 2.5 µM concentration effectively permeabilized E. coli JM83 cell membranes. It is worth mentioning that the concentration of the peptide used during the experiments corresponded to the cecropin D concentration determined in the hemolymph of naive G. mellonella larvae (Mak et al. 2010). Thus, our results indicate that, to some extent, the peptide can effectively protect the larvae against Gram-negative bacteria invading the hemocoel. In another study, the concentration below 2.5 µM of synthetic G. mellonella cecropin D completely inhibited the growth of the E. coli Re mutant (Correa et al. 2014a), whereas the growth of E. coli ATCC 25,922 was inhibited only at the 40 µM concentration of this peptide (Oñate-Garzón et al. 2017b). On the other hand, it was reported that 8.6 µM native G. mellonella cecropin D killed E. coli D31 cells (Cytryńska et al. 2007). The differences in the susceptibility of the E. coli strains to G. mellonella cecropin D may result from the different assays and growth conditions, e.g., different initial numbers of bacterial CFU (1 × 105, 5 × 105, 2 × 107 CFU/ml) used for determination of antimicrobial activity. However, there is strong literature evidence that the most important factor contributing to such differences is the properties of the bacterial cell envelope. In accordance with this, E. coli SB1004, D21e7, and D22 mutants with increased hydrophobicity of the surface layer were more susceptible to H. cecropia cecropin D than wild-type strains (Sidén and Boman 1983). These authors suggested that hydrophobicity alterations in the bacterial surface facilitated the better penetration of cecropin D through the outer membrane, which is limited in comparison with cecropins A and B due to the dissimilar properties of cecropin D.
In the present study, using FLIM imaging, we provided direct evidence on G. mellonella cecropin D interactions with live E. coli cells that occur almost immediately after addition of the peptide to the bacterial suspension. During the incubation time, the amount of cecropin D molecules bound to the bacterial cells increased considerably, indicating high affinity of this neutral peptide to the bacterial cell surface. Despite being neutral, effective binding of cecropin D to the cells was observed, which indicates an important role of other physicochemical parameters of the peptide, e.g., the amphipathic character or hydrophobic interactions. However, as it can be seen in Fig. 9, the two putative α-helices of G. mellonella cecropin D, besides being amphipathic, expose small positively charged patches on their polar surface. These regions may be involved in initial electrostatic interactions with negatively charged bacterial membranes. Interestingly, a preference of synthetic cecropin D-like peptide to anionic phospholipids, i.e., phosphatidylserine and phosphatidylglycerol, has been recently demonstrated using calorimetric analysis and fluorescence spectroscopy (Patiño-Márquez et al. 2018). Although an increase in the overall positive charge enhanced considerably the antibacterial activity of peptides ∆M1 and ∆M2 derived from G. mellonella cecropin D (Oñate-Garzón et al. 2017b), our study clearly indicates that the high positive charge is not necessary at all for effective binding of cecropin D to bacterial cells and for the peptide antibacterial function. It is worth mentioning here that another G. mellonella cecropin D-derived peptide named ∆Gm1 with 5 positive charges exhibited much lower activity against S. enterica, P. mirabilis, and E. coli Re than the synthetic parent peptide (Correa et al. 2014a). Interestingly, it was demonstrated that the Trp2 and Phe5 residues located at the N-terminus of the cecropin-like peptides were important for interaction with phospholipid model membranes (Lee et al. 2013). G. mellonella cecropin D contains two Phe residues at positions 3 and 4 (Cytryńska et al. 2007), suggesting that they may play a similar role in binding of this neutral peptide to the bacterial cell surface.
Prediction of α-helices formed by G. mellonella cecropin D a, helical wheel projections b, and distribution of electrostatic potential on the surface c of the putative α-helices. The α-helices were predicted using a protein secondary structure prediction server Jpred4 available at http://www.compbio.dundee.ac.uk/jpred/ (Drozdetskiy et al. 2015). In b the circles, diamonds, triangles, and pentagons represent, respectively, hydrophilic, hydrophobic, potentially negatively charged, and potentially positively charged residues. Hydrophobic residues are coded green (the most hydrophobic) to yellow (zero hydrophobicity). Hydrophilic uncharged and potentially charged residues are coded red and light blue, respectively (http://rzlab.ucr.edu/scripts/wheel/wheel.cgi). In c pictures represent rotations of 90° around z-axis (vertical) of putative N-terminal (left column) and C-terminal (right column) α-helix. Positively and negatively charged areas are indicated in blue and red, respectively. The most exposed positively charged regions are encircled by yellow ellipses. Non-polar regions are in gray. The electrostatic potential was analyzed using 3D-HM: The 3D Hydrophobic Moment Vector Calculator available as a web application on http://www.ibg.kit.edu/HM/ (Reißer et al. 2014)
There are studies indicating that the LPS layer plays a crucial role in the attraction and control of AMPs binding to the surface of Gram-negative bacteria (Ding et al. 2003; Papo and Shai 2005; Agrawal and Weisshaar 2018). Ebbensgaard et al. (2015) demonstrated protective effect of LPS against antimicrobial activity of many AMPs (including cecropin B and cecropin P1) manifested by a higher susceptibility of the E. coli LPS defective mutants in comparison with their parental strains: O6 serotype E. coli ATCC 25,922 and rough E. coli BW25113K-12. Moreover, comparison of the two parental E. coli strains revealed less susceptibility of smooth E. coli ATCC 25,922 to selected AMPs than E. coli K-12, further supporting important role of LPS structure in interactions with antimicrobial peptides (Ebbensgaard et al. 2015). Furthermore, Agrawal and Weisshaar (2018) using single-cell fluorescence microscopy showed that deletion of sugar layers or/and phosphate groups from the core oligosaccharide in E. coli K-12 LPS accelerated permeabilization of the outer membrane and cytoplasmic membrane by cecropin A. They postulated that the core oligosaccharide part of LPS functions as a kinetic barrier slowing cecropin A passage to the lipid A leaflet (Agrawal and Weisshaar 2018).
As revealed by the FTIR analyses, G. mellonella cecropin D interacted with E. coli LPS and exhibited an ordering effect with regard to lipid chains. No changes in the carbohydrate region of the spectrum were observed, suggesting that the sugar chain had no influence on the peptide binding. A similar effect was detected when the interactions of the peptide with live E. coli cells were analyzed, indicating that G. mellonella cecropin D interacted especially with the lipid part of LPS and probably with other lipid components of the bacterial cell envelope. It is known that the energy (spectral position) of the νsCH2 band reflects the conformational order and mobility of the lipid acyl chains. Owing to this dependency, sharp changes in a position of this particular band on the energy (wavenumber) scale, accompanying phase transitions, can be applied to monitor the gel to liquid phase transitions induced by temperature (Lewis and McElhaney 2002, 2013). On the other hand, any other factors affecting the dynamic properties of a lipid phase, such as interaction with sterols, proteins, flavonoids, etc. (Pawlikowska-Pawlęga et al. 2013), influence remarkably energy of the νsCH2 vibrations and therefore vibrational spectroscopy analysis appears a sensitive tool in monitoring fine effects of different modifiers on the dynamic and structural properties of the hydrophobic core of the lipid bilayers. A consequence of the ordering effect observed can be disturbance in membrane fluidity and, finally, loss of its integrity. Given that ordering of lipid acyl chains leads to stiffening of the membrane structure (Frolov et al. 2011; Fowler et al. 2016), these changes also contributed to the demonstrated increase in DMT modulus of E. coli cell surface upon cecropin D treatment. Using synthetic G. mellonella cecropin D named Gm1 and the rough type LPS of P. mirabilis and S. enterica, Correa et al. (2014b) demonstrated that the acyl chains of LPS became disordered upon Gm1 binding, leading to fluidization of the structure. The ordering effect observed in our study versus the disordering action described by Correa et al. (2014b) can be explained by the differences in the structure of the LPS used. These authors reported that the lipid A moiety was the most important part of the LPS for interaction with the peptide, whereas the length of the LPS sugar chain had no effect on the cecropin binding. Their results also suggested that cecropin D molecules were located in the lipid head group regions and then underwent intercalation into the P. mirabilis LPS aggregates (Correa et al. 2014b). The lipid A moiety of LPS was also important for synthetic H. cecropia cecropin A binding that occurred due to hydrophobic interactions between C-terminal helices of the peptide and the lipid A (De Lucca et al. 1995). Dissociation of LPS aggregates upon H. cecropia cecropin A binding was also reported (Lee et al. 2015). Interestingly, using circular dichroism Avitabile et al. (2014) demonstrated that folding of α-helical peptides, cecropin A and magainin 2, was induced by interaction with LPS present in the outer membrane of E. coli cells.
Correa et al. (2014a) studied the morphological changes in the S. enterica and P. mirabilis cell surface caused by synthetic G. mellonella cecropin D used at a 10-µM concentration. These alterations were described as indentations, leakage of cytoplasm, clustering of the cells, and fusion of the membranes. As demonstrated by AFM analysis in the present work, binding of G. mellonella cecropin D (applied at a four-times lower concentration than that used in the above study) to E. coli cells resulted in loss of characteristic cell surface topography and showed significant changes in nanomechanical properties. The alterations in the E. coli cell envelope were undoubtedly confirmed by TEM imaging, which revealed loss of the multilayer structure of the cell envelope, formation of membrane vesicles, and enlargement of the periplasmic space. Nevertheless, the E. coli cells exposed to G. mellonella cecropin D seemed to retain their shape to some extent, as no disintegrated cells were observed. In contrast, the cell membrane of E. coli treated with Heliothis virescens cecropin B exhibited formation of pores that eventually led to disruption of membrane continuity and cell disintegration (Lockey and Ourth 1996). A comparison of TEM images obtained in this study and those presented by Lockey and Ourth (1996) suggests that the mode of G. mellonella cecropin D action against E. coli cells may be based on a mechanism other than pore formation. On the other hand, Vaara and Vaara (1994) demonstrated that the mode of the antibacterial action of synthetic H. cecropia cecropin B resembled that of cationic amphiphilic detergents, i.e., benzalkonium chloride and cetylpyridinium chloride, known to disorganize the outer membrane resulting in lysis of Gram-negative bacteria cells. In this context, considering the TEM images of E. coli cells that exhibited symptoms of partial membrane dissolution, the action of G. mellonella cecropin D might be explained by detergent-like activity; however, this hypothesis needs further studies. Interestingly, it was postulated that cecropins A and B used at sub-inhibitory concentrations might induce a stress response in viable E. coli cells by activation of the hyperosmotic stress gene osmY (Oh et al. 1998).
In summary, the antibacterial activity of native G. mellonella cecropin D against E. coli is determined by initial interaction with LPS in the outer membrane and other lipid components of the cell envelope. This interaction affects the membrane fluidity and integrity by an ordering effect in regard to lipid chains. The effective binding demonstrated by FLIM leads to damage to the cell envelope reflected by disruption of the multilayered structure, changes in the nanomechanical properties, and loss of characteristic surface topography. Disturbance in the membrane integrity results in permeabilization of the cell membrane and leakage of the intracellular content, as confirmed by the cytoplasmic β-galactosidase activity assay. Although they retained their shape to some extent, E. coli cells experiencing these changes eventually die. Considering the neutral character of G. mellonella cecropin D, our results provide further evidence that the overall cationic character is not a necessary factor for antibacterial activity of AMPs and for effective binding to bacterial cells.
References
Agrawal A, Weisshaar JC (2018) Effects of alterations of the E. coli lipopolysaccharide layer on membrane permeabilization events induced by cecropin A. Biochim Biophys Acta 1860:1470–1479. https://doi.org/10.1016/j.bbamem.2018.04.009
Avitabile C, D’Andrea LD, Romanelli A (2014) Circular dichroism studies on the interactions of antimicrobial peptides with bacterial cells. Sci Rep 4:4293. https://doi.org/10.1038/srep04293
Bocheńska O, Rąpała-Kozik M, Wolak N, Braś G, Kozik A, Dubin A, Aoki W, Ueda M, Mak P (2013) Secreted aspartic peptidases of Candida albicans liberate bactericidal hemocidins from human hemoglobin. Peptides 48:49–58. https://doi.org/10.1016/j.peptides.2013.07.023
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Brandenburg K, Seydel U (1998) Infrared spectroscopy of glycolipids. Chem Phys Lipids 96:23–40
Brown SE, Howard A, Kasprzak AB, Gordon KH, East PD (2008) The discovery and analysis of a diverged family of novel antifungal moricin-like peptides in the wax moth Galleria mellonella. Insect Biochem Mol Biol 38:201–212. https://doi.org/10.1016/j.ibmb.2007.10.009
Brown SE, Howard A, Kasprzak AB, Gordon KH, East PD (2009) A peptidomics study reveals the impressive antimicrobial peptide arsenal of the wax moth Galleria mellonella. Insect Biochem Mol Biol 39:792–800. https://doi.org/10.1016/j.ibmb.2009.09.004
Bulet P, Stöcklin R (2005) Insect antimicrobial peptides: structures, properties and gene regulation. Protein Pept Lett 12:3–11
Correa W, Manrique-Moreno M, Behrends J, Patiño E, Marella C, Peláez-Jaramillo C, Garidel P, Gutsmann T, Brandenburg K, Heinbockel L (2014a) Galleria mellonella native and analogue peptides Gm1 and ΔGm1. II) Anti-bacterial and anti-endotoxic effects. Biochim Biophys Acta 1838:2739–2744. https://doi.org/10.1016/j.bbamem.2014.07.005
Correa W, Manrique-Moreno M, Patiño E, Peláez-Jaramillo C, Kaconis Y, Gutsmann T, Garidel P, Heinbockel L, Brandenburg K (2014b) Galleria mellonella native and analogue peptides Gm1 and ΔGm1. I) Biophysical characterization of the interaction mechanisms with bacterial model membranes. Biochim Biophys Acta 1838:2728–2738. https://doi.org/10.1016/j.bbamem.2014.07.006
Cytryńska M, Zdybicka-Barabas A (2015) Defense peptides: recent developments. Biomol Concepts 6:237–251. https://doi.org/10.1515/bmc-2015-0014
Cytryńska M, Mak P, Zdybicka-Barabas A, Suder P, Jakubowicz T (2007) Purification and characterization of eight peptides from Galleria mellonella immune hemolymph. Peptides 28:533–546
De Lucca AJ, Jacks TJ, Brogden KA (1995) Binding between lipopolysaccharide and cecropin A. Mol Cell Biochem 151:141–148
Ding L, Yang L, Weiss TM, Waring AJ, Lehrer RI, Huang HW (2003) Interaction of antimicrobial peptides with lipopolysaccharides. Biochemistry 42:12251–12259
Drozdetskiy A, Cole C, Procter J, Barton GJ (2015) JPred4: a protein secondary structure prediction server. Nucleic Acids Res 43(W1):W389–W394. https://doi.org/10.1093/nar/gkv332
Ebbensgaard A, Mordhorst H, Overgaard MT, Nielsen CG, Aarestrup FM, Hansen EB (2015) Comparative evaluation of the antimicrobial activity of different antimicrobial peptides against a range of pathogenic bacteria. PLoS one 10:e0144611. https://doi.org/10.1371/journal.pone.0144611
Fowler PW, Hélie J, Duncan A, Chavent M, Koldsø H, Sansom MS (2016) Membrane stiffness is modified by integral membrane proteins. Soft Matter 12:7792–7803
Frolov VA, Shnyrova AV, Zimmerberg J (2011) Lipid polymorphisms and membrane shape. Cold Spring Harb Perspect Biol 3:a004747. https://doi.org/10.1101/cshperspect.a004747
Gudmundsson GH, Lidholm DA, Asling B, Gan R, Boman HG (1991) The cecropin locus. Cloning and expression of a gene cluster encoding three antibacterial peptides in Hyalophora cecropia. J Biol Chem 266:11510–11517
Hassan M, Ilev I (2014) Grazing incidence angle based sensing approach integrated with fiber-optic Fourier transform infrared (FO-FTIR) spectroscopy for remote and label-free detection of medical device contaminations. Rev Sci Instrum 85:103108. https://doi.org/10.1063/1.4897247
Heinbockel L, Palacios-Chaves L, Alexander C, Rietschel E, Behrends J, Correa W, Fukuoka S, Gutsmann T, Ulmer AJ, Brandenburg K (2015) Mechanism of Hbγ-35-induced an increase in the activation of the human immune system by endotoxins. Innate Immun 21:305–313. https://doi.org/10.1177/1753425914535957
Hultmark D, Engström A, Bennich H, Kapur R, Boman HG (1982) Insect immunity: isolation and structure of cecropin D and four minor antibacterial components from Cecropia pupae. Eur J Biochem 127:207–217
Kim CH, Lee JH, Kim I, Seo SJ, Son SM, Lee KY, Lee IH (2004) Purification and cDNA cloning of a cecropin-like peptide from the great wax moth, Galleria mellonella. Mol Cells 17:262–266
Lee E, Jeong KW, Lee J, Shin A, Kim JK, Lee J, Lee DG, Kim Y (2013) Structure-activity relationships of cecropin-like peptides and their interactions with phospholipid membrane. BMB Rep 46:282–287
Lee E, Shin A, Kim Y (2015) Anti-inflammatory activities of cecropin A and its mechanism of action. Arch Insect Biochem Physiol 88:31–44. https://doi.org/10.1002/arch.21193
Lewis RN, McElhaney RN (2002) Vibrational spectroscopy of lipids. In: Chalmers JM, Griffiths PR (eds) Handbook of vibrational spectroscopy. John Wiley & Sons, Ltd., Chichester, pp 3447–3464
Lewis RN, McElhaney RN (2013) Membrane lipid phase transitions and phase organization studied by Fourier transform infrared spectroscopy. Biochim Biophys Acta 1828:2347–2358. https://doi.org/10.1016/j.bbamem.2012.10.018
Lockey TD, Ourth DD (1996) Formation of pores in Escherichia coli cell membranes by a cecropin isolated from hemolymph of Heliothis virescens larvae. Eur J Biochem 236:263–271
Luchowski R, Gryczynski Z, Sarkar P, Borejdo J, Szabelski M, Kapusta P, Gryczynski I (2009a) Instrument response standard in time-resolved fluorescencje. Rev Sci Instrum 80:033109. https://doi.org/10.1063/1.3095677
Luchowski R, Kapusta P, Szabelski M, Sarkar P, Borejdo J, Gryczynski Z, Gryczynski I (2009b) Forster resonance energy transfer (FRET)-based picosecond lifetime reference for instrument response evaluation. Meas Sci Technol 20:095601
Mak P, Chmiel E, Gacek G (2001) Antibacterial peptides of the moth Galleria mellonella. Acta Biochim Pol 48:1191–1195
Mak P, Siwek M, Pohl J, Dubin A (2007) Menstrual hemocidin HbB115-146 is an acidophilic antibacterial peptide potentiating the activity of human defensins, cathelicidin and lysozyme. Am J Reprod Immunol 57:81–91. https://doi.org/10.1111/j.1600-0897.2006.00456.x
Mak P, Zdybicka-Barabas A, Cytryńska M (2010) A different repertoire of Galleria mellonella antimicrobial peptides in larvae challenged with bacteria and fungi. Dev Comp Immunol 34:1129–1136. https://doi.org/10.1016/j.dci.2010.06.005
Nguyen LT, Haney EF, Vogel HJ (2011) The expanding scope of antimicrobial peptide structures and their modes of action. Trends Biotechnol 29:464–472. https://doi.org/10.1016/j.tibtech.2011.05.001
Oh JT, Cajal Y, Dhurjati PS, Van Dyk TK, Jain MK (1998) Cecropins induce the hyperosmotic stress response in Escherichia coli. Biochim Biophys Acta 1415:235–245
Oñate-Garzón J, Ausili A, Manrique-Moreno M, Torrecillas A, Aranda FJ, Patiño E, Gomez-Fernández JC (2017a) The increase in positively charged residues in cecropin D-like Galleria mellonella favors its interaction with membrane models that imitate bacterial membranes. Arch Biochem Biophys 629:54–62. https://doi.org/10.1016/j.abb.2017.07.008
Oñate-Garzón J, Manrique-Moreno M, Trier S, Leidy C, Torres R, Patiño E (2017b) Antimicrobial activity and interactions of cationic peptides derived from Galleria mellonella cecropin D-like peptide with model membranes. J Antibiot (Tokyo) 70:238–245. https://doi.org/10.1038/ja.2016.134
Otvos L Jr (2000) Antibacterial peptides isolated from insects. J Pept Sci 6:497–511. https://doi.org/10.1002/1099-1387(200010)6:10<497::AID-PSC277>3.0.CO;2-W
Palusińska-Szysz M, Zdybicka-Barabas A, Pawlikowska-Pawlęga B, Mak P, Cytryńska M (2012) Anti-Legionella dumoffii activity of Galleria mellonella defensin and apolipophorin III. Int J Mol Sci 13:17048–17064. https://doi.org/10.3390/ijms131217048
Papo N, Shai Y (2005) A molecular mechanism for lipopolysaccharide protection of Gram-negative bacteria from antimicrobial peptides. J Biol Chem 280:10378–10387
Patiño-Márquez IA, Manrique-Moreno M, Patiño-González E, Jemioła-Rzemińska M, Strzałka K (2018) Effect of antimicrobial peptides from Galleria mellonella on molecular models of Leishmania membrane. Thermotropic and fluorescence anisotropy study. J Antibiot (Tokyo) 71:642–652. https://doi.org/10.1038/s41429-018-0050-2
Pawlikowska-Pawlęga B, Misiak LE, Zarzyka B, Paduch R, Gawron A, Gruszecki WI (2013) FTIR, (1)H NMR and EPR spectroscopy studies on the interaction of flavone apigenin with dipalmitoylphosphatidylcholine liposomes. Biochim Biophys Acta 1828:518–527. https://doi.org/10.1016/j.bbamem.2012.10.013
Rahnamaeian M, Cytryńska M, Zdybicka-Barabas A, Dobslaff K, Wiesner J, Twyman RM, Zuchner T, Sadd BM, Regoes RR, Schmid-Hempel P, Vilcinskas A (2015) Insect antimicrobial peptides show potentiating functional interactions against Gram-negative bacteria. Proc Biol Sci 282:20150293. https://doi.org/10.1098/rspb.2015.0293
Rahnamaeian M, Cytryńska M, Zdybicka-Barabas A, Vilcinskas A (2016) The functional interaction between abaecin and pore-forming peptides indicates a general mechanism of antibacterial potentiation. Peptides 78:17–23. https://doi.org/10.1016/j.peptides.2016.01.016
Reißer S, Strandberg E, Steinbrecher T, Ulrich AS (2014) 3D hydrophobic moment vectors as a tool to characterize the surface polarity of amphiphilic peptides. Biophys J 106:2385–2394. https://doi.org/10.1016/j.bpj.2014.04.020
Schägger H, von Jagow G (1987) Tricine-sodium dodecyl sulfate-polyacrylamide gel electrophoresis for the separation of proteins in the range from 1 to 100 kDa. Anal Biochem 166:368–379
Scocchi M, Tossi A, Gennaro R (2011) Proline-rich antimicrobial peptides: converging to a non-lytic mechanism of action. Cell Mol Life Sci 68:2317–2330. https://doi.org/10.1007/s00018-011-0721-7
Sidén I, Boman HG (1983) Escherichia coli mutants with an altered sensitivity to cecropin D. J Bacteriol 154:170–176
Vaara M, Vaara T (1994) Ability of cecropin B to penetrate the enterobacterial outer membrane. Antimicrob Agents Chemother 38:2498–2501
Vogel H, Altincicek B, Glöckner G, Vilcinskas A (2011) A comprehensive transcriptome and immune-gene repertoire of the lepidopteran model host Galleria mellonella. BMC Genomics 12:308. https://doi.org/10.1186/1471-2164-12-308
Yi HY, Chowdhury M, Huang YD, Yu XQ (2014) Insect antimicrobial peptides and their applications. Appl Microbiol Biotechnol 98:5807–5822. https://doi.org/10.1007/s00253-014-5792-6
Zdybicka-Barabas A, Januszanis B, Mak P, Cytryńska M (2011) An atomic force microscopy study of Galleria mellonella apolipophorin III effect on bacteria. Biochim Biophys Acta 1808:1896–1906. https://doi.org/10.1016/j.bbamem.2011.03.013
Zdybicka-Barabas A, Mak P, Klys A, Skrzypiec K, Mendyk E, Fiołka MJ, Cytryńska M (2012) Synergistic action of Galleria mellonella anionic peptide 2 and lysozyme against Gram-negative bacteria. Biochim Biophys Acta 1818:2623–2635. https://doi.org/10.1016/j.bbamem.2012.06.008
Zdybicka-Barabas A, Palusińska-Szysz M, Gruszecki WI, Mak P, Cytryńska M (2014) Galleria mellonella apolipophorin III - an apolipoprotein with anti-Legionella pneumophila activity. Biochim Biophys Acta 1838:2689–2697. https://doi.org/10.1016/j.bbamem.2014.07.003
Acknowledgements
The work was financially supported by grant No 2012/05/B/NZ1/00,033 from the National Science Centre (Poland). The research was carried out with the equipment purchased thanks to the financial support of the European Regional Development Fund in the framework of the Polish Innovation Economy Operational Program (contract no. POIG.02.01.00-06-024/09 Centre of Functional Nanomaterials) as well as European Union structural funds (grants POIG.02.01.00-12-064/08 and POIG.02.01.00-12-167/08). The Faculty of Biochemistry, Biophysics and Biotechnology of Jagiellonian University in Krakow is a partner of the Leading National Research Centre (KNOW) supported by the Ministry of Science and Higher Education (Warszawa, Poland).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed.
Additional information
Handling Editor: J. Metcalf.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Zdybicka-Barabas, A., Stączek, S., Pawlikowska-Pawlęga, B. et al. Studies on the interactions of neutral Galleria mellonella cecropin D with living bacterial cells. Amino Acids 51, 175–191 (2019). https://doi.org/10.1007/s00726-018-2641-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-018-2641-4