Abstract
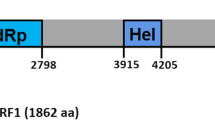
The complete genome sequence of a positive-sense single-stranded RNA (+ ssRNA) virus, Rhizoctonia beny-like virus 1 (RBLV1), isolated from binucleate Rhizoctonia AG-A strain A46, was determined. The RBLV1 genome is 10,280 nt in length and contains a short stretch of adenines at the 3′ terminus. It contains a single open reading frame (ORF) encoding a 376.30-kDa protein with viral helicase and RNA-dependent RNA polymerase (RdRp) motifs. The encoded protein exhibited the highest sequence similarity to Rhizoctonia cerealis beny-like virus 0928-1 (RcBeLV 0928-1, 45.25%), with a sequence coverage of 63%. Phylogenetic analysis based on ORF protein sequences revealed that RBLV1 is a novel unclassified mycovirus.


Similar content being viewed by others

References
Sneh B, Burpee L, Ogoshi A (1991) Identification of Rhizoctonia species. The American Phytopathological Society Press, St Paul, pp 55121–52097
Sharon M, Sneh B, Kuninaga S, Hyakumachi M (2006) The advancing identification and classification of Rhizoctonia spp. using molecular and biotechnological methods compared with the classical anastomosis grouping. Mycoscience 47:299–316
Sharon M, Sneh B, Kuninaga S, Hyakumachi M, Naito S (2008) Classification of Rhizoctonia spp. using rDNA-ITS sequence analysis supports the genetic basis of the classical anastomosis grouping. Mycoscience 49:93–114
Dong W, Li Y, Duan C, Li X, Naito S, Conner RL, Yang G, Li C (2017) Identification of AG-V, a new anastomosis group of binucleate Rhizoctonia spp. from taro and ginger in Yunnan province. Eur J Plant Pathol 148:895–906
Jian JH, Lakshman DK, Tavantzis SM (1998) A virulence-associated, 6.4-kb, double-stranded RNA from Rhizoctonia solani is phylogenetically related to plant bromoviruses and electron transport enzymes. Mol Plant Microbe In 11:601–609
Strauss EE, Lakshman DK, Tavantzis SM (2000) Molecular characterization of the genome of a partitivirus from the basidiomycete Rhizoctonia solani. J Gen Virol 81:549–555
Zheng L, Liu H, Zhang M, Cao X, Zhou E (2013) The complete genomic sequence of a novel mycovirus from Rhizoctonia solani AG-1 IA strain B275. Arch Virol 158:1609–1612
Bartholomaus A, Wibberg D, Winkler A, Puhler A, Schluter A, Varrelmann M (2017) Identification of a novel mycovirus isolated from Rhizoctonia solani (AG 2–2 IV) provides further information about genome plasticity within the order Tymovirales. Arch Virol 162:555–559
Chen Y, Su JE, Qin XY, Fan ZY, Zhang XH, Yu Q, Xia ZY, Zou CM, Zhao GK, Lin ZL (2020) A novel putative betapartitivirus isolated from the plant-pathogenic fungus Rhizoctonia solani. Arch Virol 165:1697–1701
Das S, Falloon RE, Stewart A, Pitman AR (2016) Novel mitoviruses in Rhizoctonia solani AG-3PT infecting potato. Fungal Biol 120:338–350
Lyu R, Zhang Y, Tang Q, Li Y, Cheng J, Fu Y, Chen T, Jiang D, Xie J (2018) Two alphapartitiviruses co-infecting a single isolate of the plant pathogenic fungus Rhizoctonia solani. Arch Virol 163:515–520
Zhang M, Zheng L, Liu C, Shu C, Zhou E (2018) Characterization of a novel dsRNA mycovirus isolated from strain A105 of Rhizoctonia solani AG-1 IA. Arch Virol 163:427–430
Zheng L, Shu C, Zhang M, Yang M, Zhou E (2019) Molecular characterization of a novel endornavirus conferring hypovirulence in rice sheath blight fungus Rhizoctonia solani AG-1 IA strain GD-2. Viruses. https://doi.org/10.3390/v11020178
Picarelli M, Forgia M, Rivas EB, Nerva L, Chiapello M, Turina M, Colariccio A (2019) Extreme diversity of mycoviruses present in isolates of Rhizoctonia solani AG2-2 LP From Zoysia japonica From Brazil. Front Cell Infect Mi 9:244
Abdoulaye AH, Foda MF, Kotta-Loizou I (2019) Viruses infecting the plant pathogenic fungus Rhizoctonia solani. Viruses https//. https://doi.org/10.3390/v11121113
Andika IB, Wei S, Cao C, Salaipeth L, Kondo H, Sun L (2017) Phytopathogenic fungus hosts a plant virus: a naturally occurring cross-kingdom viral infection. Proc Natl Acad Sci USA 114:12267–12272
Sun Y, Li Y, Dong W, Sun A, Chen N, Zhao Z, Li Y, Li C, Yang G (2021) Molecular characterization of a novel mycovirus isolated from Rhizoctonia solani AG-1 IA strain 9–11. Arch Virol 166:3229–3232
Li Y, Chen W, Niu Y, Xu P, Zhang L, Yu S, Yang G, Mo X (2022) Complete nucleotide sequence of a novel alphapartitivirus from Rhizoctonia solani AG-4 HG III isolate SM03. Arch Virol 167:953–957
Charlton ND, Carbone I, Tavantzis SM, Cubeta MA (2008) Phylogenetic relatedness of the M2 double-stranded RNA in Rhizoctonia fungi. Mycologia 100:555–564
Li W, Zhang T, Sun H, Deng Y, Zhang A, Chen H, Wang K (2014) Complete genome sequence of a novel endornavirus in the wheat sharp eyespot pathogen Rhizoctonia cerealis. Arch Virol 159:1213–1216
Li W, Zhang H, Shu Y, Cao S, Sun H, Zhang A, Chen H (2021) Genome structure and diversity of novel endornaviruses from wheat sharp eyespot pathogen Rhizoctonia cerealis. Virus Res 297:198368
Li Y, Sun Y, Xu P, Zhang L, Chen Z, Niu Y, Yang G, Mo X (2022) A novel alphapartitivirus from binucleate Rhizoctonia fumigata AG-Ba isolate C-314 Baishi. Arch Virol 167:255–259
Li Y, Xu P, Zhang L, Xia Z, Qin X, Yang G, Mo X (2015) Molecular characterization of a novel mycovirus from Rhizoctonia fumigata AG-Ba isolate C-314 Baishi. Arch Virol 160:2371–2374
Li Y, Xu P, Zhang L, Xia Z, Qin X, Yang G, Mo X (2015) Molecular characterization of a novel mycovirus from Rhizoctonia fumigata AG-Ba isolate C-314 Baishi. Arch Virol 160:2371–2374
Li Y, Sun Y, Yu L, Chen W, Liu H, Yin L, Guang Y, Yang G, Mo X (2022) Complete genome sequence of a novel mitovirus from binucleate Rhizoctonia AG-K strain FAS2909W. Arch Virol 167:271–276
Deakin G, Dobbs E, Bennett JM, Jones IM, Grogan HM, Burton KS (2017) Multiple viral infections in Agaricus bisporus - characterisation of 18 unique RNA viruses and 8 ORFans identified by deep sequencing. Sci Rep 7:2469
Nerva L, Turina M, Zanzotto A, Gardiman M, Gaiotti F, Gambino G, Chitarra W (2019) Isolation, molecular characterization and virome analysis of culturable wood fungal endophytes in esca symptomatic and asymptomatic grapevine plants. Environ Microbiol 21:2886–2904
Silvestri A, Turina M, Fiorilli V, Miozzi L, Venice F, Bonfante P, Lanfranco L (2020) Different genetic sources contribute to the small RNA population in the arbuscular mycorrhizal fungus gigaspora margarita. Front Microbiol 11:395
Sutela S, Forgia M, Vainio EJ, Chiapello M, Daghino S, Vallino M, Martino E, Girlanda M, Perotto S, Turina M (2020) The virome from a collection of endomycorrhizal fungi reveals new viral taxa with unprecedented genome organization. Virus Evol 6:1–39
Turina M, Ghignone S, Astolfi N, Silvestri A, Bonfante P, Lanfranco L (2018) The virome of the arbuscular mycorrhizal fungus Gigaspora margarita reveals the first report of DNA fragments corresponding to replicating non-retroviral RNA viruses in fungi. Environ Microbiol 20:2012–2025
Marzano SL, Nelson BD, Ajayi-Oyetunde O, Bradley CA, Hughes TJ, Hartman GL, Eastburn DM, Domier LL (2016) Identification of diverse mycoviruses through metatranscriptomics characterization of the viromes of five major fungal plant pathogens. J Virol 90:6846–6863
Li Y, Lyu R, Hai D, Jia J, Jiang D, Fu Y, Cheng J, Lin Y, Xie J (2022) Two novel rhabdoviruses related to hypervirulence in a phytopathogenic fungus. J Virol 96:e0001222
Solovyev AG, Morozov SY (2022) Uncovering plant virus species forming novel provisional taxonomic units related to the family Benyviridae. Viruses 14:2680
Huang HJ, Ye ZX, Wang X, Yan XT, Zhang Y, He YJ, Qi YH, Zhang XD, Zhuo JC, Lu G, Lu JB, Mao QZ, Sun ZT, Yan F, Chen JP, Zhang CX, Li JM (2021) Diversity and infectivity of the RNA virome among different cryptic species of an agriculturally important insect vector: whitefly Bemisia tabaci. NPJ Biofilms Microbi 43:1–15
Li W, Sun H, Cao S, Zhang A, Zhang H, Shu Y, Chen H (2023) Extreme diversity of mycoviruses present in single strains of Rhizoctonia cerealis, the pathogen of wheat sharp eyespot. Microbiol Spectr 11:1–19
Zhu JZ, Zhu HJ, Gao BD, Zhou Q, Zhong J (2018) Diverse, novel mycoviruses from the virome of a hypovirulent Sclerotium rolfsii strain. Front Plant Sci 9:1738
Sidharthan VK, Kalaivanan NS, Baranwal VK (2021) Discovery of putative novel viruses in the transcriptomes of endangered plant species native to India and China. Gene 786:145626
Kwon SJ, Choi GS, Choi B, Seo JK (2018) Molecular characterization of an unusual new plant RNA virus reveals an evolutionary link between two different virus families. PLoS ONE 13:e0206382
Liu S, Valencia-Jiménez A, Darlington M, Vélez AM, Bonning BC (2020) Diabrotica undecimpunctata virus 2, a novel small RNA virus discovered from southern Corn Rootworm, Diabrotica undecimpunctata howardi Barber (Coleoptera Chrysomelidae). Microbiol Resour Ann 9:1–3
Gilmer D, Ratti C, Ictv Report C (2017) ICTV virus taxonomy profile: Benyviridae. J Gen Virol 98:1571–1572
Nerva L, Ciuffo M, Vallino M, Margaria P, Varese GC, Gnavi G, Turina M (2016) Multiple approaches for the detection and characterization of viral and plasmid symbionts from a collection of marine fungi. Virus Res 219:22–38
Darissa O, Willingmann P, Adam G (2010) Optimized approaches for the sequence determination of double-stranded RNA templates. J Virol Methods 169:397–403
Attoui H, Billoir F, Cantaloube JF, Biagini P, de Micco P, de Lamballerie X (2000) Strategies for the sequence determination of viral dsRNA genomes. J Virol Methods 89:147–158
Scotto-Lavino E, Du G, Frohman MA (2006) 3′ end cDNA amplification using classic RACE. Nat Protoc 1:2742–2745
Tamura K, Stecher G, Kumar S (2021) MEGA11: molecular evolutionary genetics analysis version 11. Mol Biol Evol 38(7):3022–3027
Acknowledgements
This research was supported by the National Natural Scientific Foundation of China (32260036, 42067009, 32001684), the Scientific Research Fund of the Education Department of Yunnan Province (2022Y696), and the Yunnan Provincial Science and Technology Department Surface Project (202201AT070020).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Johannes Wittmann
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
Below is the link to the electronic supplementary material
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Shi, R., Mo, X., Chen, Z. et al. Molecular characterization of a novel mycovirus from binucleate Rhizoctonia AG-A strain A46. Arch Virol 169, 31 (2024). https://doi.org/10.1007/s00705-024-05963-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00705-024-05963-2