Abstract
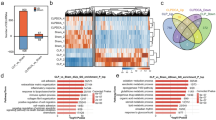
The hepatitis B virus (HBV) whole-X gene comprises the HBV X gene and the 168-bp region immediately upstream. Although the functions of HBx in hepatocarcinogenesis are well known, the activity of the HBV whole-X protein (HBwx), with 56 additional amino acids, has not yet been explored. In this study, proteomic and bioinformatic analysis was done to determine the protein interaction profiles of HBwx and HBx and to describe their functions in carcinogenesis. A total of 203 proteins were identified that interacted with HBwx, of which 149 were unique, the rest interacting also with HBx, and 73 % (148/203) of these proteins are involved in carcinogenesis. Gene ontology (GO) analysis showed that HBwx- and HBx-interacting proteins are involved in different processes, the former mainly in biosynthetic processes (glycolysis, cell-cycle functions, and protein folding), and the latter mainly in localization, viral transcription, biological adhesion and angiogenesis. Pathway networks analysis revealed that proteins interacting with HBx participate mainly in oxidative phosphorylation, localization, the cytoskeleton, and cell adhesion. In contrast, more-specific functional analysis showed that proteins interacting with HBwx are involved in apoptosis and survival, cell-cycle functions, glycolysis, and gluconeogenesis (Pathway Maps); to cellular macromolecular complex assembly, protein folding and mRNA metabolic process (GO Processes); and to regulation of protein folding in the endoplasmic reticulum and cytoplasm, transcription, cell cycle G2-M and cytoskeleton rearrangement (Process Networks). In conclusion, this study shows that HBwx functions in carcinogenesis in a way that is different from that of HBx.




Similar content being viewed by others
References
Bosch FX, Ribes J, Díaz M, Cléries R (2004) Primary liver cancer: worldwide incidence and trends. Gastroenterology 127(5 Suppl 1):S5–S16. doi:10.1053/j.gastro.2004.09.011
Kew MC (2010) Epidemiology of chronic hepatitis B virus infection, hepatocellular carcinoma, and hepatitis B virus-induced hepatocellular carcinoma. Pathol Biol (Paris) 58(4):273–277. doi:10.1016/j.patbio.2010.01.005
Michielsen P, Ho E (2011) Viral hepatitis B and hepatocellular carcinoma. Acta Gastroenterol Belg 74(1):4–8
Fares N, Peron JM (2013) Epidemiology, natural history, and risk factors of hepatocellular carcinoma. Rev Prat 63 (2):216–217, 220–212
Ng SA, Lee C (2011) Hepatitis B virus X gene and hepatocarcinogenesis. J Gastroenterol 46(8):974–990. doi:10.1007/s00535-011-0415-9
Kew MC (2011) Hepatitis B virus x protein in the pathogenesis of hepatitis B virus-induced hepatocellular carcinoma. J Gastroenterol Hepatol 26(Suppl 1):144–152. doi:10.1111/j.1440-1746.2010.06546.x
Doria M, Klein N, Lucito R, Schneider RJ (1995) The hepatitis B virus HBx protein is a dual specificity cytoplasmic activator of Ras and nuclear activator of transcription factors. EMBO J 14(19):4747–4757
Murakami S (2001) Hepatitis B virus X protein: a multifunctional viral regulator. J Gastroenterol 36(10):651–660
Loncarevic IF, Zentgraf H, Schroder CH (1990) Sequence of a replication competent hepatitis B virus genome with a preX open reading frame. Nucleic Acids Res 18(16):4940
Loncarevic IF, Schranz P, Zentgraf H, Liang XH, Herrmann G, Tang ZY, Schroder CH (1990) Replication of hepatitis B virus in a hepatocellular carcinoma. Virology 174(1):158–168
Takahashi SK (1995) A unique set of mutations in the ‘preX’ region of hepatitis B virus DNA frequently found in patients but not in asymptomatic carriers: implication for a novel variant. Int Hepatol Commun 3(3):131–138
Takahashi K, Akahane Y, Hino K, Ohta Y, Mishiro S (1998) Hepatitis B virus genomic sequence in the circulation of hepatocellular carcinoma patients: comparative analysis of 40 full-length isolates. Arch Virol 143(12):2313–2326
Yang Q, Dong J, Cheng J, Liu Y, Hong Y (2003) Definition of pre-X promoter sequence from hepatitis B virus genome and characterization of its transcription activity. JiefangJun YiXue Zazhi 28:763–765
Dong JCJ (2003) Study on definition of pre-X region in hepatitis B virus genome. World Chin J Digestol 11(8):1097–1101
Faure E (2006) Alternative peptide-fusion proteins generated by out-of-frame mutations, just upstream ORFs or elongations in mutants of human hepatitis B viruses. Virus Res 117(2):185–201. doi:10.1016/j.virusres.2005.10.023
Yang Q, Cheng J, Dong J, Zhang J, Zhang SL (2005) Molecular epidemiological study on pre-X region of hepatitis B virus and identification of hepatocyte proteins interacting with whole-X protein by yeast two-hybrid. World J Gastroenterol 11(22):3473–3478
Wisniewski JR, Zougman A, Mann M (2009) Combination of FASP and StageTip-based fractionation allows in-depth analysis of the hippocampal membrane proteome. J Proteome Res 8(12):5674–5678. doi:10.1021/pr900748n
Sun D, Zhang H, Guo D, Sun A, Wang H (2013) Shotgun proteomic analysis of plasma from dairy cattle suffering from footrot: characterization of potential disease-associated factors. PLoS One 8(2):e55973. doi:10.1371/journal.pone.0055973
Dai J, Shieh CH, Sheng QH, Zhou H, Zeng R (2005) Proteomic analysis with integrated multiple dimensional liquid chromatography/mass spectrometry based on elution of ion exchange column using pH steps. Anal Chem 77(18):5793–5799. doi:10.1021/ac050251w
Zhang T, Xie N, He W, Liu R, Lei Y, Chen Y, Tang H, Liu B, Huang C, Wei Y (2013) An integrated proteomics and bioinformatics analyses of hepatitis B virus X interacting proteins and identification of a novel interactor apoA-I. J Proteomics 84:92–105. doi:10.1016/j.jprot.2013.03.028
Lara-Pezzi E, Roche S, Andrisani OM, Sanchez-Madrid F, Lopez-Cabrera M (2001) The hepatitis B virus HBx protein induces adherens junction disruption in a src-dependent manner. Oncogene 20(26):3323–3331. doi:10.1038/sj.onc.1204451
Lee JO, Kwun HJ, Jung JK, Choi KH, Min DS, Jang KL (2005) Hepatitis B virus X protein represses E-cadherin expression via activation of DNA methyltransferase 1. Oncogene 24(44):6617–6625. doi:10.1038/sj.onc.1208827
Liu J, Lian Z, Han S, Waye MMY, Wang H, Wu MC, Wu K, Ding J, Arbuthnot P, Kew M, Fan D, Feitelson MA (2006) Downregulation of E-cadherin by hepatitis B virus X antigen in hepatocellullar carcinoma. Oncogene 25(7):1008–1017. doi:10.1038/sj.onc.1209138
Liu HO, Xu L, He HY, Zhu Y, Liu J, Wang SS, Chen L, Wu Q, Xu JJ, Gu JX (2012) Hepatitis B virus X protein promotes hepatoma cell invasion and metastasis by stabilizing Snail protein. Cancer Sci 103(12):2072–2081. doi:10.1111/cas.12017
Chen L, Hu L, Li L, Liu Y, Tu Q-Q, Chang Y-X, Yan H-X, Wu M-C, Wang H-Y (2010) Dysregulation of beta-catenin by hepatitis B virus X protein in HBV-infected human hepatocellular carcinomas. Front Med China 4(4):399–411. doi:10.1007/s11684-010-0170-y
Williams GH, Stoeber K (2012) The cell cycle and cancer. J Pathol 226(2):352–364. doi:10.1002/path.3022
Krisher RL, Prather RS (2012) A role for the Warburg effect in preimplantation embryo development: metabolic modification to support rapid cell proliferation. Mol Reprod Dev 79(5):311–320. doi:10.1002/mrd.22037
Acknowledgments
This work was supported by the Natural Scientific Foundation of China (No. 31100129).
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhang, Y., Liu, J., Liu, H. et al. Comparative study of the different activities of hepatitis B virus whole-X protein and HBx in hepatocarcinogenesis by proteomics and bioinformatics analysis. Arch Virol 160, 1645–1656 (2015). https://doi.org/10.1007/s00705-015-2421-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-015-2421-3