Summary
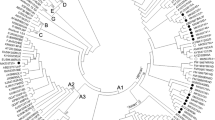
We determined full-length nucleotide sequence of hepatitis B virus (HBV) genome in sera from 40 Japanese patients with HBsAg-positive hepatocellular carcinoma (HCC), in order to obtain information on HCC-specific characteristics, if any, of the HBV genome. Direct sequencing of the long distance PCR products starting from 50 μl of serum samples revealed that 95% of our isolates were of genotype C, and that mutations and deletions/insertions were very common. With respect to envelope protein genes, deletions and missense mutations were frequent in preS2, and the determinant a domain of HBsAg was rich in “antibody-escape” mutations. Within the precore/core region, the most remarkable mutation was the replacement of proline of wild type by other amino acids at codon 130 of the core gene, which was found in 58% of our isolates, while precore-stop mutation was found in 45%. Most interestingly, however, about 90% of our isolates had mutations at nt positions 1762 (A-to-T) and 1764 (G-to-A) within the core promoter, which had been implicated in “e-suppressive” phenotype of HBV genome. G-to-A at nt 1613 and C-to-T at nt 1653 within enhancer II and T-to-C/A at nt 1753 within core promoter were also evident: 38%, 53%, and 40%, respectively. It was interesting that some of the characteristics observed in our isolates form HCC patients had been previously implicated in fulminant hepatitis and/or acute exacerbation of chronic hepatitis.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received May 18, 1998 Accepted July 18, 1998
Rights and permissions
About this article
Cite this article
Takahashi, K., Akahane, Y., Hino, K. et al. Hepatitis B virus genomic sequence in the circulation of hepatocellular carcinoma patients: comparative analysis of 40 full-length isolates. Arch. Virol. 143, 2313–2326 (1998). https://doi.org/10.1007/s007050050463
Published:
Issue Date:
DOI: https://doi.org/10.1007/s007050050463