Abstract
Ulmus laevis Pallas is a rare and endangered tree species in the northern part of Belgium, restricted to a minimal amount of natural populations that mainly consist of a strongly reduced number of individuals. Probably isolated for several generations, random phenotypic variation in the strongly declined populations can be hypothesised. We analysed U. laevis trees grown in a field trial with a single-tree-plot design (completely randomised). The plants were vegetatively propagated through cuttings from nearly all known natural relict populations in Flanders. We analysed three short-shoot leaves from different ramets from different genotypes from eleven natural populations. Leaf size and shape variables were computed using landmark and procruste methods. We visualised the variability in leaf morphology in a principal component analysis (PCA) including shape, size, and categorical morphological variables. The variance structure of the morphological variables was studied applying mixed modelling methods. The overall PCA distinguished two deviating natural populations. The Zandhoven population showed a deviating simple leaf margin toothing, correlated with larger numbers of split secondary veins. The Halle population is characterised by absence of pubescence at the lower side of the leaves. Both relict populations with deviating morphological leaf characteristics may point to an ecotype evolution putatively due to isolation combined with greatly diminished population sizes. The variance structure analysis of individual leaf variables showed that the deviating categorical leaf characters are under stronger genetic control (specialist characters) compared to the more plastic variables.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Habitat reduction and fragmentation have become major threats to many native species in Western Europe, reducing their adaptive and evolutionary potential in the face of climate change (Aguilar et al. 2006; Leimu et al. 2010). Especially in regions which suffered already in historical times from expanding human activities such as in Flanders, the northern part of Belgium, populations may express symptoms that can be related to isolation, population decline and inbreeding. As habitat fragmentation may lead to pollen limitation and limited gene flow in plants, trees and shrubs are as likely to experience negative genetic responses to habitat fragmentation (Vranckx et al. 2012). In addition, a wide range of species-specific responses to fragmentation can be observed limiting extrapolation from one species to another (Debinski and Holt 2000). Here we focus on Ulmus laevis Pallas (white elm), a European riparian tree species that is wind pollinated and self-incompatible. Seeds are dispersed by wind and by water, while natural clonal reproduction is infrequent to absent (Goodall-Copestake et al. 2005; Venturas et al. 2014). The centre of its distribution area is limited to Central and Eastern Europe, whereas populations in Western Europe have probably been less abundant naturally and have been further reduced by habitat loss. The nearly total disappearance of its natural habitat through diking of the main river systems and draining of flood plains, together with a highly impacted landscape due to intensive agriculture and urbanisation in Flanders, resulted in extremely reduced natural relict populations (Vander Mijnsbrugge et al. 2005). Although susceptible for the alien hypervirulent pathogen Ophiostoma ulmi s.l., agent of the Dutch elm disease (Solla et al. 2005), U. laevis is less attractive to the elm bark beetles acting as vector for the fungal pathogen (Sacchetti et al. 1990; Webber 2000). Also in other parts of western Europe relict populations suffer from varying human pressures making them more prone to climate change impacts (Hemery et al. 2010) and urging conservation programs (Goodall-Copestake et al. 2005; Fuentes-Utrilla et al. 2014). Relict populations do not only suffer from declined population sizes. Several molecular studies have detected low genetic diversities within natural populations together with high population differentiations (Vakkari et al. 2009; Venturas et al. 2013) and Central European populations show a demographic signature of recent bottlenecks (Nielsen and Kjaer 2010).
Flowers and fruits from U. laevis are stalked, which unambiguously distinguishes this species from the related U. minor and U. glabra (Vander Mijnsbrugge et al. 2005; Maes 2006; Lambinon et al. 2012). The leaves of U. laevis have a single tip and are often characterised by a very asymmetric base. The leaf lamina margin is strongly toothed (multiserrate) with the teeth being curled and pointing towards the leaf tip. The lower side of the leaf is softly pubescent. Secondary veins that run from the central vein to the leaf margin rarely split and, if so, only at the base of the leaf. As elms flower and fructify in spring, botanists often have to rely on leaf characteristics for identification. In addition, the flowers and fruits in the elm genus are small and simple, offering few characters of taxonomic value (Melville 1978). On the contrary, leaves of adult short shoots are highly diversified and can replace flowers for taxonomic purposes (Melville 1978). Short shoots form the lateral spurs on branches of adult trees and are characterised by a determinate growth, producing only a determinate number of leaves that are relatively stable in shape and size (Melville 1978). Very few studies have focussed on morphometry of elm populations and published results in English. Richens (1955) studied elm morphology in England using eight leaf characters, aiming to describe objectively the variation encountered in the genus. He distinguished the three main species U. minor (synonym of U. carpinifolia), U. glabra and U. procera, and suggested the presence of many clonal genotypes within the taxon U. minor. Petrokas and Baliuckas (2012) examined the complex U. minor–U. x hollandica–U. glabra by means of up to 21 measured, counted or derived leaf morphological variables. The application of a shape analysis based on geometric morphometrics (Punyasena and Smith 2014) can substantially reduce this amount of variables. The mean advantages of geometric morphometrics is that shape of an organ can be studied independent of its size and that complex shapes can be dissected into a discrete amount of quantifiable shape variations according to a limited amount of principal component axes explaining the larger part of the shape variability (Viscosi and Cardini 2011). Although leaves are plant organs suiting well geometric morphometric studies, botanists did not yet take as much advantage of this set of methods in comparison to zoologists, mostly restricted to taxonomic questions (Viscosi and Cardini 2011). For trees, several geometric morphometric studies have been conducted on the hybrid swarms of oaks (Jensen et al. 1993; Viscosi et al. 2009a, b, 2012; Penaloza-Ramirez et al. 2010) and other tree species (Jensen et al. 2002; Vit et al. 2012; Vander Mijnsbrugge 2015) but, to our knowledge, not yet on elm species.
Because of their rarity, the relicts of U. laevis in Flanders were hardly known before the end of the nineties of last century, when Flanders was inventoried specifically for autochthonous relict populations of woody species (Vander Mijnsbrugge et al. 2005). Our central aim was to quantify and visualise the morphological variation in leaves of the relict populations of U. laevis, as this can lead to better insights into the evolutionary and ecological processes of phenotypic diversification (Fu et al. 2013; Bo et al. 2014). More specifically, our objective was to look at the consequences of habitat fragmentation in quantitative genetic characters, as habitat fragmentation is expected to erode genetic variability of plant populations and to increase interpopulation genetic divergence due to genetic drift, inbreeding and reduced gene flow (Aguilar et al. 2008). We analysed U. laevis trees in a completely randomised field trial that were vegetatively propagated from eleven natural relict populations in Flanders. We (1) computed leaf size and leaf shape variables using landmark and procruste methods, (2) visualised the variability in leaf morphology in a PCA analysis including shape, size and categorical morphological variables and (3) partitioned the variance of separate morphological variables using mixed modelling techniques.
Materials and methods
Field trial and plant material
As an ex situ conservation measure, most of the known relict populations of Ulmus laevis in Flanders were visited in the late spring of 2003, and genotypes were vegetatively propagated by spring cuttings and grown in plant pots using standard nursery practices at the Research Institute for Nature and Forest, Geraardsbergen, Belgium (Table 1; Fig. 1). Potted planting stocks of successful cuttings with a maximum of four ramets per genotype were planted in a field trial at Malgedem (Fig. 1), at an interspacing of 7 × 7 m in early spring of 2006. The field trial followed a single-tree-plot design (ramets individually and randomly intermingled). In the summer of 2014, two to three healthy short shoots per tree in the crown of the young trees were sampled from all trees. In this way, leaves were sampled that were fully exposed to light on a well-developed branch. Three healthy, damage-free and well-developed leaves per tree were carefully selected from these short shoots for further analysis. In the majority of cases, these were the 4th to 5th leaf of a short shoot counting from the top.
Map of Flanders showing the original natural Ulmus laevis populations (dots) and the location of the field trial (cross). Abbreviations of populations are in Table 1
Morphological variables
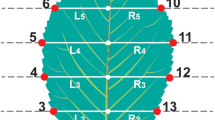
Six categorical characters were observed on each leaf (Table 2; Fig. 2): two were counted (MT and SSV) while the other four describe the density of pubescence using Kissling’s grading system (Kissling 1977). Furthermore, each leaf was digitised, together with a measuring stick, and 13 landmarks were placed on each image (Fig. 3) using the free tpsDig software (Rohlf 2010). As this is a first report, to our knowledge, on an application of geometric morphometrics on elm leaves, the configuration of the landmarks was original, aiming to capture as much as possible the shape variation present in the sampled leaves. Procrustes coordinates were calculated for these landmarks and were superimposed using the free CoordGen software (Sheets 2004). Procrustes rescale the original coordinates of the landmarks on a leaf around a central point. Because in a biological context shapes usually vary along all landmarks, the procrustes method is preferred over the bookstein method where two chosen landmarks are kept stable and variability in shape is assigned to all other landmarks. Superimposed procrustes coordinates can be considered as being points on the surface of a sphere. Transformation of these coordinates to fit in a rectangular coordinate system allows further processing in conventional statistics. The transformed coordinates are called partial warp scores. Partial warp scores were calculated from the superimposition data, and a PCA analysis was performed on the covariance matrix derived from the partial warp scores using the free PCAGen software (Sheets 2004). Because the first three axes explain a larger part of the variability in the dataset (proportional explained variance: 23.2, 19.1 and 12.2 %), the first three PCA axis scores were used as shape variables in further analysis. Applying the free Morpheus software (Slice 2013) and taking into account the scale factor (amount of pixels per measuring unit), the actual lengths between the landmarks 2–8 and 5–11 (Fig. 3) were calculated and further treated as two size variables, abbreviated as LL (lamina length) and LW (lamina width), respectively.
Observation places of the categorical morphological characters on a schematic Ulmus laevis leaf. Abbreviations are in Table 2
Principal component analysis and variance analysis
A principal component analysis was performed on the six categorical, two size and three shape variables using the command prcomp in R (R Core Team 2013). The hierarchical structure (population, genotype and ramet) was tested for each morphological variable, including the principle component scores of the axes explaining the larger part of the variation, applying ANOVA (population, genotype and ramet as categorical explanatory variables) using the R command aov. The contribution of the different populations, the different genotypes within the populations and the different ramets within the genotypes to the total variance in the dataset were estimated using mixed effect modelling. Mixed effect modelling provides a robust analytical approach for addressing hierarchical nested data (Zuur et al. 2009). In addition, mixed models can cope with unbalanced data (Zuur et al. 2009). The lme4 package in R (Bates et al. 2014) was applied to partition the variance of continuous variables (command lmer). Each continuous morphological variable was modelled as a response variable with population, genotype nested in population and ramet nested in genotype all in the random part (no co-variables in the fixed part). For the analysis of the variance structure for the categorical count variables (SSV and MT, Table 2), a general linear mixed model with a poisson distribution was applied using the command glmer of the package lme4. The variance of the ordinal categorical variables (LPV and LPM, Table 2) was studied using cumulative logistic regression in the package ordinal in R (Christensen 2013). For modelling of the categorical variables, a similar model structure was applied as for the continuous variables, with no co-variables in the fixed part and the same random part (population, genotype and ramet). General linear and cumulative logistic mixed models have no error term and are, therefore, lacking a residual variance component.
Results
Leaf shape analysis of Ulmus laevis
We studied the leaf shape variability of the sampled U. laevis trees applying landmark and procruste methods. The relative positions of the landmarks on each leaf were analysed minimising relative distances between the landmarks among the leaves while excluding any size differences. A principal component analysis quantified the variability in shape among the leaves (proportional explained variance of first three PCA axes: 23.2, 19.1 and 12.2 %). Leaves close to the middle of the PCA plots of the first three axes (scores close to zero) were considered approaching the average leaf shape of U. laevis in Flanders (Fig. 4). Allometric relationships with other morphological variables were tested with Spearman correlations (Table 3). As lamina length (LL) correlated with the first PCA axis and lamina width (LW) with the third, this mean leaf shape also approximated the mean leaf size. To visualise the shape variability among the leaves, the contours of two leaves with extreme scores along the first axis (positive and negative) while remaining close to score zero along the second and third axis, were drawn from the digitised images (Fig. 5a). Similarly, leaves were chosen with extreme scores along the second or third axis combined with scores close to zero along the other axes and contours were drawn (Fig. 5b, c). To exclude artefacts, these outlines of leaves at the extremities of the axes were checked by comparing with quiver plots that visualise shape variability along the different PCA axes and that are calculated by the PCAGen software (Online Resource 1, 2 and 3). Along the first axis, leaves varied from narrower shapes (− scores, full line drawing in Fig. 5a) to wider shapes (+ scores, dashed line drawing in Fig. 5a). As the size variable LL was negatively correlated with the first PCA axis (Table 3), the wider leaves according to the shape analysis tended to smaller lamina lengths in absolute terms. From – to + scores along the second axis (from full line to dashed line in Fig. 5b), leaves evolved towards a smaller width of the leaf base at the longest side of the leaf and tended towards more acuminate tips. The third axis fluctuated between asymmetric leaves with a longer leaf base of the longest side of the leaf (− scores, full line in Fig. 5c) and more symmetric leaves with a less elongated leaf base at the longest side of the leaf (+ scores, dashed line in Fig. 5c). The categorical variable SSV correlated with the first two PCA axes (+ with PC1 and − with PC2) whereas the categorical variable MT correlated with the second and the third PCA axis (+ and − respectively, Table 3). This is exemplified for SSV in Fig. 6: leaves counting a high number of split secondary veins (SSV) tend to have a higher score for PC1 and a lower score for PC2 and are thus concentrated in the lower corner at the right hand side of the plot. Taking into account these allometric relationships and the fact that SSV and MT are correlated (−) (Table 3), it can be deduced that these leaves that are characterised by a higher number of split secondary veins (SSV) also tend to a lower number of subteeth per tooth at the leaf margin (MT), relative wider leaves (PC1), a wider leaf base at the longest side of the leaf together with a less acuminate tip (PC2) and more symmetrical leaves (PC3). From the raw data, it is clear that these leaves mainly belong to the Zandhoven population (Online Resource 4).
Largest variation in Ulmus laevis leaf shapes along the three first principal component axes of the shape analysis. Contours were drawn from digitised leaves close to the two extremities (positive and negative, dashed and full line, respectively) along each axis. a PCA axis 1, b PCA axis 2, c PCA axis 3
Plot of first and second axis of the shape PCA indicating leaves according to the pooled levels of the categorical variable SSV (number of split secondary veins, Table 2). Proportional explained variance PC1: 23.2 %, PC2: 19.1 %
Morphological variability on population level
A PCA analysis was performed on a dataset consisting of the first three PCA axes from the shape analysis, the two size variables LL and LW, and the six categorical variables MT, SSV, UPM, UPV, LPM and LPV (Table 2). The proportional explained variances of the first three axes were 22.1, 19.4 and 15 %. In the biplot of the first two axes, one group of leaves diverged from a central group (Fig. 7a). According to the first and third axis (Fig. 7b), two deviating groups could be noticed. A first group deviated along SSV (+) and MT (−), and to a lesser extent (shorter vectors) along PC1 (+), PC2 (−) and PC3 (+) from the shape analysis, and corresponded mainly to the Zandhoven population. The other group departed from the central cluster of leaves according to mainly LPM and LPV (both −). The leaves in this group did not substantially deviate in shape, nor in size, but in the pubescence at the lower side of the leaves. They belonged to the Halle population.
Biplots of a PCA on all morphological shape, size and categorical variables. PC1 to PC3 are the PCA scores of the shape analysis, LL and LW are lamina length and lamina width, and categorical variable abbreviations are in Table 2. Shown are biplots of the first two axes (a) and of the first and third axis (b). Proportional variance explained PC1: 22.1 %, PC2: 19.4 % and PC3: 15 %
Variance structure of ramets, genotypes and populations
An ANOVA indicated that all three levels of the hierarchical structure in the data (population, genotype and ramet) contributed significantly to the variation in the different morphological variables (Online Resource 5). This analysis included the principal component axes explaining the larger part of the variability in the dataset from the shape analysis and from the PCA including all leaf morphological variables. Because of the nested nature of the data, variance could be partitioned into a part attributable to ramets within genotypes, a part to genotypes within populations and a part to the between-population variance (Table 4), applying mixed effect modelling. Variance is attributable to population and genotype points to genetic control, whereas the variance between genetically identical individuals (variance at the ramet level) indicates plasticity (different phenotypes for the same genotype). For both size variables LL and LW and the second axis of the shape analysis (shPC2), the proportional variance explained by the clonal nature of the ramets was larger than the sum of the genotype and population variance (20 % > 5 % + 7 %, 22 % > 10 % + 8 % and 22 % > 9 % + 10 %, respectively), indicating a more plastic nature for these variables in comparison to the other axes of the shape analysis (shPC1: 11 % < 2 % + 30 % and shPC3: 11 % < 4 % + 28 %). The higher variance for population and genotype compared to ramet for the latter variables suggested a relative stronger genetic control. When considering the PCA analysis on all morphological variables, all first three axes displayed a larger proportional population plus genotype variance compared to ramet (totPC1: 48 % + 22 % > 14 %, totPC2: 16 % + 11 % > 19 % and totPC3: 33 % + 16 % > 20 %) with mainly the first and third axis showing a large difference, suggesting a relative stronger genetic control. The variance of LM and SSV could mainly be assigned to the population level (100 and 86 % respectively) indicating a strong genetic control with little variation between genotypes within populations and large differences between populations. For the pubescence variables LPM and LPV the higher variance for population and genotype compared to ramet suggested again a relative stronger genetic control (56 % + 10 % > 34 % and 28 % + 25 % > 46 % respectively).
Discussion
Leaf shape variation
An in-depth investigation of leaf shape through application of geometric morphometrics as described in this paper, partitions shape variability into separate discrete and size independent shape changes along different PCA axes. One main advantage is that description of these shape changes would need more linear measurements on the leaf in comparison to the geometric morphometric approach reducing the number of variables to the first few PCA axes that explain a larger part of the variability in the dataset. Both the contour from a digitised leaf close to the zero point of the PCA axes, as the shape deviations through leaf contours at the extremities of separate PCA axes, allow picturing the average leaf shape and the deviations from it. Specifically for taxonomic treatment of highly variable species this type of imaging can be valuable. Remarkably, the variation along the first two axes of the shape analysis corresponds to the two main sources of variation in leaf shape of Ulmus minor, a related elm species, described by Melville already in 1946 (Melville 1946). In this early study, the main source of variation is attributed to the width of the elm leaf when taking the length of the leaf constant, which matches with our first PCA axis of the shape analysis. The second source of variation is credited to the width of the base of the leaf only at the side of the leaf that is longest, while keeping the other side of the leaf constant. This corresponds, at least partly, with the second axis of our shape analysis. In addition, Melville (1946) suggests that inheritance of the width of the leaf base at the longest side of the leaf in U. minor occurs independent of the leaf width, which is again in agreement with the fact that the PCA axes in our shape analysis are by definition independent of each other.
Genetic control of leaf morphological variables
The hierarchical structure of the sampled U. laevis trees in the field trial allowed assessment of the relative weight of genetic control of the separate morphological variables through a variance analysis. Morphological studies generally use samples from natural populations without a biological hierarchical structure. Biological hierarchically structured data as can be obtained from greenhouse or field trials (this study) or by assignment of parent trees to recruits through parentage analysis, allows discrimination between genetic and non-genetic components of variation (Lopez de Heredia et al. 2009; Vander Mijnsbrugge 2015). In this way, distinction is possible between more genetic and more plastic variables (Leimar 2009). Especially in taxonomy, systematics and evolutionary studies the relative magnitude of genetic control versus phenotypic plasticity of leaf morphological traits in trees can be of significance (Bresson et al. 2011; Thomas 2011). In our study, the variance structure of the first and third PCA axis of the shape analysis suggests that the shape variations along the first (relative width of the leaf) and third axis (leaf asymmetry and length of the leaf base at the longest side of the leaf) are under a relative stronger genetic control (relative larger contributions from population and genotype compared to ramet), thus more specialist, compared to the shape variation expressed by the second axis (width of the leaf base at the longest side of the leaf and acumination of leaf tip) and to the size variables, which are all relatively more plastic in nature (a comparably larger part of the variance is assigned to ramet than to genotype and population). The variance among the natural populations in our study contributes substantially to the total variance of the first and third axis of the overall PCA, likely attributable mainly to a few categorical variables explaining larger parts of the axes’ variabilities. When examining the partitioning of the variance per categorical variable, the among-population variance constitutes the total and the largest part of the variance of margin toothing and split secondary veins respectively, while it explains more than half of the proportional variance structure of the leaf mesophyll pubescence at the lower side of the leaf (Table 4). As these categorical variables are among the total set of variables used in the PCA on all morphological variables, it can be deduced that their strong among-population variance contributes substantially to the relative large among-population variance of the first and third axis of the PCA on all morphological variables. The residual variance that cannot be explained by the hierarchical structure (population, genotype and ramet) is relatively high in the second axis of the PCA on all morphological variables compared to the first and third axis, indicating more random variation.
Deviating populations
Incorporation of shape variables, the most explanatory PCA axes from the shape analysis, size variables, directly calculated from the landmarks, and the observed categorical variables in an overall PCA analysis renders two deviating groups of leaves, both attributable to two natural populations Zandhoven and Halle. From the orientation and the length of the vectors in the biplot of the first and third axis, it can be deduced that mainly leaf margin toothing, split secondary veins and pubescence at the lower side of the leaves separate the two groups. These two obviously deviating populations, Zandhoven and Halle, are both relatively isolated relict populations with strongly diminished number of trees. It is known that decreases in effective population sizes due to habitat fragmentation have strong negative effects on within-population genetic diversity in outcrossing species and that the fewer individuals remaining after fragmentation, the more severe the genetic bottleneck (Honnay and Jacquemyn 2007; Aguilar et al. 2008). This isolation and diminution may already exist for several generations, as Flanders is characterised by massive riparian habitat destruction, by a dynamic land-use history involving successive phases of deforestation and reforestation (Vandekerkhove et al. 2011; De Keersmaeker et al. 2013), and by a lack of U. laevis in reforestation programs in the past as deduced from its absence in (historical) plantation forests. Additionally, Flanders is a part of the Lower Countries characterised by little variation in topography or climate conditions (homogeneous habitat) minimising the probability of natural selection on a small geographical scale as a cause of the observed morphological deviations, notwithstanding the fact that natural selection is known to determine to a large extent the population genetic structure for quantitative traits in plants (Merila and Crnokrak 2001). For instance, although variation in pubescence can be related to water deficiency (Johnson 1975), negligible differences in precipitation regimes among the origins of the sampled populations cannot explain the deviation of the Halle population for this character. This all suggests that the morphological deviation of the two relict populations is caused “randomly”, putatively triggered by the phenomenon of genetic drift that has been observed for U. laevis (Vakkari et al. 2009), rather than by local adaptation, natural selection or epigenetic polymorphism (that would result from environmental heterogeneity). Hybridisation with taxa of the U. minor–U. glabra complex is not expected as U. laevis is reported not to hybridise (Mittempergher and Laporta 1991). The larger part of the other populations in our study is also strongly reduced in numbers of individuals (Table 1) but do not display a clear morphological deviation, indicating again that the divergence of Halle and Zandhoven is likely random in nature. The detected morphological deviation is mainly expressed in the categorical variables SSV and MT for Zandhoven and LPV and LPM for Halle. Several discontinuous morphological phenotypes within single plant species have been attributed to one or two genes with major effects (Gottlieb 1984). Especially SSV and MT, and to a lesser extent LPM are characterised by a larger proportional summed variance component attributable to genotype and population, compared to the variance accredited to ramets, suggesting a more specialist genetic base for the morphological deviations. Ecotypes have been defined as phenotypes arising from a genotypical response of a population to a particular habitat (Turesson 1922) and are interpreted as resulting from habitat heterogeneity (variable climatic and edaphic conditions) combined with natural selection (Hufford and Mazer 2003). Considering habitat fragmentation and subsequent isolation of populations together with reduced population sizes as a particular habitat, ecotype evolution in the isolated and greatly reduced U. laevis populations in Flanders can be hypothesised. This evolution occurred on a micro-geographical scale and putatively quick in evolutionary terms as the detected morphological deviations are likely driven “randomly” rather than by natural selection, and for a larger part expressed through specialist rather than plastic characters.
Conclusion
We applied geometric morphometrics to address leaf shape variation and combined this study with continuous size and categorical morphological variables. The hierarchical structure of the sampled Ulmus laevis trees allowed partitioning of variance in the dataset using mixed modelling methods. This enabled us to distinguish between more genetic specialist and more generalist plastic variables. The deviating nature of the two isolated and greatly reduced U. laevis populations prompted us to hypothesise “random phenotypic drift” which is expressed in more specialist morphological characters.
References
Aguilar R, Ashworth L, Galetto L, Aizen MA (2006) Plant reproductive susceptibility to habitat fragmentation: review and synthesis through a meta-analysis. Ecol Letters 9:968–980
Aguilar R, Quesada M, Ashworth L, Herrerias-Diego Y, Lobo J (2008) Genetic consequences of habitat fragmentation in plant populations: susceptible signals in plant traits and methodological approaches. Molec Ecol 17:5177–5188
Bates D, Maechler M, Bolker B, Walker S (2014) lme4: linear mixed-effects models using Eigen and S4. R package version 1.1–7. Available at: http://CRAN.R-project.org/package=lme4. Accessed 1 Nov 2014
Bo WH, Wang Z, Xu F, Fu GF, Sui YH, Wu WM, Zhu XL, Yin DN, Yan Q, Wu RL (2014) Shape mapping: genetic mapping meets geometric morphometrics. Brief Bioinform 15:571–581
Bresson CC, Vitasse Y, Kremer A, Delzon S (2011) To what extent is altitudinal variation of functional traits driven by genetic adaptation in European oak and beech? Tree Physiol 31:1164–1174
Christensen RHB (2013) Ordinal: regression models for ordinal data. R package version 2013.10–31. Available at: http://CRAN.R-project.org/package=ordinal. Accessed 1 Nov 2014
De Keersmaeker L, Rogiers N, Vandekerkhove K, De Vos B, Roelandt B, Cornelis J, De Schrijver A, Onkelinx T, Thomaes A, Hermy M, Verheyen K (2013) Application of the ancient forest concept to potential natural vegetation mapping in flanders, a strongly altered landscape in Northern Belgium. Folia Geobot 48:137–162
Debinski DM, Holt RD (2000) A survey and overview of habitat fragmentation experiments. Conservation Biol 14:342–355
Fu G, Bo W, Pang X, Wang Z, Chen L, Song Y, Zhang Z, Li J, Wu R (2013) Mapping shape quantitative trait loci using a radius-centroid-contour model. Heredity 110:511–519
Fuentes-Utrilla P, Venturas M, Hollingsworth PM, Squirrell J, Collada C, Stone GN, Gil L (2014) Extending glacial refugia for a European tree: genetic markers show that Iberian populations of white elm are native relicts and not introductions. Heredity 112:105–113
Goodall-Copestake WP, Hollingsworth ML, Hollingsworth PM, Jenkins GI, Collin E (2005) Molecular markers and ex situ conservation of the European elms (Ulmus spp.). Biol Conservation 122:537–546
Gottlieb LD (1984) Genetics and morphological evolution in plants. Amer Naturalist 123:681–709
Hemery GE, Clark JR, Aldinger E, Claessens H, Malvolti ME, O’Connor E, Raftoyannis Y, Savill PS, Brus R (2010) Growing scattered broadleaved tree species in Europe in a changing climate: a review of risks and opportunities. Forestry 83:65–81
Honnay O, Jacquemyn H (2007) Susceptibility of common and rare plant species to the genetic consequences of habitat fragmentation. Conservation Biol 21:823–831
Hufford KM, Mazer SJ (2003) Plant ecotypes: genetic differentiation in the age of ecological restoration. Trends Ecol Evol 18:147–155
Jensen RJ, Hokanson SC, Isebrands JG, Hancock JF (1993) Morphometric variation in oaks of the Apostle Islands in Wisconsin—evidence of hybridization between Quercus rubra and Q. ellipsoidalis (Fagaceae). Amer J Bot 80:1358–1366
Jensen RJ, Ciofani KM, Miramontes LC (2002) Lines, outlines, and landmarks: morphometric analyses of leaves of Acer rubrum, Acer saccharinum (Aceraceae) and their hybrid. Taxon 51:475–492
Johnson HB (1975) Plant pubescence: an ecological perspective. Bot Rev 41:233–258
Kissling P (1977) Les poils des quatre espèces de chênes du Jura (Quercus pubescens, Q. petraea, Q. robur et Q. cerris). Ber Schweiz Bot Ges 87:1–18
Lambinon J, de Langhe J, Delvosalle L, Duvigneaud J (2012) Flora van België, het Groothertogdom Luxemburg, Noord-Frankrijk en de aangrenzende gebieden. National Botanic Garden of Belgium, Meise
Leimar O (2009) Environmental and genetic cues in the evolution of phenotypic polymorphism. Evol Ecol 23:125–135
Leimu R, Vergeer P, Angeloni F, Ouborg NJ (2010) Habitat fragmentation, climate change, and inbreeding in plants. Ann New York Acad Sci 1195:84–98
Lopez de Heredia L, Valbuena-Carabaña M, Cordoba M, Gil L (2009) Variation components in leaf morphology of recruits of two hybridising oaks [Q. petraea (Matt.) Liebl. and Q. pyrenaica Willd.] at small spatial scale. Eur J Forest Res 128:543–554
Maes N (2006) Inheemse bomen en struiken in Nederland en Vlaanderen. Boom, Amsterdam
Melville R (1946) Typification of variation in the smooth-leaved elm, Ulmus carpinifolia Gleditsch. J Linn Soc Bot 53:83–90
Melville R (1978) On the discrimination of species in hybrid swarms with special reference to Ulmus and the nomenclature of U. minor Mill. and U. carpinifolia Gled. Taxon 27:345–351
Merila J, Crnokrak P (2001) Comparison of genetic differentiation at marker loci and quantitative traits. J Evol Biol 14:892–903
Mittempergher L, Laporta N (1991) Hybridization studies in the Eurasian species of elm (Ulmus spp.). Silvae Genet 40:237–243
Nielsen LR, Kjaer ED (2010) Fine-scale gene flow and genetic structure in a relic Ulmus laevis population at its northern range. Tree Genet Genomes 6:643–649
Penaloza-Ramirez JM, Gonzalez-Rodriguez A, Mendoza-Cuenca L, Caron H, Kremer A, Oyama K (2010) Interspecific gene flow in a multispecies oak hybrid zone in the Sierra Tarahumara of Mexico. Ann Bot (Oxford) 105:389–399
Petrokas R, Baliuckas V (2012) Natural Hybridisation within Elms (Ulmus L.) in Lithuania. Baltic Forest 18:237–246
Punyasena SW, Smith SY (2014) Bioinformatic and biometric methods in plant morphology. Appl Pl Sci. doi:10.3732/apps.1400071
R Core Team (2013) R: A language and environment for statistical computing. R Foundation for Statistical Computing. Vienna, Austria. Available at: http://www.R-project.org/. Accessed 1 Nov 2014
Richens RH (1955) Studies on Ulmus 1. The range of variation of East Anglian elms. Watsonia 3:138–153
Rohlf J (2010) tpsDig v 2.16 (computer program) New York: Stony Brook University
Sheets D (2004) CoordGen6f and PCAGen6n (computer programs). Canisius college, New York
Sacchetti P, Tiberi R, Mittempergher L (1990) Preference of Scolytus multistriatus (Marsham) during the gonad maturation phase between two species of elm. Redia 73:347–354
Slice D (2013) Morpheus (computer program). Department of Scientific Computing, The Florida state University, Florida
Solla A, Bohnens J, Collin E, Diamandis S, Franke A, Gil L, Buron M, Santini A, Mittempergher L, Pinon J, Broeck AV (2005) Screening European elms for resistance to Ophiostoma novo-ulmi. Forest Sci 51:134–141
Thomas SC (2011) Genetic vs. phenotypic responses of trees to altitude. Tree Physiol 31:1161–1163
Turesson G (1922) The species and the variety as ecological units. Hereditas 3:100–113
Vakkari P, Rusanen M, Karkkainen K (2009) High genetic differentiation in marginal populations of European White elm (Ulmus laevis). Silva Fenn 43:185–196
Vandekerkhove K, De Keersmaeker L, Walleyn R, Kohler F, Crevecoeur L, Govaere L, Thomaes A, Verheyen K (2011) Reappearance of Old-growth elements in lowland woodlands in Northern Belgium: do the associated species follow? Silva Fenn 45:909–935
Vander Mijnsbrugge K (2015) Morphological dissection of leaf, bud and infructescence traits of the interfertile native A. glutinosa and non-native A. incana in Flanders (northern part of Belgium). Trees. doi:10.1007/s00468-015-1247-7
Vander Mijnsbrugge K, Vanden Broeck A, Van Slycken J (2005) A survey of Ulmus laevis in Flanders (Northern Belgium). Belg J Bot 138:199–204
Venturas M, Fuentes-Utrilla P, Ennos R, Collada C, Gil L (2013) Human-induced changes on fine-scale genetic structure in Ulmus laevis Pallas wetland forests at its SW distribution limit. Pl Ecol 214:317–327
Venturas M, Nanos N, Gil L (2014) The reproductive ecology of Ulmus laevis Pallas in a transformed habitat. Forest Ecol Managem 312:170–178
Viscosi V, Cardini A (2011) Leaf morphology, taxonomy and geometric morphometrics: a simplified protocol for beginners. PLoS ONE. doi:10.1371/journal.pone.0025630
Viscosi V, Fortini P, Slice DE, Loy A, Blasi C (2009a) Geometric morphometric analyses of leaf variation in four oak species of the subgenus Quercus (Fagaceae). Pl Biosyst 143:575–587
Viscosi V, Lepais O, Gerber S, Fortini P (2009b) Leaf morphological analyses in four European oak species (Quercus) and their hybrids: a comparison of traditional and geometric morphometric methods. Pl Biosyst 143:564–574
Viscosi V, Antonecchia G, Lepais O, Fortini P, Gerber S, Loy A (2012) Leaf shape and size differentiation in white oaks: assessment of allometric relationships among three sympatric species and their hybrids. Int J Pl Sci 173:875–884
Vit P, Lepsi M, Lepsi P (2012) There is no diploid apomict among Czech Sorbus species: a biosystematic revision of S. eximia and discovery of S. barrandienica. Preslia 84:71–96
Vranckx G, Jacquemyn H, Muys B, Honnay O (2012) Meta-analysis of susceptibility of woody plants to loss of genetic diversity through habitat fragmentation. Conservation Biol 26:228–237
Webber J (2000) Insect vector behavior and the evolution of Dutch elm disease. In: Dunn CP (ed) The elms: breeding, conservation and disease management. Kluwer Academic Publishers, Boston, pp 47–60
Zuur A, Ieno E, Walker N, Saveliev A, Smith G (2009) Mixed effects models and extensions in ecology with R. Springer, New York
Acknowledgments
We thank Stefaan Moreels for the vegetative propagation of the plants, André Meersman for preparation of the herbarium and Marc Spanhove for maintenance of the field trial.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Handling editor: Jürg Schönenberger.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Vander Mijnsbrugge, K., Le Clercq, R. & Michiels, B. Dissection of leaf morphological traits from isolated and declined relict populations of Ulmus laevis reveals putative random ecotype evolution. Plant Syst Evol 302, 219–229 (2016). https://doi.org/10.1007/s00606-015-1255-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00606-015-1255-5