Abstract
A colorimetric sensor array designed on a paper substrate with a microfluidic structure has been developed. This array is capable of detecting COVID-19 disease by tracking metabolites of urine samples. In order to determine minor metabolic changes, various colorimetric receptors consisting of gold and silver nanoparticles, metalloporphyrins, metal ion complexes, and pH-sensitive indicators are used in the array structure. By injecting a small volume of the urine sample, the color pattern of the sensor changes after 7 min, which can be observed visually. The color changes of the receptors (recorded by a scanner) are subsequently calculated by image analysis software and displayed as a color difference map. This study has been performed on 130 volunteers, including 60 patients infected by COVID-19, 55 healthy controls, and 15 cured individuals. The resulting array provides a fingerprint response for each category due to the differences in the metabolic profile of the urine sample. The principal component analysis-discriminant analysis confirms that the assay sensitivity to the correctly detected patient, healthy, and cured participants is equal to 73.3%, 74.5%, and 66.6%, respectively. Apart from COVID-19, other diseases such as chronic kidney disease, liver disorder, and diabetes may be detectable by the proposed sensor. However, this performance of the sensor must be tested in the studies with a larger sample size. These results show the possible feasibility of the sensor as a suitable alternative to costly and time-consuming standard methods for rapid detection and control of viral and bacterial infectious diseases and metabolic disorders.
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
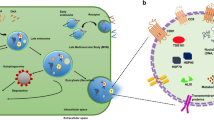
The uncontrollable and rapid spread of infectious diseases may involve a high percentage of the world’s population, which is known as a pandemic [1]. Examples of common types of viral diseases include the cold, influenza, HIV, HPV, Ebola, and COVID-19 [2]. The last one has emerged since 2019, attacking the respiratory tract, resulting in lung disorders and has infected or killed millions of people [3, 4].
The virus infection can be detected by conventional methods with virus cell culture [5]. Although these methods provide accurate and reliable results, their corresponding analyses are time-consuming, and need to propagate virus particles [6]. The nucleic acid-based detection methods such as the real-time quantitative polymerase chain reaction (RT-qPCR), loop-mediated isothermal amplification (LAMP), recombinase polymerase amplification (RPA), helicase-dependent amplification (HDA), and rolling circle amplification (RCA) are capable of directly and sensitively detecting the virus genetic material in a shorter period of time [7]. The diagnostic tests are conducted by trained operators in a laboratory equipped with sophisticated devices. However, the nucleic acid-based methods may provide false-negative results due to an improper sampling process, sample contamination, and changes in genome sequencing [8]. Antibody-based biosensors can recognize infection by trapping proteins associated with viruses [9]. These sensors can be fabricated based on lateral-flow or enzyme-linked immunosorbent assays [10]. Compared to previous methods, biosensors are more inexpensive and have a shorter detection time. Moreover, performing the analysis process does not require special skills. Nevertheless, the efficiency of biosensors is limited due to the high cost of antibodies, and their special storage conditions [11].
Since the immune system of body responds to viral pathogens by releasing, eliminating, or altering the concentration of chemical species (known as metabolites) in its biological fluids, the diagnosis of a disease can be realized by examining the metabolic profiles [12]. In this regard, previous studies have shown that the spread of viruses such as COVID-19 can affect the concentration of chemical markers in exhaled breath [13], serum [14], plasma [15], saliva [16], and urine samples [17]. The human urine contains more than 3000 types of metabolites, and is one of the most widely applied and available biological fluids for the diagnosis of various diseases, including urinary tract infections, kidney, gastrointestinal, liver, and cancer diseases, and metabolic disorders such as diabetes [18]. The healthy and coronavirus-infected urine samples have 1033 different chemical species from each other [17]. These compounds fall into the categories of amines, esters, carbohydrates, phenols, alcohols, and sulfur-containing substances [12]. In addition, the infected urine is more alkaline and has a lower specific gravity [17].
On the other hand, chromatographic methods such as gas and liquid chromatography are combined with mass spectrometry (MS), FT-IR, and nuclear magnetic resonance spectrometers provide unique results for minor changes in the type and concentration of components of urine samples [19,20,21]. However, the extraction of information is costly and time-consuming, and requires special laboratory conditions and skilled manpower.
The miniature sensing devices based on non-bioreceptors are able to qualitatively and quantitatively analyze one or more chemical markers in a urine sample [22]. While numerous chemical compounds, including organic dyes, inorganic complexes, and nanostructures, can be candidates in the fabrication of receptors, the selective identification of analytes depends on the specific structural and optical properties of the receptors [23]. These can participate in electrostatic, nucleophilic, and H-bonding interactions, or respond to changes in the pH of the reaction medium [24, 25]. It is possible to fabricate sensing devices in the form of an array structure, thus increasing the sensor’s ability to selectively detect chemical species with similar structures and determine them at low concentrations [26].
The sensor arrays can be assembled on a paper substrate in order to save cost and consumption of reactants, inject low sample volumes, reduce analysis time, and help run tests at the sampling site [27]. Based on previous reports, we can use the paper-based arrays for monitoring biogenic amines [28], glucose [29], tuberculosis [30], and urinary tract infections [31] in urine samples. However, the performance of these diagnostic tools has not been evaluated for detecting the viral infections by tracking the metabolic changes of urine.
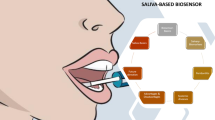
The potential of sensor arrays embedded on a paper substrate was previously assessed for non-invasive detection of COVID-19 disease using saliva [32] and exhaled breath [33] metabolites by our group. Here, a study is presented to introduce a sensor array-based microfluidic structure with 16 detection zones filled by color compounds, including gold and silver metal nanoparticles, porphyrin dyes, metal ion complexes, and organic dyes modified by a carbonyl species-sensitive reagent. The sensor is expected to have a high efficiency to react with the metabolites after receiving a low volume of the urine sample, find metabolic changes between infected and healthy samples in the shortest time, and be able to recognize the cured person after the treatment process. If promising results are obtained, this sensor can be employed as a rapid and cost-effective alternative to conventional viral infection detection methods.
Experimental section
Instruments and software
The microfluidic pattern was drawn on the paper substrate by HP LaserJet printer 1200. The receptors were poured on the detection zones using micropipette (BRAND Transferpette® S, Germany).The pH of metal ion complex solution was regulated by Metrohm 632 pH-meter (Model 780 pH lab). A canon scanner (CanoScan LiDE 220) was applied to receive the response of sensor. The design of microfluidic pattern was conducted by AutoCAD 2016. The changes in the color of receptors were verified by ImageJ (1.51n, National Institutes of Health, USA). The pattern recognition analysis was performed in the MATLAB R2015 environment. SPSS (Version 22; Chicago, IL, USA) was used to process the statistical analysis.
Chemicals
Silver nitrate (AgNO3), gold (III) chloride trihydrate (HAuCl4.3H2O), iron(III) nitrate nonahydrate)Fe(NO3)3.9H2O(, copper(II) nitrate trihydrate (Cu(NO3)2.3H2O), bromocresol purple (R1), ethanol (EtOH), boric acid (H3BO3), sodium hydroxide (NaOH), sulfuric acid (H2SO4) (18.4 mol.L−1), and sodium borohydride (NaBH4) were bought from Merck chemical company. Tyrosine (Tyr), tannic acid (TA), bromophenol red (R2), acridine orange (R3), malachite green (R4), phenol red (R5), pararosaniline hydrochloride (R6), pyrocatechol violet (Py), vanadyl sulfate pentahydrate (VOSO4.5H2O), and 2,4-dinitrophenylhydrazine were obtained from Sigma Aldrich. Note that these materials were pure and used in the analytical grade. The other materials such as meso-tetrakis(4-chlorophenyl) porphyrin-manganese (III) acetate (Mn(III)T(4-Cl)PP(OAc)), meso-tetraphenylporphyrin]Iron(III) chloride (Fe(III)TPPCl), and meso-tetrakis(4-hydroxyphenyl) porphyrin-manganese (III) acetate (Mn(III)T(4-OH)PP(OAc)) were provided from Shargi research group, Shiraz, Iran. The preparation methods for synthesis of porphyrin dyes were found in the previous reports [34,35,36]. The microfluidic sensor was fabricated on the Whatman® Grade NO.2 filter paper.
Experimental procedure for synthesis of nanoparticles
Two types of metallic nanoparticles containing gold and silver nanoparticles (AuNPs and AgNPs) were prepared by following procedures:
To synthesize the AuNPs capped by Tyr, 100 mL of a mixture included 1.0 × 10−4 mol.L−1 tyrosine and 1.0 × 10−3 mol.L−1 potassium hydroxide was poured in a volumetric flask. This solution was heated to boil under stirring and reflux condition subsequently mixed by 1.0 × 10−3 mol.L−1 HAuCl4 drop by drop. A 12 kDa dialysis membrane was used to discriminate the synthesized NPs from unreacted initial materials. The same procedure was used to prepare Tyr-modified AgNPs but the solution of AgNO3 (1.0 × 10−4 mol.L−1) was used instead of HAuCl4 in the synthesis procedure [37].
TA functionalized AuNPs was provided by addition of 2.0 mL of 6.0 × 10−3 mol.L−1 tannic acid solution to 50.0 mL of 1.3 × 10−4 mol.L−1 HAuCl4 boiling solution (pH 6.0) under stirring condition. The solution was allowed to cool and kept on the stirrer for 10 min [38]. To have AgNPs with the same capping agent, a solution containing 5.0 mL of 3.0 × 10−3 mol.L−1 of AgNO3 and 2.0 mL of 6.0 × 10−2 mol.L−1 tannic acid was stirred at room temperature for 12 h to appear the yellow color of the solution. Note that the pH of the tannic acid solution used in this study was equal to 8.0 [39].
All prepared NPs were dried by freeze drier and convert to fine powder using a laboratory mortar.
The solution of receptors
A specified amount of NPs were re-dispersed in 1.0 mL deionized water, while a certain powder of porphyrin and organic dyes (R1 to R6) was dissolved in EtOH. The organic dyes was added to DWES solution which was created by mixing 3.0 mL of deionized water with the solution of 0.4 g 2, 4-dinitrophenylhydrazine, and 10.0 mL of EtOH followed by addition of 2.0 mL of H2SO4 [39]. For formation of the metal ion complexes, 50.0 μL of the V (IV), Fe (III), and Cu (II) ion solutions was separately added to a mixing solution of 100.0 μL of borate buffer (0.1 mol.L−1) and 50 μL of pyrocatechol violet (Py), subsequently the reaction was continued on the stirrer for 10 min [40].
Selection of the studied population
One hundred fifteen males with the age range of 21 to 80 years volunteered to participate in this study [32, 33]. Of these, 60 participants were categorized into patients, and the rest were considered healthy controls. The healthy persons who had not been infected by COVID-19 before the experiment, were invited. The patients were selected from the people hospitalized at the emergency department of Baqiyatallah Hospital between the years 2020 and 2021 [32, 33]. They had the symptoms of the disease, and had not received medical care prior to this study. The positive results of chest imaging and rRT-PCR tests of these patients were approved by a pulmonologist. After 2 months of the disease treatment process, the urine samples of 15 cured participants were analyzed in a separate experiment [32, 33]. Demographic data and other medical information of the studied population have reported in our previous studies [32, 33]
Paper-based microfluidic pattern
The pattern shown in Fig. 1a was plotted by AutoCAD software, consisting of three parts: a large central circle for the sample injection, and 16 small circles to immobilize the receptors, and the channels that connect the two parts. The microfluidic pattern was printed on the substrate, followed by keeping it in an oven at 200 °C. The paper was heated for 45 min in order to allow the ink (black areas) to penetrate into the paper texture, thereby providing a hydrophobic space [41]. To form the colorimetric sensor array (Fig. 1b), 0.2 μL of the receptors was then added to the detection zones based on the order shown in Fig. 1c. Then, the sensor was transferred to a desiccator and the sensing elements were allowed to dry completely in 1 h at room temperature. In this condition, the sensor was stable for 20 days before usage (Fig. S1). Note that the sensor was used without any further pre-preparation in the detection step.
Detection steps
The microfluidic device was kept between the holders which are hand-made and fabricated by poly (methyl methacrylate). Fifty microliters of the urine sample was injected into the central circle through a hole embedded in the upper layer of the holder. Note that the turbidity of the urine sample did not impress on the response of sensor; therefore, the sample was not diluted or centrifuged before usage. The capillary nature of the paper substrate facilitated the flow of the sample through the channel, and its transport to the detection areas (small circles) in order to react with the chemical receptors. Then, the sensor was dried at ambient temperature and its images were recorded by a scanner. The color changes of the receptors before and after the injection of the urine sample were compared using ImageJ software. For each receptor, three mean values were obtained corresponding to three color elements (red, green, and blue). Moreover, the software provided a dataset with 48 numeric values (16 receptors × 3 color elements) for each sample studied (Scheme 1).
Data collection steps
A matrix with dimensions of (130 × 48) were created and inserted as input into the principal component analysis-discriminant analysis (PCA-DA) algorithm, thereby evaluating the potential of the proposed device to differentiate between the members of patient, healthy, and cure categories, respectively.
The response of the sensor array, known as the Euclidean norm, was then quantified by Eq. (1):
where \({x}_{i}\) is the numerical value of the ith component of the data vector for each urine sample. Finally, the average results obtained for healthy and patient categories were compared using two independent sample t-test in SPSS environment.
In the optimization test, the selection of optimal values was performed by calculating the discrimination ability function (DAF) parameter, followed by determining the ratio of between-class variations to within-class variations [42]. The numerical value of this parameter can be acquired through Eq. (2):
where N denotes the total number of categories, \({Y}_{ij}\) is the Euclidean norm value for the jth sample of ith category, \(\overline{Y }\) is the average of the measurements of each category, and \(\overline{\overline{Y}}\) is the average of the total data obtained for the two categories (patients and healthy ones).
Results and discussion
According to previous studies, changes in the type and amount of metabolites in urine samples can be a characteristic of a clinical failure in the body that can be assessed by instrumental methods and sensory diagnostic devices [12, 17, 30, 31]. Here, a microfluidic-based sensor array is proposed in order to investigate the effect of COVID-19 infection on the chemical species released in the urine sample. The advantage of this sensor is using the sensing potential of a wide range of receptors in its structure. For example, gold and silver nanoparticles (AuNPs and AgNPs) whose surface was modified by tyrosine and tannic acid coating agents formed four receptors. The reactivity of these sensing elements depends on the inherent properties of the metal core, and the chemical structure, size, and electrical charge of the coating chemicals [43, 44]. Tannic acid is a large polyphenol, and tyrosine is an amino acid containing amino, carboxylic, and hydroxyl functional groups [45]. Compared to NPs synthesized with tannic acid, tyrosine-NPs are smaller in size and have a negative electric charge distribution on their surface [45]. All of these factors cause NPs to react differently with various chemical species through electrostatic, H-bonding, and nucleophilic interactions [43]. However, these interactions can be inhibited by the size of NPs due to steric hindrance. Among the other sensing elements, three receptors were made of metalloporphyrins, each of which was composed of a specific type of central metal or functional group attached to a macro-heterocyclic ring that can have Lewis acid or base properties [46]. The sensing ability of complexes of the Py dye and metal ions such as V (IV), Fe (III), and Cu (II) was used for identifying the urinary metabolites. These metal ions tend to interact with compounds containing amino and carboxylic groups [40, 47]. The formation of a new ion-analyte complex leads to the release of ligands and discoloration of the receptor. This reaction is known as the indicator displacement assay [48]. The last category of receptors was organic dyes combined with DWES. The additive detects and interacts with the compounds consisting of the carbonyl group, changing pH of the reaction media [49]. The organic dye then monitors the changes in the concentration of hydronium ions. The fabricated array is shown in Fig. 1.
The optimal conditions
To increase the sensor performance in finding the difference between the metabolites of the urine samples of patients with COVID-19 and healthy participants, the effective parameters in the fabrication of the sensing elements such as the concentration of color materials or the volume ratio of the additive to the organic dye as well as the response time of sensor should be optimized. In this respect, the optimization test was run with five healthy and five infected samples for three times. For each sample, the Euclidean norm value was calculated. The optimal value for each parameter was obtained using the DAF equation (see “Data collection steps” section).
Initially, four different sets of concentrations were designed for each receptor (Fig. 2a). The lowest and highest concentrations were observed in the first and last sets, respectively. The results obtained from DAF (Fig. 2b) indicated that the best response of sensor was achieved using receptors made with the concentrations provided in the second set. The sensing potential of the receptors was reduced due to the lack of active sites to interact with the analyte at the low concentrations. Furthermore, the changes taking place due to the interaction between the receptor and the analyte were not well observed. This is because the intensity of the initial color of the receptors at higher concentrations led to a decrease in the DAF value [50].
The results for optimization: a The designing sets for the concentration of sensing elements and b the respective DAF responses. c The designing sets for the volume ratio of the organic dyes and the additive in the mixing solution and d the corresponding DAF responses. e The DAF values for different incubation times
In the following procedure, five individual volume ratios were employed for the combination of the additive and the organic dye (Fig. 2c). These ratios should be adjusted in such a way that the additive can receive the aldehyde and ketone species with high accuracy, while also avoiding a reduction in the efficiency of the organic dye when detecting hydronium ions. Figure 2d reveals that the best discrimination was achieved using a mixture of the organic dye and additive (4:1) in the fabrication of the receptor. Finally, the discriminatory ability of the proposed sensor was evaluated for healthy and infected samples at different times. As seen in Fig. 2e, a significant difference between the studied groups was obtained after 7 min. Therefore, the color changes of the receptors were recorded at this time.
Colorimetric patterns
The sensor was fabricated under the optimal conditions, and exposed to 50.0 μL of the urine sample belonging to two groups of healthy and patient volunteers. The color changes of each receptor after the interaction with urinary metabolites are shown in Fig. 3a. It was observed that metal ion complexes (S8-S10) and malachite green combined with DWES (S14) did not help differentiate between the two studied groups because they responded to the metabolites of both infected and healthy urine samples. In addition, the color variations were not recorded for Ty-AuNPs (S1), TA-AgNPs (S4), Mn (III) T (4-Cl) PP (OAC) (S5), and DWES-acridine orange. Of the receptors, a nanoparticle Ty-AgNPs (S2), a metalloporphyrin Mn (III) T (4-OH) PP (OAC) (S7), and an organic dye (DWES-pararosaniline) (S16) interacted with the metabolites of healthy samples, whereas the compounds such as TA-AuNPs (S3), Fe (III) TPPCl (S6), and DWES mixed by the organic dyes including bromophenol red (S11), bromocresol purple (S12), and phenol red (S15) were sensitive to the chemical species of the patient samples.
a The responses and b the difference patterns of developed sensor for patient infected by COVID-19 (P) and healthy control (H), cured volunteers (C), and the participants having viral infection and the other disease consist of diabetes (DM), chronic kidney (KD), and liver disorder (LD). The data was collected in the optimum conditions (Fig. 2) after 7 min. The difference patterns help to find a better description for color changes of the sensor related to each sample
In a separate experiment, a color pattern of the sensor was prepared for the urine sample of the cured group in order to evaluate the disease treatment process (Fig. 3a). Cured people are participants who have been treated for 2 months. This test determines the effectiveness of the treatment process, the consumption of drugs, and their dosage. Also, the changes in the color of sensor show that the type and the concentration of the metabolites related to patient gradually turn towards the metabolic profile of the healthy person. As can be inferred, there was a significant difference between the responses obtained for this group and those for the healthy and patient ones. It was clear that the metabolite profile of the patient sample gradually changed to that of the healthy sample. These changes were reflected in the array results by turning off the response of receptors S3, S6, S11, and S15 (relating to the patient sample) or turning on the sensing element S7 (corresponding to the healthy sample). However, the receptor S12 (for the patient sample) responded to the metabolites of the cured sample, and the sensors S2 and S16 (for the healthy sample) were reluctant to interact with these chemical species, indicating that the body was still affected by the viral infection.
Figure 3b represents the color difference obtained from the image analysis process as a color difference map. These maps not only confirmed the previous information but also provided the user with enhanced color changes of the receptor. Notably, the color changes of S7 and S11 were difficult to be detected with the naked eye, whereas the color of these sensing elements was well displayed in the respective maps. In this case, 130 urine samples were examined, and the colorimetric responses and corresponding color difference maps collected.
For each studied sample, the Euclidean norm values of receptors are presented in Fig. S2. Among the receptors responding to all samples, while S8 and S14 were more inclined to interact with metabolites of the healthy sample, S9 and S10 tended more to capture the compounds of the patient sample. As expected, the response intensity of the receptor S12 decreased for the cured sample in the wake of the progression of the treatment. Of course, an upward trend was observed in the result of receptor S7 under this condition.
Discrimination results
The responses of the proposed array to chemical species of urine samples were gathered in the form of data vectors. In this case, 60, 55, and 15 data vectors were obtained for the patient, healthy, and cured participants, respectively. These vectors were arranged in a data matrix with a size of 130 × 48. The data matrix was investigated by the PCA-DA algorithm to verify the sensor performance, creating a difference between the three studied categories. The distribution of data in the space of the first two principal components is depicted in Fig. 4. The score plot showed that PC 1 and PC 2 contained 80.2% and 14.6% of the explained variance, respectively. According to this diagram, the three categories were well separated. Table 1 provides the statistical parameters for this classification analysis. As can be seen, 73.3% (44 participants) of patients, 74.5% (41 participants) of healthy, and 66.6% (10 participants) of cured individuals were correctly diagnosed by the proposed method. However, the sensor response for 35 studied samples differed from the demographic information. The assay accuracy (with a 95% confidence interval level) was calculated to be 73.9%, 72.0%, and 72.8% for the distinction between the patient-healthy, patient-cured, and healthy-cured categories, respectively.
The discrimination of 130 studied population (60 patients (P), 55 healthy (H), and 15 cured (C)) by PCA-DA algorithm. The data was collected in the optimum conditions (Fig. 2) after 7 min
Sensor responses to comorbidities
Based on the filled questionnaires, some participants may have other diseases such as cardiovascular, chronic kidney, diabetic, chronic liver, and hypertension diseases as well as non-COVID-19 pulmonary disorders. By monitoring the colorimetric pattern of 115 patient and healthy samples, it was clear that the response of some receptors changed in the presence of the urine sample infected by a specific disease (Fig. 3). For example, Ty-AuNPs (S1) responded to urinary chemicals in the participants with kidney failure, whereas diabetic urine metabolites tended to interact with TA-AgNPs (S4). Moreover, acridine orange discoloration indicated liver disorders. The proposed method can identify the individuals suffered from kidney, liver, and diabetic diseases with 80.0%, 83.3%, and 70.0% accuracy, respectively. As illustrated in Fig. 3, the color changes of receptors S3, S6, S11, S12, and S15 repeated in the color pattern of the patients infected only by COVID-19, and that of the participants with both viral diseases and the other disorders. In turn, this evidenced that the chemical species related to the other diseases did not affect the response of the sensor to urinary metabolites of COVID-19.
Statistical point
The Euclidean norm values of the urine samples of 115 healthy and patient participants were calculated. The total average of these data was equal to 388.98 (± 28.3). For each of patient and healthy groups, the mean values of the determinations were found to be 379.90 (± 25.4) and 398.89 (± 28.2). As inferred, the healthy group had a higher (19 units) mean value than the patient group. This difference was statistically large and significant (P-value < 0.001). Accordingly, for an unknown sample, the Euclidean norms being higher and lower than the total average can be attributed to the healthy and patient groups, respectively.
In the next experiment, the dependency of the total sensor response on the age of population was statistically investigated for both patient and healthy categories. For this study, the corresponding Pearson coefficients were calculated as 0.249 (P-value = 0.075) and 0.230 (P-value = 0.092) indicating a weak and meaningless relationship between these two studied variables (total response and age). Therefore, the color changes of sensor were not affected by the age of population.
Evaluation of the reproducibility of sensor responses
To have a reliable result on the separation between the studied samples, the sensor should provide reproducible responses for the infected and healthy urine samples. This experiment was performed by adding 50.0 μL of a urine sample to five individual sensor arrays. The color changes of the sensor were calculated for each analysis, and their numerical value was determined by the Euclidean norm equation (Fig. S3). After five measurements, a relative standard deviation (RSD %) of 3.1 was obtained for the patient sample. Meanwhile, an error of 3.4 was observed for the healthy sample. This low error value indicated the high reproducibility of the sensor responses.
In contrast with the rapid detection kits fabricating biological receptors, colorimetric sensor arrays use chemical compounds as sensing elements which are easily provided and printed on a substrate without any pre-preparation steps. They do not require special temperature conditions for storage and their final cost is less than one dollar, which is far more affordable than standard methods such as chest imaging, PCR test, and diagnosis kit based on immunoassay. However, the use of medications related to other diseases such as diabetes, hypertension, or bacterial infections, the selective-cross reactive response of sensing elements to analytes, and the lower number of them in the array structure limit the achievement of an acceptable sensitivity/specificity for discriminating between patient and healthy groups. A reason is the inability of the sensor to detect the trace changes in the type and concentration of chemical markers (related to a certain disease) in a complex mixture such as urine containing 3000 metabolites. This problem can be overcome by choosing a larger number of sensing elements that show a better selectivity response to a specific set of metabolites.
Conclusions
The feasibility of the colorimetric sensor array was investigated for detecting the changes in the chemical species concentration of the urine sample of COVID-19 patients. By combining four different groups of receptors (having different optical and structural properties), it was possible to provide reliable responses for the discrimination of the patient and healthy participants. The statistical parameters confirmed that the assay was more sensitive than the standard methods. The array was able to track the treatment process of the disease, while also creating a unique colorimetric pattern for the cured individuals. The specific receptors of the sensor can interact with metabolites of kidney disease, diabetes, and liver failure, providing detection of these comorbidities. However, the performance of sensor must be tested in the studies with a larger sample size. The array was developed by designing a microfluidic structure on the paper substrate, which required a small volume of the urine sample and can be used by the public as a rapid and cost-effective diagnostic/screening tool.
References
DiMaio D, Enquist LW, Dermody TS (2020) A new coronavirus emerges, this time causing a pandemic. Annu Rev Virol 7:iii–v. https://doi.org/10.1146/annurev-vi-07-070920-100001
Rai KR, Shrestha P, Yang B, Chen Y, Liu S, Maarouf M and Chen J-L (2021) Acute infection of viral pathogens and their innate immune escape. Front Microbiol.12:672026. https://doi.org/10.3389/fmicb.2021.672026
Giovannini G, Haick H, Garoli D (2021) Detecting COVID-19 from breath: a game changer for a big challenge. ACS Sensors 6:1408–1417. https://doi.org/10.1021/acssensors.1c00312
World Health Organization (WHO) (20201) WHO coronavirus (COVID-19) dashboard. In: World Heal. Organ. https://covid19.who.int/table
Hematian A, Sadeghifard N, Mohebi R et al (2016) Traditional and modern cell culture in virus diagnosis. Osong Public Heal Res Perspect 7:77–82. https://doi.org/10.1016/j.phrp.2015.11.011
Cassedy A, Parle-McDermott A, O’Kennedy R (2021)Virus detection: a review of the current and emerging molecular and immunological methods. Front Mol Biosci 8:637559. https://doi.org/10.3389/fmolb.2021.637559
Maiti B, Anupama KP, Rai P et al (2021) Isothermal amplification-based assays for rapid and sensitive detection of severe acute respiratory syndrome coronavirus 2: Opportunities and recent developments. Rev Med Virol. https://doi.org/10.1002/rmv.2274
Jansson L, Hedman J (2019) Challenging the proposed causes of the PCR plateau phase. Biomol Detect Quantif 17:100082. https://doi.org/10.1016/j.bdq.2019.100082
Singh B, Datta B, Ashish A, Dutta G (2021) A comprehensive review on current COVID-19 detection methods: from lab care to point of care diagnosis. Sensors Int 2:100119. https://doi.org/10.1016/j.sintl.2021.100119
Mohit E, Rostami Z, Vahidi H (2021) A comparative review of immunoassays for COVID-19 detection. Expert Rev Clin Immunol 17:573–599. https://doi.org/10.1080/1744666X.2021.1908886
Sheini A (2020) Colorimetric aggregation assay based on array of gold and silver nanoparticles for simultaneous analysis of aflatoxins, ochratoxin and zearalenone by using chemometric analysis and paper based analytical devices. Microchim Acta 187:167. https://doi.org/10.1007/s00604-020-4147-5
Bouatra S, Aziat F, Fau - Mandal R, Mandal R Fau - Guo AC et al (2013) The human urine metabolome. PLoS ONE 8:e73076
Grassin-Delyle S, Roquencourt C, Moine P et al (2021) Metabolomics of exhaled breath in critically ill COVID-19 patients: a pilot study. EBioMedicine 63:103154. https://doi.org/10.1016/j.ebiom.2020.103154
Shen B, Yi X, Sun Y et al (2020) Proteomic and metabolomic characterization of COVID-19 patient sera. Cell 182:59-72.e15. https://doi.org/10.1016/j.cell.2020.05.032
Zheng H, Jin S, Li T et al (2021) Metabolomics reveals sex-specific metabolic shifts and predicts the duration from positive to negative in non-severe COVID-19 patients during recovery process. Comput Struct Biotechnol J 19:1863–1873. https://doi.org/10.1016/j.csbj.2021.03.039
Frampas CF, Longman K, Spick MP et al (2021) Untargeted saliva metabolomics reveals COVID-19 severity. Preprint-medRxiv, ID: ppmedrxiv-21260080. https://doi.org/10.1101/2021.07.06.21260080
Bi X, Liu W, Ding X et al (2022) Proteomic and metabolomic profiling of urine uncovers immune responses in patients with COVID-19. Cell Rep 38:110271. https://doi.org/10.1016/j.celrep.2021.110271
Sarosiek I, Schicho R, Blandon P, Bashashati M (2016) Urinary metabolites as noninvasive biomarkers of gastrointestinal diseases: a clinical review. World J Gastrointest Oncol 8:3674–3685. https://doi.org/10.4251/wjgo.v8.i5.459
Struck-Lewicka W, Kordalewska M, Bujak R et al (2015) Urine metabolic fingerprinting using LC-MS and GC-MS reveals metabolite changes in prostate cancer: a pilot study. J Pharm Biomed Anal 111:351–361. https://doi.org/10.1016/j.jpba.2014.12.026
Sarigul N, Kurultak İ, Uslu Gökceoğlu A, Korkmaz F (2021) Urine analysis using FTIR spectroscopy: a study on healthy adults and children. J Biophotonics 14. https://doi.org/10.1002/jbio.202100009
Kim ER, Kwon HN, Nam H et al (2019) Urine-NMR metabolomics for screening of advanced colorectal adenoma and early stage colorectal cancer. Sci Rep 9:4786. https://doi.org/10.1038/s41598-019-41216-y
Tai WC, Chang YC, Chou D, Fu LM (2021) Lab-on-paper devices for diagnosis of human diseases using urine samples—a review. Biosensors 11:260. https://doi.org/10.3390/bios11080260
Diehl KL, Anslyn EV (2013) Array sensing using optical methods for detection of chemical and biological hazards. Chem Soc Rev 42:8596–8611. https://doi.org/10.1039/c3cs60136f
Bordbar MM, Hemmateenejad B, Tashkhourian J, Nami-Ana SF (2018) An optoelectronic tongue based on an array of gold and silver nanoparticles for analysis of natural, synthetic and biological antioxidants. Microchim Acta 185:493. https://doi.org/10.1007/s00604-018-3021-1
Askim JR, Mahmoudi M, Suslick KS (2013) Optical sensor arrays for chemical sensing: the optoelectronic nose. Chem Soc Rev 42:8649–8682. https://doi.org/10.1039/c3cs60179j
Bordbar MM, Tashkhourian J, Hemmateenejad B (2019) Structural elucidation and ultrasensitive analyses of volatile organic compounds by paper-based nano-optoelectronic noses. ACS Sensors 4:1442–1451. https://doi.org/10.1021/acssensors.9b00680
Bordbar MM, Sheini A, Hashemi P et al (2021) Disposable paper-based biosensors for the point-of-care detection of hazardous contaminations—a review. Biosensors 11:316. https://doi.org/10.3390/bios11090316
Shojaeifard Z, Bordbar MM, Aseman MD et al (2021) Collaboration of cyclometalated platinum complexes and metallic nanoclusters for rapid discrimination and detection of biogenic amines through a fluorometric paper-based sensor array. Sensors Actuators, B Chem 334:129582. https://doi.org/10.1016/j.snb.2021.129582
Mohammadifar M, Choi S (2018) A paper-based enzymatic sensor array for visual detection of glucose levels in urine. 2018 IEEE 12th International Conference on Nano/Molecular Medicine and Engineering (NANOMED) 244–247 . https://doi.org/10.1109/NANOMED.2018.8641594
Sandlund J, Lim S, Queralto N et al (2018) Development of colorimetric sensor array for diagnosis of tuberculosis through detection of urinary volatile organic compounds. Diagn Microbiol Infect Dis 92:299–304. https://doi.org/10.1016/j.diagmicrobio.2018.06.014
Bordbar MM, Tashkhourian J, Tavassoli A et al (2020) Ultrafast detection of infectious bacteria using optoelectronic nose based on metallic nanoparticles. Sensors Actuators, B Chem 319:128262. https://doi.org/10.1016/j.snb.2020.128262
Bordbar MM, Samadinia H, Sheini A et al (2022) A colorimetric electronic tongue for point-of-care detection of COVID-19 using salivary metabolites. Talanta 246:123537. https://doi.org/10.1016/j.talanta.2022.123537
Bordbar MM, Samadinia H, Sheini A et al (2022) Mask assistance to colorimetric sniffers for detection of COVID-19 diseases using exhaled breath metabolites. Sensors Actuators, B Chem 369:132379. https://doi.org/10.1016/j.snb.2022.132379
Sharghi H, Nejad AH (2004) Novel synthesis of meso-tetraarylporphyrins using CF3SO2Cl under aerobic oxidation. Tetrahedron 60:1863–1868. https://doi.org/10.1002/chin.200424113
Mansuy D (1993) Activation of alkanes: the biomimetic approach. Coord Chem Rev 125:129–141. https://doi.org/10.1016/0010-8545(93)85013-T
Bernadou J, Meunier B, Fabiano AS, Robert A (1994) “Redox Tautomerism” in high-valent metal-oxo-aquo complexes. Origin of the oxygen atom in epoxidation reactions catalyzed by water-soluble metalloporphyrins. J Am Chem Soc 116:9375–9376. https://doi.org/10.1021/ja00099a083
Dubey K, Anand BG, Badhwar R et al (2015) Tyrosine- and tryptophan-coated gold nanoparticles inhibit amyloid aggregation of insulin. Amino Acids 47:2551–2560. https://doi.org/10.1007/s00726-015-2046-6
Aswathy Aromal S, Philip D (2012) Facile one-pot synthesis of gold nanoparticles using tannic acid and its application in catalysis. Physica E 44:1692–1696. https://doi.org/10.1016/j.physe.2012.04.022
Alam MF, Laskar AA, Ahmed S et al (2017) Colorimetric method for the detection of melamine using in-situ formed silver nanoparticles via tannic acid. Spectrochim Acta - Part A Mol Biomol Spectrosc 183:17–22. https://doi.org/10.1016/j.saa.2017.04.021
Sheini A, Khajehsharifi H, Shahbazy M, Kompany-Zareh M (2017) A chemosensor array for the colorimetric identification of some carboxylic acids in human urine samples. Sensors Actuators, B Chem 242:288–298. https://doi.org/10.1016/j.snb.2016.11.008
Sheini A, Aseman MD, Bordbar MM (2021) Origami paper analytical assay based on metal complex sensor for rapid determination of blood cyanide concentration in fire survivors. Sci Rep 11:3521. https://doi.org/10.1038/s41598-021-83186-0
Brereton RG (2003) Chemometrics: data analysis for the laboratory and chemical plant. Wiley, New York
Bordbar MM, Nguyen TA, Arduini F, Bagheri H (2020) A paper-based colorimetric sensor array for discrimination and simultaneous determination of organophosphate and carbamate pesticides in tap water, apple juice, and rice. Microchim Acta 187:621. https://doi.org/10.1007/s00604-020-04596-x
Bordbar MM, Tashkhourian J, Hemmateenejad B (2022) Paper-based optical nose made with bimetallic nanoparticles for monitoring ignitable liquids in gasoline. ACS Appl Mater Interfaces 14:8333–8342. https://doi.org/10.1021/acsami.1c24194
Bordbar MM, Nguyen TA, Tran AQ, Bagheri H (2020) Optoelectronic nose based on an origami paper sensor for selective detection of pesticide aerosols. Sci Rep 10:17302. https://doi.org/10.1038/s41598-020-74509-8
Suslick K, Hulkower K, Sen A, Sroka M, McNamara W (2005) Method and apparatus for detecting ammonia from exhaled breath. Patent NO. US20050171449A1, United States
Wang Y, Huo D, Wu H et al (2019) A visual sensor array based on an indicator displacement assay for the detection of carboxylic acids. Microchim Acta 186:496. https://doi.org/10.1007/s00604-019-3601-8
Khajehsharifi H, Bordbar MM (2015) A highly selective chemosensor for detection and determination of cyanide by using an indicator displacement assay and PC-ANN and its logic gate behavior. Sensors Actuators, B Chem 209:1015–1022. https://doi.org/10.1016/j.snb.2014.10.053
Li J, Hou C, Huo D et al (2014) Development of a colorimetric sensor Array for the discrimination of aldehydes. Sensors Actuators, B Chem 196:10–17. https://doi.org/10.1016/j.snb.2014.01.054
Mirzaei Y, Gholami A, Bordbar MM (2021) A distance-based paper sensor for rapid detection of blood lactate concentration using gold nanoparticles synthesized by Satureja hortensis. Sensors Actuators, B Chem 345:130445. https://doi.org/10.1016/j.snb.2021.130445
Acknowledgements
The authors gratefully acknowledge the support from the Research Councils of Baqiyatallah University of Medical Sciences. Also, the authors would like to thank the Clinical Research Development Unit of Baqiyatallah Hospital, for all their supports during carrying out this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethics approval
The research ethics committee of Baqiyatallah University of Medical Sciences has approved the project (Approval ID: IR.BMSU.REC.1399.508).
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bordbar, M.M., Samadinia, H., Sheini, A. et al. Non-invasive detection of COVID-19 using a microfluidic-based colorimetric sensor array sensitive to urinary metabolites. Microchim Acta 189, 316 (2022). https://doi.org/10.1007/s00604-022-05423-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-022-05423-1