Abstract
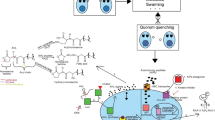
A graphene oxide (GO)–based fluorescent bioassay was developed to quantify agrA gene transcription (its mRNA) in methicillin-resistant Staphylococcus aureus (MRSA). This method is based on the use of Klenow fragment (KF)–assisted target recycling amplification and hybridization chain reaction (HCR). A triple complex was designed that contained a capture probe (CP), a trigger probe (TP), and a help probe (HP), which were partially complementary to one another. In the absence of the target, all the oligonucleotides labeled with carboxyfluorescein (FAM) are adsorbed onto the surface of GO by π-stacking interactions. This adsorption quenches the FAM signal. On the contrary, the target RNA causes the triple complex to disintegrate and initiates strand-displacement polymerization reaction (SDPR) and HCR in the presence of the appropriate raw materials, including the primer, KF, dNTPs, hairpin 1 (H1), and hairpin 2 (H2), generating double-stranded DNA (dsDNA) products. These dsDNA products are repelled by GO and produce strong fluorescence, measured at excitation/emission wavelengths of 480/514 nm. The fluorescent signal is greatly amplified by SYBR Green I (SGI) due to the synergistic effect of dsDNA-SGI. The target was assayed with this method at concentrations in the range 10 fM to 100 pM, and the detection limit (LOD) was 10 fM. This method also displayed good applicability in the analysis of real samples. It provides a new way of monitoring biofilm formation and studying the mechanisms of drug actions.

Schematic representation of the graphene oxide–based fluorescent bioassay for agrA gene transcription in methicillin-resistant Staphylococcus aureus by using strand-displacement polymerization recycling and hybridization chain reaction.





Similar content being viewed by others
References
Sowash MG, Uhlemann AC (2014) Community-associated methicillin-resistant Staphylococcus aureus case studies. Methods Mol Biol 1085:25–69
Petinaki E, Spiliopoulou I (2015) Methicillin-resistant Staphylococcus aureus colonization and infection risks from companion animals: current perspectives. Vet Med 6:373–382
Vaishampayan A, Jong AD, Wight DJ, Kok J, Grohmann E (2018) A novel antimicrobial coating represses biofilm and virulence-related genes in methicillin-resistant Staphylococcus aureus. Front Microbiol 9:221
LaSarre B, Federle MJ (2013) Exploiting quorum sensing to confuse bacterial pathogens. Microbiol Mol Biol Rev 77(1):73–111
Li YH, Tian XL (2012) Quorum sensing and bacterial social interactions in biofilms. Sensors 12(3):2519–2538
Gray B, Hall P, Gresham H (2013) Targeting agr- and agr-like quorum sensing systems for development of common therapeutics to treat multiple gram-positive bacterial infections. Sensors 13(4):5130–5166
Tan L, Li SR, Jiang B, Hu XM, Li S (2018) Therapeutic targeting of the Staphylococcus aureus accessory gene regulator (agr) system. Front Microbiol 9:55
Zia MF, Flynt AS (2017) Detection and verification of mammalian mirtrons by northern blotting. Methods Mol Biol 1823:209–219
Wen SX, Chen XL, Xu FZ, Sun HL (2016) Validation of reference genes for real-time quantitative PCR (qPCR) analysis of Avibacterium paragallinarum. PLoS One 11(12):e0167736
Winn ME, Zapala MA, Hovatta I, Risbrough VB, Lillie E, Schork NJ (2010) The effects of globin on microarray-based gene expression analysis of mouse blood. Mamm Genome 21(5–6):268–275
Ning Y, Zou L, Gao Q, Hu J, Lu FG (2018) Graphene oxide-based fluorometric determination of methicillin-resistant Staphylococcus aureus by using target-triggered chain reaction and deoxyribonuclease-assisted recycling. Microchim Acta 185(3):183
Xiong HT, Zheng XW (2017) Electrochemiluminescence based determination of micro-RNA using target-guided assembly of gold nanoparticles on an electrode modified with Nafion, carbon nanotubes and polyvinylpyrrolidone. Microchim Acta 184(6):1781–1789
Zhang DC, Yan YR, Cheng W, Zhang W, Li YH, Ju HX, Ding SJ (2013) Streptavidin-enhanced surface plasmon resonance biosensor for highly sensitive and specific detection of microRNA. Microchim Acta 180(5–6):397–403
He QZ, Luo HQ, Chen LL, Dong J, Chen KK, Ning Y (2020) Nanographite-based fluorescent biosensor for detecting microRNA using duplex-specific nuclease-assisted recycling. Luminescence 35(3):347–354
Song Y, Li WK, Duan YF, Deng L (2014) Nicking enzyme-assisted biosensor for Salmonella enteritidis detection based on fluorescence resonance energy transfer. Biosens Bioelectron 55:400–404
Ning Y, Gao Q, Zhang XQ, Wei K, Chen LL (2016) A Graphene oxide–based sensing platform for the determination of methicillin-resistant Staphylococcus aureus based on strand-displacement polymerization recycling and synchronous fluorescent signal amplification. J Biomol Screen 21(8):851–857
Liu K, Yan X, Mao B, Wang S, Deng L (2016) Aptamer-based detection of salmonella enteritidis using double signal amplification by klenow fragment and dual fluorescence. Microchim Acta 183(2):643–649
Liao DL, Jiao HP, Wang B, Lin Q, Yu C (2012) KF polymerase-based fluorescence aptasensor for the label-free adenosine detection. Analyst 137(4):978–982
Ling M, Peng ZH, Cheng LJ, Deng L (2015) Rapid fluorescent detection of enterotoxigenic Escherichia coli (ETEC) K88 based on graphene oxide-dependent nanoquencher and Klenow fragment-triggered target cyclic amplification. Appl Spectrosc 69(10):1175–1181
Li RM, Zou L, Luo YW, Zhang MJ, Ling LS (2017) Ultrasensitive colorimetric detection of circulating tumor DNA using hybridization chain reaction and the pivot of triplex DNA. Sci Rep 7:44212
Wang YH, Jiang L, Leng QG, Wu YH, He XX, Wang KM (2016) Electrochemical sensor for glutathione detection based on mercury ion triggered hybridization chain reaction signal amplification. Biosens Bioelectron 77:914–920
Li ZB, Miao XM, Zhu AH, Ling LS (2015) Hybridization chain reaction and gold nanoparticles dual signal amplification for sensitive glucose detection. Biochem Eng J 103:205–210
Wang XZ, Jiang AW, Hou T, Li HY, Li F (2015) Enzyme-free and label-free fluorescence aptasensing strategy for highly sensitive detection of protein based on target-triggered hybridization chain reaction amplification. Biosens Bioelectron 70:324–329
Ning Y, Duan YF, Feng YY, Deng L (2014) Label-free fluorescent aptasensor based on a Graphene oxide self-assembled probe for the determination of adenosine triphosphate. Anal Lett 47:2350–2360
Ning Y, Wei K, Cheng LJ, Hu J, Xiang Q (2017) Fluorometric aptamer based determination of adenosine triphosphate based on deoxyribonuclease I-aided target recycling and signal amplification using graphene oxide as a quencher. Microchim Acta 184(6):1847–1854
Joo M, Baek SH, Cheon SA, Chun HS, Choi SW, Park TJ (2017) Development of aflatoxin B 1 aptasensor based on wide-range fluorescence detection using graphene oxide quencher. Colloid Surface B 154:27–32
Wu W, Fang ZY, Zhao SM, Lu XW, Yu LX, Mei T, Zeng LW (2014) A simple aptamer biosensor for Salmonellae enteritidis based on fluorescence-switch signaling graphene oxide. RSC Adv 4(42):22009–22012
Jiang HY, Li FR, Li W, Lu XD, Ling K (2018) Multiplexed determination of intracellular messenger RNA by using a graphene oxide nanoprobe modified with target-recognizing fluorescent oligonucleotides. Microchim Acta 185(12):552
Zhang H, Liu Y, Fu X, Yuan LH, Zhu ZJ (2015) Microfluidic bead-based assay for microRNAs using quantum dots as labels and enzymatic amplification. Microchim Acta 182(3–4):661–669
Li ZJ, Guo H, Xu FY, Tang W, Duan XR (2019) Sensitive monitoring of RNA transcription by optical amplification of cationic conjugated polymers. Talanta 203:314–321
He CM, Chen SY, Zhao JJ, Tian JN, Zhao SL (2019) Ultrasensitive detection of microRNA-21 based on electrophoresis assisted cascade chemiluminescence signal amplification for the identification of cancer cells. Talanta 209:120505
Yu W, Li JQ, Zuo C, Tai YY, Bai SL, Li JL, Zhang Z, Xie GM (2019) Specific discrimination and universal signal amplification for RNA detection by coupling toehold exchange with RCA through nucleolytic conversion of a structure-switched hairpin probe. Anal Chim Acta 1068:96–103
Xu FZ, Luo L, Shi H, He XX, Lei YL, Tang JL, He DG, Qian ZZ, Wang KM (2018) Label-free and sensitive microRNA detection based on a target recycling amplification-integrated superlong poly(thymine)-hosted copper nanoparticle strategy. Anal Chim Acta 1010:54–61
Bai WQ, Cui AP, Liu MZ, Qiao XZ, Li Y, Wang T (2019) Signal-off electrogenerated chemiluminescence biosensing platform based on the quenching effect between ferrocene and Ru (bpy)32+-functionalized metal-organic frameworks for the detection of methylated RNA. Anal Chem 91(18):11840–11847
Chen YX, Zhang WJ, Huang KJ, Zheng MB, Mao YC (2017) An electrochemical microRNA sensing platform based on tungsten diselenide nanosheets and competitive RNA–RNA hybridization. Analyst 142(24):4843–4851
Acknowledgments
We would like to thank Janine Miller, PhD, from Liwen Bianji, Edanz Editing China (www.liwenbianji.cn/ac), for editing the English text of a draft of this manuscript.
Funding
We would like to thank National Key R&D Program of China (2018YFC1603300), National Natural Science Foundation (81803964, 81774126), China Postdoctoral Science Foundation (2018M630906, 2019T120707), Doctor start-up Foundation of Hunan University of Chinese Medicine (9982-1001019), Key Subjects of Hunan University of Chinese Medicine《pathogenic biology》(No. 1) and Basic Medicine Construction Project of Hunan University of Traditional Chinese Medicine (No. 6) for the financial support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOC 29350 kb)
Rights and permissions
About this article
Cite this article
Ning, Y., Chen, S., Hu, J. et al. Fluorometric determination of agrA gene transcription in methicillin-resistant Staphylococcus aureus with a graphene oxide–based assay using strand-displacement polymerization recycling and hybridization chain reaction. Microchim Acta 187, 372 (2020). https://doi.org/10.1007/s00604-020-04347-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-020-04347-y