Abstract
Aedes albopictus is a highly invasive mosquito species that has become widespread across the globe. In addition, it is an efficient vector of numerous pathogens of medical and veterinary importance, including dengue, chikungunya and Zika viruses. Among others, the vector potential of mosquitoes is influenced by their microbiome. However, this influence is very dynamic and can vary between individuals and life stages. To obtain a rough overview on the microbiome of Ae. albopictus populations in Germany, pooled female and pooled male individuals from seven German locations were investigated by total RNA sequencing. The mosquito specimens had been collected as larvae in the field and processed immediately after adult emergence, i.e. without females having fed on blood. RNA fragments with high degrees of identity to a large number of viruses and microorganisms were identified, including, for example, Wolbachia pipientis and Acinetobacter baumannii, with differences between male and female mosquitoes. Knowledge about the natural occurrence of microorganisms in mosquitoes may be translated into new approaches to vector control, for example W. pipientis can be exploited to manipulate mosquito reproduction and vector competence. The study results show how diverse the microbiome of Ae. albopictus can be, and the more so needs to be adequately analysed and interpreted.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Aedes albopictus is a thermophilic mosquito species native to the Asian-Pacific region. Due to globalisation and its high ecological and physiological plasticity, it has become widespread in many regions in the world. Presently, Ae. albopictus is considered the most invasive mosquito species of the world (Benedict et al. 2007; Bonizzoni et al. 2013). Climate warming and the resulting mild winters favour the establishment, reproduction and spread of Ae. albopictus in temperate climes, such as Central Europe (e.g., Walther et al. 2017; Fălcuţă et al. 2020).
Aedes albopictus is highly vector-competent for numerous arboviruses, including dengue, chikungunya, yellow fever, Zika, West Nile and various encephalitis viruses (Paupy et al. 2009; Martinet et al. 2019). It thus has a major impact on human and veterinary health. The vector competence, i.e. the ability of a haematophagous arthropod to transmit a pathogen, can be influenced by the arthropod’s microbiome (Engel and Moran 2013; Jupatanakul et al. 2014) which is defined by Berg et al. (2020) as ‘a characteristic microbial community occupying a reasonable well-defined habitat which has distinct physio-chemical properties’.
It has been shown that the microbiome may have a general impact on the development, reproduction and physiology of an invertebrate (Minard et al. 2013; Coon et al. 2014, 2016, 2017). For example, the endosymbiont Wolbachia pipientis is known to be widely distributed in invertebrates (Yang 2000; Hilgenboecker et al. 2008). Bourtzis and O’Neill (1998) and Ahmad et al. (2017) have demonstrated that W. pipientis can affect both the reproduction of insects and the replication and dissemination of pathogenic viruses in an insect vector. These effects are major reasons why the study of the microbiome of mosquitoes has become so popular in recent years. However, the microbiome is not static, but may change during development and can be influenced by many factors such as sex, age and life stage of the host, geographic location, breeding habitat characteristics and food supply (Wang et al. 2011; Zouache et al. 2011; Boissière et al. 2012; Terenius et al. 2012; Jupatanakul et al. 2014; Chen et al. 2020). Water temperature and nutrient content of the breeding habitat, for example, can strongly influence its bacterial community and thus have an impact on the microbiome of developing mosquito larvae (Hörtnagl et al. 2010; Onchuru et al. 2016). In turn, the microbiome ingested from the breeding habitat may considerably influence larval growth and development (Coon et al. 2014, 2016, 2017).
Insects take up a variety of microorganisms from their environment (Strand 2018). In the case of mosquitoes, this mainly occurs in the larval stage, when individuals are confronted with large numbers of microorganisms in their aquatic habitats during feeding. Larval nutrition can therefore have a major impact on the composition of the microbiome (Wang et al. 2011; Boissiere et al. 2012; Coon et al. 2016). By contrast, occasions to take up microorganisms in the adult stage are limited: both sexes feed on sugary plant juices and only females feed on blood, with the latter occasionally facilitating the uptake of disease agents. There is evidence that the insect host can exert some control over its microbiome via the innate immune response (Douglas 2015; Smith et al. 2015).
In recent years, the microbiome of several mosquito species has been studied, among them Ae. albopictus, Aedes japonicus, Anopheles gambiae and Culex pipiens, with the focus of most studies being on the midgut microbiota of adult mosquitoes (Wang et al. 2011; Gimonneau et al. 2014). It turned out that the microbiome of some species is extremely diverse, and a variety of bacterial phyla such as Actinobacteria, Proteobacteria, Bacteroidetes or Firmicutes could be detected (Moro et al 2013; Zotzmann et al. 2017; Wang et al. 2018).
The extent to which the microbiome of a species differs between populations and individuals is largely unexplored. Furthermore, nothing is known about microorganisms naturally occurring in Ae. albopictus in Germany, the influence they have on their host and whether they pose a threat to humans or may be exploited to their benefit. This study presents the first preliminary insights into the microbial RNA metagenome of Ae. albopictus from Germany which can be considered to represent the mosquito’s microbiome. When interpreting the results, however, it is essential to keep in mind that RNA reads similar to a certain microbial species vary considerably in number, and contigs generated from them are of various lengths and have various degrees of probability to be identical to a certain microbial species.
Materials and methods
Mosquito origin
The Ae. albopictus specimens investigated in this study had been collected as larvae in the field at seven sites in Germany in 2020 (Fig. 1): Mengen, Freiburg-Waldsee, Freiburg-Zähringen, Kernen, Munich, Fürth and Jena. These were the locations successfully checked for the presence of Ae. albopictus aquatic stages from all German cities known to possess established populations at the time of the study. Individuals were obtained by sieving potential breeding containers in cemeteries (Mengen, Freiburg-Waldsee, Freiburg-Zähringen, Munich, Jena), in gardens of a settlement (Kernen) and an allotment garden complex (Fürth).
Collected larvae were kept in water from their breeding habitat until adult emergence while being fed ad libitum with ground TabiMin fish food pellets (Tetra, Melle, Germany). Shortly after hatching, adults were killed by freezing at − 20 °C, without being offered blood or a sugar solution. They were morphologically identified on a chilling table using a stereomicroscope according to the determination key by Becker et al. (2010) and stored in 75% ethanol until further processing. Except for the Freiburg-Zähringen site, from where only one female and no male were available, two females and two males per site were analysed.
Nucleic acid extraction and sequencing
For nucleic acid extraction, the mosquitoes were removed from the ethanol and dried for about 1 min at room temperature for the alcohol to evaporate. Subsequently, 13 female and 12 male mosquitoes were pooled by sex and then completely homogenised in 500 µl serum-free ZB5d medium (FLI-intern cell culture medium = Eagle’s minimal essential medium with Earle’s and Hank’s salts plus non-essential amino acids) containing 5 µl of a ready-to-use mixture of penicillin–streptomycin and 1 µl of a ready-to-use mixture of gentamicin-amphotericin (Thermo Fisher Scientific, Dreieich, Germany). Three steel beads with a diameter of 3 mm (TIS GmbH, Gauting, Germany) were added and the samples agitated for 2 min at 30 Hz in a TissueLyser II (Qiagen, Hilden, Germany). Nucleic acid was then extracted from 200 µl of the supernatant using the NucleoMag VET kit (Macherey–Nagel, Düren, Germany) according to the manufacturer’s instructions, but without the addition of carrier RNA. The concentration of extracted RNA (12.2 ng/µl for the female sample, 11.3 ng/µl for the male sample) was measured using a NanoDrop Lite (Thermo Fisher Scientific).
Further processing of the sample for total RNA sequencing with Ion Torrent technology, including manual library preparation, was performed following the protocol described by Wylezich et al. (2018). Briefly, the extracted RNA was transcribed into double-stranded cDNA using the cDNA Synthesis System Kit (Roche, Mannheim, Germany), then fragmented by an M220 Focused-ultrasonicator (Covaris Ltd., Brighton, UK) and prepared for Ion Torrent-compatible library generation by means of the GeneRead L Core Kit (Qiagen) and Ion Xpress barcode adapters (Thermo Fisher Scientific). The resulting library was subjected to quality control in a 2100 Bioanalyzer (Agilent Technologies, Santa Clara, USA), using the High Sensitivity DNA Kit (Agilent Technologies) and quantification with a KAPA Library Quantification Kit (Roche). The library was then sequenced on an Ion Torrent S5 XL System (Thermo Fisher Scientific) according to the manufacturer’s instructions.
Data analysis
Sequencing results were edited and analysed using Geneious Prime version 2021.0.1 (Biomatters, Auckland, New Zealand). For this, sequences trimmed to a minimum length of 25 bp by the Geneious BBDuk tool were merged by the BBMerge tool, using a merge rate set to ‘high’. Both merged data and data that could not be merged were assembled de novo to cluster all closely related sequences into contigs, based on a ‘custom sensitivity’ setting. The obtained consensus sequences were sorted according to their length, resulting in groups of contigs with similar base pair length. Each contig of these groups was subsequently aligned with GenBank entries (www.ncbi.nlm.nih.gov), with equal results being summarised.
Results
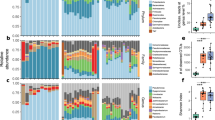
A total of 5,000,504 RNA reads were generated for the female Ae. albopictus pool and 4,653,856 reads for the male pool. In the microbial RNA metagenome of female mosquitoes, RNA fragments with high identities to 42 different microorganismal species from 37 different families were detected, whereas in males RNAs with high identities to a total of 38 different species from 36 families were found (Table 1, Supplementary Tables 1 and 2). In the pool of female mosquitoes, there were a total of 213 contigs (in the range of 69.91–100% percent identity (p.i.), 91–100% query cover and 32–1674 bp), 20 (9.35%) of which matched with eukaryotic, 136 (64.02%) with bacterial and 57 (26.63%) with viral species. In the male mosquito pool, a total of 1380 contigs (82.99–100% p.i., 96–100% query cover, 28—1702 bp) was analysed, 19 (1.87%) of which could be assigned to eukaryotes, 338 (24.29%) to bacteria and 1023 (73.83%) to viruses (Fig. 2). Often, an identification at the species level was not possible and only the genus could be determined.
Contigs with high identities to 13 species and 12 genera of viruses/microorganisms were identified in mosquitoes of both sexes (Fig. 3). Contigs arguing for seven microbial species formerly described in Ae. albopictus were found in both female and male mosquitoes. Four additional microbial species suggested by the contigs had been found in other mosquito species previously. Another two species of bacterial contigs female Ae. albopictus were suggestive of had also been found in other mosquito species, but were not present in our pool of male mosquitoes. Instead, a bacterial species indicated by contigs found in the male pool, but not in the female one of this study, had previously been detected in other mosquito species.
Table 1 shows all viruses and microorganisms whose RNA was identified in Ae. albopictus with a p.i. of at least 97% and a query cover of 100%, representing a minimum contig length of 56 bp. RNAs with less p.i. and query cover, shorter lengths or RNAs that could not be assigned to a species, but only to a genus or higher systematic level, are listed in Supplementary Tables 1 and 2. Only the first group of RNAs is discussed in the following.
RNAs with identity to Aedes albopictus anphevirus (89.23–100% p.i., 42–370 bp, 281 contigs) and Aedes phasmavirus (96.57–100% p.i., 30–350 bp, 13 contigs) were detected in both female and male Ae. albopictus (Table 1). Both viruses are considered insect-specific and had been isolated from adult Ae. albopictus before, with Aedes phasmavirus being detected in all life stages of the mosquito (Manni and Zdobnov 2020; Shi et al. 2020). In addition, both Ae. albopictus sexes harboured RNA similar to Barstukas virus (97.91–100% p.i., 83–524 bp, 5 contigs) and Guapiaçu virus (98.50–98.83% p.i., 133–170 bp, 2 contigs) which had previously been identified in adult Aedes mosquitoes (Batson et al. 2021). RNA suggesting High Island virus (97.69–98.70% p.i., 120–386 bp, 3 contigs), Usinis virus (99.74–100%, p.i., 118–390 bp, 6 contigs) and Wenzhou sobemo–like virus (96.47–100% p.i., 31–1677 bp, 750 contigs) was also found in both female and male Ae. albopictus. High Island virus had formerly been detected in adult mosquitoes of the species Psorophora ciliata (Sadeghi et al. 2017), Usinis virus in adult Ae. aegypti and Ae. albopictus (Batson et al. 2021) and Wenzhou sobemo-like virus in adult Ae. albopictus (Kubacki et al. 2020).
In addition to the RNA of these viruses, RNA with high identities to various bacteria was identified. Among others, RNA largely matching Acinetobacter baumannii (97.50–100% p.i., 40–170 bp, 3 contigs), A. johnsonii (98.76–100% p.i., 150–188 bp, 7 contigs), Elizabethkingia anophelis (99.12–100% p.i., 90–480 bp, 7 contigs), Escherichia coli (97.37–100% p.i., 53–167 bp; 14 contigs) and W. pipientis (96.91–100% p.i., 33–1052 bp, 204 contigs) was found in both female and male Ae. albopictus. Acinetobacter baumannii and A. johnsonii are human pathogens and had previously been reported from adult Ae. albopictus (Seifert et al. 1993; Minard et al. 2013), A. baumannii also from lice and fleas (Kempf et al. 2012). The ubiquitous E. coli is a potentially pathogenic (Kaper et al. 2004), widely spread intestinal bacterium of humans and other vertebrates, which had been recognised in adult Anopheles funestus and An. gambiae before (Straif et al. 1989). Elizabethkingia anophelis has emerged as a human pathogen in Africa and Asia (Lau et al. 2016) and had previously been detected in adult An. gambiae (Kämpfer et al. 2011). Wolbachia pipientis is a widely distributed essential bacterial symbiont of mosquitoes, which had frequently been described from Ae. albopictus (e.g. Wiwatanaratanabutr 2013; Park et al. 2016) and other mosquito species (e.g. Kittayapong et al. 2000).
Finally, RNA corresponding to that of Candida sake (100% p.i., 104–189 bp, 3 contigs) was detected in both Ae. albopictus females and males in this study. The fungus belongs to a genus widely distributed in arthropods but has also been extracted from the oral cavity of HIV-positive humans (Hoegl et al. 1998).
In addition to RNA fragments suggesting the same microbial species to occur in both Ae. albopictus females and males, RNA fragments referring to some bacteria were found in one mosquito sex only (Supplementary Tables 1 and 2). These include those of the bacteria A. dispersus (99.72% p.i., 422 bp, 1 contig), A. oleivorans (100% p.i., 1378 bp, 2 contigs), Chryseobacterium aureum (99.11% p.i., 676 bp, 1 contig), C. indoltheticum (100% p.i., 602–686 bp, 3 contigs), C. scophthalmum (99.79% p.i., 470 bp, 1 contig), Leclercia adecarboxylata (100% p.i., 96–546 bp, 6 contigs), Limnobacter humi (100% p.i., 106 bp, 1 contig), Serratia marcescens (100% p.i., 80 bp, 1 contig) and Zooglea resiniphila (100% p.i., 56 bp, 1 contig) in the males (Table 1). Acinetobacter dispersus can be frequently found on human skin and in water and soil (Kang et al. 2011; Nemec et al. 2016). Acinetobacter oleivorans had been detected in soil (Kang et al. 2011) and C. aureum in river water in Korea (Lee et al. 2019). Chryseobacterium indoltheticum is a widespread bacterium occurring in soil and water which may be pathogenic to humans (Calderón et al. 2011), and C. scophthalmum is a fish pathogen (Shahi et al. 2018). Leclercia adecarboxylata had previously been documented in other insects such as the potato beetle Leptinotarsa decemlineata (Muratoglu et al. 2009) and is also considered potentially pathogenic for humans (Hess et al. 2008). By contrast, L. humi had been recognised from humus soil (Nguyen and Kim 2017). Another human pathogen similar to RNA which was found in male Ae. albopictus was S. marcescens (Hejazi and Falkiner 1997). This bacterium had been detected in adult An. sinensis mosquitoes previously (Bai et al. 2019) and might become a problem in mosquito laboratory colonies (Seitz et al. 1987). Zooglea resiniphila had been found in activated sludge (Gao et al. 2018).
RNAs with similarity to some bacterial species were identified in the female Ae. albopictus of this study but not in the males (Table 1). These include Acidovorax avena (100% p.i., 121 bp, 1 contig), Acinetobacter tandoii (100% p.i., 558 bp, 1 contig), Aeromonas hydrophila (100% p.i., 33–1012 bp, 5 contigs), Arthrobacter woluwensis (100% p.i., 120–411 bp, 4 contigs), Hydrogenophaga pseudoflava (100% p.i., 176–189 bp, 2 contigs), Micrococcus luteus (100% p.i., 118–140 bp, 2 contigs), Paracoccus yeei (100% p.i., 126–168 bp, 2 contigs) and Pseudomonas luteola (100% p.i., 137–198 bp, 2 contigs). Acidovorax avenea is a plant-pathogenic bacterium (Walcott and Gitaitis 2000), whereas A. tandoii had been detected in termites (van Dexter and Boopathy 2019). Aeromonas hydrophila is a pathogen of many different vertebrates including humans (Emerson and Norris 1905; Wohlgemut et al. 1970; Agger et al. 1985), which can naturally be found in water habitats (Hazen et al. 1978). Arthrobacter woluwensis is a potential human pathogen, which can cause endocarditis, among other symptoms (Bernasconi et al. 2004; Li et al. 2021). Hydrogenophaga pseudoflava had previously been detected in the midgut of adult An. gambiae (Straif et al. 1989). RNA fragments suggesting another potential human pathogen, which had led to human meningitis in the past, is M. luteus (Fosse et al. 1985). Paracoccus yeei, on the other hand, is a human bacterial pathogen, which had formerly been isolated from the salivary glands of adult Ae. aegypti (Balaji et al. 2021) and can lead to human dialysis-related peritonitis (Arias and Clark 2019). Pseudomonas luteola is another fish pathogen, which can cause, for example, meningitis and wound infection in immunocompromised humans (Kostmann et al. 1990; Altinok et al. 2007).
In addition to RNAs with high identities to the above bacteria found in only one sex of Ae. albopictus, RNA with a high identity to the fungus Conidiobolus coronatus, which has a human-pathogenic potential (Fischer et al. 2008), could be detected in the female mosquitoes.
Discussion
The mosquitoes in this study were pooled from seven sites within Germany known to be populated by Ae. albopictus. Since the tiger mosquito is controlled in Germany by Bti (Bacillus thuringiensis israelensis) larvicide as soon as local reproduction is detected (Becker et al. 2017, 2022), the finding of larvae is difficult and was limited in the framework of this study. Due to the pooling of the collected samples, no statement can be made about the geographical origin of the microorganisms or the individual colonisation of Ae. albopictus specimens. Moreover, all viruses and microorganisms referred to in this study were identified exclusively by their RNAs and the alignment of those with sequences in the used databases. Therefore, it is not known whether exactly these species were present or other (unknown) species with closely related RNA sequences, whether they were viable viruses, living symbionts or similar and able to replicate/multiply in Ae. albopictus, and whether they were arbitrarily taken up from the environment and had no additional correlation to the mosquitoes.
As adult mosquitoes emerged from collected larvae were tested here, and no food sources whatsoever had been offered to the investigated adults, the detected RNAs, or the microbes characterised by them, must be supposed to have been transmitted transstadially from mosquito larva to pupa and through metamorphosis to adult. During metamorphosis, the midgut of mosquitoes is transformed and the digestive cells are histolised (Fernandes et al. 2014). The survival of microorganisms or the persistence of RNA, respectively, must therefore be supposed to be possible only intracellularly or with certain mechanisms of adapted symbionts. To clarify this and check for the viability of microorganisms, instead of mere RNA, cultivation attempts are necessary, but were not carried out in this study.
As we studied the microbial RNA metagenome of complete mosquitoes, it cannot be determined in which organ or tissue the found RNAs had been localised. Principally, the composition of the bacterial fauna in different organs of a mosquito can be very variable, and some bacteria colonise several organs in the mosquito at the same time (Gao et al. 2020). The tropism of microbes might give information about their migration paths in the mosquito, or about transmissibility from mosquito parent to offspring or mosquito female to blood host. For the latter, the emergence of the microorganism in the mosquito salivary glands would be requisite (Anderson et al. 2010). Whether this was the case for the potential human pathogens RNAs found were suggestive of, such as A. baumannii, E. coli or A. hydrophila, cannot be determined retrospectively. In addition to the localisation of the microorganisms in the mosquito, the pathogen load which was also not determined in this study might be decisive for a mosquito to become a vector. Thus, the mere presence of a pathogen in the mosquito might not be sufficient for transmission (Beerntsen et al. 2000).
The presence in the microbiome of certain bacteria is beneficial to the mosquito. For example, A. baumannii and A. johnsonii improve blood digestion and nectar assimilation in Ae. albopictus (Minard et al. 2013). However, the influence on mosquito development, reproduction and physiology of most microorganisms found in the microbiome is largely unknown.
The most common phyla ever found in the microbiome of adult Ae. albopictus include Proteobacteria, Bacteroidetes, Firmicutes and Actinobacteria (Mancini et al. 2018). The most common bacterial genera found in Aedes, Anopheles and Culex species are Enterobacter, Escherichia, Klebsiella, Pseudomonas and Serratia as detected in mosquitoes from the USA, England and India (Demaio et al. 1996; Touré et al. 2000; Pidiyar 2002). RNA with identities to all four bacterial phyla as well as to all five genera were found in the Ae. albopictus samples from Germany. In addition, RNA fragments indicating viruses and fungi, such as Riboviria and Ascomycota, were identified. In summary, RNAs with identities to a high number of microorganisms were detected in the German Ae. albopictus some of which represent microorganisms already described from this mosquito species previously, such as W. pipientis, A. baumannii or Usinis virus (Minard et al. 2013; Wiwatanaratanabutr 2013; Batson et al. 2021). Some other viruses or microbial species suggested by the RNA analysis in this study, such as High Island virus, Guapiaçu virus and E. anophelis, have not been detected in Ae. albopictus before, but in other mosquito species and other invertebrates (Kämpfer et al. 2011; Sadeghi et al. 2017; Batson et al. 2021; Oliveira Ribeiro et al. 2021). Furthermore, RNA fragments suggesting microorganisms previously not described from mosquitoes at all, such as L. humi, Z. resiniphila and C. aureum, were found.
It has also been shown that the midgut microbiome of adult mosquitoes may reduce a mosquito’s susceptibility to pathogens (Dong et al. 2009; Bahia et al. 2014) and have a general influence on its vector competence (Dodson et al. 2014; Jupatanakul et al. 2014). It can thus be harnessed by manipulating its microorganisms to artificially reduce vector competence. In culture, for example, E. coli was genetically modified in order to express two surface molecules that suppress the development of Plasmodium berghei. Unfortunately, E. coli had difficulties in colonising the mosquitoes and disappeared from their midgut shortly after infection (Riehle et al. 2007). In addition, there are studies that show that an infection of mosquitoes with Wolbachia leads to a strong inhibition of the development of potential pathogens. The infection with Wolbachia of the wMel strain, for example, leads to a highly reduced replication of dengue virus in Ae. aegypti. The reason for this seems to be the Wolbachia-linked upregulation of the immune system of the mosquito (Blagrove et al. 2012). Such a way of influencing vector competence is certainly also possible with the help of other organisms from the microbiome.
Since insecticides and physical measures are often inefficient tools for mosquito control (Bourtzis et al. 2016; Pang et al. 2017; Flores and O'Neill 2018), W. pipientis is also exploited for innovative biological control by manipulating mosquito reproduction. Wolbachia infection can lead to the feminisation of genetically male insects, to the killing of male siblings by females or cytoplasmic incompatibility, which ensures that females can successfully mate only with males harbouring the same Wolbachia strain (Werren et al. 2008).
Efficient control tools are particularly important in areas where mosquitoes serve as vectors of human disease agents such as Zika virus, yellow fever virus and malaria parasites. Knowledge about the natural occurrence of microorganisms in mosquitoes can therefore contribute to developing and designing new forms of vector control.
It is difficult to explain the differences in the RNA presence between female and male Ae. albopictus in this study. Individuals of both genders were collected from the same breeding sites of the seven locations, so differences between the microbial RNA cannot be attributed to developmental conditions. However, RNA with high identities to 25 species of microorganisms were found in the males but not in the females, and RNA with high identities to 29 species of microorganisms were found in the females but not in the males, with clear differences in the distribution of microorganism to kingdoms (in females, most contigs were assigned to bacteria, whereas in males most contigs were assigned to viruses). For example, RNA fragments arguing for plants were found in the pool of males, but not in the pool of females. That sex has an influence on the microbiome has been shown in previous studies (Chen et al. 2020). Also, Rani et al. (2009) detected Chryseobacterium, Pseudomonas and Serratia species only in females of Anopheles stephensi. One possible explanation for the differences might be owing to the limited number of individuals examined, with such differences becoming smaller the more individuals are studied per site. Another explanation might be that female and male larvae in fact have different food preferences and therefore take up different microorganisms with their food, but this is mere speculation and cannot be substantiated. Although it is known that larvae consume a wide range of food (Gimnig et al. 2002; Ye-Ebiyo et al. 2003) and have different feeding preferences depending on species (Merritt et al. 1992), no data exist about different feeding preferences of female and male larvae of the same species.
Especially in cases of a low percent identity and a low query coverage of the found contig sequences with sequences in GenBank, results have to be considered carefully with regard to the occurrence of the respective microorganisms in German Ae. albopictus. This applies, for example, to Kocuria rhizophila. RNA found in female Ae. albopictus matched this soil bacterium with a percent identity of 85.43% only and was registered with only one read. Due to the low percent identity, it is unlikely that the detected RNA belonged to exactly this species. Also, in the case of very short sequence lengths, the linked microorganisms must be viewed critically. This was the case for Wuchereria bancrofti-RNA where the sequence length was 45 bp only. Although percent identity and query coverage were both close to 100%, the probability of the RNA belonging to another organism, possibly a filarial species not described so far, is very high. Hitherto, W. bancrofti has not been reported from Ae. albopictus, and its geographic distribution range is restricted to subtropical and tropical regions (Service 2001).
Possible contamination must also be considered and, in fact, has already been described in similar studies. Genera such as Flavobacterium, Micrococcus, Microbacterium, Chryseobacterium, Neskia and Acidovorax have been found contaminating laboratory reagents like DNA extraction kits (Salter et al. 2014). In this study, Flavobacterium, Micrococcus, Chryseobacterium and Acidovorax could be detected in both sexes of German Ae. albopictus, Neskia only in females.
Conclusion
The microbiome of vector species such as Ae. albopictus holds much potential for the development of efficient control measures and the reduction of vector competence. However, the influence of many microorganisms on the mosquito is still largely unexplored.
Studying the composition of a mosquito’s microbiome is difficult, as it is influenced by many factors and can vary considerably both within a species and between species. It is necessary to investigate the influences of the microbiome in more detail and to examine its diversity in a large number of mosquito microbiomes.
To clarify the question of which influence the microorganisms of the microbiome have on the mosquito, it would be helpful to cultivate the microorganisms. However, cultivation is only possible with a small number of species of the microorganisms detected so far. Only with detailed knowledge about the composition and influence of the microbiome on the mosquito can these microorganisms be used for innovative approaches to vector and disease management. The study demonstrates that the microbiome of German Ae. albopictus is comprehensive and might be worth further investigations.
Data availability
Data supporting the conclusions of this article are included within the article and its supplementary tables.
References
Agger WA, McCormick JD, Gurwith MJ (1985) Clinical and microbiological features of Aeromonas hydrophila-associated diarrhea. J Clin Microbiol 21:909–913. https://doi.org/10.1128/jcm.21.6.909-913.1985
Ahmad NA, Vythilingam I, Lim YAL, Zabari NZAM, Lee HL (2017) Detection of Wolbachia in Aedes albopictus and their effects on chikungunya virus. Am J Trop Med Hyg 96:148–156. https://doi.org/10.4269/ajtmh.16-0516
Allen TD, Lawson PA, Collins MD, Falsen E, Tanner RS (2006) Cloacibacterium normanense gen. nov., sp. nov., a novel bacterium in the family Flavobacteriaceae isolated from municipal wastewater. Int J Syst Evol Microbiol 56:1311–1316. https://doi.org/10.1099/ijs.0.64218-0
Altinok I, Balta F, Capkin E, Kayis S (2007) Disease of rainbow trout caused by Pseudomonas luteola. Aquaculture 273:393–397. https://doi.org/10.1016/j.aquaculture.2007.10.025
An W, Guo F, Song Y, Gao N, Bai S, Dai J, Wei J, Zhang L, Yu D, Xia M, Yu Y, Qi M, Tian C, Chen H, Wu Z, Zhang T, Qui D (2016) Comparative genomics analyses on EPS biosynthesis genes required for floc formation of Zoogloea resiniphila and other activated sludge bacteria. Water Res 102:494–504. https://doi.org/10.1016/j.watres.2016.06.058
Anderson SL, Richards SL, Smartt CT (2010) A simple method for determining arbovirus transmission in mosquitoes. J Am Mosq Control Assoc 26:108–111. https://doi.org/10.2987/09-5935.1
Arias MA, Clark J (2019) Paracoccus yeei as a cause of peritoneal dialysis peritonitis in the United Kingdom. Idcases 15:e00486. https://doi.org/10.1016/j.idcr.2019.e00486
Bahia AC, Dong Y, Blumberg BJ, Mlambo G, Tripathi A, BenMarzouk-Hidalgo OJ, Chandra R, Dimopoulos G (2014) Exploring Anopheles gut bacteria for Plasmodium blocking activity. Environ Microbiol 16:2980–2994. https://doi.org/10.1111/1462-2920.12381
Bai L, Wang L, Vega-Rodríguez J, Wang G, Wang S (2019) A gut symbiotic bacterium Serratia marcescens renders mosquito resistance to Plasmodium infection through activation of mosquito immune responses. Front Microbiol 10:1580. https://doi.org/10.3389/fmicb.2019.01580
Balaji S, Shekaran SG, Prabagaran SR (2021) Cultivable bacterial communities associated with the salivary gland of Aedes aegypti. Int J Trop Sci 41:1203–1211. https://doi.org/10.1007/s42690-020-00310-9
Batson J, Dudas G, Haas-Stapleton E, Kistler AL, Li LM, Logan P, Ratnasiri K, Retallack H (2021) Single mosquito metatranscriptomics identifies vectors, emerging pathogens and reservoirs in one assay. eLife 10:e68353. https://doi.org/10.7554/eLife.68353
Becher PG, Jensen RE, Natsopoulou ME, Verschut V, De Fine Licht HH (2018) Infection of Drosophila suzukii with the obligate insect-pathogenic fungus Entomophthora muscae. J Pest Sci 91:781–787. https://doi.org/10.1007/s10340-017-0915-3
Becker N, Petrić D, Zgomba M, Boase C, Madon M, Dahl C, Kaiser A (2010) Mosquitoes and their Control, 2nd edn. Springer, Berlin Heidelberg
Becker N, Schön S, Klein AM, Ferstl I, Kizgin A, Tannich E, Kuhn C, Pluskota B, Jöst A (2017) First mass development of Aedes albopictus (Diptera: Culicidae) – its surveillance and control in Germany. Parasitol Res 116:847–858. https://doi.org/10.1007/s00436-016-5356-z
Becker N, Langentepe-Kong SM, Tokatlian Rodriguez A, Oo TT, Reichle D, Lühken R, Schmidt-Chanasit J, Lüthy P, Puggioli A, Bellini R (2022) Integrated control of Aedes albopictus in Southwest Germany supported by the Sterile Insect Technique. Parasit Vectors 15:9. https://doi.org/10.1186/s13071-021-05112-7
Beerntsen BT, James AA, Christensen BM (2000) Genetics of mosquito vector competence. Microbiol Mol Biol Rev 64:115–137. https://doi.org/10.1128/MMBR.64.1.115-137.2000
Benedict MQ, Levine RS, Hawley WA, Lounibos LP (2007) Spread of the tiger: global risk of invasion by the mosquito Aedes albopictus. Vector Borne Zoonot Dis 7:76–85. https://doi.org/10.1089/vbz.2006.0562
Bennett AJ, Bushmaker T, Cameron K, Ondzie A, Niama FR, Parra HJ, Mombouli JV, Olson SH, Munster VJ, Goldberg TJ (2019) Diverse RNA viruses of arthropod origin in the blood of fruit bats suggest a link between bat and arthropod viromes. Virology 528:64–72. https://doi.org/10.1016/j.virol.2018.12.009
Berg G, Rybakova D, Fischer D, Cernava T, Champomier Vergès MC, Charles T, Chen X, Cocolin L, Eversole K, Herrero Corral G, Kazou M, Kinkel L, Lange L, Lima N, Loy A, Macklin JA, Maguin E, Mauchline T, McClure R, Mitter B, Ryan M, Sarand I, Smidt H, Schelkle B, Roume H, Kiran SG, Selvin J, Correa S, de Souza R, van Overbeek L, Singh BK, Wagner M, Walsh A, Sessitsch A, Schloter M (2020) Microbiome definition re-visited: old consepts and new challenges. Microbiome 8:103. https://doi.org/10.1186/s40168-020-00875-0
Bernasconi E, Valsangiacomo C, Peduzzi R, Carota A, Moccetti T, Funke G (2004) Arthrobacter woluwensis subacute infective endocarditis: case report and review of the literature. Clin Infect Dis 38:27–31. https://doi.org/10.1086/381436
Blagrove MSC, Arias-Goeta C, Failloux A-B, Sinkins SP (2012) Wolbachia strain wMel induces cytoplasmic incompatibility and blocks dengue transmission in Aedes albopictus. Proc Natl Acad Sci USA 109:255–260. https://doi.org/10.1073/pnas.1112021108
Boissière A, Tchioffo MT, Bachar D, Abate L, Marie A, Nsango SE, Shahbazkia HR, Awono-Ambene PH, Levashina EA, Christen R, Morlais I (2012) Midgut microbiota of the malaria mosquito vector Anopheles gambiae and interactions with Plasmodium falciparum infection. PLoS Pathog 8:e1002742. https://doi.org/10.1371/journal.ppat.1002742
Bonizzoni M, Gasperi G, Chen X, James AA (2013) The invasive mosquito species Aedes albopictus: current knowledge and future perspectives. Trends Parasitol 29:460–468. https://doi.org/10.1016/j.pt.2013.07.003
Bourtzis K, O’Neill S (1998) Wolbachia infections and arthropod reproduction. Bioscience 48:287–293. https://doi.org/10.2307/1313355
Bourtzis K, Lees RS, Hendrichs J, Vreysen MJB (2016) More than one rabbit out of the hat: radiation, transgenic and symbiont-based approaches for sustainable management of mosquito and tsetse fly populations. Acta Trop 157:115–130. https://doi.org/10.1016/j.actatropica.2016.01.009
Calderón G, García E, Rojas P, García E, Rosso M, Losada A (2011) Chryseobacterium indologenes infection in a newborn: a case report. J Med Case Rep 5:10. https://doi.org/10.1186/1752-1947-5-10
Carr EL, Kämpfer P, Patel BKC, Gürtler V, Seviour RJ (2003) Seven novel species of Acinetobacter isolated from activated sludge. Int J Syst Evol Microbiol 53:953–963. https://doi.org/10.1099/ijs.0.02486-0
Cerezales M, Xanthopoulou K, Ertel J, Nemec A, Bustamante Z, Seifert H, Gallego L, Higgins PG (2018) Identification of Acinetobacter seifertii isolated from Bolivian hospitals. J Med Microbiol 67:834–837. https://doi.org/10.1099/jmm.0.000751
Chen WM, Cho NT, Huang WC, Young CC, Sheu SY (2013) Description of Gemmobacter fontiphilus sp. nov., isolated from a freshwater spring, reclassification of Catellibacterium nectariphilum as Gemmobacter nectariphilus comb. nov., Catellibacterium changlense as Gemmobacter changlensis comb. nov., Catellibacterium aquatile as Gemmobacter aquaticus nom. nov., Catellibacterium caeni as Gemmobacter caeni comb. nov., Catellibacterium nanjingense as Gemmobacter nanjingensis comb. nov., and emended description of the genus Gemmobacter and of Gemmobacter aquatilis. Int J Syst Evol Microbiol 63:470–478. https://doi.org/10.1099/ijs.0.042051-0
Chen S, Zhang D, Augustinos A, Doudoumis V, Bel Mokhtar N, Maiga H, Tsiamis G, Bourtzis K (2020) Multiple factors determine the structure of bacterial communities associated with Aedes albopictus under artificial rearing conditions. Front Microbiol 11:605. https://doi.org/10.3389/fmicb.2020.00605
Coon KL, Vogel KJ, Brown MR, Strand MR (2014) Mosquitoes rely on their gut microbiota for development. Mol Ecol 23:2727–2739. https://doi.org/10.1111/mec.12771
Coon KL, Brown MR, Strand MR (2016) Mosquitoes host communities of bacteria that are essential for development but vary greatly between local habitats. Mol Ecol 25:5806–5826. https://doi.org/10.1111/mec.13877
Coon KL, Valzania L, McKinney DA, Vogel KJ, Brown MR, Strand MR (2017) Bacteria-mediated hypoxia functions as a signal for mosquito development. Proc Natl Acad Sci USA 114:5362–5369. https://doi.org/10.1073/pnas.1702983114
Dahal RH, Kim J (2018) Fluviicola kyonggii sp. nov., a bacterium isolated from forest soil and emended description of the genus Fluviicola. Int J Syst Evol Microbiol 68:1885–1889. https://doi.org/10.1099/ijsem.0.002759
Demaio J, Pumpuni CB, Kent M, Beier JC (1996) The midgut bacterial flora of wild Aedes triseriatus, Culex pipiens and Psorophora columbiae mosquitoes. Am J Trop Med Hyg 54:219–223. https://doi.org/10.4269/ajtmh.1996.54.219
Derelle E, Monier A, Cooke R, Worden AZ, Grimsley NH, Moreau H (2015) Diversity of viruses infecting the green microalga Ostreococcus lucimarinus. J Virol 89:5812–5821. https://doi.org/10.1128/JVI.00246-15
Dodson BL, Hughes GL, Paul O, Matacchiero AC, Kramer LD, Rasgon JL (2014) Wolbachia enhances West Nile virus (WNV) infection in the mosquito Culex tarsalis. PLoS Negl Trop Dis 8:2965. https://doi.org/10.1371/journal.pntd.0002965
Dong Y, Manfredini F, Dimopoulos G (2009) Implication of the mosquito midgut microbiota in the defense against malaria parasites. PLoS Pathog 5:e1000423. https://doi.org/10.1371/journal.ppat.1000423
Douglas AE (2015) Multiorganismal insects: diversity and function of resident microorganisms. Annu Rev Entomol 60:17–34. https://doi.org/10.1146/annurev-ento-010814-020822
Emerson H, Norris C (1905) “Red-Leg” – an infectious disease of frogs. J Exp Medicine 7:32–57. https://doi.org/10.1084/jem.7.1.32
Engel P, Moran NA (2013) The gut microbiota of insects – diversity in structure and function. FEMS Microbiol Rev 37:699–735. https://doi.org/10.1111/1574-6976.12025
Fălcuţă E, Prioteasa LF, Horváth C, Păstrav IR, Schaffner F, Mihalca AD (2020) The invasive Asian tiger mosquito Aedes albopictus in Romania: towards a country-wide colonization? Parasitol Res 119:841–845. https://doi.org/10.1007/s00436-020-06620-8
Fernandes MK, Neves CA, Serrão JE, Martins GF (2014) Aedes aegypti midgut remodeling during metamorphosis. Parasitol Int 63:506–512. https://doi.org/10.1016/j.parint.2014.01.004
Fischer N, Ruef C, Ebnöther C, Bächli EB (2008) Rhinofacial Conidiobolus coronatus infection presenting with nasal enlargement. Infection 36:594–596. https://doi.org/10.1007/s15010-008-8056-5
Flores HA, O’Neill SL (2018) Controlling vector-borne diseases by releasing modified mosquitoes. Nat Rev Microbiol 16:508–518. https://doi.org/10.1038/s41579-018-0025-0
Fosse T, Peloux Y, Granthil C, Toga B, Bertrando J, Sethian M (1985) Meningitis due to Micrococcus luteus. Infection 13:280–281. https://doi.org/10.1007/BF01645439
Gao N, Xia M, Dai J, Yu D, An W, Li S, Liu S, He P, Zhang L, Wu Z, Bi X, Chen S, Haft DH, Qiu D (2018) Both widespread PEP-CTERM proteins and exopolysaccharides are required for floc formation of Zoogloea resiniphila and other activated sludge bacteria. Environ Microbiol 20:1677–1692. https://doi.org/10.1111/1462-2920.14080
Gao H, Cui C, Wang L, Jacobs-Lorena M, Wang S (2020) Mosquito microbiota and implications for disease control. Trends Parasitol 36:98–111. https://doi.org/10.1016/j.pt.2019.12.001
Gimnig JE, Ombok M, Otieno S, Kaufman MG, Vulule JM, Walker ED (2002) Density-dependent development of Anopheles gambiae (Diptera: Culicidae) larvae in artificial habitats. J Med Entomol 39:162–172. https://doi.org/10.1603/0022-2585-39.1.162
Gimonneau G, Tchioffo MT, Abate L, Boissière A, Awono-Ambéné PH, Nsango SE, Christen R, Morlais I (2014) Composition of Anopheles coluzzii and Anopheles gambiae microbiota from larval to adult stages. Infect Genet Evol 28:715–724. https://doi.org/10.1016/j.meegid.2014.09.029
Hazen TC, Fliermans CB, Hirsch RP, Esch GW (1978) Prevalence and distribution of Aeromonas hydrophila in the United States. Appl Environ Microbiol 36:731–738. https://doi.org/10.1128/aem.36.5.731-738.1978
Hejazi S, Falkiner FR (1997) Serratia marcescens. J Med Microbiol 46:903–912. https://doi.org/10.32388/HXQR6V
Hess B, Burchett A, Huntington MK (2008) Leclercia adecarboxylata in an immunocompetent patient. J Med Microbiol 57:896–898. https://doi.org/10.1099/jmm.0.47673-0
Hilgenboecker K, Hammerstein P, Schlattmann P, Telschow A, Werren JH (2008) How many species are infected with Wolbachia? A statistical analysis of current data. FEMS Microbiol Letters 281:215–220. https://doi.org/10.1111/j.1574-6968.2008.01110.x
Hoegl L, Schönian G, Ollert M, Korting HC (1998) Candida sake: a relevant species in the context of HIV-associated oropharyngeal candidosis? J Mol Med 76:70–73. https://doi.org/10.1007/s001090050192
Holmes B, Steigerwalt AG, Nicholson AC (2013) DNA-DNA hybridization study of strains of Chryseobacterium, Elizabethkingia and Empedobacter and of other usually indole-producing non-fermenters of CDC groups IIc, IIe, IIh and IIi, mostly from human clinical sources, and proposals of Chryseobacterium bernardetii sp. nov., Chryseobacterium carnis sp. nov., Chryseobacterium lactis sp. nov., Chryseobacterium nakagawai sp. nov. and Chryseobacterium taklimakanense comb. nov. Int J Syst Evol Microbiol 63:4639–4662. https://doi.org/10.1099/ijs.0.054353-0
Hörtnagl P, Pérez MT, Zeder M, Sommaruga R (2010) The bacterial community composition of the surface microlayer in a high mountain lake. FEMS Microbiol Ecol 73:458–467. https://doi.org/10.1111/j.1574-6941.2010.00904.x
Hunter WJ, Kuykendall LD, Manter DK (2007) Rhizobium selenireducens sp. nov.: a selenite-reducing alpha-Proteobacteria isolated from a bioreactor. Curr Microbiol 55:455–460. https://doi.org/10.1007/s00284-007-9020-9
Joseph SW, Carnahan AM, Brayton PR, Fanning GR, Almazan R, Drabick C, Trudo EWJR, Colwell RR (1991) Aeromonas jandaei and Aeromonas veronii dual infection of a human wound following aquatic exposure. J Clin Microbiol 29:565–569. https://doi.org/10.1128/jcm.29.3.565-569.1991
Jupatanakul N, Sim S, Dimopoulos G (2014) The insect microbiome modulates vector competence for arboviruses. Viruses 6:4294–4313. https://doi.org/10.3390/v6114294
Kämpfer P, Matthews H, Glaeser SP, Martin K, Lodders N, Faye I (2011) Elizabethkingia anophelis sp. nov., isolated from the midgut of the mosquito Anopheles gambiae. Int J Syst Evol Microbiol 61:2670–2675. https://doi.org/10.1099/ijs.0.026393-0
Kang YS, Jung J, Jeon CO, Park W (2011) Acinetobacter oleivorans sp. nov. is capable of adhering to and growing on diesel-oil. J Microbiol 49:29–34. https://doi.org/10.1007/s12275-011-0315-y
Kanyuka K, Ward E, Adams MJ (2003) Polymyxa graminis and the cereal viruses it transmits: a research challenge. Mol Plant Pathol 4:393–406. https://doi.org/10.1046/j.1364-3703.2003.00177.x
Kaper JB, Nataro JP, Mobley HL (2004) Pathogenic Escherichia coli. Nat Rev Microbiol 2:123–140. https://doi.org/10.1038/nrmicro818
Kempf M, Abdissa A, Diatta G, Trape J-F, Angelakis E, Mediannikov O, La Scola B, Raoult D (2012) Detection of Acinetobacter baumannii in human head and body lice from Ethiopia and identification of new genotypes. Int J Infect Dis 16:e680–e683. https://doi.org/10.1016/j.ijid.2012.05.1024
Kim D, Baik KS, Kim MS, Park SC, Kim SS, Rhee MS, Kwak YS, Seon CN (2008) Acinetobacter soli sp. nov., isolated from forest soil. J Microbiol 46:396–401. https://doi.org/10.1007/s12275-008-0118-y
Kittayapong P, Baisley KJ, Baimai V, O’Neill SL (2000) Distribution and diversity of Wolbachia infections in Southeast Asian mosquitoes (Diptera: Culicidae). J Med Entomol 37:340–345. https://doi.org/10.1093/jmedent/37.3.340
Kostmann JR, Solomon F, Fekete T (1990) Infections with Chryseomonas luteola (CDC Group Ve-1) and Flavimonas oryzhabitans (CDC Group Ve-2) in neurosurgical patients. Rev Infect Dis 13:233–236. https://doi.org/10.1093/clinids/13.2.233
Kubacki J, Flacio E, Qi W, Guidi V, Tonolla M, Fraefel C (2020) Viral metagenomic analysis of Aedes albopictus mosquitos from southern Switzerland. Viruses 12:929. https://doi.org/10.3390/v12090929
Larkin JM, Williams PM (1978) Runella slithyformis gen. nov., sp. nov., a curved, nonflexible, pink bacterium. Int J Syst Bacteriol 28:32–36. https://doi.org/10.1099/00207713-28-1-32
Lau SKP, Chow WN, Foo CH, Curreem SOT, Lo GCS, Teng JLL, Chen JHK, Ng RHY, Wu AKL, Cheung IYY, Chau SKY, Lung DC, Lee RA, Tse CWS, Fung KSC, Que TL, Woo PCY (2016) Elizabethkingia anophelis bacteremia is associated with clinically significant infections and high mortality. Sci Rep 6:26045. https://doi.org/10.1038/srep26045
Lee JE, Hwang EM, Cha CJ, Kim GB (2019) Chryseobacterium aureum sp. nov., isolated from the Han River, Republic of Korea. Int J Syst Evol Microbiol 69:1628–1633. https://doi.org/10.1099/ijsem.0.003370
Li SY, Kao CC, Hu YC, Lai CH, Jiang YP, Mao YC, Huang YT, Liu PY (2021) Arthrobacter woluwensis bacteremia: a clinical and genomic report. Pathogens 10:443. https://doi.org/10.3390/pathogens10040443
Ling LL, Schneider T, Peoples AJ, Spoering AL, Engels I, Conlon BP, Mueller A, Schäberle TF, Hughes DE, Epstein S, Jones M, Lazarides L, Steadman VA, Cohen DR, Felix CR, Fetterman KA, Millett WP, Nitti AG, Zullo AM, Chen C, Lewis K (2015) A new antibiotic kills pathogens without detectable resistance. Nature 517:455–459. https://doi.org/10.1038/nature14098
Mancini MV, Damiani C, Accoti A, Tallarita M, Nunzi E, Cappelli A, Bozic J, Catanzani R, Rossi P, Valzano M, Serrao A, Ricci I, Spaccapelo R, Favia G (2018) Estimating bacteria diversity in different organs of nine species of mosquito by next generation sequencing. BMC Microbiol 18:126. https://doi.org/10.1186/s12866-018-1266-9
Manni M, Zdobnov EM (2020) A novel anphevirus in Aedes albopictus mosquitoes is distributed worldwide and interacts with the host RNA interference pathway. Viruses 12:1264. https://doi.org/10.3390/v12111264
Martinet JP, Ferté H, Failloux AB, Schaffner F, Depaquit J (2019) Mosquitoes of north-western Europe as potential vectors of arboviruses: a review. Viruses 11:1059. https://doi.org/10.3390/v11111059
Merritt RW, Dadd RH, Walker ED (1992) Feeding behavior, natural food, and nutritional relationships of larval mosquitoes. Annu Rev Entomol 37:349–376. https://doi.org/10.1146/annurev.en.37.010192.002025
Minard G, Tran FH, Raharimalala FN, Hellard E, Ravelonandro P, Mavingui P, Valiente Moro C (2013) Prevalence, genomic and metabolic profiles of Acinetobacter and Asaia associated with field-caught Aedes albopictus from Madagascar. FEMS Microbiol Ecol 83:63–73. https://doi.org/10.1111/j.1574-6941.2012.01455.x
Moissenet D, Becker K, Mérens A, Ferroni A, Dubern B, Vu-Thien H (2012) Persistent bloodstream infection with Kocuria rhizophila related to a damaged central catheter. J Clin Microbial 50:1495–1498. https://doi.org/10.1128/JCM.06038-11
Moreira JS, Riccetto AGL, Da Silva MTN, Vilela MMDS (2015) Endocarditis by Kocuria rosea in an immunocompetent child. Braz J Infect Dis 19:82–84. https://doi.org/10.1016/j.bjid.2014.09.007
Moro CV, Tran FH, Raharimalala FN, Ravelonandro P, Mavingui P (2013) Diversity of culturable bacteria including Pantoea in wild mosquito Aedes albopictus. BMC Microbiol 13:70. http://www.biomedcentral.com/1471-2180/13/70.
Muratoglu H, Kati H, Demirbag Z, Sezen K (2009) High insecticidal activity of Leclercia adecarboxylata isolated from Leptinotarsa decemlineata (Col.: Chrysomelidae). Afr J Biotechnol 24:7111–7115. http://www.academicjournals.org/AJB
Nemec A, Radolfova-Krizova L, Maixnerova M, Vrestiakova E, Jezek P, Sedo O (2016) Taxonomy of haemolytic and/or proteolytic strains of the genus Acinetobacter with the proposal of Acinetobacter courvalinii sp. nov. (genomic species 14 sensu Bouvet & Jeanjean), Acinetobacter dispersus sp. nov. (genomic species 17), Acinetobacter modestus sp. nov., Acinetobacter proteolyticus sp. nov. and Acinetobacter vivianii sp. nov. Int J Syst Evol Microbiol 66:1673–1685. https://doi.org/10.1099/ijsem.0.000932
Nguyen TM, Kim J (2017) Limnobacter humi sp. nov., a thiosulfate-oxidizing, heterotrophic bacterium isolated from humus soil, and emended description of the genus Limnobacter Spring et al. 2001. J Microbiol 55:508–513. https://doi.org/10.1007/s12275-017-6645-7
Nogueira MM, Neves E, Johnsson R (2021) Effects of habitat structure on the mollusc assemblage in Mussismilia corals: evaluation of the influence of different coral growth morphology. J Mar Biol Assoc UK 101:61–69. https://doi.org/10.1017/S0025315421000023
Oliveira Ribeiro Gd, Da Costa AC, Gill DE, Ribeiro ESD’A, Da Rego MOS, Monteiro FJC, Villanova F, Nogueira JS, Maeda AY, de Preira Souza R, Tahmasebi R, Morais VS, Pandey RP, Raj VS, Scandar SAS, da Silva Vasami FG, D’Agostino LG, Maiorka PC, Deng X, Nogueira ML, Sabino EC, Delwart ED, Leal ÈL, Cunha MS (2021) Guapiaçu virus, a new insect-specific flavivirus isolated from two species of Aedes mosquitoes from Brazil. Sci Rep 11:4674. https://doi.org/10.1038/s41598-021-83879-6
Onchuru TO, Ajamma YU, Burugu M, Kaltenpoth M, Masiga D, Villinger J (2016) Chemical parameters and bacterial communities associated with larval habitats of Anopheles, Culex and Aedes mosquitoes (Diptera: Culicidae) in western Kenya. Int J Trop Insect Sci 36:146–160. https://doi.org/10.1017/S1742758416000096
Pang T, Mak TK, Gubler DJ (2017) Prevention and control of dengue – the light at the end of the tunnel. Lancet Infect Dis 17:e79–e87. https://doi.org/10.1016/S1473-3099(16)30471-6
Park CH, Lim HW, Kim HW, Lee WGL, Roh JY, Park MY, Shin EH (2016) High prevalence of Wolbachia infection in Korean populations of Aedes albopictus (Diptera: Culicidae). J Asia-Pacific Entomol 19:191–194. https://doi.org/10.1016/j.aspen.2015.12.014
Paupy C, Delatte H, Bagny L, Corbel V, Fontenille D (2009) Aedes albopictus, an arbovirus vector: from the darkness to the light. Microbes Infect 11:1177–1185. https://doi.org/10.1016/j.micinf.2009.05.005
Pękala A, Paździor E, Antychowicz J, Bernad A, Głowacka H, Więcek B, Niemczuk W (2018) Kocuria rhizophila and Micrococcus luteus as emerging opportunist pathogens in brown trout (Salmo trutta Linnaeus, 1758) and rainbow trout (Oncorhynchus mykiss Walbaum, 1792). Aquaculture 486:285–289. https://doi.org/10.1016/j.aquaculture.2017.12.028
Pidiyar V (2002) Aeromonas culicicola sp. nov., from the midgut of Culex quinquefasciatus. Int J Syst Evol Microbiol 52:1723–1728. https://doi.org/10.1099/ijs.0.02019-0
Pitt A, Schmidt J, Koll U, Hahn MW (2019) Aquirufa antheringensis gen. nov., sp. nov. and Aquirufa nivalisilvae sp. nov., representing a new genus of widespread freshwater bacteria. Int J Syst Evol Microbiol 69:2739–2749. https://doi.org/10.1099/ijsem.0.003554
Pladdies T, Babenzien HD, Cypionka H (2004) Distribution of Nevskia ramosa and other rosette-forming neustonic bacteria. Microb Ecol 47:218–223. https://doi.org/10.1007/s00248-003-1070-3
Ponte R, Jenkis SG (1992) Fatal Chromobacterium violaceum infections associated with exposure to stagnant waters. Pediatr Infect Dis J 11:583–586
Rani A, Sharma A, Rajagopal R, Adak T, Bhatnagar RK (2009) Bacterial diversity analysis of larvae and adult midgut microflora using culture-dependent and culture-independent methods in lab-reared and field-collected Anopheles stephensi – an Asian malarial vector. BMC Microbiol 9:96. https://doi.org/10.1186/1471-2180-9-96
Reiss I, Borkhardt A, Füssle R, Sziegoleit A, Gortner L (2000) Disinfectant contaminated with Klebsiella oxytoca as a source of sepsis in babies. Lancet 356:310. https://doi.org/10.1016/S0140-6736(00)02509-5
Riehle MA, Moreira CK, Lampe D, Lauzon C, Jacobs-Lorena M (2007) Using bacteria to express and display anti-Plasmodium molecules in the mosquito midgut. Int J Parasitol 37:595–603. https://doi.org/10.1016/j.ijpara.2006.12.002
Romano-Bertrand S, Bourdier A, Aujoulat F, Michon AL, Masnou A, Parer S, Marchandin H, Jumas-Bilak E (2016) Skin microbiota is the main reservoir of Roseomonas mucosa, an emerging opportunistic pathogen so far assumed to be environmental. Clin Microbiol Infect 22:e1-7. https://doi.org/10.1016/j.cmi.2016.05.024
Sadeghi M, Popov V, Guzman H, Phan TG, Vasilakis N, Tesh R, Delwart E (2017) Genomes of viral isolates derived from different mosquito species. Virus Res 242:49–57. https://doi.org/10.1016/j.virusres.2017.08.012
Salter SJ, Cox MJ, Turek EM, Caslus ST, Cookson WO, Moffatt MF, Turner P, Parkhill J, Lohman NJ, Walker AW (2014) Reagent and laboratory contamination can critically impact sequences-based microbiome analyses. BMC Biol 12:87. https://doi.org/10.5194/gi-2016-11-RC2
Schmidt R, Battaglia V, Scow K, Kane S, Hristova KR (2008) Involvement of a novel enzyme, MdpA, in methyl tert-butyl ether degradation in Methylibium petroleiphilum PM1. Appl Environ Microbial 74:6631–6638. https://doi.org/10.1128/AEM.01192-08
Seifert H, Strate A, Schulze A, Pulverer G (1993) Vascular catheter-related bloodstream infection due to Acinetobacter johnsonii (formerly Acinetobacter calcoaceticus var. lwoffii): report of 13 cases. Clin Infect Dis 17:632–636. https://doi.org/10.1093/clinids/17.4.632
Seitz HM, Maier WA, Rottok M, Becker-Feldmann H (1987) Concomitant infections of Anopheles stephensi with Plasmodium berghei and Serratia marcescens: additive detrimental effects. Zentralbl Bakt Mikrobiol Hyg A 266:155–166. https://doi.org/10.1016/s0176-6724(87)80029-9
Service MW (2001) The encyclopedia of arthropod-transmitted infections. CABI Publishing, Liverpool School of Tropical Medicine, UK
Shahi N, Sharma P, Pandey J, Bisht I, Mallik SK (2018) Characterization and pathogenicity study of Chryseobacterium scophthalmum recovered from gill lesions of diseased golden mahseer, Tor putitora (Hamilton, 1822) in India. Aquaculture 485:81–92. https://doi.org/10.1016/j.aquaculture.2017.11.018
Shi C, Zhao L, Atoni E, Zeng W, Hu X, Matthijnssens J, Yuan Z, Xia H (2020) Stability of the virome in lab- and field-collected Aedes albopictus mosquitoes across different developmental stages and possible core viruses in the publicly available virome data of Aedes mosquitoes. mSystems 5:e00640-20. https://doi.org/10.1128/mSystems.00640-20
Shil RK, Mojumder S, Sadida FF, Uddin M, Sikdar D (2014) Isolation and identification of cellulolytic bacteria from the gut of three phytophagus insect species. Braz Arch Biol Technol 57:927–932. https://doi.org/10.1590/S1516-8913201402620
Smith AH, Łukasik P, O’Connor MP, Lee A, Mayo G, Drott MT, Doll S, Tuttle R, Disciullo RA, Messina A, Oliver KM, Russel JA (2015) Patterns, causes and consequences of defensive microbiome dynamics across multiple scales. Mol Ecol 24:1135–1149. https://doi.org/10.1111/mec.13095
Straif SC, Mbogo CNM, Toure AM, Walker ED, Kaufman M, Toure YT, Beier JC (1989) Midgut bacteria in Anopheles gambiae and An. funestus (Diptera: Culicidae) from Kenya and Mali. J Med Entomol 35:222–226. https://doi.org/10.1093/jmedent/35.3.222
Strand MR (2018) Composition and functional roles of the gut microbiota in mosquitoes. Curr Opin Insect Sci 28:59–65. https://doi.org/10.1016/j.cois.2018.05.008
Terenius O, Lindh JM, Eriksson-Gonzales K, Bussière L, Laugen AT, Bergquist H, Titanji K, Faye I (2012) Midgut bacterial dynamics in Aedes aegypti. FEMS Microbiol Ecol 80:556–565. https://doi.org/10.1111/j.1574-6941.2012.01317.x
Touré AM, Mackey AJ, Wang ZX, Beier JC (2000) Bactericidal effects of sugar-fed antibiotics on resident midgut bacteria of newly emerged anopheline mosquitoes (Diptera: Culicidae). J Med Entomol 37:246–249. https://doi.org/10.1093/jmedent/37.2.246
Uniyal S, Paliwal R, Verma M, Sharma RK, Rai JPN (2016) Isolation and characterization of fipronil degrading Acinetobacter calcoaceticus and Acinetobacter oleivorans from rhizospheric zone of Zea mays. Bull Environ Contam Toxicol 96:833–838. https://doi.org/10.1007/s00128-016-1795-6
van Dexter S, Boopathy R (2019) Biodegradation of phenol by Acinetobacter tandoii isolated from the gut of the termite. Environ Sci Pollut Res Int 26:34067–34072. https://doi.org/10.1007/s11356-018-3292-4
van Pham HT, Jeong SW, Kim J (2015) Aquabacterium olei sp. nov., an oil-degrading bacterium isolated from oil-contaminated soil. Int J Syst Evol Microbial 65:3597–3602. https://doi.org/10.1099/ijsem.0.000458
Walcott RR, Gitaitis RD (2000) Detection of Acidovorax avenae subsp. citrulli in watermelon seed using immunomagnetic separation and the polymerase chain reaction. Plant Dis 84:470–474. https://doi.org/10.1094/PDIS.2000.84.4.470
Walther D, Scheuch DE, Kampen H (2017) The invasive Asian tiger mosquito Aedes albopictus (Diptera: Culicidae) in Germany: local reproduction and overwintering. Acta Trop 166:186–192. https://doi.org/10.1016/j.actatropica.2016.11.024
Wang Y, Gilbreath TM, Kukutla P, Yan G, Xu J (2011) Dynamic gut microbiome across life history of the malaria mosquito Anopheles gambiae in Kenya. PLoS One 6:e24767. https://doi.org/10.1371/journal.pone.0024767
Wang X, Liu T, Wu Y, Zhong D, Zhou G, Su X, Xu J, Sotero CF, Sadruddin AA, Wu K, Chen X-G, Yan G (2018) Bacterial microbiota assemblage in Aedes albopictus mosquitoes and ist impacts on larval development. Mol Ecol 27:2972–2985. https://doi.org/10.1111/mec.14732
Watson DW, Mullens BA, Petersen JJ (1993) Behavioral fever response of Musca domestica (Diptera: Muscidae) to infection by Entomophthora muscae (Zygomycetes: Entomophthorales). J Invert Pathol 61:10–16. https://doi.org/10.1006/jipa.1993.1003
Werren JH, Baldo L, Clark ME (2008) Wolbachia: master manipulators of invertebrate biology. Nat Rev Microbiol 6:741–751. https://doi.org/10.1038/nrmicro1969
Wiklund T, Bylund G (1990) Pseudomonas anguilliseptica as a pathogen of salmonid fish in Finland. Dis Aquat Org 8:13–19
Wiwatanaratanabutr I (2013) Geographic distribution of wolbachial infections in mosquitoes from Thailand. J Invert Pathol 114:337–340. https://doi.org/10.1016/j.jip.2013.04.011
Wohlgemut K, Pierce RL, Kirkbride CA (1970) Bovine abortion associated with Aeromonas hydrophila. J Am Med Assoc 160:1001–1002
Wylezich C, Papa A, Beer M, Höper D (2018) A versatile sample processing workflow for metagenomic pathogen detection. Sci Rep 8:13108. https://doi.org/10.1038/s41598-018-31496-1
Yang M (2000) Long PCR improves Wolbachia DNA amplification: wsp sequences found in 76% of sixty-three arthropod species. Insect Mol Biol 9:393–405. https://doi.org/10.1046/j.1365-2583.2000.00203.x
Yang CH, Li YH (2011) Chromobacterium violaceum infection: a clinical review of an important but neglected infection. J Chin Med Assoc 74:435–441. https://doi.org/10.1016/j.jcma.2011.08.013
Ye-Ebiyo Y, Pollack RJ, Kiszewski A, Spielman A (2003) Enhancement of development of larval Anopheles arabiensis by proximity to flowering maize (Zea Mays) in turbid water and when crowded. Am J Trop Med Hyg 68:748–752. https://doi.org/10.4269/ajtmh.2003.68.748
Yoon JH, Oh TK, Park YH (2004) Kangiella koreensis gen. nov., sp. nov. and Kangiella aquimarina sp. nov., isolated from a tidal flat of the Yellow Sea in Korea. Int J Syst Evol Microbiol 54:1829–1835. https://doi.org/10.1099/ijs.0.63156-0
Zamora L, Vela AI, Palacios MA, Sánchez-Porro C, Svensson-Stadler LA, Domínguez L, Moore ERB, Ventosa A, Fernández-Garayzábal JF (2012) Chryseobacterium viscerum sp. nov., isolated from diseased fish. Int J Syst Evol Microbiol 62:2934–2940. https://doi.org/10.1099/ijs.0.036699-0
Zhu D, Xie C, Huang Y, Sun J, Zhang W (2014) Description of Comamonas serinivorans sp. nov., isolated from wheat straw compost. Int J Syst Evol Microbiol 64:4141–4146. https://doi.org/10.1099/ijs.0.066688-0
Zotzmann S, Steinbrink A, Schleich K, Frantzmann F, Xoumpholphakdy C, Spaeth M, Moro CV, Mavingui P, Klimpel S (2017) Bacterial diversity of cosmopolitan Culex pipiens and invasive Aedes japonicus from Germany. Parasitol Res 116:1899–1906. https://doi.org/10.1007/s00436-017-5466-2
Zouache K, Raharimalala FN, Raquin V, van Tran-Van R, Lala HR, Ravelonandro P, Mavingui P (2011) Bacterial diversity of field-caught mosquitoes, Aedes albopictus and Aedes aegypti, from different geographic regions of Madagascar. FEMS Microbiol Ecol 75:377–389. https://doi.org/10.1111/j.1574-6941.2010.01012.x
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
JR, DW, MB and HK designed the study; DW and HK collected and identified the mosquitoes; JR and DH carried out the molecular work; JR analysed and interpreted the data; JR and HK wrote the main manuscript text; JR prepared the figure; and all authors reviewed and edited the manuscript. All authors approved the final manuscript version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Handling Editor: Una Ryan
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rau, J., Werner, D., Beer, M. et al. The microbial RNA metagenome of Aedes albopictus (Diptera: Culicidae) from Germany. Parasitol Res 121, 2587–2599 (2022). https://doi.org/10.1007/s00436-022-07576-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00436-022-07576-7