Abstract
Main conclusion
Some salt-stress responsive DEGs, mainly involved in ion transmembrane transport, hormone regulation, antioxidant system, osmotic regulation, and some miRNA jointly regulated the salt response process in allotriploid Populus cathayana.
Abstract
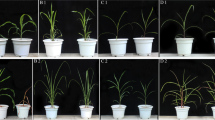
The molecular mechanism of plant polyploid stress resistance has been a hot topic in biological research. In this study, Populus diploids and first division restitution (FDR) and second division restitution (SDR) triploids were selected as research materials. All materials were treated with 70 mM NaCl solutions for 30 days in the same pot environment. We observed the growth state of triploids and diploids and determined the ratio of potassium and sodium ions, peroxidase (POD) activity, proline content, and ABA and jasmonic acid (JA) hormone content in leaves in the same culture environment with the same concentration of NaCl solution treatment. In addition, RNA-seq technology was used to study the differential expression of mRNA and miRNA. The results showed that triploid Populus grew well and the K+ content and the K+/Na+ ratio in the salt treatment were significantly lower than those in the control. The contents of ABA, JA, POD, and proline were increased compared with contents in diploid under salt stress. The salt-stress responsive DEGs were mainly involved in ion transport, cell homeostasis, the MAPK signaling pathway, peroxisome, citric acid cycle, and other salt response and growth pathways. The transcription factors mainly included NAC, MYB, MYB_related and AP2/ERF. Moreover, the differentially expressed miRNAs involved 32 families, including 743 miRNAs related to predicted target genes, among which 22 miRNAs were significantly correlated with salt-stress response genes and related to the regulation of hormones, ion transport, reactive oxygen species (ROS) and other biological processes. Our results provided insights into the physiological and molecular aspects for further research into the response mechanisms of allotriploid Populus cathayana to salt stress. This study provided valuable information for the salt tolerance mechanism of allopolyploids.








Similar content being viewed by others
Abbreviations
- DEGs:
-
Differentially expressed genes
- FDR:
-
First division restitution
- JA:
-
Jasmonic acid
- POD:
-
Peroxidase
- SDR:
-
Second division restitution
- TF:
-
Transcription factor
References
Ali MA, Azeem F, Nawaz MA, Acet T, Abbas A, Imran QM, Shah KH, Rehman HM, Chung G, Yang SH, Bohlmann H (2018) Transcription factors WRKY11 and WRKY17 are involved in abiotic stress responses in Arabidopsis. J Plant Physiol 226:12–21
An J, Cheng C, Hu Z, Chen H, Cai W, Yu B (2018) The Panax ginseng PgTIP1 gene confers enhanced salt and drought tolerance to transgenic soybean plants by maintaining homeostasis of water, salt ions and ROS. Environ Exp Bot 155:45–55
Asif MA, Zafar Y, Iqbal J, Iqbal MM, Rashid U, Ali GM, Arif A, Nazir F (2011) Enhanced expression of AtNHX1, in transgenic groundnut (Arachis hypogaea L.) improves salt and drought tolerence. Mol Biotechnol 49(3):250–256
Chakraborty K, Chattaopadhyay K, Nayak L, Ray S, Sarkar RK (2019) Ionic selectivity and coordinated transport of Na+ and K+ in flag leaves render differential salt tolerance in rice at the reproductive stage. Planta 250:1637–1653
Chao DY, Dilkes B, Luo H, Douglas A, Yakubova E, Lahner B, Salt DE (2013) Polyploids exhibit higher potassium uptake and salinity tolerance in Arabidopsis. Science 341(6146):658–659
Cheng SP, Huang Z, Li Y, Liao T, Suo YJ, Zhang PD, Wang J, Kang XY (2015) Differential transcriptome analysis between Populus and its synthesized allotriploids driven by second-division restitution. J Integr Plant Biol 57(12):1031–1045
Cui MH, Yoo KS, Hyoung S (2013) An Arabidopsis R2R3-MYB transcription factor, AtMYB20, negatively regulates type 2C serine/threonine protein phosphatases to enhance salt tolerance. FEBS Lett 587(12):1773–1778
Deinlein U, Stephan AB, Horie T, Luo W, Schroeder JI (2014) Plant salt-tolerance mechanisms. Trends Plant Sci 19(6):371–379
Diray-Arce J, Clement M, Gul B, Khan MA, Nielsen BL (2015) Transcriptome assembly, profiling and differential gene expression analysis of the halophyte Suaeda fruticosa provides insights into salt tolerance. BMC Genomics 16:353
Du XL, Wang G, Ji J, Shi LP, Guan CF, Jin C (2017) Comparative transcriptome analysis of transcription factors in different maize varieties under salt stress conditions. Plant Growth Regul 81(1):183–195
Du Q, Wang K, Zou C, Xu L, Wen X (2018) The PILNCR1-miR399 regulatory module is important for low phosphate tolerance in maize. Plant Physiol 177:1743–1753
Frazier TP, Sun G, Burklew CE, Zhang B (2011) Salt and drought stresses induce the aberrant expression of microRNA genes in tobacco. Mol Biotechnol 49(2):159–165
Giri B, Kapoor R, Mukerji KG (2007) Improved tolerance of Acacia niloticato salt stress by arbuscular mycorrhiza, Glomus fasciculatum may be partly related to elevated K/Na ratios in root and shoot tissues. Microb Ecol 54(4):753–760
Guan X, Song Q, Chen ZJ (2014) Polyploidy and small RNA regulation of cotton fiber development. Trends Plant Sci 19:516–528
He Z, Li Z, Lu H, Huo L, Wang Z, Wang Y, Ji X (2019) The NAC protein from Tamarix hispida, ThNAC7, confers salt and osmotic stress tolerance by increasing reactive oxygen species scavenging capability. Plants 8(7):221–239
Hu BQ, Wang B, Wang CG, Song WQ, Chen CB (2012) Microarray analysis of gene expression in triploid black poplar. Silvae Genet 61(4–5):148–157
Jasinski S, Piazza P, Craft J, Hay A, Tsiantis M (2005) KNOX action in Arabidopsis is mediated by coordinate regulation of cytokinin and gibberellin activities. Curr Biol 15(17):1560–1565
Keita M, Jun F, Haniyeh B, Masashi A, Shinjiro Y, Shinobu S (2014) Gibberellin-induced expression of Fe uptake-related genes in Arabidopsis. Plant Cell Physiol 55(1):87–98
Kim W, Ahn HJ, Chiou TJ, Ahn JH (2011) The role of the miR399-PHO2 module in the regulation of flowering time in response to different ambient temperatures in Arabidopsis thaliana. Mol Cells 32(1):83–88
Lang J, Genot B, Hirt H, Colcombet J (2017) Constitutive activity of the ArabidopsisMAP kinase 3 confers resistance to Pseudomonas syringae and drives robust immune responses. Plant Signal Behav 12:1559–2324
Laohavisit A, Richards SL, Shabala L, Chen C, Colaco RDDR, Swarbreck SM, Shaw E, Dark A, Shabala S, Shang Z, Davies JM (2013) Salinity-induced calcium signaling and root adaptation in Arabidopsis require the calcium regulatory protein annexin1. Plant Physiol 163(1):253–262
Li DL, Kang XY, Chen HW, Wang J, Zhang PD (2010) Induction of diploid eggs with colchicine during embryo sac development in Populus. Silvae Genet 59:40–48
Li A, Liu D, Wu J, Zhao X, Hao M (2014) mRNA and small RNA transcriptomes reveal insights into dynamic homoeolog regulation of allopolyploid heterosis in nascent hexaploid wheat. Plant Cell 26(5):1870–1900
Li J, Gao Z, Zhou L, Li LZ, Zhang JH, Liu Y, Chen HY (2019) Comparative transcriptome analysis reveals K+ transporter gene contributing to salt tolerance in eggplant. BMC Plant Biol 19(1):67
Liu BB, Sun GL (2017) microRNAs contribute to enhanced salt adaptation of the autopolyploid Hordeum bulbosum compared with its diploid ancestor. Plant J 91(1):57–69
Liu M, Song X, Jiang Y (2018) Growth, ionic response, and gene expression of shoots and roots of perennial ryegrass under salinity stress. Acta Physiol Plant 40(6):112–120
Ma X, Zhang J, Huang B (2016) Cytokinin-mitigation of salt-induced leaf senescence in perennial ryegrass involving the activation of antioxidant systems and ionic balance. Environ Exp Bot 125:1–11
Maryamsadat V, Hee LS, Zwiazek JJ (2018) Water transport properties of root cells contribute to salt tolerance in halophytic grasses Poa juncifolia and Puccinellia nuttalliana. Plant Sci 276:54–62
Mehlmer N, Wurzinger B, Stael S, Hofmann-Rodrigues D, Csaszar E, Pfister B, Bayer R, Teige M (2010) The Ca2+-dependent protein kinase CPK3 is required for MAPK-independent salt-stress acclimation in Arabidopsis. Plant J 63(3):484–498
Meng F, Luo Q, Wang Q, Zhang X, Qi Z, Xu F, Lei X, Cao Y, Chow WS, Sun G (2016) Physiological and proteomic responses to salt stress in chloroplasts of diploid and tetraploid black locust (Robinia pseudoacacia L.). Sci Rep-UK 6:23098
Nawrocki EP, Burge SW, Bateman A, Daub J, Eberhardt RY, Eddy SR, Floden EW, Gardner PP, Jones TA, Tate J, Finn RD (2015) Rfam 12.0: updates to the RNA families database. Nucleic Acids Res 43(D1):130–137
Noman A, Fahad S, Aqeel M, Ali U, Amanullah A, Sumera B, Shahbaz K, Zainab M (2017) miRNAs: major modulators for crop growth and development under abiotic stresses. Biotechnol Lett 39(5):685–700
Peng M, Meng FJ, Jiang MQ, Xu FL, Huang FL (2016) Methyl jasmonate regulated diploid and tetraploid black locust (Robinia pseudoacacia L.) tolerance to salt stress. Acta Physiol Plant 38(4):106–119
Qin Z, Chen J, Jin L, Duns GJ, Ouyang P (2015) Differential expression of miRNAs under salt stress in Spartina alterniflora leaf tissues. J Nanosci Nanotechnol 15(2):1554–1561
Ruiz M, Quiñones A, Martínez-Alcántara B, Aleza P, Morillon R, Navarro L, Primo-Millo E, Martínez-Cuenca MR (2016) Effects of salinity on diploid (2x) and doubled diploid (4x) Citrus macrophylla genotypes. Sci Hortic 207:33–40
Sah SK, Reddy KR, Li JX (2016) Abscisic acid and abiotic stress tolerance in crop plants. Front Plant Sci 7:571–597
Soliman MH, Omar HS, El-Awady MA, Al-Assal S, Gamal El-Din AY (2009) Transformation and expression of Na+/H+ antiporter vacuolar (AtNHX1) gene in tobacco plants under salt stress. Arab J Biotechnol 12:99–108
Sunarpi Horie T, Motoda J, Kubo M, Yang H (2005) Enhanced salt tolerance mediated by AtHKT1 transporter–induced Na+ unloading from xylem vessels to xylem parenchyma cells. Plant J 44:928–938
Takasaki H, Maruyama K, Kidokoro S, Ito Y, Fujita Y, Shinozaki K, Yamaguchi-Shinozaki K, Nakashima K (2010) The abiotic stress-responsive NAC-type transcription factor Os-NAC5 regulates stress-inducible genes and stress tolerance in rice. Mol Genet Genomics 284(3):173–183
Tan FQ, Hong T, Liang WJ, Long JM, Wu XM, Zhang HY, Guo WW (2015) Comparative metabolic and transcriptional analysis of a doubled diploid and its diploid citrus rootstock (C. junos cv. Ziyang xiangcheng) suggests its potential value for stress resistance improvement. BMC Plant Biol 15(1):89–103
Thilagavathy A, Devaraj VR (2016) Evaluation of appropriate reference gene for normalizationof microRNA expression by real-time PCR in Lablab purpureusunder abiotic stress conditions. Biologia 71(6):660–668
Upadhyay U, Singh P, Verma OP (2019) Role of microRNAs in regulating drought stress tolerance in maize. J Pharmacog Phytochem 8(3):328–331
Vieira SM, Silva TM, Glória MBA (2010) Influence of processing on the levels of amines and proline and on the physico-chemical characteristics of concentrated orange juice. Food Chem 119(1):7–11
Volkov V, Wang B, Dominy PJ, Fricke W, Amtmann A (2004) Thellungiella halophila, a salt-tolerant relative of Arabidopsis thaliana, possesses effective mechanisms to discriminate between potassium and sodium. Plant Cell Environ 27(1):1–14
Wang J, Li DL, Kang XY (2012) Induction of unreduced megaspores with high temperature during megasporogenesis in Populus. Ann For Sci 69:59–67
Wang X, Ma X, Wang H, Li BJ, Tang WQ (2015) Proteomic study of microsomal proteins reveals a key role for Arabidopsis annexin 1 in mediating heat stress-induced increase in intracellular calcium levels. Mol Cell Proteomics 14(3):686–694
Wang J, Ye Y, Xu M, Feng L, Xu L (2019a) Roles of the SPL gene family and miR156 in the salt stress responses of tamarisk (Tamarix chinensis). BMC Plant Biol 19:370–381
Wang Z, Zhao Z, Fan G, Dong Y, Deng M, Xu E, Zhai X, Cao H (2019b) A comparison of the transcriptomes between diploid and autotetraploid Paulownia fortunei under salt stress. Physiol Mol Biol Plant 25:1–11
Wang W, Chen Q, Xu S, Liu W, Zhu X, Song C (2020) Trehalose-6-phosphate phosphatase E modulates ABA-controlled root growth and stomatal movement in Arabidopsis. J Integr Plant Biol 62(10):1518–1534
Wu HJ, Ma YK, Chen T, Wang M, Wang XJ (2012) PsRobot: a web-based plant small RNA meta-analysis toolbox. Nucleic Acids Res 40(W1):W22–W28
Xia K, Wang R, Ou X, Fang Z, Tian C, Duan J, Wang Y, Mi Z, Zhang B (2012) OsTIR1 and OsAFB2 down regulation via osmiR393 over expression leads to more tillers, early flowering and less tolerance to salt and drought in rice. PLoS ONE 7:e30039
Xing LT, Yue Z, Hua L, Ting WU, Bin LW, Xia ZH (2010) Stable expression of Arabidopsis vacuolar Na+/H+antiporter gene AtNHX1, and salt tolerance in transgenic soybean for over six generations. Chinese Sci Bull 55:1127–1134
Xu EK, Fan GQ, Niu SY, Zhao ZL, Deng MJ, Dong YP, Wu KQ (2014) Transcriptome-wide profiling and expression analysis of diploid and autotetraploid Paulownia tomentosa × Paulownia fortunei under drought stress. PLoS ONE 9(11):e113313
Yang C, Zhao L, Zhang H, Yang Z, Wang H, Wen S, Zhang C, Rustgi S, von Wettstein D, Liu B (2014) Evolution of physiological responses to salt stress in hexaploid wheat. Proc Natl Acad Sci USA 111(32):11882–11887
Yang L, Han Y, Wu D, Yong W, Liu M, Su T, Wen X, Lu M, Wei Y, Sun J (2017) Salt and cadmium stress tolerance caused by overexpression of the Glycine Max Na+/H+ Antiporter (GmNHX1) gene in duckweed (Lemna turionifera 5511). Aquatic Toxicol Amsterdam 192:127–135
Yi M, Zhao Q, Tang J, Wang C (2011) A theoretical modeling for frequency modulation of Ca2+ signal on activation of MAPK cascade. Biophys Chem 157(1–3):33–42
Zelm EV, Zhang Y, Testerink C (2020) Salt tolerance mechanisms of plants. Annu Rev Plant Biol 71(24):1–31
Zhang X, Zou Z, Gong P, Zhang J, Ziaf K, Li H, Xiao F, Ye Z (2011) Over-expression of microRNA169 confers enhanced drought tolerance to tomato. Biotechnol Lett 33(2):403–409
Zhang X, Deng M, Fan G (2014) Differential transcriptome analysis between Paulownia fortunei and its synthesized autopolyploid. Int J Mol Sci 15(3):5079–5093
Zhang H, Cui F, Wu Y, Lou L, Xie Q (2015a) The RING finger ubiquitin E3 ligase SDIR1 targets SDIR1-INTERACTING PROTEIN1 for degradation to modulate the salt stress response and ABA signaling in Arabidopsis. Plant Cell 27(1):214–227
Zhang HY, Li W, Mao XG, Jing RL (2015b) Characterization of genomic sequence of a drought-resistant gene TaSnRK2.7 in wheat species. J Genet 94(2):299–304
Zhao B, Ge L, Liang R, Li W, Ruan K, Lin H, Jin Y (2009) Members of miR-169 family are induced by high salinity and transiently inhibit the NF-YA transcription factor. BMC Mol Biol 10(1):29–39
Acknowledgements
The research was supported by National Natural Science Foundation of China of "Molecular basis of the vegetative growth advantage in allotriploid poplar (31530012) and the Program of the Co-Construction with Beijing Municipal Commission of Education of China.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Dorothea Bartels.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.

Fig. S1
Sequencing of small RNA and distribution of conserved miRNA family. DC, FC and SC are diploid, FDR triploid and SDR triploid in the control group. DS, FS and SS are diploids, FDR triploid and SDR triploid in the salt-treatment group (GIF 40 KB)
Rights and permissions
About this article
Cite this article
Qiu, T., Du, K., Jing, Y. et al. Integrated transcriptome and miRNA sequencing approaches provide insights into salt tolerance in allotriploid Populus cathayana. Planta 254, 25 (2021). https://doi.org/10.1007/s00425-021-03600-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-021-03600-9