Abstract
A cytological comparative analysis of male meiocytes was performed for Arabidopsis wild type and the ahp2 (hop2) mutant with emphasis on ahp2’s largely uncharacterized prophase I. Leptotene progression appeared normal in ahp2 meiocytes; chromosomes exhibited regular axis formation and assumed a typical polarized nuclear organization. In contrast, 4′,6′-diamidino-2-phenylindole-stained ahp2 pachytene chromosome spreads demonstrated a severe reduction in stabilized pairing. However, transmission electron microscopy (TEM) analysis of sections from meiocytes revealed that ahp2 chromosome axes underwent significant amounts of close alignment (44% of total axis). This apparent paradox strongly suggests that the Ahp2 protein is involved in the stabilization of homologous chromosome close alignment. Fluorescent in situ hybridization in combination with Zyp1 immunostaining revealed that ahp2 mutants undergo homologous synapsis of the nucleolus-organizer-region-bearing short arms of chromosomes 2 and 4, despite the otherwise “nucleus-wide” lack of stabilized pairing. The duration of ahp2 zygotene was significantly prolonged and is most likely due to difficulties in chromosome alignment stabilization and subsequent synaptonemal complex formation. Ahp2 and Mnd1 proteins have previously been shown, “in vitro,” to form a heterodimer. Here we show, “in situ,” that the Ahp2 and Mnd1 proteins are synchronous in their appearance and disappearance from meiotic chromosomes. Both the Ahp2 and Mnd1 proteins localize along the chromosomal axis. However, localization of the Ahp2 protein was entirely foci-based whereas Mnd1 protein exhibited an immunostaining pattern with some foci along the axis and a diffuse staining for the rest of the chromosome.








Similar content being viewed by others
References
Armstrong SJ, Caryl AP, Jones GH, Franklin FCH (2002) Asy1, a protein required for meiotic chromosome synapsis, localizes to axis-associated chromatin in Arabidopsis and Brassica. J Cell Sci 115:3645–3655
Armstrong SJ, Franklin FCH, Jones GH (2003) A meiotic time-course for Arabidopsis thaliana. Sex Plant Reprod 16:141–149
Azumi Y, Liu D, Zhao D, Li W, Wang G, Hu Y, Ma H (2002) Homolog interaction during meiotic prophase I in Arabidopsis requires the SOLO DANCERS gene encoding a novel cyclin-like protein. EMBO J 21:3081–3095
Bleuyard J-Y, White CI (2004) The Arabidopsis homologue of Xrcc3 plays an essential role in meiosis. EMBO J 23:439–449
Borde V (2007) The multiple roles of the Mre11 complex for meiotic recombination. Chomosom Res 15:551–563
Cai X, Dong F, Edelmann RE, Makaroff CA (2003) The Arabidopsis SYN1 cohesin protein is required for sister chromatid arm cohesion and homologous chromosome pairing. J Cell Sci 116:2999–3007
Chen C, Zhang W, Timofejeva L, Gerardin Y, Ma H (2005) The Arabidopsis ROCK-N- ROLLERS gene encodes a homolog of the yeast ATP-dependent DNA helicase MER3 and is required for normal meiotic crossover formation. Plant J 43:321–334
Chi P, Filippo JS, Sehorn MG, Petukhova GV, Sung P (2007) Bipartite stimulatory action of the Hop2–Mnd1 complex on the Rad51 recombinase. Genes Dev 21:1747–1757
Couteau F, Belzile F, Horlow C, Grandjean O, Vezon D, Doutriaux M (1999) Random chromosome segregation without meiotic arrest in both male and female meiocytes of a dmc1 mutant of Arabidopsis. Plant Cell 11:1623–1634
Dernburg AF, McDonald K, Moulder G, Barstead R, Dresser M, Villeneuve AM (1998) Meiotic recombination in C. elegans initiates by a conserved mechanism and is dispensable for homologous chromosome synapsis. Cell 94:387–398
Enomoto R, Kinebuchi T, Sato M, Yagi H, Kurumizaka H, Yokoyama S (2006) Stimulation of DNA strand exchange by the human TBPIP/Hop2–Mnd1 complex. J Biol Chem 281:5575–5581
Fiala JC (2005) Reconstruct: a free editor for serial section microscopy. J Microscopy 218:52–61
Fransz P, Armstrong S, Alonso-Blanco C, Fischer TC, Torres-Ruiz RA, Jones G (1998) Cytogenetics for the model system Arabidopsis thaliana. Plant J 13:867–876
Henderson KA, Keeney S (2004) Tying synaptonemal complex initiation to the formation and programmed repair of DNA double-strand breaks. PNAS 101:4519–4524
Higgins JD, Armstrong SJ, Franklin FCH, Jones GH (2004) The Arabidopsis MutS homolog AtMSH4 functions at an early step in recombination: evidence for two classes of recombination in Arabidopsis. Genes Dev 18:2557–2570
Higgins JD, Sanchez-Moran E, Armstrong SJ, Jones GH, Franklin FCH (2005) The Arabidopsis synaptonemal complex protein ZYP1 is required for chromosome synapsis and normal fidelity of crossing over. Genes Dev 19:2488–2500
Jackson N, Sanchez-Moran E, Buckling E, Armstrong SJ, Jones GH, Franklin FCH (2006) Reduced meiotic crossovers and delayed prophase I progression in AtMLH3-deficient Arabidopsis. EMBO 25:1315–1323
Keeney S, Giroux CN, Kleckner N (1997) Meiosis-specific DNA double strand breaks are catalyzed by Spo11, a member of a widely conserved protein family. Cell 88:375–384
Kerzendorfer C, Vignard J, Pedrosa-Harand A, Siwiec T, Akimcheva S, Jolivet S, Sablowski R, Armstrong S, Schweizer D, Mercier R, Schlogelhofer P (2006) The Arabidopsis thaliana MND1 homologue plays a key role in meiotic homologous pairing, synapsis and recombination. J Cell Sci 119:2486–2496
Lam WS, Yang X, Makaroff CA (2005) Characterization of Arabidopsis thaliana SMC1 and SMC3: evidence that AtSMC3 may function beyond chromosome cohesion. J Cell Sci 118:3037–3048
Leu J-Y, Chua PR, Roeder GS (1998) The meiosis-specific Hop2 protein of S. cerevisiae ensures synapsis between homologous chromosomes. Cell 94:375–386
Li W, Chen C, Markmann-Mulisch U, Timofejeva L, Schmelzer E, Ma H, Reiss B (2004) The Arabidopsis AtRAD51 gene is dispensable for vegetative development but required for meiosis. PNAS 101:10596–10601
Liu Z, Makaroff CA (2006) Arabidopsis separase AESP is essential for embryo development and the release of cohesin during meiosis. Plant Cell 18:1213–1225
Lysak M, Fransz P, Schubert I (2006) In: Salinas J, Sanchez-Serrano J (eds) Arabidopsis protocols, vol 323, 2nd edn, Methods in molecular biology. Humana, Totowa, pp 173–186
MacQueen AJ, Phillips CM, Bhalia N, Weiser P, Villeneuve AM, Dernburg AF (2005) Chromosome sites play dual roles to establish homologous synapsis during meiosis in C. elegans. Cell 123:1037–1050
Maniatus T, Fritsch EF, Sambrook J (1989) Molecular cloning: a laboratory manual, vol 3, 2nd edn. Cold Spring Harbor Laboratory, Cold Spring Harbor, pp 18-18–18-19
McKee BD (1996) The license to pair: identification of meiotic pairing sites in Drosophila. Chromosoma 105:135–141
McKee BD, Karpen GH (1990) Drosophila ribosomal RNA genes function as an X–Y pairing site during male meiosis. Cell 61:61–72
McKee BD, Habera L, Vrana JA (1992) Evidence that intergenic spacer repeats of Drosophila melanogaster rDNA genes function as XY pairing sites in male meiosis, and a general model for achiasmatic pairing. Genetics 132:529–544
Osman K, Sanchez-Moran E, Higgins JD, Jones GH, Franklin FCH (2006) Chromosome synapsis in Arabidopsis: analysis of the transverse filament protein ZYP1 reveals novel functions for the synaptonemal complex. Chromosoma 115:212–219
Ottaviani D, Lever E, Takousis P, Sheer D (2008) Anchoring the genome. Genome Biol 9:201–206
Petukhova GV, Romanienko PJ, Camerini-Otero RD (2003) The Hop2 protein has a direct role in promoting interhomolog interactions during mouse meiosis. Dev Cell 5:927–936
Petukhova GV, Pezza RJ, Vanevski F, Ploquin M, Masson JY, Camerini-Otero RD (2005) The Hop2 and Mnd1 proteins act in concert with Rad51 and Dmc1 in meiotic recombination. Nat Struct Mol Biol 12:449–453
Pezza RJ, Voloshin ON, Vanevski F, Camerini-Otero RD (2007) Hop2/Mnd1 acts on two critical steps in Dmc1-promoted homologous pairing. Genes Dev 21:1758–1766
Phillips CM, Dernburg AF (2006) A family of zinc-finger proteins is required for chromosome-specific pairing and synapsis during meiosis in C. elegans. Dev Cell 11:817–829
Phillips CM, Wong C, Bhalla N, Carlton PM, Meneely P, Dernburg AF (2005) HIM-8 binds to the X chromosome pairing center and mediates chromosome-specific meiotic synapsis. Cell 123:1051–1063
Richardson C, Horikoshi N, Pandita TK (2004) The role of the DNA double-strand break response network in meiosis. DNA Repair 3:1149–1164
Rockmill B, Sym M, Scherthan H, Roeder GS (1995) Roles for two RecA homologs in promoting meiotic chromosome synapsis. Genes Dev 9:2684–2695
Roque A, Orrego M, Ponte I, Suau P (2004) The preferential binding of histone H1 to DNA scaffold-associated regions is determined by its C-terminal domain. Nucleic Acids Res 32:6111–6119
Sanchez-Moran E, Armstrong S, Santos JL, Franklin FC, Jones GH (2001) Chiasma formation in Arabidopsis thaliana accession Wassileskija and in two meiotic mutants. Chromosome Res 9:121–128
Schommer C, Beven A, Lawrenson T, Shaw P, Sablowski R (2003) Ahp2 is required for bivalent formation and segregation of homologous chromosomes in Arabidopsis meiosis. Plant J 36:1–11
Schwacha A, Kleckner N (1997) Interhomolog bias during meiotic recombination: meiotic functions promote a highly differentiated interhomolog-only pathway. Cell 90:1123–1135
Sherizen D, Jang JK, Bhagat R, Kato N, McKim KS (2005) Meiotic recombination in Drosophila females depends on chromosome continuity between genetically defined boundaries. Genetics 169:767–781
Shimada M, Nabeshima K, Tougan T, Nojima H (2002) The meiotic recombination checkpoint is regulated by checkpoint rad (+) genes in fission yeast. EMBO J 21:2807–2818
Stevens R, Grelon M, Vezon D, Oh J, Meyer P, Perennes C, Domenichini S, Bergounioux C (2004) A CDC45 homolog in Arabidopsis is essential for meiosis, as shown by RNA interference-induced gene silencing. Plant Cell 16:99–113
Stronghill P, Hasenkampf C (2007) Analysis of substage associations in prophase I of meiosis in floral buds of wild-type Arabidopsis thaliana (Brassicaceae). Am J Bot 94:2063–2067
Tsubouchi H, Roeder S (2002) The Mnd1 protein forms a complex with Hop2 to promote homologous chromosome pairing and meiotic double strand break repair. Mol Cell Biol 22:3078–3088
Vignard J, Siwiec T, Chelysheva L, Vrielynck N, Gonord F, Armstrong SJ, Schlogelhofer P, Mercier R (2007) The interplay of RecA-related proteins and the MND1–HOP2 complex during meiosis in Arabidopsis thaliana. PloS Genet 3:1894–1905
von Wettstein D, Rasmussen SW, Holm PB (1984) The synaptonemal complex in genetic segregation. Ann Rev Genet 18:331–413
Wijeratne A, Chen C, Zhang W, Timofejeva L, Ma H (2006) The Arabidopsis thaliana PARTING DANCERS gene encoding a novel protein is required for normal meiotic homologous recombination. Mol Biol Cell 17:1331–1343
Zhao D, Ni W, Feng B, Han T, Petrasek MG, Ma H (2003) Members of the Arabidopsis- SKP1-like gene family exhibit a variety of expression patterns and may play diverse roles in Arabidopsis. Plant Physiol 133:203–217
Zickler D (2006) From early homologue recognition to synaptonemal complex formation. Chromosoma 115:158–174
Zickler D, Kleckner N (1999) Meiotic chromosomes: integrating structure and function. Annu Rev Genet 33:603–754
Acknowledgements
We would like to thank the reviewers for their insightful comments. We thank Dr. Jones for providing antibodies to Zyp1 and Asy1, Dr. Mercier for antibodies to Mnd1, and Dr. Makaroff for antibodies to Syn1 and Smc3. We would like to thank both Dr. Riggs and Dr. Siddiqui for their invaluable assistance in generating an antibody to the Ahp2 protein. We are also grateful to Dr. Sablowski for sending us seeds heterozygous for the ahp2 mutation. We also wish to thank Dr. Lysak for both FISH probes and valuable advice. This work was supported by a grant from the National Science and Engineering Research Council to CAH.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by: P. Shaw
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplemental Figs. 1–3
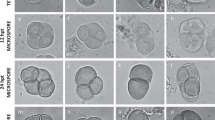
Supplemental Fig. 1. Representative negative control images from Ahp2 immunocytology experiments. Note: There was no labeling of wild-type and ahp2 chromosome spreads with preimmune serum and Ahp2 antibody, respectively. Scale bar 10 μm. Supplemental Fig. 2. Representative negative control images from Zyp1, Asy1, Syn1, and Smc3 immunocytochemistry experiments. Scale bar 10 μm. Supplemental Fig. 3. DAPI-stained Arabidopsis chromosome spreads from male meiocytes: a–c, g–i, and m, n wild type, d–f, j–l, and o, p ahp2 mutant. The prophase I substages are as follows: a, d mid-leptotene, b, e mid-zygotene, c, f mid-pachytene, g, j mid-diplotene, h, k diakinesis, i, l metaphase I, m, o anaphase I, and n, p anaphase II. Note: the ahp2 spreads were generated from buds of increasing size from the same inflorescence. Short white arrows indicate unpaired centromeres; long white arrows indicate paired centromeres and the arrowhead indicates the NOR. Scale bar 10 μm (GIF 2 kb)
Supplemental Figs. 4–6
Supplemental Fig. 4. Examples of chromatin bridging during anaphase II in an ahp2 meiocyte (see arrows). The chromosome spread was stained with DAPI. Scale bar 10 μm. Supplemental Fig. 5. S-phase BrdU pulse label time course experiment results for wild-type and ahp2 meiocytes: a A schematic summarizing both the average times for BrdU label to appear in tetrads and the average duration of meiosis in wild-type and ahp2 meiocytes. The bracketed times apply to the ahp2 meiocytes. L leptotene; TII telophase II (tetrads). b A table of the average times and the range of times measured from the mid-point of the BrdU pulse until peak BrdU signal was observed in the tetrads from the three wild-type and three ahp2 time course experiments performed. c A table of the raw data from the three wild-type BrdU experiments. d A table of the raw data from the three ahp2 mutant BrdU experiments. NTF no tetrads found, NA not assayed. Supplemental Fig. 6. Additional images demonstrating Mnd1 labeling of Arabidopsis ahp2 chromosome spreads and the nucleolus. Mnd1 antibody did not label mnd1 chromosome spreads. Note: All experimental parameters were kept constant for wild type and ahp2and mnd1 mutants; this included primary/secondary antibody dilutions and incubation times/conditions. Scale bar 10 μm (GIF 2 kb)
Supplemental Figs. 7–9
Supplemental Fig. 7. Fluorescent immunolocalization of Asy1 protein to Arabidopsis chromosome spreads: a–d wild type, e–h ahp2 meiocytes. The images shown are the result of merging the image of Asy1 protein (green) localization signal and the DAPI-counterstained (white) image of the same chromosome spread. The prophase I substages are as follows: a early leptotene, e late leptotene, b, f zygotene, c, g pachytene, and d, h diplotene. Scale bar 10 μm. TEM images of wild-type (i) and ahp2 (j) meiocyte sections provide representative examples of unpaired chromosomal axis (black arrows). Scale bar 1 μm. Supplemental Fig. 8 DAPI-stained pachytene ahp2 chromosome spread images. Arrows indicate the region of pairing that extends from the centromere to the NOR. Scale bar 10 μm. Supplemental Fig. 9. Fluorescent immunolocalization of the meiotic cohesin proteins Syn1 (a–j) and Smc3 (k–t) on Arabidopsis wild-type and ahp2 chromosome spreads counterstained with DAPI (white). The Syn1 antibody (green) was localized to both wild-type (a–e) and ahp2 nuclei (f–j) of the following substages: a, f leptotene, b, g mid-zygotene, c, h mid-pachytene, d, i diplotene, and e, j postdiakinesis. Scale bar 10 μm. The micrographs shown are merged DAPI and Syn1 images. The Smc3 antibody (green) was localized to both wild-type (k–o) and ahp2 nuclei (p–t) of the following substages: k, p leptotene, l, q mid-zygotene, m, r mid-pachytene, n, s diakinesis, and o, t postdiakinesis. The micrographs shown are merged DAPI and Smc3 images. Scale bar 10 μm (JPEG 258 kb)
Supplemental Fig. 10
A clustalW alignment of Ahp2 protein orthologs from several plant and animal species. Bolded sequence denotes the DNA minor groove binding motif (GIF 153 kb)
Rights and permissions
About this article
Cite this article
Stronghill, P., Pathan, N., Ha, H. et al. Ahp2 (Hop2) function in Arabidopsis thaliana (Ler) is required for stabilization of close alignment and synaptonemal complex formation except for the two short arms that contain nucleolus organizer regions. Chromosoma 119, 443–458 (2010). https://doi.org/10.1007/s00412-010-0270-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-010-0270-0