Abstract
Conventional genetic approaches and computational strategies have converged on immune-inflammatory pathways as key events in the pathogenesis of late onset sporadic Alzheimer’s disease (LOAD). Mutations and/or differential expression of microglial specific receptors such as TREM2, CD33, and CR3 have been associated with strong increased risk for developing Alzheimer’s disease (AD). DAP12 (DNAX-activating protein 12)/TYROBP, a molecule localized to microglia, is a direct partner/adapter for TREM2, CD33, and CR3. We and others have previously shown that TYROBP expression is increased in AD patients and in mouse models. Moreover, missense mutations in the coding region of TYROBP have recently been identified in some AD patients. These lines of evidence, along with computational analysis of LOAD brain gene expression, point to DAP12/TYROBP as a potential hub or driver protein in the pathogenesis of AD. Using a comprehensive panel of biochemical, physiological, behavioral, and transcriptomic assays, we evaluated in a mouse model the role of TYROBP in early stage AD. We crossed an Alzheimer’s model mutant APP KM670/671NL /PSEN1 Δexon9 (APP/PSEN1) mouse model with Tyrobp −/− mice to generate AD model mice deficient or null for TYROBP (APP/PSEN1; Tyrobp +/− or APP/PSEN1; Tyrobp −/−). While we observed relatively minor effects of TYROBP deficiency on steady-state levels of amyloid-β peptides, there was an effect of Tyrobp deficiency on the morphology of amyloid deposits resembling that reported by others for Trem2 −/− mice. We identified modulatory effects of TYROBP deficiency on the level of phosphorylation of TAU that was accompanied by a reduction in the severity of neuritic dystrophy. TYROBP deficiency also altered the expression of several AD related genes, including Cd33. Electrophysiological abnormalities and learning behavior deficits associated with APP/PSEN1 transgenes were greatly attenuated on a Tyrobp-null background. Some modulatory effects of TYROBP on Alzheimer’s-related genes were only apparent on a background of mice with cerebral amyloidosis due to overexpression of mutant APP/PSEN1. These results suggest that reduction of TYROBP gene expression and/or protein levels could represent an immune-inflammatory therapeutic opportunity for modulating early stage LOAD, potentially leading to slowing or arresting the progression to full-blown clinical and pathological LOAD.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Conventional wisdom has held that the chronic neuroinflammation associated with LOAD may be a secondary or even protective event that occurs in response to Aβ deposition and may occur only in late stages of AD. However, recent genetic and genomic approaches, as well as computational strategies, have converged on immune-inflammatory pathways as risk factors and as key events in the pathogenesis of late-onset sporadic Alzheimer’s disease (LOAD) [19]. Moreover, correlation between inflammatory genes and clinical presentation of previously asymptomatic cerebral amyloidosis (ACA) indicates a role for inflammation and microglia in the progression from ACA to the earliest stages of mild cognitive impairment (MCI) and/or mild clinical AD. Among the genes implicated by the largest available genome-wide association studies [43], one-third is either unique to, or enriched in, microglia. Recently identified mutations and variants in genes encoding important immune receptors including CD33, CR3 (Complement Receptor 3), and TREM2 (Triggering Receptor Expressed On Myeloid Cells 2), have been genetically linked to LOAD risk, highlighting the potential role of a dysregulated immune response in an early, and perhaps causative role in AD pathogenesis. Unlike autosomal dominant familial Alzheimer’s mutations that promote elevation of the Aβ42:40 ratio or of other variant hyperaggregatable Aβ species, these AD risk factors specify some of the cell surface signal transduction pathways that modulate the phagocytic machinery of microglia.
TYROBP (TYROsine kinase Binding Protein) (also known as DAP12), is a microglial transmembrane signaling polypeptide that contains an immunoreceptor phosphotyrosine-based activation motif (ITAM) in its cytoplasmic domain and is a direct partner/adaptor for immune receptors, including TREM2, CR3, and SIRPβ1 (Signal Regulatory Protein β1) all of which are independently linked to, or associated with, LOAD [5, 7, 23, 55, 86]. Interaction of TYROBP with its partners forms phagocytosis “active zones” (known as phagocytic synapses) on the surface of microglia. In preparation for phagocytosis, there is a respiratory burst that generates reactive oxygen species (ROS) and appears to involve an interaction between TYROBP and CR3, which in turn interacts with complement component C3 associated with nearby neurites. Mice lacking the complement receptor CR3 or expressing defective TYROBP show reduced ROS production and apoptosis [77]. A recent report demonstrates that the complement pathway can mediate the toxic effects of soluble Aβ on synapses, and that overactivation of this pathway in AD leads to excessive synapse pruning and early synapse loss [25]. Since the discovery of a link between mutations of TREM2 and AD, several studies have emerged regarding the role of a loss of function of TREM2 in AD. While these studies have some conflicting results, the most consistent observation is that either Trem2 deficiency or Tyrobp deficiency can cause reduced recruitment of microglial cells around Aβ plaques. The impact of this reduction in microglia per plaque was interpreted as deleterious in Trem2 haploinsufficient and Trem2 deficient mice.
Through a multi-scale integrated computational approach, we and two other independent groups [12, 48, 86] have previously reported TYROBP as a network hub or driver gene in LOAD. Additionally, missense mutations in TYROBP have been recently reported as risk factors for AD [61]. Evidence associating TYROBP to LOAD notwithstanding, it is important to recognize that most TYROBP mutations (as well as TREM2 mutations) represent loss-of-function mutations that result not in AD but in an osteopathy/encephalopathy known as Nasu–Hakola disease (NHD) [59]. One formulation of these data is that the pathogenic mechanism(s) of loss-of-function (nonsense) mutations in TYROBP associated with NHD may cause molecular events that differ from those associated with missense polymorphisms that increase the risk for AD.
Herein, we report the effects of a constitutively null mutation in Tyrobp on the phenotype of an APP/PSEN1 mouse model of AD. In the Tyrobp-null mouse, there is a deletion of exons 3 and 4 resulting in loss of function of the TYROBP protein by deletion of the transmembrane region and part of the cytoplasmic region including the first tyrosine of the ITAM motif [2]. The APP/PSEN1 mouse model [29] expresses APP KM670/671NL /PSEN1 Δexon9 in neurons and accumulates in the interstitial spaces of the brain fibrillar amyloid that goes on to form typical amyloid plaques accompanied by neuritic dystrophy, age-dependent synaptic loss without neuronal loss, and abnormalities in spatial memory [25, 29, 40, 41]. Since TYROBP expression is increased in the LOAD brain [86], we hypothesized that the APP/PSEN1 phenotype may be improved in the presence of reduced TYROBP levels. Since Tyrobp is not expressed in neurons, our observations in this report describe non-cell autonomous effects wherein signals arising from microglia perturb the homeostasis of nearby neurons or nerve terminals or the pathophysiology of evolving structural intraneuronal or extracellular Alzheimer’s pathology.
Methods
Mouse husbandry
The experimental procedures were conducted in accordance with NIH guidelines for animal research and were approved by the Institutional Animal Care and Use Committee (IACUC) at Icahn School of Medicine at Mount Sinai. APP KM670/671NL /PSEN1 Δexon9 (APP/PSEN1) and Tyrobp knockout (KO) mice were obtained from Jackson Laboratories and Taconic/Merck Laboratory, respectively. APP/PSEN1 mice were crossed with Tyrobp KO mice to obtain APP/PSEN1 mice heterozygous or KO for Tyrobp. Four-month-old male and female mice were killed by decapitation. One hemisphere was collected for immunohistochemical analysis. The second hemisphere was collected for transcriptomic and biochemical analyses.
Immunohistochemical and biochemical analyses
Immunohistochemical and biochemical characterization were performed as previously described [40, 41, 44, 76]. For biochemical analysis, hemibrains were processed via differential detergent solubilization to produce TBS-soluble, Triton-X-soluble, and formic-acid soluble Aβ fractions. For analysis of native oligomeric Aβ peptides, 2 μl protein samples from the TBS-soluble fraction were spotted onto activated/pre-wetted PVDF membrane (0.22 μm; Millipore, Billerica, MA). Membranes were incubated with rabbit pAb A11 (anti-prefibrillar oligomers, 0.5 μg/ml), rabbit pAb OC (anti-fibrillar oligomers and fibrils; 0.25 μg/ml), and mouse mAb Nu-4 (anti-oligomers; 1 μg/ml) [44, 76]. Normalization to total APP/Aβ signal was achieved by detection of human APP transgene metabolites with the mouse pAb 6E10 antibody (1:1000; Covance, Princeton, NJ). To quantify total Aβ levels, human/rat Aβ 1–40/1–42 ELISA kits (Wako) were used according to the manufacturer’s instructions.
For immunohistochemistry, 30 µm thick sagittal sections were incubated with the following antibodies: rabbit anti-Iba1 (1:500; Wako, Richmond, VA), mouse anti-6E10 (1:1000; Covance, Princeton, NJ), and rat anti-CD68 (1:200, mca1957, AbD Serotec BioRad). Sections were then incubated with the appropriate secondary antibody: anti-rabbit Alexa Fluor 488 or Alexa Fluor 568 (1:400; Invitrogen, Carlsbad, CA), anti-mouse Alexa Fluor 568 (1:400; Invitrogen, Carlsbad, CA), and anti-rat Alexa Fluor 488 (1:400; Invitrogen, Carlsbad, CA) antibodies. ThioflavinS (Sigma-Aldrich, T1892, 1% w/v stock solution) was used for labeling amyloid deposits.
For measuring microglia number, Iba1-immunolabeled sections were thresholded and particles analyzed with Fiji (v2.0.0). Sizes of 6E10 immunoreactive plaques and fluorescent intensities were analyzed with Fiji (v2.0.0). The regions of interest were determined by manual tracing. Thioflavin S fluorescence intensity and circularity were analyzed as described [85].
For immunoblotting, membranes were incubated with either anti-CD68 (1:1000, mca1957, AbD Serotec BioRad), anti-phospho-Tau pSer202/Thr205 (1:1000; MN1020, Thermo Fisher Scientific, Waltham, MA), anti-Tau (1:1000; MN1000, Thermo Fisher Scientific, Waltham, MA), anti-Synaptophysin (1:200; ab16659, Abcam, Cambridge, MA), anti-Lamp1 (1:200; ab24170, Abcam, Cambridge, MA), anti-C3 (1:50; ab11862, Abcam, Cambridge, MA), and anti-GAPDH (1:5000; sc32233, Santa Cruz, Dallas, TX) antibodies. Integrated density of immunoreactive bands was measured using MultiGauge Software (FujiFilm). At least two independent western blot analyses were performed and normalized using APP/PSEN1 female mice as controls.
Behavior analysis
The Barnes Maze test was performed using a standard apparatus [3, 74]. Four-month-old mice were transported from their cage to the center of the platform via a closed starting chamber where they remained for 10 s prior to exploring the maze for 3 min. Mice failing to enter the escape box within 3 min were guided to the escape box by the experimenter, and the latency was recorded as 180 s. Mice were allowed to remain in the escape box for 1 min before the next trial. Two trials per day during 4 consecutive days were performed. The platform and the escape box were wiped with 70% ethanol after each trial to eliminate the use of olfactory cues to locate the target hole. All trials were recorded by video camera and analyzed with ANY-maze video tracking software (Stoelting Co, Wood Dale, USA).
Field electrophysiology
Coronal brain slices containing the hippocampal formation were prepared as previously described [17]. Following anesthesia with isoflurane, brains were rapidly removed and cut into 400 µm thick coronal sections using a vibratome VT1000S (Leica Microsystems, Germany). Brain slices were incubated at room temperature for ≥3 h in a physiologic ACSF containing 120 mM NaCl, 3.3 mM KCl, 1.2 mM Na2HPO4, 26 mM NaHCO3, 1.3 mM MgSO4, 1.8 mM CaCl2, 11 mM Glucose (pH 7.4) and then transferred to a recording chamber perfused with ACSF at a flow rate of ~2 mL/min; experiments were performed at 28.0 ± 0.1 °C. Recordings were acquired with a GeneClamp 500B amplifier (Axon Instruments, Union City, CA) and Digidata 1440A (Molecular Devices, Sunnyvale, CA). All signals were low-pass filtered at 2 kHz and digitized at 10 kHz. For extracellular field recordings, a patch-type pipette was filled with ACSF and placed in the middle third of stratum radiatum in area CA1. Field excitatory postsynaptic potentials (fEPSPs) were evoked by activating Shaffer Collaterals with a Concentric Bipolar Electrode stimulator (FHC, St Bowdoin, ME) placed in the middle third of stratum radiatum 150–200 µm away from the recording pipette. Square-wave current pulses (60 ms pulse width) were delivered through a stimulus isolator (Isoflex, AMPI). Input–output curves were generated by a series of stimuli in 0.1 mA steps. Paired-pulse facilitation was measured by delivering two stimuli at 20, 50, and 100 ms inter-stimulus intervals. Each inter-stimulus interval was repeated three times and the resulting potentials were averaged. The paired-pulse ratio was calculated by dividing the slope of the second EPSP by the slope of the first EPSP. All results were analyzed by ANOVAs followed by Tukey post hoc tests. Baseline recordings (stable for 20 min) were made every 30 s using stimuli that yielded a response equal to 50% of spike threshold. LTD was induced using a 1-Hz train of 900 bursts, each burst containing three stimuli delivered at 20 Hz, using stimulus strength just superthreshold for evoking a population spike during baseline.
Molecular biological analyses
RNA isolation, library preparation, differential expression analysis and gene set enrichment analyses were performed as described [6, 26, 27, 64, 66, 67].
Computational screen of TYROBP regulating compounds
Drug-induced gene expression fold change was obtained from the Connectivity Map database [42], which consists of 6100 individual experiments, representing 1309 unique compounds. The 6100 individual expression profiles were merged into a single representative signature for the 1309 unique compounds, according to the prototype-ranked list method [28]. Each compound was scored according to the rank of Tyrobp expression fold change within its signature. Compounds were ranked in descending order of Tyrobp expression fold change and used for a secondary enrichment analysis of drug-target associations. For each compound in the drug signature library, referenced drug–target associations [45, 83] and predicted off-targeting [36, 37] were collected. For each of these features, we calculated a running sum enrichment score, reflecting whether that feature was over-represented among the compounds at the top (associated with Tyrobp upregulation) or at the bottom (associated with Tyrobp down-regulation). Two-tailed p values were based on comparison with 10,000 permuted null scores, generated from randomized drug target sets that contain an equivalent number of compounds to the true set under evaluation, and adjusted using the Benjamini–Hochberg method [6]. Computational screening and chemogenomic enrichment analysis were performed using the R project for statistical computing version 3.2.5 [62].
Data and software availability
Gene expression data generated contributing to the described study will be deposited electronically to the Synapse Web Portal (https://www.synapse.org) in accordance with data sharing policies established by the NIH Accelerating Medicine Partnership (AMP) AD consortium. Specific software will also be made available upon request.
Results
TYROBP deficiency or absence does not modify the number and size of Aβ plaque depositions nor the number of microglial cells in prefrontal cortex and hippocampus of APP/PSEN1 mice
We assessed whether TYROBP deficiency or absence modulates Aβ deposition in APP/PSEN1 mice. We did not observe differences in number or size of 6E10 immunoreactive plaques in cortices or in the hippocampi of APP/PSEN1 mice heterozygous or KO for Tyrobp as compared to APP/PSEN1 mice with normal levels of TYROBP (Fig. 1a–c). It is important to note that 4-month-old APP/PSEN1 mice represent an early time point of AD pathology, and all genotypes presented very little Aβ deposition in the hippocampus as compared to the cortex.
A decrease in TYROBP protein impairs Aβ deposits compaction, microglial activation, and recruitment around Aβ deposits in 4-month-old APP/PSEN1 mice. a Images of Iba1-immunostained microglia (green) and 6E10-immunoreactive plaques (red) in frontal cortices and hippocampi of APP/PSEN1, APP/PSEN1; Tyrobp +/− and APP/PSEN1; Tyrobp −/− mice. Arrows indicate location of the plaques. Scale bar 500 µm. b Quantification of the number of 6E10-immunoreactive Aβ deposits in cortices and hippocampi (Hip) of APP/PSEN1, APP/PSEN1; Tyrobp +/− and APP/PSEN1; Tyrobp −/− mice. c Measurements of the size of 6E10-immunoreactive Aβ deposits in cortices of APP/PSEN1 mice WT, deficient or null for Tyrobp. d Quantification of the number of Iba1-immunostained microglia in frontal cortices and hippocampi of APP/PSEN1, APP/PSEN1; Tyrobp +/− and APP/PSEN1; Tyrobp −/− mice. e, f Images of Iba1-immunostained microglia (green) and 6E10-immunoreactive plaques (red) and quantification of numbers of cortices plaque-associated microglia located on or within 30 µm radius of 6E10 immunoreactive Aβ plaques in APP/PSEN1, APP/PSEN1; Tyrobp +/− and APP/PSEN1; Tyrobp −/− mice. n = 3–4 mice per group. Scale bar 10 µm. g–i Images of thioflavin S-labeled amyloid plaques (g), circularity (h) and quantification of fluorescence intensity (i) of thioflavin S-labeled amyloid plaques from APP/PSEN1, APP/PSEN1; Tyrobp +/− and APP/PSEN1; Tyrobp −/− mice. n = 3–4 mice per group. Scale bar 5 µm. j, k Images of phagocytic microglial marker CD68 (green) and Iba1 (red) co-immunostaining (j) and quantification of fluorescence intensity of CD68 (k) in APP/PSEN1, APP/PSEN1; Tyrobp +/− and APP/PSEN1; Tyrobp −/− mice. n = 3–4 mice per group. Scale bar 30 µm. i Western blot analysis of CD68 in brain protein homogenates from APP/PSEN1, APP/PSEN1; Tyrobp +/− and APP/PSEN1; Tyrobp −/− mice. n = 3–6 mice per group. At least two independent western blot analyses were performed. Representative immunoreactive bands from the same western blot are shown on the right. One-way ANOVA corrected for multiple comparisons (Tukey) was used for (c, h, i, k) and Two-way ANOVA corrected for multiple comparisons (Tukey) was used for (b, d, f, l), *p < 0.05; ***p < 0.001; ****p < 0.0001. Data presented as mean ± SEM
Reduction in total number of microglia has been observed in older TREM2 KO mice with Aβ pathology [30, 81], most likely due to a reduction of microglia proliferation. No differences were observed in younger mice. In our hands, Iba1 immunostaining in 4-month-old APP/PSEN1 mice deficient or null for Tyrobp did not show differences in the total number of microglia in (pre)frontal cortices (PC) nor in hippocampi as compared to APP/PSEN1 mice with normal levels of TYROBP (Fig. 1a, d). Similar results were observed in WT mice with normal or absent TYROBP (see Suppl. Figure 1).
Loss of TYROBP reduces plaque compaction, microglia clustering, and phagocytosis
When 5XFAD mice, which develop rapid and aggressive amyloid pathology and neuronal loss [56], were rendered deficient or null for TYROBP or TREM2, microglial clustering around plaques and plaque compaction were reduced at 4 months of age [85]. We observed decreased microglial recruitment on and around antibody 6E10-immunoreactive Aβ deposits in the PC of 4-month-old APP/PSEN1;Tyrobp −/− mice as compared to APP/PSEN1 mice with a normal level of TYROBP (Fig. 1e, f). We also observed reduced compaction and fluorescence intensity of thioflavin S reactive plaques (Fig. 1g–i).
We next assessed by immunostaining the level of a phagocytic marker CD68 [88] in the PC of APP/PSEN1 mice WT, deficient or KO for TYROBP. In APP/PSEN1;Tyrobp −/− mice, we observed a decreased expression of CD68 in microglial Iba1-positive cells as compared to APP/PSEN1 mice (Fig. 1j–k). APP/PSEN1 mice heterozygous for Tyrobp did not present a statistically significant reduction of CD68 expression. Accordingly, the level of CD68 in hemibrain protein homogenates was lower in APP/PSEN1;Tyrobp −/− as compared with that observed in APP/PSEN1 mice (Figure i, j). These data support an interpretation that microglial phagocytic activity was reduced in AD mice in the absence of TYROBP (Table 1 for a summary of results).
Aβ levels and oligomeric Aβ in APP/PSEN1 mice deficient or null for TYROBP
We assessed whether TYROBP deficiency or absence modulates levels of Aβ species in APP/PSEN1 mice. We measured levels of Aβ40 and Aβ42 in TBS, Triton-X, and formic acid-soluble Aβ fractions from brains of 4-month-old male and female APP/PSEN1 mice on a Tyrobp heterozygous or null background (Suppl. Figure 2). In males, deletion of one or both Tyrobp alleles did not alter levels of Aβ40, Aβ42 or Aβ42/40 ratio in any of the three fractions as compared to male APP/PSEN1 mice (Suppl. Figure 2a–i). Female APP/PSEN1;Tyrobp −/− mice exhibited lower levels of Aβ40 in Triton-X and formic acid fractions relative to APP/PSEN1 mice, resulting in an increase in the Aβ42/40 ratio in the Triton-X fraction. This was not observed in the formic acid fraction (see Suppl. Figure 2a–i). Notably, female APP/PSEN1 mice WT, heterozygous or knockout (KO) for Tyrobp had higher levels of Aβ40 and 42 in the Triton-X and formic acid fractions when compared to genotype-matched males.
We next assayed oligomeric Aβ peptides using antibodies NU-4, A11, and OC antibodies to distinguish among various Aβ conformers (Fig. 2a–c). Higher levels of oligomeric Aβ reactive with these antibodies have been correlated with impaired cognitive performances in humans and mice [49]. We and others [40, 73] have reported an association of excess levels of NU4-epitope-containing oligomeric Aβ with deficits in learning behavior in AD mouse models. NU-4 reactive oligomer levels were reduced in both male and female APP/PSEN1 mice with deficiencies in TYROBP as compared to levels observed in APP/PSEN1 mice with normal TYROBP (Fig. 2a). A11 reactive oligomer levels were also reduced in female mice with reduced TYROBP as compared to APP/PSEN1 mice WT for Tyrobp (Fig. 2b). TYROBP level played no obvious role in determining levels of OC epitope-containing oligomeric Aβ in this system (Fig. 2c). (See Table 1 for a summary of results).
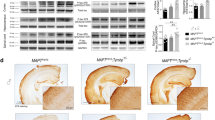
A decrease in TYROBP protein decreases oligomeric Aβ levels and alters phospho-TAU, synaptophysin, LAMP1, and complement C3 levels in 4-month-old APP/PSEN1 mice. a–c Hemibrains of male and female APP/PSEN1 (n = 4–6), APP/PSEN1; Tyrobp +/− (n = 3–8) and APP/PSEN1; Tyrobp −/− (n = 3–4) mice were processed via differential detergent solubilization to produce fractions of TBS soluble, Triton-X soluble, and formic acid soluble Aβ. Oligomeric Aβ was assessed from the TBS-soluble fraction via dot blot analyses using NU-4 (a), A11 (b) and OC (c) antibodies. d–i Western blot analysis in brain protein homogenates from 4-month-old male and female mice WT, Tyrobp +/−, Tyrobp −/− and APP/PSEN1, APP/PSEN1; Tyrobp +/− and APP/PSEN1; Tyrobp −/− mice. d, e Phospho-tau (AT8 epitope)/total tau ratio for d WT, Tyrobp +/−, Tyrobp −/− mice and e APP/PSEN1, APP/PSEN1; Tyrobp +/− and APP/PSEN1; Tyrobp −/− mice. f, g Synaptophysin level for f WT, Tyrobp +/−, Tyrobp −/− mice and g APP/PSEN1, APP/PSEN1; Tyrobp +/− and APP/PSEN1; Tyrobp −/− mice. h Marker of dystrophic neurites (Lamp1) in APP/PSEN1, APP/PSEN1; Tyrobp +/− and APP/PSEN1; Tyrobp −/− mice. i Complement C3 in APP/PSEN1, APP/PSEN1; Tyrobp +/− and APP/PSEN1; Tyrobp −/− mice. At least two independent western blot analyses were performed. Representative immunoreactive bands from the same western blot are shown on the right. n = 3–6 mice per group. Two-way ANOVA corrected for multiple comparisons (Tukey) was used for all statistical comparisons in male and female mice, *p < 0.05; **p < 0.01; ***p < 0.001, ****p < 0.0001. Data presented as mean ± SEM
Phospho-TAU, synaptophysin, LAMP1, and C3 levels are altered in APP/PSEN1 and WT mice with reduced or absent TYROBP
In addition to amyloid deposition, APP/PSEN1 mice develop hyperphosphorylated microtubule-associated protein TAU (MAPT). We assayed the phosphorylation status of MAPT in WT (nontransgenic) and APP/PSEN1 mice with normal, reduced, or absent TYROBP (Fig. 2d, e). We observed an apparent increased stoichiometry of TAU phosphorylation in male mice deficient for TYROBP as compared to WT mice (Fig. 2d). Females deficient for TYROBP demonstrated a trend toward increased phosphorylation of TAU as compared to WT mice (p = 0.07). In the presence of mutant APP/PSEN1 transgenes, there was a reduction in the stoichiometry of TAU phosphorylation in female mice with reduced or absent TYROBP, but no difference in male mice (Fig. 2e).
To examine synaptic integrity, we measured the levels of the presynaptic neuronal marker, synaptophysin (Fig. 2f, g). Synaptophysin was increased in male and female Tyrobp −/− mice as compared to WT mice (Fig. 2f), but no difference was observed between groups in APP/PSEN1 mice heterozygous-null or homozygous-null for Tyrobp (Fig. 2g). Notably, however, LAMP1, a lysosomal protein enriched in dystrophic neurites [15, 21], was decreased in both male and female APP/PSEN1; Tyrobp −/− mice relative to APP/PSEN1 alone (Fig. 2h). Excessive activation of the complement system is an early event in AD leading to synapse loss. The level of complement C3 was decreased in female APP/PSEN1 homozygous-null for Tyrobp relative to those expressing APP/PSEN1 alone (Fig. 2i). No difference was observed in corresponding male mice (see Table 1 for a summary of results). Despite the relatively minor effect size, likely due to the early stage of AD pathology in 4-month-old APP/PSEN1 mice, when taken together, these results are consistent with a conclusion that decreased expression of Tyrobp may have beneficial effects in the proteinopathy of AD.
Electrophysiological changes in APP/PSEN1 mice deficient for TYROBP
TYROBP, amyloid, and presenilin proteins play important roles in excitatory synaptic transmission at Shaffer collateral-CA1 pyramidal cell synapses [52, 68]. All of the recombinant mouse models tested in this report showed either altered basal synaptic function, reduced plasticity, or both. Basal synaptic efficiency, as measured by the slope of the input–output relationship, was normal in APP/PSEN1 and APP/PSEN1; Tyrobp −/− as compared to WT mice (Fig. 3a). Interestingly, the slope of the input–output relationship was increased in Tyrobp −/− mice compared to WT, APP/PSEN1, and APP/PSEN1; Tyrobp −/− suggesting an increased basal synaptic activity in absence of TYROBP (Fig. 3a).
A decrease in TYROBP protein alters excitatory synaptic transmission in the hippocampus in 4-month-old APP/PSEN1 mice and interacts with the APP/PSEN1 genotype. In all panels, summary graphs are shown on the left and representative traces on the right. a Basal synaptic function is increased in Tyrobp −/− mice, but is unaffected by other transgenic genotypes. The slope of the input/output relationship was steeper for the Tyrobp −/− mice than for all other genotypes (p < 0.05), which did not differ among themselves. b APP/PSEN1 mice showed reduced paired-pulse facilitation (PPF) relative to other genotypes, which did not differ among themselves. c Synaptically induced long-term depression (LTD) was impaired in all recombinant mice. Analysis over the final 5 min of the recordings showed the most profound deficits for Tyrobp −/− and APP/PSEN1;Tyrobp −/− mice, both of which were significantly less depressed than in APP/PSEN1 mice. Wild-type (WT) controls showed significantly greater depression than the other genotypes. One-way ANOVA corrected for multiple comparisons (Tukey) was used for statistical comparisons, *p < 0.05, **p < 0.01; ***p < 0.001. Data presented as mean as mean ± SEM
We tested the possibly that this effect of Tyrobp deletion was presynaptically mediated using paired-pulse facilitation (PPF), a short-term form of synaptic plasticity that is sensitive to the probability of transmitter release [9]. PPF was normal in Tyrobp −/− mice (Fig. 3b) suggesting that the increase in basal efficiency observed in Tyrobp −/− mice most likely reflects postsynaptic regulation such as increased expression and/or function of synaptic AMPA-type glutamate receptors (AMPARs). These data also raise the possibility of impaired endocytosis of AMPARs [87]. In contrast, PPF was depressed in APP/PSEN1 mice relative to WT controls, indicating an increase in transmitter release probability. Importantly, Tyrobp deletion reversed the deleterious effect of APP/PSEN1 on presynaptic function, since PPF was normal in slices from APP/PSEN1; Tyrobp −/− mice.
We also examined the effects of the different APP/PSEN1 and Tyrobp genotypes on long-term depression (LTD), a persistent form of plasticity whose expression depends on endocytosis of postsynaptic AMPARs [38]. For these experiments, we used a synaptic induction protocol that induces a prominent protein synthesis-dependent “late” phase of LTD [69]. Slices from the APP/PSEN1 mice showed impaired LTD (Fig. 3c), similar to that reported in older APP/PSEN1 mice. Similar results were observed following a weaker induction protocol [13] or when late LTD was induced by metabotropic glutamate receptor activation (mGluR-LTD) [84]. LTD was even more impaired in Tyrobp −/− and APP/PSEN1; Tyrobp −/− mice. Thus, unlike the phenotypes for basal efficiency and PPF, superimposition of TYROBP deficiency on the APP/PSEN1 mutations failed to restore normal LTD.
Barnes maze
We next probed the effect of Tyrobp deletion on the modulation of spatial learning and memory using the Barnes Maze Test (Fig. 4a–d). The escape latency and distance traveled of Tyrobp heterozygous- and homozygous-null mice were identical to WT littermates (Fig. 4a, b). In the presence of APP/PSEN1 mutations, deficiency of TYROBP improved learning and memory relative to APP/PSEN1 with normal TYROBP levels (Fig. 4c, d). This improvement was associated with a reduction in the time spent finding the hidden location (target hole) and a smaller distance traveled in all quadrants. These behavioral data are consistent with a beneficial effect of the Tyrobp deletion on the APP/PSEN1 phenotype.
A decrease in TYROBP protein improves spatial learning and memory in the Barnes Maze Test in 4-month-old APP/PSEN1 mice. a–d 6 groups of 4-month-old mice were used: wild-type (WT) (n = 14), Tyrobp +/− (n = 9), Tyrobp −/− (n = 10), APP/PSEN1 (n = 5), APP/PSEN1; Tyrobp +/− (n = 9) or APP/PSEN1; Tyrobp −/− (n = 11). a, b Mean latencies to enter the target hole for a APP/PSEN1 negative mice and b APP/PSEN1 positive mice. c, d Mean distances traveled for c APP/PSEN1 negative mice and d APP/PSEN1 positive mice. Two-way ANOVA corrected for multiple comparisons (Tukey) was used for all statistical comparisons, *p < 0.05; **p < 0.01; ***p < 0.001. Data presented as mean ± SEM
Differential gene expression analysis of Tyrobp −/− and Tyrobp +/− mice relative to WT mice
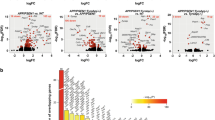
Given our extensive database on the regional and disease-stage-specific transcriptomic changes in human LOAD [64, 79, 86], we began by investigating how Tyrobp deletion perturbed brain regional transcriptomes. We generated transcriptomic profiles from (pre)frontal cortices (PC) and dentate gyri (DG) (n = 24) for 4-month-old female and male Tyrobp −/− and Tyrobp +/− mice and compared with WT mice, including sex as a variable. In comparison with non-recombinant (WT) mice, we identified 10 differentially expressed genes (DEG) in the PC, and 28 DEG in the DG of Tyrobp −/− mice [false discovery rate (FDR) <0.1] (Fig. 5a). We also identified 2 DEG in the PC and DG of Tyrobp +/− mice vs. WT mice. Tyrobp was the top DEG among the different models and brain areas (logFC = −1.1 and −4.8 in Tyrobp +/− and Tyrobp −/−, respectively) (Fig. 5b, c). Overall, we observed strong overlap of the DEG between the different brain areas in mice KO for Tyrobp (Fig. 5b, c for top 10 DEG in PC and DG, see Suppl. 3 for full DEG results). Thus, eight out of ten DEG in the PC of Tyrobp −/− were also differentially expressed in the DG of Tyrobp −/−. Among them are two genes that have been proposed as early biomarkers of AD: biliverdin reductase B (Blvrb) (logFC = 0.9) [54] and Nudix motif 19 (Nudt19) (logFC −1.2) [1]. We also noted a strong trend toward down-regulation of Cd33 in the DG of Tyrobp −/− mice (log FC = −0.9, FDR = 0.077). Recent genome-wide studies identified CD33 as a late-onset AD susceptibility variant [24, 55]. Moreover, CD33 protein is elevated in AD brain and has been associated with amyloid pathology and disease progression [10, 22, 78]. As expected with such small DEG signatures, we did not observe significantly dysregulated Gene Ontology (GO) term enrichments in Tyrobp heterozygous- or homozygous-null mice using DAVID, Ingenuity Pathway Analysis (IPA) or gene set enrichment analysis (GSEA).
Differential gene expression analysis suggests potential molecular mechanisms associated with TYROBP deficiency. a–c Differential gene expression analysis in dentate gyrus and prefrontal cortex of Tyrobp −/− , Tyrobp +/− and WT mice. a Number of up- and down-regulated genes in Tyrobp −/− and Tyrobp +/− vs. WT and Tyrobp −/− vs. Tyrobp +/−. b Top 10 differentially expressed genes in b dentate gyrus of Tyrobp −/− vs. WT and c prefrontal cortex of Tyrobp −/− vs. WT. Bolding highlights differentially expressed genes shared across dentate gyrus and prefontal cortex. RNA sequencing was performed on a total of 47 samples (Tyrobp −/− n = 8 males and 8 females, Tyrobp +/− n = 8 males and 8 females, and WT n = 7–8 samples, eight males and eight females, for each brain regions). All analyses were corrected for sex effect. Differential gene expression threshold was set at fold change ≥1.2 and adjusted p value <0.1. DG dentate gyrus, PC prefrontal cortex. d–g Differential gene expression analysis in prefrontal cortex of APP/PSEN1;Tyrobp −/− , APP/PSEN1; Tyrobp +/− and APP/PSEN1 mice at 4-months-old. d Number of up- and down-regulated genes in APP/PSEN1;Tyrobp −/− and APP/PSEN1;Tyrobp +/− vs. APP/PSEN1 and APP/PSEN1; Tyrobp −/− vs. APP/PSEN1; Tyrobp +/−. e Top 10 differentially expressed genes in APP/PSEN1; Tyrobp +/−vs. APP/PSEN1; f APP/PSEN1; Tyrobp −/− vs. APP/PSEN1 and g APP/PSEN1; Tyrobp −/− vs. APP/PSEN1; Tyrobp +/−. RNA sequencing was performed on a total of 23 samples comprising of both male and female mice (n = 7–8 samples per genotype). All analyses were corrected for sex effect. Differential gene expression threshold was set at fold change ≥1.2 and false discovery rate (FDR) <0.1. (See Suppl. 3 and 4 for full list of differentially expressed genes)
Differential gene expression and enrichment analysis of APP/PSEN1;Tyrobp −/−, APP/PSEN1;Tyrobp +/− and APP/PSEN1 mice
We generated transcriptomic profiles from 23 PC samples from 4-month-old male and female APP/PSEN1 mice that were either heterozygous- or homozygous-null, or WT for Tyrobp. Sex effect was taken into account for all analyses. In comparison to APP/PSEN1 mice, we identified 84 DEG in APP/PSEN1;Tyrobp −/− and 51 DEG in APP/PSEN1; Tyrobp +/− (FDR <0.1) (Fig. 5d–g. See Suppl.4 for full DEG results). All of the ten DEG detected in the PC of Tyrobp −/− vs. WT mice were also differentially expressed in the PC of APP/PSEN1; Tyrobp −/− vs. APP/PSEN1 comparison. The increased signature size in Tyrobp −/− in the APP/PSEN1 background provides strong independent support for the conclusion that TYROBP is relevant not only in human AD [86] but also in the amyloid-depositing mouse brain AD model.
As above, Tyrobp was the top DEG in both APP/PSEN1 deficient and KO for Tyrobp (logFC = −1.2 and −4.8, respectively). Comparison of APP/PSEN1; Tyrobp −/− vs. APP/PSEN1; Tyrobp +/− highlighted 32 DEG. Interestingly, we found evidence in APP/PSEN1 mice that were heterozygous- or homozygous-null for Tyrobp for several DEG associated with AD and/or memory loss. TYROBP deficiency produced changes in Cd33 expression providing independent confirmation of similar phenomena observed by others using different approaches [12]. Also, Sirt2 expression was increased in APP/PSEN1; Tyrobp −/− (logFC = 0.4). A SIRT2 polymorphism has been associated with increased LOAD susceptibility [82] and its level of expression is linked with neurodegenerative disease, likely due to its role in lysosome-mediated authophagic turnover [18, 51, 58]. Igfbp2 expression was decreased in APP/PSEN1; Tyrobp +/− (logFC = −0.6). Several proteomic studies aiming to identify AD markers in human sera have reported an increased level of IGFBP2 in AD patients [39, 57]. Moreover, Pedrós et al. have shown an increased expression level of IGFBP2 in hippocampi of an APP/PSEN1 mouse model similar to that which we used [60]. These data suggest that Tyrobp-related modulation of the expression of several AD-related genes only appears in the presence of cerebral amyloidosis and/or APP/PSEN1 mutations.
To identify biological pathways that may be dysregulated, we performed GSEA using DAVID and IPA (Fig. 6a). Comparisons between APP/PSEN1 KO for Tyrobp and WT for Tyrobp highlighted common themes between DAVID and IPA analyses for perturbation of neurotransmission and ion transport (Fig. 6b and Suppl.5). These included potassium transport, general regulation of transmembrane ion transport, and depolarization and action potential of neurons. Other overlapping themes included neuronal, axonal fasciculation, and synapse assembly. Enrichment analysis with IPA comparing APP/PSEN1; Tyrobp +/− against APP/PSEN1 detected dysregulation of immune function, including migration, movement, and activation of immune and phagocyte cells (Fig. 6c). Perturbation of metabolic functions was also detected. Using DAVID analysis, we noted a pervasive perturbation of protein phosphorylation signal transduction in APP/PSEN1 mice either heterozygous- or homozygous-null for Tyrobp in comparison to APP/PSEN1 mice on a Tyrobp wild-type background. This is potentially relevant to the role of TYROBP as a phosphotyrosine-signal-based sensor of extracellular debris and instigator and/or organizer of phagocytosis. This is also interesting in light of the evidence that Aβ cerebral amyloidosis in humans and mice is accompanied by hyperphosphorylation of cytoskeletal proteins. Comparisons between APP/PSEN1; Tyrobp −/− and APP/PSEN1; Tyrobp +/− showed a unique GO term involving stress-activated protein kinase signaling cascade (Fig. 6d). Pathways identified by IPA included production of superoxide, apoptosis, and differentiation/polarization of macrophages. Excessive production of superoxide can induce an uncontrolled oxidative stress leading to increased microglia activation and neuronal apoptosis [80]. Oxidative stress may also promote production and deposition of Aβ and formation of neurofibrillary tangles [8, 14, 20, 46, 53].
Gene enrichment analysis summary for prefrontal cortex of 4-month-old APP/PSEN1;Tyrobp −/− , APP/PSEN1; Tyrobp +/− and APP/PSEN1 mice at suggests potential molecular mechanisms associated with TYROBP deficiency. a Schematic overview of comparisons between mouse groups. b Enrichment analysis and selected GO terms (DAVID) and diseases and functions (Ingenuity Pathway Analysis) in APP/PSEN1; Tyrobp −/− vs. APP/PSEN1, c in APP/PSEN1; Tyrobp +/− vs. APP/PSEN1, d in APP/PSEN1; Tyrobp −/− vs. in APP/PSEN1; Tyrobp +/−. Enrichments shown were selected for known or suspected relevance to AD pathophysiology. Gene set enrichment threshold was set at p value <0.05. (See Suppl. 5 for full list of enrichments)
Gene regulatory network analysis
Gene regulatory network analysis is a powerful tool in identifying gene modules pathologically related to complex human diseases including AD [86]. To test if the DE signatures detected in the present study were enriched for any AD networks, we collected the co-expression network modules from our two AD cohorts and overlaid the DEG onto the co-expression network modules. We had previously constructed transcriptome-wide gene co-expression networks in different brain regions of postmortem samples from two AD cohorts, the Mount Sinai Brain Bank (MSBB) AD cohort [79] and the Harvard Brain Bank (HBB) AD cohort [86]. Importantly, the age of the mice in this study corresponds to an early stage of AD while our human postmortem co-expression networks from HBB correspond to later disease stages. Although we did not observe enrichments in APP/PSEN1;Tyrobp −/−, we found that the DEG down-regulated in PC of APP/PSEN1;Tyrobp +/− (FDR <0.2) were enriched for several sub-networks from both the MSBB and the HBB AD cohorts, including insulin-like growth factor binding, skeletal development, immune system process, anion transport, and particularly the extracellular matrix (see Suppl.6). The up-regulated genes in APP/PSEN1;Tyrobp +/− mice showed enrichment for nucleobase nucleoside and nucleotide metabolic process sub-network (see Suppl. 6).
Drug repurposing
Through the experiments described above, we identified benefits of TYROBP deficiency on multiple aspects of the phenotype in APP/PSEN1 mice. When those AD model mice were also deficient in TYROBP, beneficial effects in gene expression, phosphorylation of tau, nerve terminal integrity, behavior, and electrophysiology were observed. The data indicate that reduction of Tyrobp gene expression could represent a novel computation- and mutation-based, immune-inflammatory therapeutic opportunity for treating or preventing LOAD. Therefore, we probed a comprehensive pharmacopeia database to determine whether safe existing medications would be predicted to reduce levels of Tyrobp mRNA or protein.
To identify small molecule compounds capable of perturbing Tyrobp expression (Fig. 7), we performed a computational screen against a library of drug-induced transcriptional profiles from Connectivity Map [42]. We scored 1309 unique compounds according to the rank of Tyrobp expression fold change (based on comparison to within-batch vehicle control assays). Top compounds associated with Tyrobp up and down regulation are summarized in Fig. 7b.
Computational analysis of a current pharmacopeia database identified compounds that would be predicted to cause up- or down-regulation of TYROBP expression. a Compounds were scored and ranked according to their associated TYROBP expression fold change, and then used as the basis for a secondary enrichment analysis to identify drug targets that associate with up- or down-regulation of TYROBP. Top 10 compounds that b up-regulate and c down-regulate TYROBP are shown. Drug targets enriched among compounds that d up-regulate, and e down-regulate TYROBP are shown
To explore the pharmacological context of these compounds, we performed a secondary enrichment analysis to identify drug targets that are associated with Tyrobp regulation (Fig. 7a, c). We found that compounds that regulate Tyrobp expression are enriched for multiple drug targets, including many with known links to Alzheimer’s pathology. These include LOAD risk-associated gene Cathepsin D (FDR = 4.9E−02) [16, 70, 72], and Akt1 (FDR = 6.8E−04), a molecule that is activated in LOAD [65] and is associated with LOAD risk in Chinese Han AD patients with type 2 diabetes [50].
Of the targets enriched among compounds predicted to suppress Tyrobp expression (and so potentially representing therapeutic candidates), RIPK1 (FDR = 1.1E−02) was most strongly implicated. Interestingly, RIPK1, a key constituent of the necrosome, was recently shown in an independent study by other investigators to regulate context-dependent regulation of programmed necrosis via formation of an amyloid signaling complex [47]. Experimental validation of these predicted repurposable drugs is underway and will be reported in detail in a future publication.
Discussion
Association of TYROBP with LOAD arose via a multiscale computational network approach [86]. The physical interaction between TREM2 and TYROBP as well as with other LOAD risk factors such as CR3, and SIRPβ1 [5, 7, 23, 55, 86] provided an important lead for our experimental strategy aiming to validate the important role of TYROBP in the pathogenesis of LOAD. We have previously defined via a multiscale computational network approach TYROBP as a strong candidate for playing the role of a key “hub” or “driver” gene in LOAD [86]. It is worth noting that two independent groups have also identified TYROBP as a driver of LOAD despite having followed different and highly idiosyncratic computational strategies [12, 48]. CD33 is a known AD risk gene and a component of the TYROBP network. The regulation of Cd33 by TYROBP reported herein as well as the regulation of TREM2 by CD33 reported by de Jager and colleagues [12] provide compelling evidence in support of the role of TYROBP as a “driver” gene in LOAD. Capping off the evidence associating TYROBP with LOAD is the recent discovery that missense mutations in the coding region of the TYROBP gene are associated with AD risk [61]. Interestingly, in the same study, in vitro overexpression of the candidate pathogenic p.D50_L51ins14 TYROBP variant led to a strong reduction of TREM2 expression [61]. We have previously shown that TYROBP expression is elevated in AD brain and mouse models [64, 86], but it was not immediately apparent whether that elevation represented a pre-existing, predisposing factor or was a secondary reaction to LOAD pathology. Based on the data presented above, a Tyrobp null mutation appears to exert effects that would be characterized as beneficial with respect to both the normal physiology of neurons and the proteinopathy of LOAD.
The effects of Tyrobp knockdown or knockout on Aβ levels and Aβ oligomer conformers as defined by epitope content were limited to the reduction of the level of NU-4 and A11 type oligomers in TYROBP deficient APP/PSEN1 mice. There were no consistent statistically significant differences on levels of total Aβ, Aβ40, Aβ42, or on levels of OC type Aβ oligomers. The relatively minor effect size notwithstanding, it is worth noting that the converging evidence from several laboratories (including our own) is that the NU-4 epitope is the signature of the Aβ oligomer strain that is most consistently neurotoxic [40, 73]. A11 and OC oligomer strains are not consistently neurotoxic. As reported above and in one of our previous studies [41], we noted sex differences in Aβ and oligomer levels suggesting an earlier progression of the disease in female than male APP/PSEN1 mice.
The difference in Aβ levels observed between the male and female mice is of importance considering the sexual dimorphism observed in the phosphorylation status of TAU in APP/PSEN1 background but not in WT background. Thus, the effect of a decreased Tyrobp expression on the stoichiometry of TAU phosphorylation appeared to be different in the presence or absence of APP/PSEN1 mutations leading to amyloid deposition. Indeed, TYROBP deficiency tends to increase the phosphorylation of TAU on a WT background, but, on a APP/PSEN1 background, loss of TYROBP decreased the phosphorylation status of TAU in female mice in the setting of higher Aβ loads as compared with males. Although the mechanism(s) by which microglia exert their effects on neuronal tau pathology remains unclear, several reports have linked TREM2 expression and hyperphosphorylated TAU [31, 33, 35]. These reports suggest that TREM2 deficiency could increase tauopathy in human tau-expressing models but could decrease tau pathology in AD mouse models displaying cerebral amyloidosis. Herein we report that a decreased expression of Tyrobp can have beneficial effects on tau pathology and neuronal injury in APP/PSEN1 mouse model of AD. In accordance with our data, Strittmatter and colleagues [75] recently reported that mouse deficient for Progranulin presented an overexpression of Tyrobp network genes correlating with an increased neuronal injury and tau pathology in the absence of amyloid pathology [75].
As mentioned above, no differences were noted in number and size of Aβ plaque depositions and the general histological impact of TYROBP deficiency on plaque morphology and microglia recruitment was identical in appearance to that reported by Colonna and colleagues in their studies of TREM2-deficient mice [85]. Indeed, Tyrobp KO mice presented fewer microglia decorating each amyloid plaque without modification in the total number of microglia, and plaques exhibited less compact morphology. However, unlike the Colonna report wherein the reduced numbers of microglia per plaque were predicted to be associated with increased severity of the phenotype [85], we observed that this histological appearance was instead associated with beneficial effects on neuritic dystrophy, TAU metabolism, learning behavior, and neuronal electrophysiology. Although beyond the scope of this study, it will be interesting to determine whether overexpression leads to opposite results. In addition, recent papers from Lamb and colleagues [30], Yu and colleagues [32, 34], and Raha-Chowdhury and colleagues [63] raise the possibility that there could be aging-related and/or disease-stage-related changes in the effect of TYROBP. These papers focused on TREM2 and suggest that reduced TREM2 may be beneficial early in life (~4 months) while reduced TREM2 late in life (~8 months) could be detrimental. We are in the process of assessing whether a similar phenomenon occurs with TYROBP.
Electrophysiological assays revealed that the loss of TYROBP normalized some of the synaptic dysfunctions caused by the APP/PSEN1 mutations. The strong increase in basal synaptic efficiency seen in the Tyrobp −/− mice is of particular interest. If observed in isolation, this phenomenon might lead to overactivation of pyramidal neurons and damage, but the same effect could prove protective in the context of LOAD-associated factors that reduce neuronal function. The protective effect of TYROBP deficiency in an early AD context is confirmed by the improvement in the behavioral performance of APP/PSEN1 mice deficient in TYROBP. The effect of the Tyrobp-null background on the electrophysiological findings and gene set enrichment (synapse assembly, ion transport, and neurotransmission) of APP/PSEN1; Tyrobp −/− vs. APP/PSEN1 are in keeping with the growing appreciation for the role of microglia in maintaining normal synaptic physiology [4]. Indeed, in addition to their pro-inflammatory and phagocytic functions, microglia release cytokines including TGFβ and interleukin-1β that acutely modulate synaptic plasticity at hippocampal synapses [11, 71].
Thus, in a comprehensive panel of transcriptomic, biochemical, electrophysiological, and behavioral paradigms, reduction or ablation of TYROBP prevented the expression of many of the corresponding APP/PSEN1 phenotypes at 4 months of age. These results would appear to argue against the possibility that early TYROBP deficiency is likely to be a predisposing factor for LOAD. Indeed, these results would indicate that a decrease in TYROBP activity could represent an important therapeutic opportunity either for treating or preventing LOAD or else for slowing or arresting the progression of MCI or early AD to full-blown clinical and pathological LOAD.
References
Arisi I, Mara D, Brandi R, Felsani A, Capsoni S, Drovandi G, Felici G, Weitschek E, Bertolazzi P, Cattaneo A (2011) Gene expression biomarkers in the brain of a mouse model for Alzheimer’s disease: mining of microarray data by logic classification and feature selection. J Alzheimers Dis 24:721–738. doi:10.3233/JAD-2011-101881
Bakker AB, Hoek RM, Cerwenka A, Blom B, Lucian L, McNeil T, Murray R, Phillips LH, Sedgwick JD, Lanier LL (2000) DAP12-deficient mice fail to develop autoimmunity due to impaired antigen priming. Immunity 13:345–353
Barnes CA (1979) Memory deficits associated with senescence: a neurophysiological and behavioral study in the rat. J Comp Physiol Psychol 93:74–104
Benarroch EE (2013) Microglia: multiple roles in surveillance, circuit shaping, and response to injury. Neurology 81:1079–1088. doi:10.1212/WNL.0b013e3182a4a577
Benitez BA, Jin SC, Guerreiro R, Graham R, Lord J, Harold D, Sims R, Lambert J-C, Gibbs JR, Bras J, Sassi C, Harari O, Bertelsen S, Lupton MK, Powell J, Bellenguez C, Brown K, Medway C, Haddick PC, van der Brug MP, Bhangale T, Ortmann W, Behrens T, Mayeux R, Pericak-Vance MA, Farrer LA, Schellenberg GD, Haines JL, Turton J, Braae A, Barber I, Fagan AM, Holtzman DM, Morris JC, Williams J, Kauwe JSK, Amouyel P, Morgan K, Singleton A, Hardy J, Goate AM, Cruchaga C (2014) Missense variant in TREML2 protects against Alzheimer’s disease. Neurobiol Aging 35:1510.e19–1510.e26. doi:10.1016/j.neurobiolaging.2013.12.010
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B Methodol 57:289–300
Bertram L, Lange C, Mullin K, Parkinson M, Hsiao M, Hogan MF, Schjeide BMM, Hooli B, Divito J, Ionita I, Jiang H, Laird N, Moscarillo T, Ohlsen KL, Elliott K, Wang X, Hu-Lince D, Ryder M, Murphy A, Wagner SL, Blacker D, Becker KD, Tanzi RE (2008) Genome-wide association analysis reveals putative Alzheimer’s disease susceptibility loci in addition to APOE. Am J Hum Genet 83:623–632. doi:10.1016/j.ajhg.2008.10.008
Bertram L, Tanzi RE (2008) Thirty years of Alzheimer’s disease genetics: the implications of systematic meta-analyses. Nat Rev Neurosci 9:768–778. doi:10.1038/nrn2494
Blatow M, Caputi A, Burnashev N, Monyer H, Rozov A (2003) Ca2+ buffer saturation underlies paired pulse facilitation in calbindin-D28k-containing terminals. Neuron 38:79–88
Bradshaw EM, Chibnik LB, Keenan BT, Ottoboni L, Raj T, Tang A, Rosenkrantz LL, Imboywa S, Lee M, Von Korff A, Morris MC, Evans DA, Johnson K, Sperling RA, Schneider JA, Bennett DA, De Jager PL (2013) CD33 Alzheimer’s disease locus: altered monocyte function and amyloid biology. Nat Neurosci 16:848–850. doi:10.1038/nn.3435
Caraci F, Gulisano W, Guida CA, Impellizzeri AAR, Drago F, Puzzo D, Palmeri A (2015) A key role for TGF-β1 in hippocampal synaptic plasticity and memory. Sci Rep 5:11252. doi:10.1038/srep11252
Chan G, White CC, Winn PA, Cimpean M, Replogle JM, Glick LR, Cuerdon NE, Ryan KJ, Johnson KA, Schneider JA, Bennett DA, Chibnik LB, Sperling RA, Bradshaw EM, De Jager PL (2015) CD33 modulates TREM2: convergence of Alzheimer loci. Nat Neurosci 18:1556–1558. doi:10.1038/nn.4126
Chang EH, Savage MJ, Flood DG, Thomas JM, Levy RB, Mahadomrongkul V, Shirao T, Aoki C, Huerta PT (2006) AMPA receptor downscaling at the onset of Alzheimer’s disease pathology in double knockin mice. Proc Natl Acad Sci USA 103:3410–3415. doi:10.1073/pnas.0507313103
Christen Y (2000) Oxidative stress and Alzheimer disease. Am J Clin Nutr 71:621S–629S
Condello C, Yuan P, Schain A, Grutzendler J (2015) Microglia constitute a barrier that prevents neurotoxic protofibrillar Aβ42 hotspots around plaques. Nat Commun 6:6176. doi:10.1038/ncomms7176
Davidson Y, Gibbons L, Pritchard A, Hardicre J, Wren J, Tian J, Shi J, Stopford C, Julien C, Thompson J, Payton A, Thaker U, Hayes AJ, Iwatsubo T, Pickering-Brown SM, Pendleton N, Horan MA, Burns A, Purandare N, Lendon CL, Neary D, Snowden JS, Mann DMA (2006) Genetic associations between cathepsin D exon 2 C→ T polymorphism and Alzheimer’s disease, and pathological correlations with genotype. J Neurol Neurosurg Psychiatry 77:515–517. doi:10.1136/jnnp.2005.063917
Fanutza T, Prete D, Ford M, Castillo P, Luciano D (2015) APP and APLP2 interact with the synaptic release machinery and facilitate transmitter release at hippocampal synapses. Elife 4:e09743. doi:10.7554/eLife.09743
Gal J, Bang Y, Choi HJ (2012) SIRT2 interferes with autophagy-mediated degradation of protein aggregates in neuronal cells under proteasome inhibition. Neurochem Int 61:992–1000. doi:10.1016/j.neuint.2012.07.010
Gandy S, Heppner F (2013) Microglia as dynamic and essential components of the amyloid hypothesis. Neuron 78:575–577. doi:10.1016/j.neuron.2013.05.007
Gella A, Durany N (2009) Oxidative stress in Alzheimer disease. Cell Adhes Migr 3:88–93
Gowrishankar S, Yuan P, Wu Y, Schrag M, Paradise S, Grutzendler J, De Camilli P, Ferguson SM (2015) Massive accumulation of luminal protease-deficient axonal lysosomes at Alzheimer’s disease amyloid plaques. Proc Natl Acad Sci USA 112:E3699–E3708. doi:10.1073/pnas.1510329112
Griciuc A, Alberto S-P, Parrado AR, Lesinski AN, Asselin CN, Mullin K, Hooli B, Choi SH, Hyman BT, Tanzi RE (2013) Alzheimer’s disease risk gene CD33 inhibits microglial uptake of amyloid beta. Neuron 78:631–643. doi:10.1016/j.neuron.2013.04.014
Guerreiro R, Wojtas A, Bras J, Carrasquillo M, Rogaeva E, Majounie E, Cruchaga C, Sassi C, Kauwe JS, Younkin S, Hazrati L, Collinge J, Pocock J, Lashley T, Williams J, Lambert J-C, Amouyel P, Goate A, Rademakers R, Morgan K, Powell J, Peter G-H, Singleton A, Hardy J, Group A (2013) TREM2 variants in Alzheimer’s disease. N Engl J Med 368:117–127. doi:10.1056/NEJMoa1211851
Hollingworth P, Harold D, Sims R, Gerrish A, Lambert J-CC, Carrasquillo MM, Abraham R, Hamshere ML, Pahwa JS, Moskvina V, Dowzell K, Jones N, Stretton A, Thomas C, Richards A, Ivanov D, Widdowson C, Chapman J, Lovestone S, Powell J, Proitsi P, Lupton MK, Brayne C, Rubinsztein DC, Gill M, Lawlor B, Lynch A, Brown KS, Passmore PA, Craig D, Bernadette M, Todd S, Holmes C, Mann D, Smith A, Beaumont H, Warden D, Wilcock G, Love S, Kehoe PG, Hooper NM, Vardy ER, Hardy J, Mead S, Fox NC, Rossor M, Collinge J, Maier W, Jessen F, Rüther E, Schürmann B, Heun R, Kölsch H, van den Bussche H, Heuser I, Kornhuber J, Wiltfang J, Dichgans M, Frölich L, Hampel H, Gallacher J, Hüll M, Rujescu D, Giegling I, Goate AM, Kauwe JS, Cruchaga C, Nowotny P, Morris JC, Mayo K, Sleegers K, Bettens K, Engelborghs S, De Deyn PP, Van Broeckhoven C, Livingston G, Bass NJ, Gurling H, Andrew M, Gwilliam R, Deloukas P, Ammar A-C, Shaw CE, Tsolaki M, Singleton AB, Guerreiro R, Mühleisen TW, Nöthen MM, Moebus S, Jöckel K-HH, Klopp N, Wichmann H-EE, Pankratz V, Sando SB, Aasly JO, Barcikowska M, Wszolek ZK, Dickson DW, Neill RG-R, Petersen RC (2011) Common variants at ABCA7, MS4A6A/MS4A4E, EPHA1, CD33 and CD2AP are associated with Alzheimer’s disease. Nat Genet 43:429–435. doi:10.1038/ng.803
Hong S, Beja-Glasser VF, Nfonoyim BM, Frouin A, Li S, Ramakrishnan S, Merry KM, Shi Q, Rosenthal A, Barres BA, Lemere CA, Selkoe DJ, Stevens B (2016) Complement and microglia mediate early synapse loss in Alzheimer mouse models. Science 352:712–716. doi:10.1126/science.aad8373
Huang DW, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4:44–57. doi:10.1038/nprot.2008.211
Huang DW, Sherman BT, Lempicki RA (2009) Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res 37:1–13. doi:10.1093/nar/gkn923
Iorio F, Bosotti R, Scacheri E, Belcastro V, Mithbaokar P, Ferriero R, Murino L, Tagliaferri R, Brunetti-Pierri N, Isacchi A, di Bernardo D (2010) Discovery of drug mode of action and drug repositioning from transcriptional responses. Proc Natl Acad Sci USA 107:14621–14626. doi:10.1073/pnas.1000138107
Jankowsky JL, Slunt HH, Ratovitski T, Jenkins NA, Copeland NG, Borchelt DR (2001) Co-expression of multiple transgenes in mouse CNS: a comparison of strategies. Biomol Eng 17:157–165
Jay TR, Hirsch AM, Broihier ML, Miller CM, Neilson LE, Ransohoff RM, Lamb BT, Landreth GE (2017) Disease progression-dependent effects of TREM2 deficiency in a mouse model of Alzheimer’s disease. J Neurosci 37:637–647. doi:10.1523/JNEUROSCI.2110-16.2017
Jay TR, Miller CM, Cheng PJ, Graham LC, Bemiller S, Broihier ML, Xu G, Margevicius D, Karlo JC, Sousa GL, Cotleur AC, Butovsky O, Bekris L, Staugaitis SM, Leverenz JB, Pimplikar SW, Landreth GE, Howell GR, Ransohoff RM, Lamb BT (2015) TREM2 deficiency eliminates TREM2+ inflammatory macrophages and ameliorates pathology in Alzheimer’s disease mouse models. J Exp Med 212:287–295. doi:10.1084/jem.20142322
Jiang T, Tan L, Zhu X-C, Zhang Q-Q, Cao L, Tan M-S, Gu L-Z, Wang H-F, Ding Z-Z, Zhang Y-D, Yu J-T (2014) Upregulation of TREM2 ameliorates neuropathology and rescues spatial cognitive impairment in a transgenic mouse model of Alzheimer’s disease. Neuropsychopharmacology 39:2949–2962. doi:10.1038/npp.2014.164
Jiang T, Tan L, Zhu X-C, Zhou J-S, Cao L, Tan M-S, Wang H-F, Chen Q, Zhang Y-D, Yu J-T (2015) Silencing of TREM2 exacerbates tau pathology, neurodegenerative changes, and spatial learning deficits in P301S tau transgenic mice. Neurobiol Aging 36:3176–3186. doi:10.1016/j.neurobiolaging.2015.08.019
Jiang T, Wan Y, Zhang Y-D, Zhou J-S, Gao Q, Zhu X-C, Shi J-Q, Lu H, Tan L, Yu J-T (2017) TREM2 overexpression has no improvement on neuropathology and cognitive impairment in aging APPswe/PS1dE9 mice. Mol Neurobiol 54:855–865. doi:10.1007/s12035-016-9704-x
Jiang T, Zhang Y-D, Chen Q, Gao Q, Zhu X-C, Zhou J-S, Shi J-Q, Lu H, Tan L, Yu J-T (2016) TREM2 modifies microglial phenotype and provides neuroprotection in P301S tau transgenic mice. Neuropharmacology 105:196–206. doi:10.1016/j.neuropharm.2016.01.028
Keiser MJ, Roth BL, Armbruster BN, Ernsberger P, Irwin JJ, Shoichet BK (2007) Relating protein pharmacology by ligand chemistry. Nat Biotechnol 25:197–206. doi:10.1038/nbt1284
Keiser MJ, Setola V, Irwin JJ, Laggner C, Abbas AI, Hufeisen SJ, Jensen NH, Kuijer MB, Matos RC, Tran TB, Whaley R, Glennon RA, Hert J, Thomas KLH, Edwards DD, Shoichet BK, Roth BL (2009) Predicting new molecular targets for known drugs. Nature 462:175–181. doi:10.1038/nature08506
Kessels HW, Malinow R (2009) Synaptic AMPA receptor plasticity and behavior. Neuron 61:340–350. doi:10.1016/j.neuron.2009.01.015
Kiddle SJ, Sattlecker M, Proitsi P, Simmons A, Westman E, Bazenet C, Nelson SK, Williams S, Hodges A, Johnston C, Soininen H, Kłoszewska I, Mecocci P, Tsolaki M, Vellas B, Newhouse S, Lovestone S, Dobson RJB (2014) Candidate blood proteome markers of Alzheimer’s disease onset and progression: a systematic review and replication study. J Alzheimers Dis JAD 38:515–531. doi:10.3233/JAD-130380
Knight E, Kim S, Kottwitz J, Hatami A, Albay R, Suzuki A, Lublin A, Alberini C, Klein W, Szabo P, Relkin N, Ehrlich M, Glabe C, Gandy S, Steele J (2016) Effective anti-Alzheimer Aβ therapy involves depletion of specific Aβ oligomer subtypes. Neurol Neuroimmunol Neuroinflam 3:e237. doi:10.1212/NXI.0000000000000237
Knight EM, Ruiz HH, Kim SH, Harte JC, Hsieh W, Glabe C, Klein WL, Attie AD, Buettner C, Ehrlich ME, Gandy S (2016) Unexpected partial correction of metabolic and behavioral phenotypes of Alzheimer’s APP/PSEN1 mice by gene targeting of diabetes/Alzheimer’s-related Sorcs1. Acta Neuropathol Commun 4:16. doi:10.1186/s40478-016-0282-y
Lamb J, Crawford ED, Peck D, Modell JW, Blat IC, Wrobel MJ, Lerner J, Brunet J-P, Subramanian A, Ross KN, Reich M, Hieronymus H, Wei G, Armstrong SA, Haggarty SJ, Clemons PA, Wei R, Carr SA, Lander ES, Golub TR (2006) The Connectivity Map: using gene-expression signatures to connect small molecules, genes, and disease. Science 313:1929–1935. doi:10.1126/science.1132939
Lambert J-C, Ibrahim-Verbaas CA, Harold D, Naj AC, Sims R, Bellenguez C, Jun G, DeStefano AL, Bis JC, Beecham GW, Grenier-Boley B, Russo G, Thornton-Wells TA, Jones N, Smith AV, Chouraki V, Thomas C, Ikram MA, Zelenika D, Vardarajan BN, Kamatani Y, Lin C-F, Gerrish A, Schmidt H, Kunkle B, Dunstan ML, Ruiz A, Bihoreau M-T, Choi S-H, Reitz C, Pasquier F, Hollingworth P, Ramirez A, Hanon O, Fitzpatrick AL, Buxbaum JD, Campion D, Crane PK, Baldwin C, Becker T, Gudnason V, Cruchaga C, Craig D, Amin N, Berr C, Lopez OL, De Jager PL, Deramecourt V, Johnston JA, Evans D, Lovestone S, Letenneur L, Morón FJ, Rubinsztein DC, Eiriksdottir G, Sleegers K, Goate AM, Fiévet N, Huentelman MJ, Gill M, Brown K, Kamboh MI, Keller L, Barberger-Gateau P, McGuinness B, Larson EB, Green R, Myers AJ, Dufouil C, Todd S, Wallon D, Love S, Rogaeva E, Gallacher J, St George-Hyslop P, Clarimon J, Lleo A, Bayer A, Tsuang DW, Yu L, Tsolaki M, Bossù P, Spalletta G, Proitsi P, Collinge J, Sorbi S, Sanchez-Garcia F, Fox NC, Hardy J, Naranjo MCD, Bosco P, Clarke R, Brayne C, Galimberti D, Mancuso M, Matthews F, European Alzheimer’s Disease Initiative (EADI), Genetic and Environmental Risk in Alzheimer’s Disease (GERAD), Alzheimer’s Disease Genetic Consortium (ADGC), Cohorts for Heart and Aging Research in Genomic Epidemiology (CHARGE), Moebus S, Mecocci P, Del Zompo M, Maier W, Hampel H, Pilotto A, Bullido M, Panza F, Caffarra P, Nacmias B, Gilbert JR, Mayhaus M, Lannfelt L, Hakonarson H, Pichler S, Carrasquillo MM, Ingelsson M, Beekly D, Alvarez V, Zou F, Valladares O, Younkin SG, Coto E, Hamilton-Nelson KL, Gu W, Razquin C, Pastor P, Mateo I, Owen MJ, Faber KM, Jonsson PV, Combarros O, O’Donovan MC, Cantwell LB, Soininen H, Blacker D, Mead S, Mosley TH Jr, Bennett DA, Harris TB, Fratiglioni L, Holmes C, de Bruijn RFAG, Passmore P, Montine TJ, Bettens K, Rotter JI, Brice A, Morgan K, Foroud TM, Kukull WA, Hannequin D, Powell JF, Nalls MA, Ritchie K, Lunetta KL, Kauwe JSK, Boerwinkle E, Riemenschneider M, Boada M, Hiltunen M, Martin ER, Schmidt R, Rujescu D, Wang L-S, Dartigues J-F, Mayeux R, Tzourio C, Hofman A, Nöthen MM, Graff C, Psaty BM, Jones L, Haines JL, Holmans PA, Lathrop M, Pericak-Vance MA, Launer LJ, Farrer LA, van Duijn CM, Van Broeckhoven C, Moskvina V, Seshadri S, Williams J, Schellenberg GD, Amouyel P (2013) Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat Genet 45:1452–1458. doi:10.1038/ng.2802
Lambert MP, Velasco PT, Chang L, Viola KL, Fernandez S, Lacor PN, Khuon D, Gong Y, Bigio EH, Shaw P, De Felice FG, Krafft GA, Klein WL (2007) Monoclonal antibodies that target pathological assemblies of Aβ. J Neurochem 100:23–35. doi:10.1111/j.1471-4159.2006.04157.x
Law V, Knox C, Djoumbou Y, Jewison T, Guo AC, Liu Y, Maciejewski A, Arndt D, Wilson M, Neveu V, Tang A, Gabriel G, Ly C, Adamjee S, Dame ZT, Han B, Zhou Y, Wishart DS (2014) DrugBank 4.0: shedding new light on drug metabolism. Nucleic Acids Res 42:D1091–D1097. doi:10.1093/nar/gkt1068
Li F, Calingasan NY, Yu F, Mauck WM, Toidze M, Almeida CG, Takahashi RH, Carlson GA, Flint Beal M, Lin MT, Gouras GK (2004) Increased plaque burden in brains of APP mutant MnSOD heterozygous knockout mice. J Neurochem 89:1308–1312. doi:10.1111/j.1471-4159.2004.02455.x
Li J, McQuade T, Siemer AB, Napetschnig J, Moriwaki K, Hsiao Y-S, Damko E, Moquin D, Walz T, McDermott A, Chan FK-M, Wu H (2012) The RIP1/RIP3 necrosome forms a functional amyloid signaling complex required for programmed necrosis. Cell 150:339–350. doi:10.1016/j.cell.2012.06.019
Li X, Long J, He T, Belshaw R, Scott J (2015) Integrated genomic approaches identify major pathways and upstream regulators in late onset Alzheimer’s disease. Sci Rep 5:12393. doi:10.1038/srep12393
Liu P, Reed MN, Kotilinek LA, Grant MKO, Forster CL, Qiang W, Shapiro SL, Reichl JH, Chiang ACA, Jankowsky JL, Wilmot CM, Cleary JP, Zahs KR, Ashe KH (2015) Quaternary structure defines a large class of amyloid-β oligomers neutralized by sequestration. Cell Rep 11:1760–1771. doi:10.1016/j.celrep.2015.05.021
Liu S-Y, Zhao H-D, Wang J-L, Huang T, Tian H-W, Yao L-F, Tao H, Chen Z-W, Wang C-Y, Sheng S-T, Li H, Zhao B, Li K-S (2015) Association between polymorphisms of the AKT1 gene promoter and risk of the Alzheimer’s disease in a chinese han population with type 2 diabetes. CNS Neurosci Ther 21:619–625. doi:10.1111/cns.12430
Luthi-Carter R, Taylor DM, Pallos J, Lambert E, Amore A, Parker A, Moffitt H, Smith DL, Runne H, Gokce O, Kuhn A, Xiang Z, Maxwell MM, Reeves SA, Bates GP, Neri C, Thompson LM, Marsh JL, Kazantsev AG (2010) SIRT2 inhibition achieves neuroprotection by decreasing sterol biosynthesis. Proc Natl Acad Sci USA 107:7927–7932. doi:10.1073/pnas.1002924107
Marchetti C, Marie H (2011) Hippocampal synaptic plasticity in Alzheimer’s disease: what have we learned so far from transgenic models? Rev Neurosci 22:373–402. doi:10.1515/RNS.2011.035
Misonou H, Morishima-Kawashima M, Ihara Y (2000) Oxidative stress induces intracellular accumulation of amyloid β-protein (Aβ) in human neuroblastoma cells. Biochemistry (Mosc) 39:6951–6959
Mueller C, Zhou W, Vanmeter A, Heiby M, Magaki S, Ross MM, Espina V, Schrag M, Dickson C, Liotta LA, Kirsch WM (2010) The heme degradation pathway is a promising serum biomarker source for the early detection of Alzheimer’s disease. J Alzheimers Dis 19:1081–1091. doi:10.3233/JAD-2010-1303
Naj A, Jun G, Beecham G, Wang L-S, Vardarajan B, Buros J, Gallins P, Buxbaum J, Jarvik G, Crane P, Larson E, Bird T, Boeve B, Neill G-R, Jager P, Evans D, Schneider J, Carrasquillo M, Nilufer E-T, Younkin S, Cruchaga C, Kauwe J, Nowotny P, Kramer P, Hardy J, Huentelman M, Myers A, Barmada M, Demirci F, Baldwin C, Green R, Rogaeva E, Peter G-H, Arnold S, Barber R, Beach T, Bigio E, Bowen J, Boxer A, Burke J, Cairns N, Carlson C, Carney R, Carroll S, Chui H, Clark D, Corneveaux J, Cotman C, Cummings J, Charles D, Steven D, Ramon D-A, Dick M, Dickson D, Ellis W, Faber K, Fallon K, Farlow M, Ferris S, Frosch M, Galasko D, Ganguli M, Gearing M, Geschwind D, Ghetti B, Gilbert J, Gilman S, Giordani B, Glass J, Growdon J, Hamilton R, Harrell L, Head E, Honig L, Hulette C, Hyman B, Jicha G, Jin L-W, Johnson N, Karlawish J, Karydas A, Kaye J, Kim R, Koo E, Kowall N, Lah J, Levey A, Lieberman A, Lopez O, Mack W, Marson D, Martiniuk F, Mash D, Masliah E, Wayne M, Susan M, Andrew M, Ann M, Mesulam M, Miller B, Miller C, Miller J, Parisi J, Perl D, Peskind E, Petersen R, Poon W, Quinn J, Rajbhandary R, Raskind M, Reisberg B, Ringman J, Roberson E, Rosenberg R, Sano M, Schneider L, Seeley W, Shelanski M, Slifer M, Smith C, Sonnen J, Spina S, Stern R, Tanzi R, Trojanowski J, Troncoso J, Deerlin V, Vinters H, Vonsattel J, Weintraub S, Kathleen W-B, Williamson J, Woltjer R, Cantwell L, Dombroski B, Beekly D, Lunetta K, Martin E, Kamboh M, Saykin A, Reiman E, Bennett D, Morris J, Montine T, Goate A, Blacker D, Tsuang D, Hakonarson H, Kukull W, Foroud T, Haines J, Mayeux R, Margaret P-V, Farrer L, Schellenberg G (2011) Common variants at MS4A4/MS4A6E, CD2AP, CD33 and EPHA1 are associated with late-onset Alzheimer’s disease. Nat Genet 43:436–441. doi:10.1038/ng.801
Oakley H, Cole SL, Logan S, Maus E, Shao P, Craft J, Guillozet-Bongaarts A, Ohno M, Disterhoft J, Eldik LV, Berry R, Vassar R (2006) Intraneuronal β-amyloid aggregates, neurodegeneration, and neuron loss in transgenic mice with five familial Alzheimer’s disease mutations: potential factors in amyloid plaque formation. J Neurosci 26:10129–10140. doi:10.1523/JNEUROSCI.1202-06.2006
O’Bryant SE, Xiao G, Barber R, Reisch J, Doody R, Fairchild T, Adams P, Waring S, Diaz-Arrastia R, Texas Alzheimer’s Research Consortium (2010) A serum protein-based algorithm for the detection of Alzheimer disease. Arch Neurol 67:1077–1081. doi:10.1001/archneurol.2010.215
Outeiro TF, Kontopoulos E, Altmann SM, Kufareva I, Strathearn KE, Amore AM, Volk CB, Maxwell MM, Rochet J-C, McLean PJ, Young AB, Abagyan R, Feany MB, Hyman BT, Kazantsev AG (2007) Sirtuin 2 inhibitors rescue α-synuclein-mediated toxicity in models of Parkinson’s disease. Science 317:516–519. doi:10.1126/science.1143780
Paloneva J, Kestilä M, Wu J, Salminen A, Böhling T, Ruotsalainen V, Hakola P, Bakker A, Phillips J, Pekkarinen P, Lanier L, Timonen T, Peltonen L (2000) Loss-of-function mutations in TYROBP (DAP12) result in a presenile dementia with bone cysts. Nat Genet 25:357–361. doi:10.1038/77153
Pedrós I, Petrov D, Allgaier M, Sureda F, Barroso E, Beas-Zarate C, Auladell C, Pallàs M, Vázquez-Carrera M, Casadesús G, Folch J, Camins A (2014) Early alterations in energy metabolism in the hippocampus of APPswe/PS1dE9 mouse model of Alzheimer’s disease. Biochim Biophys Acta BBA Mol Basis Dis 1842:1556–1566. doi:10.1016/j.bbadis.2014.05.025
Pottier C, Ravenscroft T, Brown P, Finch N, Baker M, Parsons M, Asmann Y, Ren Y, Christopher E, Levitch D, Blitterswijk M, Cruchaga C, Campion D, Nicolas G, Richard A-C, Guerreiro R, Bras J, Zuchner S, Gonzalez M, Bu G, Younkin S, Knopman D, Josephs K, Parisi J, Petersen R, Nilüfer E-T, Neill G-R, Boeve B, Dickson D, Rademakers R (2016) TYROBP genetic variants in early-onset Alzheimer’s disease. Neurobiol Aging. doi:10.1016/j.neurobiolaging.2016.07.028
R Development Core Team (2015) R: a language and environment for statistical computing. In: GBIF.ORG. http://www.gbif.org/resource/81287
Raha AA, Henderson JW, Stott SRW, Vuono R, Foscarin S, Friedland RP, Zaman SH, Raha-Chowdhury R (2017) Neuroprotective effect of TREM-2 in aging and Alzheimer’s disease model. J Alzheimers Dis 55:199–217. doi:10.3233/JAD-160663
Readhead B, Haure-Mirande J-V, Zhang B, Haroutunian V, Gandy S, Schadt EE, Dudley JT, Ehrlich ME (2016) Molecular systems evaluation of oligomerogenic APPE693Q and fibrillogenic APPKM670/671NL/PSEN1Δexon9 mouse models identifies shared features with human Alzheimer’s brain molecular pathology. Mol Psychiatry 21:1099–1111. doi:10.1038/mp.2015.167
Rickle A, Bogdanovic N, Volkman I, Winblad B, Ravid R, Cowburn RF (2004) Akt activity in Alzheimer’s disease and other neurodegenerative disorders. NeuroReport 15:955–959
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK (2015) limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43:e47. doi:10.1093/nar/gkv007
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics (Oxf Engl) 26:139–140. doi:10.1093/bioinformatics/btp616
Roumier A, Pascual O, Béchade C, Wakselman S, Poncer J-C, Réal E, Triller A, Bessis A (2008) Prenatal activation of microglia induces delayed impairment of glutamatergic synaptic function. PLoS One 3:e2595. doi:10.1371/journal.pone.0002595
Sajikumar S, Frey JU (2004) Late-associativity, synaptic tagging, and the role of dopamine during LTP and LTD. Neurobiol Learn Mem 82:12–25. doi:10.1016/j.nlm.2004.03.003
Sayad A, Noruzinia M, Zamani M, Harirchian MH, Kazemnejad A (2014) Association study of cathepsin D gene polymorphism in Iranian patients with sporadic late-onset Alzheimer’s disease. Dement Geriatr Cogn Disord 37:257–264. doi:10.1159/000347128
Schneider H, Pitossi F, Balschun D, Wagner A, del Rey A, Besedovsky HO (1998) A neuromodulatory role of interleukin-1beta in the hippocampus. Proc Natl Acad Sci USA 95:7778–7783
Schuur M, Ikram MA, van Swieten JC, Isaacs A, Vergeer-Drop JM, Hofman A, Oostra BA, Breteler MMB, van Duijn CM (2011) Cathepsin D gene and the risk of Alzheimer’s disease: a population-based study and meta-analysis. Neurobiol Aging 32:1607–1614. doi:10.1016/j.neurobiolaging.2009.10.011
Sebollela A, Mustata G-M, Luo K, Velasco PT, Viola KL, Cline EN, Shekhawat GS, Wilcox KC, Dravid VP, Klein WL (2014) Elucidating molecular mass and shape of a neurotoxic Aβ oligomer. ACS Chem Neurosci 5:1238–1245. doi:10.1021/cn500156r
Sunyer B, Patil S, Höger H, Luber G (2007) Barnes maze, a useful task to assess spatial reference memory in the mice. Protoc Exch. doi:10.1038/nprot.2007.390
Takahashi H, Klein ZA, Bhagat SM, Kaufman AC, Kostylev MA, Ikezu T, Strittmatter SM, Alzheimer’s Disease Neuroimaging Initiative (2017) Opposing effects of progranulin deficiency on amyloid and tau pathologies via microglial TYROBP network. Acta Neuropathol (Berl). doi:10.1007/s00401-017-1668-z
Tomic JL, Pensalfini A, Head E, Glabe CG (2009) Soluble fibrillar oligomer levels are elevated in Alzheimer’s disease brain and correlate with cognitive dysfunction. Neurobiol Dis 35:352–358. doi:10.1016/j.nbd.2009.05.024
Wakselman S, Béchade C, Roumier A, Bernard D, Triller A, Bessis A (2008) Developmental neuronal death in hippocampus requires the microglial CD11b integrin and DAP12 immunoreceptor. J Neurosci 28:8138–8143. doi:10.1523/JNEUROSCI.1006-08.2008
Walker DG, Whetzel AM, Serrano G, Sue LI, Beach TG, Lue L-FF (2015) Association of CD33 polymorphism rs3865444 with Alzheimer’s disease pathology and CD33 expression in human cerebral cortex. Neurobiol Aging 36:571–582. doi:10.1016/j.neurobiolaging.2014.09.023
Wang M, Roussos P, McKenzie A, Zhou X, Kajiwara Y, Brennand KJ, De Luca GC, Crary JF, Casaccia P, Buxbaum JD, Ehrlich M, Gandy S, Goate A, Katsel P, Schadt E, Haroutunian V, Zhang B (2016) Integrative network analysis of nineteen brain regions identifies molecular signatures and networks underlying selective regional vulnerability to Alzheimer’s disease. Genome Med 8:104. doi:10.1186/s13073-016-0355-3
Wang X, Michaelis EK (2010) Selective neuronal vulnerability to oxidative stress in the brain. Front Aging Neurosci 2:12. doi:10.3389/fnagi.2010.00012
Wang Y, Ulland TK, Ulrich JD, Song W, Tzaferis JA, Hole JT, Yuan P, Mahan TE, Shi Y, Gilfillan S, Cella M, Grutzendler J, DeMattos RB, Cirrito JR, Holtzman DM, Colonna M (2016) TREM2-mediated early microglial response limits diffusion and toxicity of amyloid plaques. J Exp Med 213:667–675. doi:10.1084/jem.20151948
Wei W, Xu X, Li H, Zhang Y, Han D, Wang Y, Yan W, Wang X, Zhang J, Liu N, You Y (2014) The SIRT2 polymorphism rs10410544 and risk of Alzheimer’s disease: a meta-analysis. NeuroMol Med 16:448–456. doi:10.1007/s12017-014-8291-0
Wishart DS, Knox C, Guo AC, Shrivastava S, Hassanali M, Stothard P, Chang Z, Woolsey J (2006) DrugBank: a comprehensive resource for in silico drug discovery and exploration. Nucleic Acids Res 34:D668–D672. doi:10.1093/nar/gkj067
Yang W, Zhou X, Zimmermann HR, Cavener DR, Klann E, Ma T (2016) Repression of the eIF2α kinase PERK alleviates mGluR-LTD impairments in a mouse model of Alzheimer’s disease. Neurobiol Aging 41:19–24. doi:10.1016/j.neurobiolaging.2016.02.005
Yuan P, Condello C, Keene CD, Wang Y, Bird TD, Paul SM, Luo W, Colonna M, Baddeley D, Grutzendler J (2016) TREM2 haplodeficiency in mice and humans impairs the microglia barrier function leading to decreased amyloid compaction and severe axonal dystrophy. Neuron 90:724–739. doi:10.1016/j.neuron.2016.05.003
Zhang B, Gaiteri C, Bodea L-G, Wang Z, Joshua M, Podtelezhnikov A, Zhang C, Xie T, Tran L, Dobrin R, Fluder E, Clurman B, Melquist S, Narayanan M, Suver C, Shah H, Mahajan M, Gillis T, Mysore J, Marcy M, Lamb J, Bennett D, Molony C, Stone D, Gudnason V, Myers A, Schadt E, Neumann H, Zhu J, Emilsson V (2013) Integrated systems approach identifies genetic nodes and networks in late-onset Alzheimer’s disease. Cell 153:707–720. doi:10.1016/j.cell.2013.03.030
Zhang J, Wang Y, Chi Z, Keuss MJ, Pai Y-ME, Kang HC, Shin J-H, Bugayenko A, Wang H, Xiong Y, Pletnikov MV, Mattson MP, Dawson TM, Dawson VL (2011) The AAA+ ATPase Thorase regulates AMPA receptor-dependent synaptic plasticity and behavior. Cell 145:284–299. doi:10.1016/j.cell.2011.03.016
Zotova E, Bharambe V, Cheaveau M, Morgan W, Holmes C, Harris S, Neal JW, Love S, Nicoll JAR, Boche D (2013) Inflammatory components in human Alzheimer’s disease and after active amyloid-β42 immunization. Brain J Neurol 136:2677–2696. doi:10.1093/brain/awt210
Acknowledgements
The authors recognize the support of the National Institute on Aging Accelerated Medicines Partnership in Alzheimer’s disease (AMP-AD) U01 AG046170 (EES corresponding Principal Investigator; multi-PI to EES, BZ, SG and MEE). The study was also supported by the Louis B. Mayer Foundation (SG), by the Cure Alzheimer’s Fund (SG), by the Alzheimer’s Disease Research Division of BrightFocus Foundation (Grant No. A2016482F) (JVHM), by the Sarah and Gideon Gartner Foundation (SG), by the Georgianne and Dr Reza Khatib Foundation (SG), by the Werber Family Foundation (SG), by the Jennifer and Scott Moskowitz Foundation (SG), and by the Jane Martin and Stuart Katz Foundation (SG).
Author information
Authors and Affiliations
Contributions
MEE, SG, JVHM designed the study, JVHM, MA, SHK performed the experiments, BZ, EES, MW, JTD, BR performed the computational analysis, TF, RB performed the electrophysiological analysis, JVHM analyzed the data, WLK and CG contributed anti-oligomer antibodies, MEE, SG, JVHM wrote the paper. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
401_2017_1737_MOESM1_ESM.tiff
Suppl. Figure 1: Deficiency or absence of TYROBP does not modify microglia number in 4-month-old WT mice. (a-b) Images of NISSL (a) and Iba1 staining (b) in prefrontal cortices (PFC) and hippocampi of 4-month-old Tyrobp(+/+), Tyrobp(+/-) and Tyrobp(-/-) mice. (c-d) Flow cytometry analysis of CD45mid and CD11bhigh cells (microglia) of 4-month-old Tyrobp(+/+) (n=3) and Tyrobp(-/-) (n=7) mouse brains. Representative flow cytometry plots after live/dead and doublet cells exclusions showing CD45mid and CD11bhigh cells (blue squares) in Tyrobp(+/+) and Tyrobp(-/-) mouse brains (c). Percentages of the microglia population are presented in (d). (TIFF 7851 kb)
401_2017_1737_MOESM2_ESM.tiff
Suppl. Figure 2: Female APP/PSEN1;Tyrobp -/- mice have lower Aß40 levels than their counterparts with normal levels of Tyrobp. (a-i) Hemibrains of male and female APP/PSEN1 (n=4-6), APP/PSEN1;Tyrobp +/- (n = 3-8) and APP/PSEN1;Tyrobp -/- (n=3-4) mice were processed via differential detergent solubilization to produce fractions of TBS soluble, Triton-X soluble, and formic acid soluble Aβ. Levels of Aß40 (a-c) and Aß42 (d-f) were determined from each fraction via ELISA. The Aβ42/40 ratio was calculated for each fraction (g-i). (TIFF 756 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Haure-Mirande, JV., Audrain, M., Fanutza, T. et al. Deficiency of TYROBP, an adapter protein for TREM2 and CR3 receptors, is neuroprotective in a mouse model of early Alzheimer’s pathology. Acta Neuropathol 134, 769–788 (2017). https://doi.org/10.1007/s00401-017-1737-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00401-017-1737-3