Abstract
Background
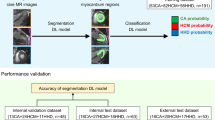
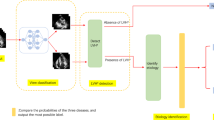
Arrhythmogenic right ventricular cardiomyopathy (ARVC) is diagnosed according to the Task Force Criteria (TFC) in which cardiovascular magnetic resonance (CMR) imaging plays an important role. Our study aims to apply an automatic deep learning-based segmentation for right and left ventricular CMR assessment and evaluate this approach for classification of the CMR TFC.
Methods
We included 227 subjects suspected of ARVC who underwent CMR. Subjects were classified into (1) ARVC patients fulfilling TFC; (2) at-risk family members; and (3) controls. To perform automatic segmentation, a Bayesian Dilated Residual Neural Network was trained and tested. Performance of automatic versus manual segmentation was assessed using Dice-coefficient and Hausdorff distance. Since automatic segmentation is most challenging in basal slices, manual correction of the automatic segmentation in the most basal slice was simulated (automatic−basal). CMR TFC calculated using manual and automatic−basal segmentation were compared using Cohen’s Kappa (κ).
Results
Automatic segmentation was trained on CMRs of 70 subjects (39.6 ± 18.1 years, 47% female) and tested on 157 subjects (36.9 ± 17.6 years, 59% female). Dice-coefficient and Hausdorff distance showed good agreement between manual and automatic segmentations (≥ 0.89 and ≤ 10.6 mm, respectively) which further improved after simulated correction of the most basal slice (≥ 0.92 and ≤ 9.2 mm, p < 0.001). Pearson correlation of volumetric and functional CMR measurements was good to excellent (automatic (r = 0.78–0.99, p < 0.001) and automatic−basal (r = 0.88–0.99, p < 0.001) measurements). CMR TFC classification using automatic−basal segmentations was comparable to manual segmentations (κ 0.98 ± 0.02) with comparable diagnostic performance.
Conclusions
Combining automatic segmentation of CMRs with correction of the most basal slice results in accurate CMR TFC classification of subjects suspected of ARVC.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
Arrhythmogenic right ventricular cardiomyopathy (ARVC) is an inherited heart disease that is characterized by ventricular dysfunction, predominantly affecting the right ventricle (RV), and potentially life-threatening ventricular arrhythmias. Accurate recognition of this disease is vital since the implantation of an implantable cardioverter defibrillator can be life-saving. ARVC is diagnosed according to the revised 2010 Task Force Criteria (TFC) [1]. Apart from electrical and family history criteria, an important role is given to the assessment of ventricular dysfunction and structural alterations. Cardiac magnetic resonance (CMR) imaging is the modality of choice for the assessment of cardiac function and dimensions in ARVC [2] since the asymmetric geometry and the position of the RV in the chest hampers visualization of the entire RV by 2D echocardiography [3].
The CMR TFC are based on RV regional wall motion abnormalities combined with cut-off values for RV ejection fraction (EF) or sex-specific cut-off values for RV indexed end-diastolic volume (EDVI)[1]. CMR can deliver one minor or two major points of the necessary four TFC points for an ARVC diagnosis. Therefore, accurate RV assessment is essential. Segmenting CMRs to measure functional and structural parameters is a laborious task, taking about 25 min to segment both ventricles in end-diastole (ED) and end-systole (ES) [4, 5]. Notably, RV segmentation takes two-thirds of this segmentation time and is prone to intra- and inter-observer variability [6]. RV segmentation difficulties can arise from the trabeculated and complex RV geometry [7, 8]. In ARVC, RV and left ventricular (LV) anatomy can be further complicated by pathological wall thinning and aneurysms due to fibrofatty replacement of the myocardial wall [2]. As a consequence, CMR misinterpretations are a key cause of over-diagnosis in ARVC [2]. The use of automatic methods for the segmentation of the ventricles may overcome these challenges. Over the last few years many state-of-the-art deep learning segmentation approaches for short-axis CMR have been developed [4, 9,10,11]. For automatic LV segmentation such methods can achieve performance level of human experts [12, 13]. However, previous studies also demonstrated that in manual and automatic segmentation of short-axis CMR, the largest disagreements and errors occur in the most basal and apical slices [8, 12,13,14,15]. Moreover, previous methods have often been evaluated on CMR datasets with limited pathology especially related to the RV. In contrast, this study included a large hospital population being evaluated for ARVC, including subjects with structurally normal hearts and those with complex structural abnormalities. In this work we apply a previously validated state-of-the-art segmentation approach [16] on a large heterogeneous hospital population of patients suspected of ARVC. The purpose of this study was to (i) evaluate our previously developed deep learning segmentation approach for RV and LV CMR assessment in patients suspected of ARVC and (ii) evaluate the clinical implication of this approach for classification of the CMR TFC in subjects suspected of ARVC.
Methods
Study population
We included a consecutive cohort of subjects suspected of ARVC who underwent CMR as part of their clinical evaluation between 2014 and 2019 at the University Medical Center (UMC) Utrecht. This yielded 241 subjects, of whom 14 were excluded because of an equivocal diagnosis (ARVC neither confirmed nor rejected) (n = 12), prior chemotherapy (n = 1) and imaging artefacts due to irregular heart rhythm (n = 1). This led to a study population of 227 subjects who were classified into three groups: (1) ARVC patients diagnosed according to the 2010 TFC (n = 53); (2) family members at-risk of developing ARVC (n = 96); and (3) control subjects initially suspected of ARVC but in whom ARVC was excluded after full clinical assessment (n = 78). Diagnosis in the control patients included RV outflow tract tachycardia (n = 45), premature ventricular contractions/non-sustained ventricular tachycardia (n = 19), mutation-negative family members of mutation-positive ARVC patients (n = 3), healthy athletes (n = 3), syncope without a cardiac cause (n = 3) repolarization abnormalities with a structurally normal heart (n = 3) and pectus excavatum (n = 2). This study was reviewed by the UMC Utrecht Institutional Review Board and was granted a waiver of informed consent.
ARVC diagnosis
ARVC diagnosis was based on the revised 2010 diagnostic TFC [1]. In short, these consensus-based criteria rely on major and minor criteria for six different categories: (1) global and regional dysfunction and structural alterations; (2) tissue characterization; (3) repolarization abnormalities; (4) depolarization/conduction abnormalities; (5) arrhythmias; and (6) family history/genetics. In each of these six categories subjects can score a minor criterium (one point), a major criterium (two points) or no criteria (0 points). A definite ARVC diagnosis was made if a subject has at least four points. The first category can be assessed by CMR, with minor criteria for regional RV wall motion abnormalities plus RVEF > 40 to ≤ 45% or RVEDVI ≥ 100 to < 110 ml/m2 (males) or ≥ 90 to < 100 mL/m2 (females) and major criteria for RV regional wall motion abnormalities plus RVEF ≤ 40% or RVEDVI ≥ 110 ml/m2 (males) or ≥ 100 ml/m2 (females) [1].
CMR dataset
All subjects underwent CMR using either 1.5 or 3 Tesla Ingenia or Achieva Philips scanners (Best, the Netherlands). The CMR dataset consisted of conventional steady-state free precession sequence short-axis and longitudinal-axis (4-chamber, 2-chamber and 3-chamber of both ventricles) cine CMR images acquired during breath holds. For this work, we only included the short-axis CMR volumes consisting of 12–18 contiguous slices covering both ventricles. The short-axis imaging parameters were as follows: each slice containing 25 to 40 phases covering one cardiac cycle with repetition time 2.6–3.4 ms and echo time 1.3–1.7 ms, flip angle 45–60 degrees. The CMR images have an in-plane resolution ranging from 1.11 to 1.45 with a slice thickness varying from 7 to 10 mm. Furthermore, reconstruction matrix of images ranges from 240 × 240 to 288 × 288 voxels. Expert radiology technicians made manual reference segmentations of the RV and LV endocardium for all CMR slices at ED and ES time frames. Both time points were manually chosen by the same experts. The CMR segmentation protocol was published previously [17] and adheres to the guidelines of the Society of Cardiovascular Magnetic Resonance (SCMR) [18]. Furthermore, the presence of RV and/or LV wall motion abnormalities was visually evaluated by an experienced cardiovascular radiologist on all available cine images and used for the calculation of the CMR TFC.
Automatic segmentation of CMR
Prior to segmentation, voxel intensities in CMR scans were normalized by rescaling the values between [0,1] based on their 1st and 99th percentiles per scan. Furthermore, voxels intensities below or above the 1st and 99th percentiles were clamped to 0 and 1, respectively.
To perform automatic segmentation of RV and LV in the 2D short-axis CMR images, we trained a Bayesian Dilated Residual Neural Network (DRN) [19] that was previously developed and evaluated by Sander et al. [16]. The Bayesian DRN was based on the original DRN from Yu et al. [19] for image segmentation. To convert the original DRN [19] into a Bayesian DRN, we implemented Monte Carlo dropout (MC dropout) introduced by Gal & Ghahramani [20]. Using a Bayesian, i.e. MC dropout approach is advantageous because multiple predictions for the same voxel can be averaged to obtain an improved final prediction per voxel [16]. Furthermore, architecture and parameters of the Bayesian DRN were identical to the model described in [16]. The network used a 2D CMR image as input and had three output channels, each providing probability for the LV, RV or background. Softmax probabilities were calculated over the three tissue classes. To train the model a combination of soft-Dice [21] and cross-entropy was used as loss function. For completeness, we provide the equations for both loss functions:
where N denotes the number of voxels in an image, \({R}_{c}\) is the binary reference image for class c and \({A}_{c}\) is the probability map for class c.
where \(p\) denotes the probability for a specific voxel \({x}_{i}\) with corresponding reference label \({y}_{i}\) for class \(c\); and \({t}_{ic}=1\) if \({y}_{i}=c\); and 0 otherwise. Hyper-parameters of the network were determined in our previous work [16] using CMR images from the MICCAI 2017 Automated Cardiac Diagnosis Challenge (ACDC) [12]. Therefore, no validation set was required in the current work.
To train the model, patches of 160 × 160 voxels were randomly chosen from the training set. Training data were augmented by 90 degree rotations, elastic deformations and gamma transformations of the images. The model was trained for 160,000 iterations using mini-batch stochastic gradient descent with batch-size 16 and Adam as optimizer [22]. Learning rate was set to 0.001 and decayed with a factor of 0.1 after every 40,000 steps. To increase generalization performance weight decay was used and set to 0.0005. Furthermore, dropout percentage was set to 0.1. Enabling MC dropout during testing, tissue class per voxel was determined using the mean softmax probabilities over 15 samples. Voxel wise segmentation may result in isolated (small clusters of) voxels. To address this, only the largest 3D connected component for each class was retained in the automatic segmentations.
Simulation of the correction of automatic segmentation
Previous research demonstrated that most segmentation inaccuracies occur in the most basal slice on the CMR [8, 12,13,14,15]. To evaluate whether these inaccuracies of our method impact TFC classification, correction of the automatic segmentation in the most basal slice of each CMR volume was simulated. This was achieved by replacing the automatic segmentation of the most basal slice with the corresponding manual reference defined by specially trained radiology technicians as a part of a regular clinical workup. We refer to this scenario as automatic−basal hereafter.
Automatic ED/ES phase selection
Accurate identification of ED and ES phase in the cardiac cycle is a prerequisite to automatically compute RVEDV and RVESV. To show the potential of the method to automatically determine the ED and ES phase, we automatically segmented all CMR volumes of the patients in the test set, and derived the RV and LV volumes for all time points of the cardiac cycle. For each patient ED was identified as the phase in which the fully automatically segmented volume was maximal and ES as the phase in which the volume was minimal. Automatically identified phases were compared with the manually selected phases using Bland–Altman analysis. In these plots (e.g. Figure 2a), the distance between automatically and manually selected phases is expressed as percentage of a complete cardiac cycle. Evaluation was performed for RV and LV separately, and for automatic and automatic−basal segmentations separately.
Evaluation of automatic segmentation
To evaluate performance of the automatic segmentation method 3D Dice-coefficient and 3D Hausdorff distance between manual and automatic segmentations were computed. For this, the 2D automatic segmentation masks were stacked into a 3D volume per patient and cardiac phase. The Dice-coefficient quantifies overlap between manual and automatic segmentation and its value ranges between 0 and 1. A higher Dice-coefficient indicates better agreement between manual and automatic segmentations. The Hausdorff distance evaluates segmentation along the boundary of the target structure by measuring the maximum distance between manual and automatic segmentation. Qualitative performance of the automatic segmentation method was visually assessed. To investigate whether segmentation errors accumulate at specific slice locations in the CMR volume the distribution of segmentation errors over slice location was computed. For this, four slice locations in a volume were distinguished: (i) most apical slice; (ii) most basal slice; (iii) mid-ventricular slices and (vi) slices located below the apex or above the base of the heart. Furthermore, to evaluate the clinical implications of our automatic CMR segmentation approach for the classification of the CMR TFC in subjects suspected of ARVC, the following CMR measurements were computed for manual, automatic and automatic−basal segmentations: LV end-diastolic volume (EDV); LV end-systolic volume (ESV); LV stroke volume (SV); LVEF; RVEDV; RVESV; RVSV and RVEF.
Statistical analysis
Statistical analysis was performed using RStudio Version 1.3.1093 (Boston, MA, USA) and IBM SPSS Statistics (version 25, USA). Continuous values were presented as mean ± standard deviation or median [interquartile range]. Categorical data were displayed as absolute frequency (n) and percentages (%). For continuous comparisons of two groups, two-tailed Student’s t test was used. For continuous comparisons of three or more groups, one-way ANOVA was used. Categorical data were compared using the chi-squared test. A p value of < 0.05 was considered significant. Comparison of automatic and manual absolute CMR measurements were assessed using Bland–Altman analysis and the Pearson correlation coefficient (r). CMR TFC was first classified using visual assessment of wall motion abnormalities and manually derived RVEDVI and RVEF, and next using visual assessment of wall motion abnormalities and automatically derived RVEDVI and RVEF.
CMR TFC classification agreement between manually vs. automatically derived CMR measurements was assessed using Cohen’s kappa (κ). Furthermore, sensitivity and specificity of CMR TFC by manual and automatic approach was determined and compared using the McNemar test.
Results
Study population
We included 70 subjects in the training set (mean age 39.6 ± 18.1 years, 47% female) and 157 subjects in the test set (mean age 36.9 ± 17.6 years, 59% female). Patient characteristics are shown in Table 1. The test set included 37 ARVC patients, 66 at-risk family members and 54 controls subjects. The distribution of subjects across the three patient categories was the same for training and test sets (34% controls, 42% at risk, 24% ARVC patients). No statistically significant difference in sex existed between the three subgroups (p = 0.37), but at-risk family members were younger than ARVC patients (p = 0.021) and controls (p < 0.001). ARVC patients had a median of 5 [4,5,6] diagnostic TFC points, while at-risk family members had a median of 2 [1,2,3] points (p < 0.001). In total, 84% of ARVC patients and 3% of at-risk family members had minor or major CMR TFC (RV wall motion abnormalities combined with abnormal RVEF or RVEDVI cut-off values). Among 103 ARVC patients and at-risk family members, 90 (87%) carried a pathogenic variant, mostly in plakophilin-2 (n = 57, 63%) followed by phospholamban (n = 26, 29%) and desmoplakin (n = 5, 6%).
Assessment of segmentation performance
Table 2 lists quantitative results of the automatic segmentation. The automatic method achieved mean Dice-coefficient for ED and ES 0.96 ± 0.01 and 0.93 ± 0.03, respectively, for the LV and 0.93 ± 0.04 and 0.89 ± 0.04, respectively, for the RV. Visual assessment of automatic segmentation results depicted in Fig. 1 reveal that performance was higher for mid-ventricular slices (second and third rows Fig. 1) compared with apical and basal slices (first and fourth row Fig. 1), while an under-segmentation of trabeculated areas occurred in the apical slices (first row Fig. 1). Furthermore, as depicted in Supplementary Fig. 1, visual assessment of the manual reference segmentation revealed a high variability of the RV shape in the basal slices in both ED and ES time points. Furthermore, as listed in Table 3, comparison of automatic with manual reference segmentations disclosed that on average 24.5% of the segmentation errors, i.e. misclassified voxels were located in the most basal slice (30.7 and 18.3% for RV and LV, respectively). In contrast, on average only 6.5% of the errors were located in an apical slice (5.4 and 7.6% for RV and LV, respectively).
example automatic segmentation vs. manual segmentation. Qualitative segmentation results for left (yellow) and right (blue) ventricles at end-systole for a patient included in the test set. Columns depict raw CMR (first column), CMR with manual reference segmentation (second column) and CMR with automatic segmentation (third column). Rows show apical, mid-ventricular and most basal slices for LV (third row) and RV (fourth row), respectively
Table 2 lists segmentation results after the simulated correction of the automatic RV and LV segmentation in the most basal slice. The results show an increased segmentation performance: mean Dice-coefficient for the ED and ES are 0.97 ± 0.01 and 0.95 ± 0.03 (vs. 0.96 ± 0.01 and 0.93 ± 0.03 uncorrected), respectively for the LV and 0.95 ± 0.02 and 0.92 ± 0.03 (vs. 0.93 ± 0.04 and 0.89 ± 0.04 uncorrected), respectively for the RV (p < 0.001 [one side Wilcoxon signed-rank test]).
Automatic ED and ES phase selection
The Bland–Altman plots shown in Fig. 2a demonstrate the comparison between automatically identified cardiac phases using the automatic segmentations with the manually selected ED and ES phases. The bias [limits of agreement] were – 0.87 [ – 6.26, 4.52]% for the ED-LV phase and – 1.64 [ – 10.28, 6.99]% for the ES-LV phase, respectively, and – 0.96 [ – 11.69, 9.76]% for the ED-RV phase and –0.05 [-7.62, 7.53]% for the ES-RV phase, respectively. Figure 2b depicts the same comparison using the automatic−basal segmentations to automatically determine the ED and ES phases. For this scenario the bias [limits of agreement] were – 0.72 [ – 5.29, 3.85]% for the ED-LV phase and – 3.03 [ – 10.08, 4.03]% for the ES-LV phase, respectively, and – 0.34 [ – 9.58, 8.89]% for the ED-RV and 0.48 [ – 7.20, 8.17]% for the ES-RV.
Bland–Altman plots with the agreement between the manually and automatically selected ED and ES phases for RV and LV, respectively, using automatic segmentations. b Bland–Altman plots with the agreement between the manually and automatically selected ED and ES phases for RV and LV, respectively, using automatic−basal segmentations. Distance between automatically and manually selected phases is expressed as percentage of a complete cardiac cycle. Evaluation was performed for RV (top row) and LV (bottom row) separately. Higher opacity of colors correlates to higher density of data points. Abbreviations: ED = end-diastolic; ES = end-systolic
Assessment of absolute CMR measurements
Automatically measured volumes (RV and LV EDV and ESV) are slightly underestimated compared to manually measured volumes (Supplementary Figs. 2 and 3). However, as shown in Table 4, the correlations of both RV and LV volumes were excellent (0.95–0.99, p < 0.001). For RV and LV EF and SV, automatic measurements seem to be slightly overestimated compared to manual measurements; nonetheless, correlations were excellent 0.82–0.89 for RV and good to excellent (0.78–0.93) for LV. After simulated manual correction of the basal slice, agreement between manual and automated measurements increased, as depicted in the Bland–Altman plots (Supplementary Figs. 2 and 3). This was also reflected in the Pearson correlation coefficient for both the volumetric (EDV, ESV) (r = 0.97–0.99, p < 0.001) as well as the functional (SV, EF) (r = 0.88–0.98, p < 0.001) CMR measurements.
Classification of ARVC TFC
Since agreement between manual and automatic measurements was higher in the automatic−basal, we used these results for the further analysis. Supplementary Table 1 depicts the mean and standard deviation of the CMR measurements stratified per subgroup. The trends between the three subgroups (ARVC, at-risk family members and controls) were comparable between manual and automated measurements: ARVC patients had significantly reduced RVEF (p < 0.001) and LVEF (p = 0.002), as well as increased RVEDVI (p < 0.001), RVESVI (p < 0.001) and LVESVI (p < 0.013) compared to at-risk family members and controls. These trends between the subgroups were also observed in the boxplots of Fig. 3.
Boxplots depicting RV and LV function and dimension. CMR measurements are given for controls, at-risk family members and ARVC patients, stratified per method (manual [orange] vs. automatic [blue]). These data represent the automatic + basal correction data, see Supplementary Fig. 4 for the boxplots of the uncorrected automatic measurements. Abbreviations: EDVI = end-diastolic volume index; EF = ejection fraction; LV = left ventricle; RV = right ventricle
We next compared CMR TFC classification using manual vs. automatic−basal CMR measurements. All but one subject (156/157, 99%) were correctly classified, showing an agreement of κ 0.98 ± 0.02. As depicted in Fig. 4, subjects who classified as no (n = 130) or minor (n = 6) CMR TFC were correctly classified using the CMR measurements computed using the automatic segmentations obtained from the deep learning segmentation model. For major TFC, all but one subject were correctly classified; with one female subject being misclassified as minor CMR TFC. This classification discrepancy was based on a 5 ml/m2 difference in RVEDVI (102 ml/m2 using manual measurements and 97 ml/m2 using automatic measurements), whereby the cutoff for major CMR TFC is set at > 100 ml/m2 in women. The total TFC in this patient went from 5 to 4, which did not change the ARVC diagnosis. Sensitivity and specificity of minor and major CMR TFC for diagnosis of ARVC were comparable for manual (minor TFC 31% | 99% and major TFC 66% | 100%) and for automatic−basal (minor TFC 35% | 100% and major TFC 65% | 100%, p = 0.32). CMR TFC classification using the uncorrected automatic measurements are depicted in Supplementary Fig. 5. This resulted in correct classification of 149/157 (95%) subjects.
Classification of CMR criteria of TFC (no, minor and major) for manual and automatic + basal correction CMR measurements. The thick blue arrows indicate the matching subjects (between manual and automatic + basal correction), the thinner blue arrows indicate the number of patients that change CMR classification category when using automated measurements. See Supplementary Fig. 5 for uncorrected automatic measurements. Abbreviations: CMR = cardiovascular magnetic resonance; N = number of subjects; TFC = Task Force Criteria
Discussion
In this study, we (i) evaluated our previously developed deep learning segmentation approach for RV and LV ventricular CMR assessment in patients suspected of ARVC and (ii) evaluated the clinical implication of this approach for classification of the CMR TFC in subjects suspected of ARVC. We demonstrated that CMR TFC classification using our automatic segmentation with limited manual correction in the most basal slice was comparable to classification using manual segmentation performed during clinical workup. Therefore, CMR TFC classification could potentially be performed using automatically measured CMR parameters with limited expert interaction.
Previous studies
Recently studies [15, 23, 24] have shown that deep learning segmentation methods outperform traditional approaches such as those exploiting level set, graph-cuts, deformable models, cardiac atlases, and statistical models [25, 26]. Many current state-of-the-art deep learning biventricular segmentation algorithms have been evaluated on publicly available cine CMR data from the MICCAI 2017 ACDC [12]. The dataset contains CMR volumes from 150 patients distributed uniformly over normal cardiac function and four disease groups: dilated cardiomyopathy, hypertrophic cardiomyopathy, ischemic cardiomyopathy, and RV abnormality (RVEDVI greater than 110 mL/m2 for men, and greater than 100 mL/m2 for women, and/or a RVEF below 40%). The ACDC challenge showed that the largest segmentation inaccuracies were located in the most basal and apical slices of the short-axis [12], which is in line with our results presented in Table 3. Comparable results were obtained in the recently held Multi-Centre, Multi-Vendor and Multi-Disease Cardiac Segmentation (M&Ms) challenge [13]. Importantly, in contrast to the ACDC and M&Ms datasets, the clinical annotation protocol used in our study adheres to the guidelines of the SCMR [18]. Segmentation of the RV, especially in basal slices, is more challenging when following SCMR guidelines compared with the protocol used for the ACDC and M&Ms datasets. For example, in the SCMR guideline the outflow tract is included as part of the RV blood volume which challenges segmentation of the basal slices due to the unclear ventricular-atrial transition.
Researchers [27, 28] have also trained and evaluated deep learning CMR segmentation algorithms on the large-scale annotated dataset from the UK Biobank [29], reaching a performance comparable with human experts. The dataset contains short-axis cine CMR volumes of 5008 subjects. As the majority of the subjects are healthy, the dataset is considered relatively homogenous [29]. In the present work, we trained and evaluated a previously developed deep learning segmentation algorithm [16] on a real-life dataset with subjects suspected of ARVC who underwent CMR as part of their clinical evaluation. Compared to the previously mentioned datasets [12, 13, 29], our dataset contains substantially more subjects with RV complexity caused by ARVC due to possible aneurysms and wall thinning and contained CMR images acquired on different field strengths (1.5 and 3 Tesla), pulse sequences and imaging parameters. Hence, the current work demonstrates that by only correcting a single slice per volume, an existing state-of-the-art segmentation method [16] is sufficiently reliable to be applied to a relevant clinical problem. Furthermore, we are the first to compare classification of the CMR TFC of subjects suspected of ARVC using manually and automatically derived CMR measurements and showing that the deep learning segmentation algorithm we use performs well in this diverse clinical environment.
Comparison to manual segmentation
We showed a good to excellent agreement of manual and automated CMR measurements, which significantly increased after simulated correction of the most basal slice of the RV and LV (automatic (r = 0.78–0.99, p < 0.001) and automatic−basal (r = 0.88–0.99, p < 0.001) measurements). This was also reflected in the significant increase of the Dice coefficients and Hausdorff distance after basal correction (p < 0.001). This is in agreement with a recent study, showing an improvement of the agreement between automatic and manual segmentation when manually adjusting the most basal slice [30].
Large intra- and inter-observer variability is currently the greatest source of error when manually segmenting CMRs [8, 31] with more variability seen for the RV compared to LV due to the RV geometrical complexity [18]. Previously published inter-observer variability ranges from 2.6 – 10.5% [32, 33] for the LV and 6.2–14.1% [33, 34] for the RV. The largest variability between manual readers also appears in the apical and basal slices [14] presumably due to low tissue contrast ratios, hypertrabeculation and unclear ventricular-atrial transition of especially the RV. The variability in contouring of the basal slice is illustrated in Supplementary Fig. 1. The corresponding manual segmentations convey the difficulty to determine the anatomical boundaries of cardiac structures in these slices. We presumed that such variability also hampers performance of the automatic segmentation method. This limitation can be alleviated by increasing the size training set. To further improve performance of deep learning segmentation approaches, especially of basal and apical short-axis slices, future work could exploit anatomical information extracted from long-axis views (2-, 3-, 4-chamber views) e.g. valve landmarks and apical point [35, 36]. Furthermore, deep learning-based CMR segmentation methods would benefit from short-axis volumes with higher through-plane resolution (e.g. using super-resolution) [10, 37, 38]. This would make application of 3D segmentation approaches more feasible and hence, those models could potentially harness any inter-slice dependencies. Finally, using explicit topological prior information [39] for model optimization is a promising training approach to prevent automatic models from generating anatomically implausible segmentation.
Clinical implementation of deep learning methods
Depending on the stage of disease, ARVC patients show a wide variety of ventricular changes that can be observed on CMR: ventricular wall motion abnormalities (e.g. aneurysms, akinesia, dyskinesia), wall thinning (due to fibrofatty replacement of the myocardium), increased trabeculations, dilatation and decreased functional measurements, that are especially present in the RV [2]. These challenges make ARVC eminently suitable to study the performance of machine learning algorithms on the RV. Previously published algorithms showed better agreement for LV than RV volumes [40]. Although limits of agreement were smaller for the LV compared to the RV, we showed comparable segmentation performance for RV and LV CMR measurements in this heterogenous study population. Furthermore, segmentation performance was comparable between structurally normal hearts and hearts affected by ARVC.
Importantly, we showed that calculation of ARVC TFC from automatically computed CMR parameters is feasible when combining automatic segmentation with correction of the most basal slice only. The diagnostic performance of the CMR TFC calculated using automatic segmentations (sensitivity 32–58%, specificity 99–100%) were comparable to manual measurements in this and previously published studies (sensitivity 13–69%, specificity 88–100%) [41, 42]. Although the correlation of manual and automatic measurements is high, the differences in CMR TFC classification without basal correction demonstrates that a fully automatic segmentation approach without human intervention is not yet reliable. However, the conducted experiments reveal that current state-of-the-art deep learning segmentation models can substantially reduce manual effort to semi-automatically segment cardiac structures in a heterogeneous dataset: manual segmentation time would be approximately 2 min instead of 25 min. Recently, Huellebrand et al.[43] proposed a human-in-the-loop approach that combines deep learning-based CMR segmentation and cardiac disease classification. The authors show that manual correction of automatic CMR segmentations by a clinical expert results in increased classification performance compared to a fully automatic segmentation approach. To identify volumes that contain segmentation failures the user can explore parallel coordinates plots that visualize CMR measurements along with cardiac shape and texture features. A similar approach was previously presented in Sander et al. [16] that combines automatic segmentation and assessment of segmentation uncertainty in CMR to automatically detect image regions containing local segmentation failures. Subsequently, detected regions are manually corrected by a clinical expert. Such a semi-automatic approach could lead to a large reduction in inter-observer variability. This is not only interesting for specialized tertiary ARVC centers, but even more for less experienced centers, since CMR misinterpretations are an important cause of over-diagnosis in ARVC and only 27% of people referred to a tertiary center with a suspected ARVC diagnosis finally meet diagnostic criteria for ARVC [44]. Our work shows that our previously developed deep learning segmentation method is able to fulfill a diagnostic purpose by simplifying accurate calculation of functional and volumetric measurements for the CMR TFC, showing opportunities to facilitate and improve individual patients health.
Limitations
Although we automated the calculation of the dimensional and functional parameters, wall motion abnormalities are also part of the CMR TFC. This was evaluated visually by experienced cardiovascular radiologists in this work, but it is subject to inter-observer variation in less experienced readers. Due to anatomical challenges of the RV a fully automatic RV strain algorithm is not yet available. Future work should focus on automatic computation of RV strain and better automatic segmentation of the basal slice, which could contribute to full automatization and standardization of the CMR TFC.
Combining automatic segmentation with manual correction of the most basal slice, 99% of the CMR TFC were correctly classified, with misclassification of only one patient from major to minor CMR TFC. Moreover, one could argue that this latter classification falls within measurement error, and it did not change the diagnosis (total TFC score went from 5 to 4). Although the absolute differences in volumetric and functional parameters were small, due to the absolute cut-off values used for the CMR TFC, differences in classification can theoretically exist when the difference is as small as 1 ml/m2, and clinical interpretation of automatic measurements remains important. Notably, CMR is no gold standard for the diagnosis of ARVC, but rather part of the diagnostic process.
Conclusions
Automatic deep learning-based CMR segmentation has the ability to provide a fast, standardized and reproducible method to measure RV and LV volumetric parameters on CMR. We demonstrate that the applied automated segmentations have a good agreement with manual segmentations. Furthermore, combining automatic segmentation with manual correction of the segmentation in the most basal slice results in accurate CMR TFC classification of subjects suspected of ARVC.
Availability of data and material
All data generated or analyzed during this study are included in this published article [and its supplementary/additional information files].
Abbreviations
- ARVC:
-
Arrhythmogenic right ventricular cardiomyopathy
- CMR:
-
Cardiovascular magnetic resonance
- DRN:
-
Dilated Residual Network
- ED:
-
End-diastole
- EDV:
-
End-diastolic volume
- EDVI:
-
End-diastolic volume index
- EF:
-
Ejection fraction
- ES:
-
End-systolic
- ESV:
-
End-systolic volume
- ESVI:
-
End-systolic volume index
- LV:
-
Left ventricle
- RV:
-
Right ventricle
- SV:
-
Stroke volume
- TFC:
-
Task force criteria
References
Marcus FI, McKenna WJ, Sherrill D, Basso C, Bauce B, Bluemke DA et al (2010) Diagnosis of arrhythmogenic right ventricular cardiomyopathy/dysplasia: proposed modification of the task force criteria. Circulation 121:1533–1541
te Riele ASJM, Tandri H, Bluemke DA (2014) Arrhythmogenic right ventricular cardiomyopathy (ARVC): cardiovascular magnetic resonance update. J Cardiovasc Magn Reson 16:50
von Knobelsdorff-Brenkenhoff F, Pilz G, Schulz-Menger J (2017) Representation of cardiovascular magnetic resonance in the AHA/ACC guidelines. J Cardiovasc Magn Reson 19:70
Bai W, Sinclair M, Tarroni G, Oktay O, Rajchl M, Vaillant G et al (2018) Automated cardiovascular magnetic resonance image analysis with fully convolutional networks. J Cardiovasc Magn Reson 20:65
Corona-Villalobos CP, Kamel IR, Rastegar N, Damico R, Kolb TM, Boyce DM et al (2015) Bidimensional measurements of right ventricular function for prediction of survival in patients with pulmonary hypertension: comparison of reproducibility and time of analysis with volumetric cardiac magnetic resonance imaging analysis. Pulm Circ 5:527–537
Grothues F, Moon JC, Bellenger NG, Smith GS, Klein HU, Pennell DJ (2004) Interstudy reproducibility of right ventricular volumes, function, and mass with cardiovascular magnetic resonance. Am Heart J 147:218–223
Haddad F, Doyle R, Murphy DJ, Hunt SA (2008) Right ventricular function in cardiovascular disease, part II: pathophysiology, clinical importance, and management of right ventricular failure. Circulation 117:1717–1731
Bonnemains L, Mandry D, Marie P-Y, Micard E, Chen B, Vuissoz P-A (2012) Assessment of right ventricle volumes and function by cardiac MRI: quantification of the regional and global interobserver variability. Magn Reson Med 67:1740–1746
Isensee F, Jaeger P, Full PM, Wolf I, Engelhardt S, Maier-Hein KH. Automatic Cardiac Disease Assessment on cine-MRI via Time-Series Segmentation and Domain Specific Features. arXiv [Internet]. 2017 [cited 2022 Jan 30]; Available from: https://arxiv.org/abs/1707.00587
Oktay O, Ferrante E, Kamnitsas K, Heinrich M, Bai W, Caballero J et al (2018) Anatomically constrained neural networks (ACNNs): application to cardiac image enhancement and segmentation. IEEE Trans Med Imaging 37:384–395
Tao Q, Yan W, Wang Y, Paiman EHM, Shamonin DP, Garg P et al (2019) Deep learning-based method for fully automatic quantification of left ventricle function from cine MR images: a multivendor multicenter study. Radiology 290:81–88
Bernard O, Lalande A, Zotti C, Cervenansky F, Yang X, Heng P-A et al (2018) Deep learning techniques for automatic MRI cardiac multi-structures segmentation and diagnosis: is the problem solved? IEEE Trans Med Imaging 37(2514):25
Campello VM, Gkontra P, Izquierdo C, Martin-Isla C, Sojoudi A, Full PM et al (2021) Multi-centre, multi-vendor and multi-disease cardiac segmentation: the M&Ms challenge. IEEE Trans Med Imaging 40:3543–3554
Suinesiaputra A, Bluemke DA, Cowan BR, Friedrich MG, Kramer CM, Kwong R et al (2015) Quantification of LV function and mass by cardiovascular magnetic resonance: multi-center variability and consensus contours. J Cardiovasc Magn Reson 17:63
Leiner T, Rueckert D, Suinesiaputra A, Baeßler B, Nezafat R, Išgum I et al (2019) Machine learning in cardiovascular magnetic resonance: basic concepts and applications. J Cardiovasc Magn Reson 21:61
Sander J, de Vos BD, Išgum I (2020) Automatic segmentation with detection of local segmentation failures in cardiac MRI. Sci Rep 10:21769
Prakken NHJ, Velthuis BK, Vonken E, Mali WP, Cramer MJM (2008) Cardiac MRI: standardized right and left ventricular quantification by briefly coaching inexperienced personnel. Open Magn Reson J 1:104–111
Schulz-Menger J, Bluemke DA, Bremerich J, Flamm SD, Fogel MA, Friedrich MG et al (2020) Standardized image interpretation and post-processing in cardiovascular magnetic resonance - 2020 update : society for cardiovascular magnetic resonance (SCMR): board of trustees task force on standardized post-processing. J Cardiovasc Magn Reson 22:19
Yu F, Koltun V, Funkhouser T. Dilated Residual Networks. 2017 IEEE Conf Comput Vis Pattern Recognit. 2017. p. 636–44.
Gal Y, Ghahramani Z (2016) Dropout as a bayesian approximation: representing model uncertainty in deep learning. In: Balcan MF, Weinberger KQ (eds) Proc 33rd int conf mach learn. New York, PMLR, pp 1050–1059
Milletari F, Navab N, Ahmadi S-A. V-Net: Fully Convolutional Neural Networks for Volumetric Medical Image Segmentation. 2016; Available from: http://arxiv.org/abs/1606.04797
Kingma DP, Ba J. Adam: A Method for Stochastic Optimization. arXiv [Internet]. 2014; Available from: http://arxiv.org/abs/1412.6980
Litjens G, Kooi T, Bejnordi BE, Setio AAA, Ciompi F, Ghafoorian M et al (2017) A survey on deep learning in medical image analysis. Med Image Anal 42:60–88
Chen C, Qin C, Qiu H, Tarroni G, Duan J, Bai W et al (2020) Deep learning for cardiac image segmentation: a review. Front Cardiovasc Med 7:25
Petitjean C, Dacher J-N (2011) A review of segmentation methods in short axis cardiac MR images. Med Image Anal 15:169–184
Peng P, Lekadir K, Gooya A, Shao L, Petersen SE, Frangi AF (2016) A review of heart chamber segmentation for structural and functional analysis using cardiac magnetic resonance imaging. MAGMA 29:155–195
Robinson R, Valindria VV, Bai W, Oktay O, Kainz B, Suzuki H et al (2019) Automated quality control in image segmentation: application to the UK biobank cardiovascular magnetic resonance imaging study. J Cardiovasc Magn Reson 21:18
Attar R, Pereañez M, Gooya A, Albà X, Zhang L, de Vila MH et al (2019) Quantitative CMR population imaging on 20,000 subjects of the UK biobank imaging study: LV/RV quantification pipeline and its evaluation. Med Image Anal 56:26–42
Petersen SE, Matthews PM, Francis JM, Robson MD, Zemrak F, Boubertakh R et al (2016) UK biobank’s cardiovascular magnetic resonance protocol. J Cardiovasc Magn Reson 18:8
Snel GJH, Poort S, Velthuis BK, van Deursen VM, Nguyen CT, Sosnovic D et al (2011) Balancing speed and accuracy in cardiac magnetic resonance function post-processing: comparing 2 levels of automation in 3 vendors to manual assessment. Diagnostics (Basel, Switzerland) 11:1758
Bhuva A, Bai W, Lau C, Davies R, Ye Y, Bulluck H et al (2019) A multicenter, scan-rescan, human and machine learning CMR study to test generalizability and precision in imaging biomarker analysis. Circ Cardiovasc Imaging 12:e009214
Moon JCC, Lorenz CH, Francis JM, Smith GC, Pennell DJ (2002) Breath-hold FLASH and FISP cardiovascular MR imaging: left ventricular volume differences and reproducibility. Radiology 223:789–797
Mooij CF, de Wit CJ, Graham DA, Powell AJ, Geva T (2008) Reproducibility of MRI measurements of right ventricular size and function in patients with normal and dilated ventricles. J Magn Reson Imaging 28:67–73
Grothues F, Smith GC, Moon JCC, Bellenger NG, Collins P, Klein HU et al (2002) Comparison of interstudy reproducibility of cardiovascular magnetic resonance with two-dimensional echocardiography in normal subjects and in patients with heart failure or left ventricular hypertrophy. Am J Cardiol 90:29–34
Xue H, Artico J, Fontana M, Moon JC, Davies RH, Kellman P (2021) Landmark detection in cardiac MRI using a convolutional neural network. Radiol Artif Intell 3:e200197
Painchaud N, Skandarani Y, Judge T, Bernard O, Lalande A, Jodoin P-M (2020) Cardiac segmentation with strong anatomical guarantees. IEEE Trans Med Imaging 39:3703–3713
Bhatia KK, Price AN, Shi W, Hajnal JV, Rueckert D. Super-resolution reconstruction of cardiac MRI using coupled dictionary learning. Proc ISBI. 2014; 947–50.
Sander J, de Vos BD, Išgum I. Autoencoding Low-Resolution MRI for Semantically Smooth Interpolation of Anisotropic MRI. Med Image Anal [Internet]. 2022; Available from: http://arxiv.org/abs/2202.09258
Clough J, Byrne N, Oksuz I, Zimmer VA, Schnabel JA, King A. A topological loss function for deep-learning based image segmentation using persistent homology. IEEE Trans Pattern Anal Mach Intell. 2020. https://doi.org/10.1109/TPAMI.2020.3013679.
Backhaus SJ, Metschies G, Billing M, Kowallick JT, Gertz RJ, Lapinskas T et al (2019) Cardiovascular magnetic resonance imaging feature tracking: Impact of training on observer performance and reproducibility. PLoS ONE 14:e0210127
Bosman LP, Cadrin-Tourigny J, Bourfiss M, Aliyari Ghasabeh M, Sharma A, Tichnell C et al (2020) Diagnosing arrhythmogenic right ventricular cardiomyopathy by 2010 task force criteria: clinical performance and simplified practical implementation. Europace 22:787–796
Aquaro GD, Barison A, Todiere G, Grigoratos C, Ait Ali L, Di Bella G et al (2016) Usefulness of combined functional assessment by cardiac magnetic resonance and tissue characterization versus task force criteria for diagnosis of arrhythmogenic right ventricular cardiomyopathy. Am J Cardiol 118:1730–1736
Huellebrand M, Ivantsits M, Tautz L, Kelle S, Hennemuth A (2022) A collaborative approach for the development and application of machine learning solutions for CMR-based cardiac disease classification. Front Cardiovasc Med 9:829512
Bomma C, Rutberg J, Tandri H, Nasir K, Roguin A, Tichnell C et al (2004) Misdiagnosis of arrhythmogenic right ventricular dysplasia/cardiomyopathy. J Cardiovasc Electrophysiol 15:300–306
Acknowledgements
We thank the ARVC patients and families who have made this work possible.
Funding
Dr. Bourfiss is supported by the Alexandre Suerman Stipend of the UMC Utrecht (2017). J. Sander is supported by the Dutch Technology Foundation (DLMedIA program (P15-26)) with participation of Pie Medical Imaging. Dr. te Riele is supported by the Dutch Heart Foundation (grant no. 2015T058), the UMC Utrecht Fellowship Clinical Research Talent, and the CVON PREDICT Young Talent Program. Dr. Asselbergs is supported by UCL Hospitals NIHR Biomedical Research Center. The Netherlands ACM Registry (www.acmregistry.nl) is supported by the Netherlands Heart Institute (project 06901).
Author information
Authors and Affiliations
Contributions
MB and JS were responsible for the conception/design of the study, acquisition/interpretation of data and drafting the manuscript. BKV and II were responsible for the conception/design of the study. BDV made substantial contributions to the method of the algorithm. BKV, II, BDV, ASJMtR and FWA were involved in revising the manuscript critically for important intellectual content.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval and consent to participate
This retrospective study was reviewed by the UMC Utrecht Institutional Review Board and was granted a waiver of informed consent.
Consent for publication
Not applicable.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bourfiss, M., Sander, J., de Vos, B.D. et al. Towards automatic classification of cardiovascular magnetic resonance Task Force Criteria for diagnosis of arrhythmogenic right ventricular cardiomyopathy. Clin Res Cardiol 112, 363–378 (2023). https://doi.org/10.1007/s00392-022-02088-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00392-022-02088-x