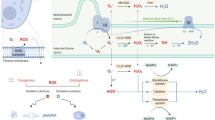
Abstract
In Saccharomyces cerevisiae, intracellular phosphate levels are maintained by the PHO pathway, activation of which is assayed by increased phosphatase activity. The PHO pathway of Schizosaccharomyces pombe upregulates phosphatase activity (encoded by pho1 +) during low extracellular phosphate levels, but the underlying mechanism is poorly understood. We utilized an alternate repressor of pho1 + expression (adenine supplementation) along with epistasis analysis to develop a model of how S. pombe PHO pathway components interact. Analyzing Pho1 activity in S. pombe PHO pathway deletion mutants during adenine starvation, we observed most mutants with a phosphatase defect in phosphate starvation also had a defect in adenine starvation. Pho7, a transcription factor in the PHO pathway, is necessary for an adenine starvation-mediated increase in Pho1 activity. Comparing adenine starvation to phosphate starvation, there are differences in the degree to which individual mutants regulate the two responses. Through epistasis studies, we identified two positive regulatory arms and one repressive arm of the PHO pathway. PKA activation is a positive regulator of Pho1 activity under both environmental conditions and is critical for transducing adenine concentrations in the cell. The synthesis of IP7 also appears critical for the induction of Pho1 activity during adenine starvation, but IP7 is not critical during phosphate starvation, which differs from S. cerevisiae. Finally, Csk1 is critical for repression of pho1 + expression during phosphate starvation. We believe all of these regulatory arms converge to increase transcription of pho1 + and some of the regulation acts through pho7 +.






Similar content being viewed by others
References
Bhoite LT, Allen JM, Garcia E, Thomas LR, Gregory ID, Voth WP, Whelihan K, Rolfes RJ, Stillman DJ (2002) Mutations in the pho2 (bas2) transcription factor that differentially affect activation with its partner proteins bas1, pho4, and swi5. J Biol Chem 277:37612–37618. doi:10.1074/jbc.M206125200
Carter-O’Connell I, Peel MT, Wykoff DD, O’Shea EK (2012) Genome-wide characterization of the phosphate starvation response in Schizosaccharomyces pombe. BMC Genom 13:697. doi:10.1186/1471-2164-13-697
Chapman AG, Atkinson DE (1977) Adenine nucleotide concentrations and turnover rates. Their correlation with biological activity in bacteria and yeast. Adv Microb Physiol 15:253–306
Conway MK, Grunwald D, Heideman W (2012) Glucose, nitrogen, and phosphate repletion in Saccharomyces cerevisiae: common transcriptional responses to different nutrient signals. G3 (Bethesda) 2:1003–1017. doi:10.1534/g3.112.002808
Elliott S, Chang CW, Schweingruber ME, Schaller J, Rickli EE, Carbon J (1986) Isolation and characterization of the structural gene for secreted acid phosphatase from Schizosaccharomyces pombe. J Biol Chem 261:2936–2941
Forsburg SL, Rhind N (2006) Basic methods for fission yeast. Yeast 23:173–183. doi:10.1002/yea.1347
Gancedo JM (2013) Biological roles of cAMP: variations on a theme in the different kingdoms of life. Biol Rev Camb Philos Soc 88:645–668. doi:10.1111/brv.12020
Gauthier S, Coulpier F, Jourdren L, Merle M, Beck S, Konrad M, Daignan-Fornier B, Pinson B (2008) Co-regulation of yeast purine and phosphate pathways in response to adenylic nucleotide variations. Mol Microbiol 68:1583–1594. doi:10.1111/j.1365-2958.2008.06261.x
Giots F, Donaton MC, Thevelein JM (2003) Inorganic phosphate is sensed by specific phosphate carriers and acts in concert with glucose as a nutrient signal for activation of the protein kinase A pathway in the yeast Saccharomyces cerevisiae. Mol Microbiol 47:1163–1181
Gupta DR, Paul SK, Oowatari Y, Matsuo Y, Kawamukai M (2011) Multistep regulation of protein kinase A in its localization, phosphorylation and binding with a regulatory subunit in fission yeast. Curr Genet 57:353–365. doi:10.1007/s00294-011-0354-2
Henry TC, Power JE, Kerwin CL, Mohammed A, Weissman JS, Cameron DM, Wykoff DD (2011) Systematic screen of Schizosaccharomyces pombe deletion collection uncovers parallel evolution of the phosphate signal transduction pathway in yeasts. Eukaryot Cell 10:198–206. doi:10.1128/EC.00216-10
Hoffman CS, Winston F (1991) Glucose repression of transcription of the Schizosaccharomyces pombe fbp1 gene occurs by a cAMP signaling pathway. Genes Dev 5:561–571
Joo YJ, Kim JA, Baek JH, Seong KM, Han KD, Song JM, Choi JY, Kim J (2009) Cooperative regulation of ADE3 transcription by Gcn4p and Bas1p in Saccharomyces cerevisiae. Eukaryot Cell 8:1268–1277. doi:10.1128/EC.00116-09
Kao RS, Morreale E, Wang L, Ivey FD, Hoffman CS (2006) Schizosaccharomyces pombe Git1 is a C2-domain protein required for glucose activation of adenylate cyclase. Genetics 173:49–61 doi:10.1534/genetics.106.055699
Karthikeyan AS, Varadarajan DK, Jain A, Held MA, Carpita NC, Raghothama KG (2007) Phosphate starvation responses are mediated by sugar signaling in Arabidopsis. Planta 225:907–918. doi:10.1007/s00425-006-0408-8
Kerwin CL, Wykoff DD (2009) Candida glabrata PHO4 Is necessary and sufficient for Pho2-independent transcription of phosphate starvation genes. Genetics 182:471–479. doi:10.1534/genetics.109.101063
Kim JH, Roy A, Jouandot D 2nd, Cho KH (2013) The glucose signaling network in yeast. Biochim Biophys Acta 1830:5204–5210. doi:10.1016/j.bbagen.2013.07.025
Lagunas R (1986) Misconceptions about the energy metabolism of Saccharomyces cerevisiae. Yeast 2:221–228. doi:10.1002/yea.320020403
Lecoq K, Belloc I, Desgranges C, Daignan-Fornier B (2001) Role of adenosine kinase in Saccharomyces cerevisiae: identification of the ADO1 gene and study of the mutant phenotypes. Yeast 18:335–342. doi:10.1002/1097-0061(20010315)18:4<335:AID-YEA674>3.0.CO;2-X
Lee YS, Mulugu S, York JD, O’Shea EK (2007) Regulation of a cyclin-CDK-CDK inhibitor complex by inositol pyrophosphates. Science 316:109–112. doi:10.1126/science.1139080
Lee YS, Huang K, Quiocho FA, O’Shea EK (2008) Molecular basis of cyclin-CDK-CKI regulation by reversible binding of an inositol pyrophosphate. Nat Chem Biol 4:25–32. doi:10.1038/nchembio.2007.52
Lenburg ME, O’Shea EK (1996) Signaling phosphate starvation. Trends Biochem Sci 21:383–387
Ljungdahl PO, Daignan-Fornier B (2012) Regulation of amino acid, nucleotide, and phosphate metabolism in Saccharomyces cerevisiae. Genetics 190:885–929. doi:10.1534/genetics.111.133306
Ma P, Goncalves T, Maretzek A, Dias MC, Thevelein JM (1997) The lag phase rather than the exponential-growth phase on glucose is associated with a higher cAMP level in wild-type and cAPK-attenuated strains of the yeast Saccharomyces cerevisiae. Microbiology 143(Pt 11):3451–3459
Markwardt DD, Garrett JM, Eberhardy S, Heideman W (1995) Activation of the Ras/cyclic AMP pathway in the yeast Saccharomyces cerevisiae does not prevent G1 arrest in response to nitrogen starvation. J Bacteriol 177:6761–6765
Mulugu S, Bai W, Fridy PC, Bastidas RJ, Otto JC, Dollins DE, Haystead TA, Ribeiro AA, York JD (2007) A conserved family of enzymes that phosphorylate inositol hexakisphosphate. Science 316:106–109. doi:10.1126/science.1139099
Pinson B, Vaur S, Sagot I, Coulpier F, Lemoine S, Daignan-Fornier B (2009) Metabolic intermediates selectively stimulate transcription factor interaction and modulate phosphate and purine pathways. Genes Dev 23:1399–1407. doi:10.1101/gad.521809
Popova Y, Thayumanavan P, Lonati E, Agrochao M, Thevelein JM (2010) Transport and signaling through the phosphate-binding site of the yeast Pho84 phosphate transceptor. Proc Natl Acad Sci USA 107:2890–2895. doi:10.1073/pnas.0906546107
Rebora K, Laloo B, Daignan-Fornier B (2005) Revisiting purine-histidine cross-pathway regulation in Saccharomyces cerevisiae: a central role for a small molecule. Genetics 170:61–70. doi:10.1534/genetics.104.039396
Schweingruber ME, Edenharter E, Zurlinden A, Stockmaier KM (1992) Regulation of pho1-encoded acid phosphatase of Schizosaccharomyces pombe by adenine and phosphate. Curr Genet 22:289–292
Secco D, Wang C, Shou H, Whelan J (2012) Phosphate homeostasis in the yeast Saccharomyces cerevisiae, the key role of the SPX domain-containing proteins. FEBS Lett 586:289–295. doi:10.1016/j.febslet.2012.01.036
Shah S, Wittmann S, Kilchert C, Vasiljeva L (2014) lncRNA recruits RNAi and the exosome to dynamically regulate pho1 expression in response to phosphate levels in fission yeast. Genes Dev 28:231–244. doi:10.1101/gad.230177.113
Tanaka K, Okayama H (2000) A pcl-like cyclin activates the Res2p-Cdc10p cell cycle “start” transcriptional factor complex in fission yeast. Mol Biol Cell 11:2845–2862
Wapinski I, Pfeffer A, Friedman N, Regev A (2007) Natural history and evolutionary principles of gene duplication in fungi. Nature 449:54–61. doi:10.1038/nature06107
Wilson MS, Livermore TM, Saiardi A (2013) Inositol pyrophosphates: between signalling and metabolism. Biochem J 452:369–379. doi:10.1042/BJ20130118
Wykoff DD, O’Shea EK (2001) Phosphate transport and sensing in Saccharomyces cerevisiae. Genetics 159:1491–1499
Acknowledgments
This work was supported by the National Science Foundation grant MCB-1121714, and MCB-1412582, the Dennis M. Cook Endowed Gregor Mendel Chair in Genetics, the Villanova College of Liberal Arts and Sciences, and the Villanova Department of Biology. We also appreciate the suggested experiments from anonymous reviewers that improved the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by C. S. Hoffman.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplemental Fig. 1 Measurement of phosphatase activity in a wild-type strain in varying phosphate and adenine conditions to demonstrate that de-repression of Pho1 activity in response to adenine starvation is not additive to phosphate starvation.
Supplemental Fig. 2 Transcript abundance of pho1 +, ade1 +, and pho84 +, after shifting from high to low adenine at time = 0 h, measured by quantitative PCR of reverse transcribed RNA. Transcripts were normalized to act1 +, and are presented as a ratio of expression. The average and standard deviation of triplicate measurements are given for each time point.
Supplemental Fig. 3 Measurement of phosphatase activity in mutants lacking ado1 +, an adenosine kinase, and asp1 +, the IP6 kinase. Phosphatase measurements suggest that Ado1 is a negative regulator of PHO1 expression, which is consistent with previous studies (Henry, et al. 2011, Lecoq, et al. 2001), and that Ado1 may be acting independently of all arms of the proposed pathway save for IP7, as the constitutive phenotype of ado1∆ is dependent on the presence of IP7.
Supplemental Fig. 4 Measurement of phosphatase activity in wild-type and pho7 + or snf5 + deletion strains and selected constitutive deletion strains. Deletion of pho7 + or snf5 + in strains deleted for aps1 + or csk1 + resulted in low phosphatase expression. Note that Pho1 expression may be more dependent on Pho7 than Snf5, suggesting that the loss of chromatin remodeling does not completely eliminate expression of PHO genes.
Supplemental Fig. 5 Measurement of phosphatase activity in strains with increased accumulation of cAMP. Mutants gpa2 R176H, cgs1-1, and cgs2-2 were generated, grown, and assayed as described in the Materials and Methods section (Hoffman and Winston 1991, Kao, et al. 2006). The cgs1-1 and cgs2-2 mutants have a slight constitutive appearance (higher level of phosphatase activity than wild-type in high adenine conditions) when grown on solid medium, but that qualitative appearance is not detectable using the quantitative liquid phosphatase assay used in this work. The discrepancy between the qualitative and quantitative results may indicate that factors in addition to cAMP are required for elevated Pho1 activity.
294_2014_466_MOESM1_ESM.pdf
Supplementary material 1 (PDF 92 kb) Supplemental Fig. 1 Measurement of phosphatase activity in a wild-type strain in varying phosphate and adenine conditions to demonstrate that de-repression of Pho1 activity in response to adenine starvation is not additive to phosphate starvation. Supplemental Fig. 2 Transcript abundance of pho1 +, ade1 +, and pho84 +, after shifting from high to low adenine at time = 0 hours, measured by quantitative PCR of reverse transcribed RNA. Transcripts were normalized to act1 +, and are presented as a ratio of expression. The average and standard deviation of triplicate measurements are given for each time point. Supplemental Fig. 3 Measurement of phosphatase activity in mutants lacking ado1 +, an adenosine kinase, and asp1 +, the IP6 kinase. Phosphatase measurements suggest that Ado1 is a negative regulator of PHO1 expression, which is consistent with previous studies (Henry, et al. 2011, Lecoq, et al. 2001), and that Ado1 may be acting independently of all arms of the proposed pathway save for IP7, as the constitutive phenotype of ado1Δ is dependent on the presence of IP7. Supplemental Fig. 4 Measurement of phosphatase activity in wild-type and pho7 + or snf5 + deletion strains and selected constitutive deletion strains. Deletion of pho7 + or snf5 + in strains deleted for aps1 + or csk1 + resulted in low phosphatase expression. Note that Pho1 expression may be more dependent on Pho7 than Snf5, suggesting that the loss of chromatin remodeling does not completely eliminate expression of PHO genes. Supplemental Fig. 5 Measurement of phosphatase activity in strains with increased accumulation of cAMP. Mutants gpa2 R176H, cgs1-1, and cgs2-2 were generated, grown, and assayed as described in the Materials and Methods section (Hoffman and Winston 1991, Kao, et al. 2006). The cgs1-1 and cgs2-2 mutants have a slight constitutive appearance (higher level of phosphatase activity than wild-type in high adenine conditions) when grown on solid medium, but that qualitative appearance is not detectable using the quantitative liquid phosphatase assay used in this work. The discrepancy between the qualitative and quantitative results may indicate that factors in addition to cAMP are required for elevated Pho1 activity.
Rights and permissions
About this article
Cite this article
Estill, M., Kerwin-Iosue, C.L. & Wykoff, D.D. Dissection of the PHO pathway in Schizosaccharomyces pombe using epistasis and the alternate repressor adenine. Curr Genet 61, 175–183 (2015). https://doi.org/10.1007/s00294-014-0466-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-014-0466-6