Abstract
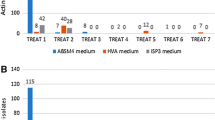
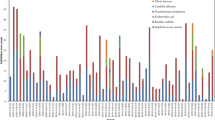
Marine actinomycetes are less investigated compared to terrestrial strains as potential sources of natural products. To date, few investigations have been performed on culturable actinomycetes associated with South China Sea sediments. In the present study, twenty-eight actinomycetes were recovered from South China Sea sediments after dereplication by traditional culture-dependent method. The 16S rRNA gene sequences analyses revealed that these strains related to five families and seven genera. Twelve representative strains possessed at least one of the biosynthetic genes coding for polyketide synthase I, II, and nonribosomal peptide synthetase. Four strains had anti-Mycobacterium phlei activities and five strains had activities against methicillin-resistant Staphylococcus aureus. 10 L-scale fermentation of strains Salinispora sp. NHF45, Nocardiopsis sp. NHF48, and Streptomyces sp. NHF86 were carried out for novel and bioactive compounds discovery. Finally, we obtained a novel α-pyrone compound from marine Nocardiopsis sp. NHF48, an analogue of paulomenol from marine Streptomyces sp. NHF86 and a new source of rifamycin B, produced by Salinispora sp. NHF45. The present study concluded that marine actinomycetes, which we isolated from South China Sea sediments, will be a suitable source for the development of novel and bioactive compounds.



Similar content being viewed by others
References
Argoudelis AD, Baczynskyj L, Haak WJ, Knoll WM, Mizsak SA, Shilliday FB (1988) New paulomycins produced by Streptomyces paulus. J Antibiot 41:157–169
Argoudelis AD, Baczynskyj L, Mizsak SA, Shilliday FB (1988) O-demethylpaulomycin-A and O-demethylpaulomycin-B, U-77,802 and U-77,803, paulomenol-A and paulomenol-B, new metabolites produced by Streptomyces paulus. J Antibiot 41:1316–1330
Baltz RH (2006) Marcel Faber Roundtable: is our antibiotic pipeline unproductive because of starvation, constipation or lack of inspiration? J Ind Microbiol Biotechnol 33:507–513. doi:10.1371/journal.pone.0149216
Berdy J (2005) Bioactive microbial metabolites—a personal view. J Antibiot 58:1–26
Chen C, Wang J, Guo H, Hou W, Yang N, Ren B, Liu M, Dai H, Liu X, Song F, Zhang L (2013) Three antimycobacterial metabolites identified from a marine-derived Streptomyces sp. MS100061. Appl Microbiol Biotechnol 97:3885–3892. doi:10.1007/s00253-012-4681-0
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791. doi:10.1111/j.1558-5646.1985.tb00420.x
Fenical W, Jensen PR (2006) Developing a new resource for drug discovery: marine actinomycete bacteria. Nat Chem Biol 2:666–673. doi:10.1038/nchembio841
Fischbach MA, Walsh CT (2006) Assembly-line enzymology for polyketide and nonribosomal peptide antibiotics: logic, machinery, and mechanisms. Chem Rev 106:3468–3496. doi:10.1007/s10295-013-1348-5
Gomez-Escribano JP, Bibb MJ (2014) Heterologous expression of natural product biosynthetic gene clusters in Streptomyces coelicolor: from genome mining to manipulation of biosynthetic pathways. J Ind Microbiol Biotechnol 41:425–431. doi:10.1007/s10295-013-1348-5
Guo X, Geng P, Bai F, Bai G, Sun T, Li X, Shi L, Zhong Q (2012) Draft genome sequence of Streptomyces coelicoflavus ZG0656 reveals the putative biosynthetic gene cluster of acarviostatin family alpha-amylase inhibitors. Lett Appl Microbiol 55:162–169. doi:10.1111/j.1472-765X.2012.03274.x
Hodges TW, Slattery M, Olson JB (2012) Unique actinomycetes from marine caves and coral reef sediments provide novel PKS and NRPS biosynthetic gene clusters. Mar Biotechnol 14:270–280. doi:10.1007/s10126-011-9410-7 (NY)
Kamjam M, Sivalingam P, Deng Z, Hong K (2017) Deep sea actinomycetes and their secondary metabolites. Front Microbiol 8:760. doi:10.3390/md15030071
Kwon HC, Kauffman CA, Jensen PR, Fenical W (2006) Marinomycins A-D, antitumor-antibiotics of a new structure class from a marine actinomycete of the recently discovered genus “Marinispora”. J Am Chem Soc 128:1622–1632. doi:10.1021/ja0558948
Lazzarini A, Cavaletti L, Toppo G, Marinelli F (2001) Rare genera of actinomycetes as potential producers of new antibiotics. Anton Leeuw Int J G 79:399–405. doi:10.1023/A:1010287600557
Liao L, Chen R, Jiang M, Tian X, Liu H, Yu Y, Fan C, Chen B (2016) Bioprospecting potential of halogenases from Arctic marine actinomycetes. BMC Microbiol 16:34. doi:10.1186/s12866-016-0662-2
Liu W, Ahlert J, Gao Q, Wendt-Pienkowski E, Shen B, Thorson JS (2003) Rapid PCR amplification of minimal enediyne polyketide synthase cassettes leads to a predictive familial classification model. Proc Natl Acad Sci USA 100:11959–11963. doi:10.1073/pnas.2034291100
Moore JM, Bradshaw E, Seipke RF, Hutchings MI, McArthur M (2012) Use and discovery of chemical elicitors that stimulate biosynthetic gene clusters in Streptomyces bacteria. Methods Enzymol 517:367–385. doi:10.1016/B978-0-12-404634-4.00018-8
Pathom-aree W, Stach JEM, Ward AC, Horikoshi K, Bull AT, Goodfellow M (2006) Diversity of actinomycetes isolated from Challenger Deep sediment (10,898 m) from the Mariana Trench. Extremophiles 10:181–189. doi:10.1007/s00792-005-0482-z
Prieto-Davó A, Dias T, Gomes SE, Rodrigues S, Parera-Valadezl Y, Borralho PM, Pereira F, Rodrigues CMP, Santos-Sanches I, Gaudencio SP (2016) The madeira archipelago as a significant source of marine-derived actinomycete diversity with anticancer and antimicrobial potential. Front Microbiol. doi:10.3389/fmicb.2016.01594
Qin S, Li J, Chen HH, Zhao GZ, Zhu WY, Jiang CL, Xu LH, Li WJ (2009) Isolation, diversity, and antimicrobial activity of rare actinobacteria from medicinal plants of tropical rain forests in Xishuangbanna, China. Appl Environ Microbiol 75:6176–6186. doi:10.1128/AEM.01034-09
Raahave D (1974) Paper disk-agar diffusion assay of penicillin in the presence of streptomycin. Antimicrob Agents Chemother 6:603–605
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Schäberle TF (2016) Biosynthesis of alpha-pyrones. Beilstein J Org Chem 12:571–588. doi:10.3762/bjoc.12.56
Song Y, Huang H, Chen Y, Ding J, Zhang Y, Sun A, Zhang W, Ju J (2013) Cytotoxic and antibacterial marfuraquinocins from the deep South China Sea-derived Streptomyces niveus SCSIO 3406. J Nat Prod 76:2263–2268. doi:10.1021/np4006025
Subramani R, Aalbersberg W (2012) Marine actinomycetes: an ongoing source of novel bioactive metabolites. Microbiol Res 167:571–580. doi:10.1016/j.micres.2012.06.005
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729. doi:10.1093/molbev/mst197
Tian XP, Zhang S, Li W (2011) Advance in marine actinobacterial research—a review. Wei sheng wu xue bao= Acta microbiologica Sinica 51:161–169
Toske SG, Jensen PR, Kauffman CA, Fenical W (1995) Elijopyrones A-D: new α-pyrones from a marine actinomycete. Nat Prod Lett 6:303–308. doi:10.1080/10575639508043175
Wawrik B, Kutliev D, Abdivasievna UA, Kukor JJ, Zylstra GJ, Kerkhof L (2007) Biogeography of actinomycete communities and type II polyketide synthase genes in soils collected in New Jersey and Central Asia. Appl Environ Microbiol 73:2982–2989. doi:10.1128/AEM.02611-06
Wiley PF, Mizsak SA, Baczynskyj L, Argoudelis AD (1984) The structure of paulomycin. J Antibiot 37:1273–1275
Yang N, Sun C (2016) The inhibition and resistance mechanisms of actinonin, isolated from marine Streptomyces sp. NHF165, against Vibrio anguillarum. Front Microbiol 7:1467. doi:10.3389/fmicb.2016.01467
Zhang W, Liu Z, Li S, Lu Y, Chen Y, Zhang H, Zhang G, Zhu Y, Zhang G, Zhang W, Liu J, Zhang C (2012) Fluostatins I-K from the South China Sea-derived Micromonospora rosaria SCSIO N160. J Nat Prod 75:1937–1943. doi:10.1021/np300505y
Zhang XM, Sun MW, Shi H, Lu CH (2017) α-Pyrone derivatives from a marine actinomycete Nocardiopsis sp. YIM M13066. Nat Prod Res. doi:10.1080/14786419.2017.1299730
Zou G, Liao XJ, Peng Q, Chen GD, Wei FY, Xu ZX, Zhao BX, Xu SH (2017) A new alpha-pyrone from the deep-sea actinomycete Nocardiopsis dassonvillei subsp. dassonvillei DSM 43111(T). J Asian Nat Prod Res. doi:10.1080/10286020.2017.1307186
Acknowledgements
This work was supported by grants from Shandong Provincial Natural Science Foundation, China (ZR2016CB13) and National Natural Science Foundation of China (31600136).
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yang, N., Song, F. Bioprospecting of Novel and Bioactive Compounds from Marine Actinomycetes Isolated from South China Sea Sediments. Curr Microbiol 75, 142–149 (2018). https://doi.org/10.1007/s00284-017-1358-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-017-1358-z