Abstract
Hydrogen is an alternative fuel for transportation vehicles because it is clean, sustainable, and highly flammable. However, the production of hydrogen from lignocellulosic biomass by microorganisms presents challenges. This microbial process involves multiple complex steps, including thermal, chemical, and mechanical treatment of biomass to remove hemicellulose and lignin, as well as enzymatic hydrolysis to solubilize the plant cell walls. These steps not only incur costs but also result in the production of toxic hydrolysates, which inhibit microbial growth. A hyper-thermophilic bacterium of Caldicellulosiruptor bescii can produce hydrogen by decomposing and fermenting plant biomass without the need for conventional pretreatment. It is considered as a consolidated bioprocessing (CBP) microorganism. This review summarizes the basic scientific knowledge and hydrogen-producing capacity of C. bescii. Its genetic system and metabolic engineering strategies to improve hydrogen production are also discussed.
Key points
• Hydrogen is an alternative and eco-friendly fuel.
• Caldicellulosiruptor bescii produces hydrogen with a high yield in nature.
• Metabolic engineering can make C. bescii to improve hydrogen production.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Currently, there are increasing concerns about serious environmental problems such as the greenhouse effect, global climate change, fine dust caused by the use of fossil fuels, and other complications caused by the depletion of fossil fuels (Fawzy et al. 2020; Manisalidis et al. 2020; Martins et al. 2019). To address these issues, there has been a growing interest in biofuels, such as bioethanol, biodiesel, and biohydrogen, which are produced through biological processes using various renewable resources (Cha et al. 2013a; Hoang et al. 2023; Jeswani et al. 2020; Martínez-Jaramillo et al. 2019). Among these biofuels, hydrogen is an attractive and promising option for two important reasons: (i) hydrogen is non-toxic and does not release the greenhouse gas CO2 when combusted (clean energy), and (ii) it carries higher energy compared to other hydrocarbon fuels (Hassan et al. 2023; Okolie et al. 2021). Therefore, hydrogen has been suggested as a major chemical energy carrier and could be utilized as a high-energy storage for transportation vehicles (Miller et al. 2021). Hydrogen can be utilized not only as a commercial transportation fuel but also in the chemical industry (chemical looping hydrogen) for the production of methanol and ammonia (Palone et al. 2023), as well as in various other industries such as electronics, metals, and food. Unfortunately, hydrogen does not exist in a free form in nature. However, it is present in water or in the main components of all living organisms, suggesting that hydrogen can be produced through biological processes (Akhlaghi and Najafpour-Darzi 2020; Lepage et al. 2021).
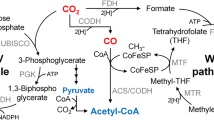
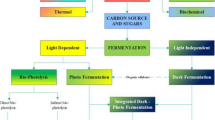
The biological production of hydrogen can be accomplished through various steps including enzymatic saccharification of renewable biomass to convert into fermentable sugars, as well as anaerobic fermentation of these sugars to hydrogen by anaerobic bacteria (Fig. 1) (Alicia Benitez et al. 2021; Cha et al. 2013a, 2016). C5 and C6 sugars, which are derived from a variety of carbohydrates like glucose and xylose found in plant biomass, are oxidized via the Embden–Meyerhof–Parnas glycolytic pathway (Fig. 1) to produce acetate, lactate, carbon dioxide, and hydrogen (Cha et al. 2013a, 2016, 2023; Chandel 2021). In terms of final fermentative products, carbon flow is directed towards lactate or acetyl-CoA, while electrons flux towards lactate and H2 from pyruvate, which serves as a major metabolic branch point (Fig. 1).
Microorganisms are crucial for achieving high yields of hydrogen, especially with the involvement of thermophiles such as Thermoanaerobacter tengcongensis (~ 4.0 mol H2/mol glucose) (Soboh et al. 2004), Thermotoga maritima (~ 4.0 mol H2/mol glucose) (Schroder et al. 1994; Singh et al. 2019), and Thermococcus kodakarensis (~ 3.3 mol H2/mol glucose) (Burkhart et al. 2019; Kanai et al. 2005). The hyperthermophile, Pyrococcus furiosus (optimal temperature 90 °C), also produced ~ 2.8 mol H2/mol glucose (Servé and Kengen 1994; Song et al. 2019) although it is smaller compared to others. The utilization of high temperatures (> 50 °C) should be beneficial for hydrogen production due to reduced viscosity, improved mixing, trace contamination, enhanced reaction rates, and the elimination of the need for reactor cooling (Shahbeik et al. 2022). Additional microorganisms capable of producing hydrogen are listed in Table 1. While most hydrogen-producing microorganisms rely on starch-based biomass, which can be easily saccharified, the direct production of hydrogen from lignocellulosic biomass is challenging for microorganisms as it requires additional steps, such as pretreatment, enzymatic saccharification, and the generation of fermentation inhibitors (Zafar et al. 2021). Therefore, the development of microbial strains that fermenting lignocellulosic biomass effectively is necessary (Kim et al. 2022).
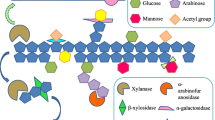
The genus Caldicellulosiruptor is a thermophilic microorganism with cellulosic activity. It can produce hydrogen at high rates from lignocellulosic biomass, with an optimal growth temperature between 75 and 80 °C (Scott et al. 2019). C. bescii can serve as a consolidated bioprocessing (CBP) organism (Fig. 2) because it can utilize both C5 and C6 sugars simultaneously and directly convert lignocellulosic biomass without conventional pretreatment steps (Fig. 2) (Cha et al. 2013a, 2016; Chung et al. 2014; Periyasamy et al. 2023). The C. bescii genome encodes many carbohydrate-active enzymes (CAZymes), which are multi-domain enzymes with cellulolytic and hemicellulolytic activity and utilize a broad range of substrates, including plant biomass, without the need for conventional pretreatment (Kim et al. 2019). Therefore, there is potential to improve the economics of biofuel production from lignocellulosic biomass by skipping thermal, chemical, and mechanical treatment steps (Cha et al. 2013a; Chung et al. 2014).
This review provides the scientific knowledge and data on C. bescii, focusing on its H2 production capacity, genetic system, and metabolic engineering strategies, which can make C. bescii a highly efficient organism for hydrogen production at high temperatures.
Biosynthetic pathway of hydrogen of C. bescii
Usually, the genus Caldicellulosiruptor produces relatively high yields of H2 (4 mol of H2/ mol of glucose) compared with other microorganisms (Cha et al. 2013a, 2016; Straub et al. 2020). Additionally, acetate should be coupled with H2 production for the reoxidation of NADH (a two-electron donor) and ferredoxin (a one-electron donor) (Buckel 2021; Cha et al. 2016; White 2012) (Fig. 1).
As shown in Fig. 1, H2 is produced through proton reduction catalyzed by hydrogenases (Cha et al. 2016; Jay et al. 2020; Lu and Koo 2019). These hydrogenases are metalloenzymes that contain iron in their active site, such as di-iron, nickel–iron, or iron-sulfur clusters (Lu and Koo 2019). Specifically, Caldicellulosiruptor spp. have only two types of hydrogenases: bifurcating [Fe–Fe] hydrogenase and [Ni–Fe] hydrogenase (Cha et al. 2013a, 2016; Zhang et al. 2021). NADH and ferredoxin are catabolized by the bifurcating [Fe–Fe] hydrogenase, resulting in the production of H2 (Cha et al. 2016; Zhang et al. 2021). On the other hand, the [Ni–Fe] hydrogenase is a membrane-bound heterodimer and is widely found in nature (Alfano and Cavazza 2020). Although the [Ni–Fe] hydrogenase also catalyzes H2 production, the bifurcating [Fe–Fe] hydrogenase is the primary enzyme for H2 production in C. bescii, while the main role of the [Ni–Fe] hydrogenase is to pump out protons across the cellular membrane to generate the “proton motive force” (Kaila and Wikstrom 2021; White 2012).
Hydrogen production from C5 and C6 sugars
In most of the studies reported, the maximum amount of hydrogen produced was 2.0–3.8 mol of H2 /mol of glucose (C6 sugars) due to the formation of co-products such as lactic acid and acetic acid (Esercizio et al. 2021). The theoretical molar yield of hydrogen from xylose (C5 sugar) fermentation is 3.3 mol of H2/mol of xylose (C5 sugar) with acetate as the sole byproduct, but the reported values were lower than 2 mol of H2/mol of xylose (Chiu-Yue Lin and Hung 2008). Hydrogen production has been reported to be between 0.5 and 4 L/L/day (Ghimire et al. 2015; Beckers et al. 2015). The nature, carbohydrate content, and biodegradability of carbon substrate play an important role in the H2 yield, production rate, and overall economics of the process (Nanqi et al. 2011). Many bacterial species have been reported to produce hydrogen from C5 and C6 sugars, including enteric bacteria such as Enterobacter aerogenes, Enterobacter cloacae, and Escherichia coli, which produce about 1–2 mol of H2/mol of glucose (Yoshida et al. 2006). Clostridium spp. also produce similar amounts (Liu et al. 2006). Caldicellulosiruptor spp. produce about 3–4 mol of H2/mol of glucose (Willquist et al. 2010). Enterobacter utilizes a formate-H2 lyase, and Clostridium spp. use a ferredoxin-dependent hydrogenase to avoid the thermodynamically unfavorable formation of H2 from NADH (Schut and Adams 2009). Based on previous studies on various microorganisms, metabolic engineering and pre-treatment of substrates can increase hydrogen production by improving the biodegradability of substrates. In summary, C. bescii can be a superior organism as it does not contain competing pathways other than lactate, offers the potential to produce maximum amounts of H2 (4 mol of H2/mol of C5 and C6 sugars), and is tractable to metabolic engineering.
Hydrogen production from various biomass by C. bescii
The most important aspect of C. bescii is its ability to decompose various monosaccharides and polysaccharides, such as glucose, xylose, crystalline cellulose, and non-pretreated plant biomass. To compare hydrogen production from real-world substrates, C. bescii wild type (JWCB001) and its mutant strains (JWCB005 and 017) were grown on 0.5% switchgrass (Cha et al. 2013a). The strains were incubated for 120 h; then, hydrogen production was measured. The hydrogen productions of JWCB001 and JWCB005 were ~ 1.8 mol/mol of glucose and ~ 1.7 mol/mol of glucose, and it is a bit lower than H2 production by Caldicellulosiruptor saccharolyticus (~ 2.5 mol/mol of glucose). However, in the case of C. saccharolyticus, yeast extracts were added to the growth medium. Even without the addition of yeast extracts, the engineered C. bescii strain JWCB017 produced significantly more hydrogen (~ 3.4 mol/mol of glucose; Table 2) (Cha et al. 2013a). Actually, C. bescii lacks the enzyme required for ethanol production. Chung et al. reported the heterologous expression of adhE to enable C. bescii to produce alcohol (Chung et al. 2014). However, this resulted in reduced hydrogen production as carbon and electrons were redirected for alcohol production (Chung et al. 2014). C. bescii can also produce hydrogen from barley straw and Miscanthus. In a previous study, JWCB018 without the chromosomal ldh gene produced 63% and 25% more hydrogen from barley straw and Miscanthus than the wild-type strain (JWCB001), respectively. It might be due to a decrease in lactate production by interrupting lactate dehydrogenase function by a native active transposon (Cha et al. 2023, 2013b). Yilmazel and Duran reported hydrogen production in co-substrate reactors, where C. bescii was grown on four different substrate mixtures of cattle manure (CM), switchgrass (SG), and biosolid (BS). C. bescii grown on BS + SG + CM (~ 15.0 mM) showed much better hydrogen production compared to BS + SG (~ 11.0 mM), exhibiting synergistic effects of co-fermentation of these feedstocks (Yilmazel and Duran 2021).
Genetic system for C. bescii
There are many interesting thermophiles that produce interesting and important chemicals. The ability to manipulate C. bescii genes is required to make the hyper-thermophilic strain more useful in the real world. However, the wild-type strain of C. bescii is not sufficient to produce biofuel, so it needs to be metabolically engineered to produce biofuel at a suitable yield. One of the most difficult aspects of studying hyperthermophiles like C. bescii is the lack of genetic tools for metabolic engineering. In order to develop a genetic tool, there are several requirements: (i) overcoming the restriction-modification (R-M) system, (ii) constructing a E. coli-C. bescii shuttle vector, (iii) establishing a transformation method, and (iv) selecting a selection marker (Chung et al. 2013a). One significant barrier to develop genetic tools for uncharacterized microorganisms, especially hyperthermophiles, is the lack of selectable markers. Antibiotics are typically used in mesophilic bacteria, but not in thermophiles because thermostable antibiotic markers are usually not available at high temperatures over 70 °C (Crosby et al. 2019). Because of their high growth temperatures, the genetics of most thermophiles depend on auxotrophic mutant strains. This method is often used for many thermophiles including Caldicellulosiruptor (Cha et al. 2013a, 2016; Chung et al. 2014; Lipscomb et al. 2016), Sulfolobus (Wagner et al. 2012; Zheng et al. 2012), and Thermotoga sp. RQ7 (Han and Xu 2017). The selection method (using an auxotroph mutant strain) for transformation in C. bescii relies on the loss of the uracil biosynthetic enzyme coding for orotidine-5′-monophosphate (OMP) decarboxylase (pyrF), which was first described in yeast (Boeke et al. 1984) and has been a useful genetic tool in both bacteria and archaea (Lucas et al. 2002). In order to generate a spontaneous pyrF mutant strain, the cells were grown on low osmolarity-defined growth medium (LOD) (Farkas et al. 2013) supplemented with uracil and 5-fluoroorotic acid (5-FOA). The strain with ΔpyrFA, C. bescii JWCB005, was obtained as a host strain for gene manipulation (Chung et al. 2013a). In order to create a shuttle vector capable of replicating in both Escherichia coli and C. bescii, the pyrF gene for uracil auxotroph was cloned and inserted into pBAS2 vector (Clausen et al. 2004), which is a small plasmid with a replication origin of the two plasmids in C. bescii. The E. coli/C. bescii shuttle vector pDCW89 was constructed by linking a low copy replication origin of E. coli, PSC101, and apramycin-resistant gene cassette (AprR) to pBAS2 vector (Clausen et al. 2004; Dam et al. 2011). Although a shuttle plasmid is available, there is still another barrier that needs to be addressed to manipulate C. bescii genes. The biggest obstacle when transforming foreign DNA for deletion/insertion of a gene is the restriction-modification (R-M) system, which recognizes the difference in DNA methylation when foreign DNA is introduced into the cells, leading to the degradation of the foreign DNA by the restriction system in the strain (Chung et al. 2013b). When the pDCW89 shuttle vector is transformed into the pyrF deleted strain by electroporation, the transformation competency is significantly low because C. bescii has its own restriction endonuclease, CbeI (Cbe_2438), which was discovered by Chung et al. (2011). CbeI has a HaeIII-like activity and is a type II restriction endonuclease that cleaves unmethylated sequences at 5′-GG/CC-3′ (Chung et al. 2013b; Han et al. 2014). The CbeI activity should be removed in the host strains for successful DNA transformation. CbeI (Cbes_2438) and a neighboring α-class N4-cytosine methyltransferase (M.CbeI, Cbes_2437) were confirmed to be the counterpart of the R-M system in C. bescii (Chung et al. 2013b). Treatment of the E. coli/C. bescii shuttle plasmid DNA with cloned M.CbeI protein resulted in efficient transformation. Chung et al. also reported a cbeI deletion (Cbes_2438) and generated a new host strain, C. bescii JWCB018 (ΔpyrAF ΔcbeI Δldh), through homologous recombination. JWCB018 (ΔpyrAF ΔcbeI Δldh) can be transformed by DNA isolated from E. coli without in vitro methylation (Chung et al. 2013b). A brief procedure of gene deletion is described in Fig. 3.
Enhanced hydrogen production by metabolically engineered C. bescii
The ultimate goal of biofuel production is to construct a microbial strain for a consolidated bioprocessing (CBP) organism (Olguin-Maciel et al. 2020), which is an organism capable of producing biofuels, such as alcohol and hydrogen gas, through a one-step process using plant biomass (Fig. 2).
Previous research has shown (Cha et al. 2013a, 2016, 2023; Farkas et al. 2013) that various stains of C. bescii were constructed using metabolic engineering techniques to enhance its capabilities as a CBP organism (Cha et al. 2013a). The first engineered C. bescii strain, obtained through a newly developed metabolic engineering technique, was a mutant strain with a deletion in the lactate dehydrogenase gene (ldh), resulting in the complete absence of lactate production. By removing the lactate production pathway, more carbon should be flowed to acetate and more electron flux carried by NADHs runs to acetate and hydrogen production from pyruvate. The ldh-deleted strain of C. bescii (JWBC017) was grown on 0.5% switchgrass supplemented as the sole carbon source and showed increased production of acetate and hydrogen but no lactate production (Fig. 1) (Cha et al. 2013a). The ldh-deleted strain producing no lactate showed a 6.5% conversion of cellobiose to acetate (9.2 mM) with 105% overall carbon recovery (Cha et al. 2013a). The metabolically engineered strain JWCB017 (ΔpyrFA, Δldh) produced more hydrogen (~ 3.4 mol/mol of glucose) compared to wild-type C. bescii (~ 1.8 mol/mol of glucose) and C. saccharolyticus (~ 2.5 mol/mol of glucose) (de Vrije et al. 2007). This indicates that C. saccharolyticus wild type produced more hydrogen than C. bescii wild type due to the use of yeast extract in the culture media, which can improve growth and yield.
As mentioned earlier, hydrogenases play a key role in microbial energy metabolism, but the exact nature and function of these enzymes remain unclear. Cha et al. (2016) reported the deletion of a gene cluster called hypABFCDE, which encodes the maturation proteins for the C. bescii [Ni–Fe] hydrogenase. The resulting mutant strain, JWCB038 (Cha et al. 2016), exhibited slower growth compared to its wild type or parent strain (JWCB005, ΔpyrFA) because the main function of the [Ni–Fe] hydrogenase may act as a proton pump generating a proton motive force (PMF) across the cellular membrane for ATP synthesis (Fig. 1). The data (Cha et al. 2016) also indicated that the mutant strain JWCB038 did not exhibit a significant reduction in hydrogen production, suggesting that the [Ni–Fe] hydrogenase may not be the main enzyme involved, and that the bifurcating [Fe–Fe] hydrogenase might be the primary enzyme responsible for hydrogen production in C. bescii.
To remove the R-M system in C. bescii to facilitate metabolic engineering, a cbeI deletion strain was generated (Chung et al. 2013b). However, the function of LDH was also disrupted by a native active transposon at the same time (Cha et al. 2013b). The resulting strain JWCB018 (ΔpyrFA ΔcbeI Δldh) (Cha et al. 2013b; Chung et al. 2013b) was grown on 10 g/L cellobiose, 20 g/L Avicel, 10 g/L barley straw, and 10 g/L Miscanthus as carbon sources and then compared to the C. bescii wild type for hydrogen production (Cha et al. 2023). Because of the interruption of the lactate producing pathway in the mutant strain JWCB018, there was an increase in NADHs carrying electrons, resulting in enhanced H2 production. This study showed that JWCB018 exhibited up to 25%, 21%, 33%, and 25% increases in H2 production on 1.0% cellobiose, 2% Avicel, 1.0% barley straw, and 1.0% Miscanthus, respectively (Cha et al. 2023). These findings clearly indicate that appropriate metabolic engineering can significantly enhance the production of H2 and other valuable chemicals.
Further strategies to improve hydrogen production by C. bescii
Although very useful gene manipulation techniques for C. bescii metabolic engineering have been developed and research for biofuel production is being intensively conducted, there is still a need for the development of more efficient genetic tools and techniques for thermophiles, especially C. bescii. For example, promoters for high expression and better replicating plasmids for heterologous gene expression should be developed. Instead of using an uracil auxotroph, new thermo-stable antibiotic screening techniques, such as thermo-stable kanamycin (Lipscomb et al. 2016), will also be needed to save time and effort. However, the current methods for deletion and insertion of multiple genes are sufficient.
There is one possible strategy to increase hydrogen production from real-world plant biomass. This strategy involves utilizing the genetic tools and the techniques developed for C. bescii. The first step is to remove the pathway for acetate production by deleting the phosphate acetyltransferase-encoding gene (pta, Cbes_1494). By deleting pta, the electrons carried by ferredoxins can be used by both types of hydrogenases such as [Fe–Fe] hydrogenases and [Ni–Fe] hydrogenases, resulting in more hydrogen production and ATP synthesis. Another potential strategy involves the manipulating of various glycosyl hydrolases by encoding genes by overexpressing the corresponding genes and manipulating regulatory genes to increase the decomposing efficiency of unpretreated plant biomass.
Conclusions
In this review, we highlighted that C. bescii can produce hydrogen directly from plant biomass without conventional pretreatment processes. Additionally, novel efficient methods for genetic modification of C. bescii have been developed through the deletion of cbeI, which is a thermostable type II restriction endonuclease. Overall, previous studies have demonstrated that C. bescii can be metabolically engineered to enhance hydrogen production. These would help C. bescii to efficiently produce hydrogen from biomass and biowaste including lignocellulosic biomass, cattle manure, and wastewater sludge (Table 2).
References
Akhlaghi N, Najafpour-Darzi G (2020) A comprehensive review on biological hydrogen production. Int J Hydrogen Energy 45(43):22492–22512. https://doi.org/10.1016/j.ijhydene.2020.06.182
Alfano M, Cavazza C (2020) Structure, function, and biosynthesis of nickel-dependent enzymes. Protein Sci 29(5):1071–1089. https://doi.org/10.1002/pro.3836
Alicia Benitez CW, Andreas de Palmenaer, Michael Lengersdorf, Tim Röding, Thomas Grube, Martin Robinius, Detlef Stolten, Wilhelm Kuckshinrichs (2021) Ecological assessment of fuel cell electric vehicles with special focus on type IV carbon fiber hydrogen tank. J Clean Prod 278 https://doi.org/10.1016/j.jclepro.2020.123277
Beckers L, Julien Masset C, Hamilton F Delvigne, Toye D, Crine M, Thonart P, Hiligsmann S (2015) Investigation of the links between mass transfer conditions, dissolved hydrogen concentration and biohydrogen production by the pure strain Clostridium butyricum CWBI1009. Biochem Eng J 98:18–28. https://doi.org/10.1016/j.bej.2015.01.008
Boeke JD, LaCroute F, Fink GR (1984) A positive selection for mutants lacking orotidine-5′-phosphate decarboxylase activity in yeast: 5-fluoro-orotic acid resistance. Mol Gen Genet 197(2):345–6. https://doi.org/10.1007/BF00330984
Buckel W (2021) Energy conservation in fermentations of anaerobic bacteria. Front Microbiol 12:703525. https://doi.org/10.3389/fmicb.2021.703525
Burkhart BW, Febvre HP, Santangelo TJ (2019) Distinct physiological roles of the three ferredoxins encoded in the hyperthermophilic archaeon Thermococcus kodakarensis. mBio 10(2) https://doi.org/10.1128/mBio.02807-18
Cha M, Chung D, Elkins JG, Guss AM, Westpheling J (2013a) Metabolic engineering of Caldicellulosiruptor bescii yields increased hydrogen production from lignocellulosic biomass. Biotechnol Biofuels 6(1):85. https://doi.org/10.1186/1754-6834-6-85
Cha M, Wang H, Chung D, Bennetzen JL, Westpheling J (2013b) Isolation and bioinformatic analysis of a novel transposable element, ISCbe4, from the hyperthermophilic bacterium, Caldicellulosiruptorbescii. J Ind Microbiol Biotechnol 40(12):1443–1448. https://doi.org/10.1007/s10295-013-1345-8
Cha M, Chung D, Westpheling J (2016) Deletion of a gene cluster for [Ni-Fe] hydrogenase maturation in the anaerobic hyperthermophilic bacterium Caldicellulosiruptor bescii identifies its role in hydrogen metabolism. Appl Microbiol Biotechnol 100(4):1823–1831. https://doi.org/10.1007/s00253-015-7025-z
Cha M, Kim JH, Choi HJ, Nho SB, Kim SY, Cha YL, Song H, Lee WH, Kim SK, Kim SJ (2023) Hydrogen production from barley straw and Miscanthus by the hyperthermophilic bacterium, Cadicellulosirupterbescii. J Microbiol Biotechnol 33(10):1–6. https://doi.org/10.4014/jmb.2305.05022
Chandel NS (2021) Glycolysis. Cold Spring Harbor Perspect Biol 13(5):a040535
Chung DH, Huddleston JR, Farkas J, Westpheling J (2011) Identification and characterization of CbeI, a novel thermostable restriction enzyme from Caldicellulosiruptor bescii DSM 6725 and a member of a new subfamily of HaeIII-like enzymes. J Ind Microbiol Biotechnol 38(11):1867–1877. https://doi.org/10.1007/s10295-011-0976-x
Chung D, Cha M, Farkas J, Westpheling J (2013a) Construction of a stable replicating shuttle vector for Caldicellulosiruptor species: use for extending genetic methodologies to other members of this genus. PLoS One 8(5):e62881. https://doi.org/10.1371/journal.pone.0062881
Chung D, Farkas J, Westpheling J (2013b) Overcoming restriction as a barrier to DNA transformation in Caldicellulosiruptor species results in efficient marker replacement. Biotechnol Biofuels 6(1):82. https://doi.org/10.1186/1754-6834-6-82
Chung D, Cha M, Guss AM, Westpheling J (2014) Direct conversion of plant biomass to ethanol by engineered Caldicellulosiruptor bescii. Proc Natl Acad Sci USA 111(24):8931–8936. https://doi.org/10.1073/pnas.1402210111
Clausen A, Mikkelsen MJ, Schroder I, Ahring BK (2004) Cloning, sequencing, and sequence analysis of two novel plasmids from the thermophilic anaerobic bacterium Anaerocellum thermophilum. Plasmid 52(2):131–138. https://doi.org/10.1016/j.plasmid.2004.06.001
Cofré O, Ramirez M, Gómez JM, Cantero D (2016) Pilot scale fed-batch fermentation in a closed loop mixed reactor for the biotransformation of crude glycerol into ethanol and hydrogen by Escherichia coli MG1655. Biomass Bioenergy 91:37–47. https://doi.org/10.1016/j.biombioe.2016.04.015
Crosby JR, Laemthong T, Lewis AM, Straub CT, Adams MWW, Kelly RM (2019) Extreme thermophiles as emerging metabolic engineering platforms. Curr Opin Biotechnol 59:55–64. https://doi.org/10.1016/j.copbio.2019.02.006
Dam P, Kataeva I, Yang SJ, Zhou F, Yin Y, Chou W, Poole FL 2nd, Westpheling J, Hettich R, Giannone R, Lewis DL, Kelly R, Gilbert HJ, Henrissat B, Xu Y, Adams MW (2011) Insights into plant biomass conversion from the genome of the anaerobic thermophilic bacterium Caldicellulosiruptor bescii DSM 6725. Nucleic Acids Res 39(8):3240–3254. https://doi.org/10.1093/nar/gkq1281
David B, Levin RI, Cicek N, Sparling R (2006) Hydrogen production by Clostridium thermocellum 27405 from cellulosic biomass substrates. Int J Hydrogen Energy 31:1496–1503. https://doi.org/10.1016/j.ijhydene.2006.06.015
de Vrije T, Mars AE, Budde MA, Lai MH, Dijkema C, de Waard P, Claassen PA (2007) Glycolytic pathway and hydrogen yield studies of the extreme thermophile Caldicellulosiruptor saccharolyticus. Appl Microbiol Biotechnol 74(6):1358–1367. https://doi.org/10.1007/s00253-006-0783-x
de Vrije T, Bude MAW, Lips SJ, Bakker RR, Mars AE, Claassen PAM (2010) Hydrogen production from carrot pulp by the extreme thermophiles Caldicellulosiruptor saccharolyticus and Thermotoga neapolitana. Int J Hydrogen Energy 35(24):13206–13213. https://doi.org/10.1016/j.ijhydene.2010.09.014
Esercizio N, Lanzilli M, Vastano M, Landi S, Xu Z, Gallo C, Nuzzo G, Manzo E, Fontana A, d’Ippolito G (2021) Fermentation of biodegradable organic waste by the family Thermotogaceae. Resources 10(4):34. https://doi.org/10.3390/resources10040034
Farkas J, Chung D, Cha M, Copeland J, Grayeski P, Westpheling J (2013) Improved growth media and culture techniques for genetic analysis and assessment of biomass utilization by Caldicellulosiruptor bescii. J Ind Microbiol Biotechnol 40(1):41–49. https://doi.org/10.1007/s10295-012-1202-1
Fawzy S, Osman AI, Doran J, Rooney DW (2020) Strategies for mitigation of climate change: a review. Environ Chem Lett 18:2069–2094. https://doi.org/10.1007/s10311-020-01059-w
Ghimire A, Frunzo L, Pontoni L, d’Antonio G, Lens PN, Esposito G, Pirozzi F (2015) Dark fermentation of complex waste biomass for biohydrogen production by pretreated thermophilic anaerobic digestate. J Environ Manage 152:43–48. https://doi.org/10.1016/j.jenvman.2014.12.049
Han D, Xu Z (2017) Development of a pyrE-based selective system for Thermotoga sp. strain RQ7. Extremophiles 21(2):297–306. https://doi.org/10.1007/s00792-016-0902-2
Han D, Xu H, Puranik R, Xu Z (2014) Natural transformation of Thermotoga sp. strain RQ7. BMC Biotechnol 14:39. https://doi.org/10.1186/1472-6750-14-39
Hassan Q, Sameen AZ, Salman HM, Jaszczur M, Al-Jiboory AK (2023) Hydrogen energy future: advancements in storage technologies and implications for sustainability. J Energy Storage 72:108404. https://doi.org/10.1016/j.est.2023.108404
Hoang AT, Sirohi R, Pandey A, Nižetić S, Lam SS, Chen W-H, Luque R, Thomas S, Arıcı M, Pham VV (2023) Biofuel production from microalgae: challenges and chances. Phytochem Rev 22:1089–1126. https://doi.org/10.1007/s11101-022-09819-y
Jay ZJ, Hunt KA, Chou KJ, Schut GJ, Maness PC, Adams MWW, Carlson RP (2020) Integrated thermodynamic analysis of electron bifurcating [FeFe]-hydrogenase to inform anaerobic metabolism and H(2) production. Biochim Biophys Acta Bioenerg 1:148087. https://doi.org/10.1016/j.bbabio.2019.148087
Jayasinghearachchi HS, Sarma PM, Singh S, Aginihotri A, Mandal AK, Lal B (2009) Fermentative hydrogen production by two novel strains of Enterobacter aerogenes HGN-2 and HT 34 isolated from sea buried crude oil pipelines. Int J Hydrogen Energy 34(17):7197–7207. https://doi.org/10.1016/j.ijhydene.2009.06.079
Jeswani HK, Chilvers A, Azapagic A (2020) Environmental sustainability of biofuels: a review. Proc Math Phys Eng Sci 476(2243):20200351. https://doi.org/10.1098/rspa.2020.0351
Kaila VRI, Wikstrom M (2021) Architecture of bacterial respiratory chains. Nat Rev Microbiol 19(5):319–330. https://doi.org/10.1038/s41579-020-00486-4
Kanai T, Imanaka H, Nakajima A, Uwamori K, Omori Y, Fukui T, Atomi H, Imanaka T (2005) Continuous hydrogen production by the hyperthermophilic archaeon, Thermococcus kodakaraensis KOD1. J Biotechnol 116(3):271–282. https://doi.org/10.1016/j.jbiotec.2004.11.002
Kim SK, Chung D, Himmel ME, Bomble YJ, Westpheling J (2019) Heterologous co-expression of two beta-glucanases and a cellobiose phosphorylase resulted in a significant increase in the cellulolytic activity of the Caldicellulosiruptor bescii exoproteome. J Ind Microbiol Biotechnol 46(5):687–695. https://doi.org/10.1007/s10295-019-02150-0
Kim J, Hwang S, Lee SM (2022) Metabolic engineering for the utilization of carbohydrate portions of lignocellulosic biomass. Metab Eng 71:2–12. https://doi.org/10.1016/j.ymben.2021.10.002
Lepage T, Kammoun M, Schmetz Q, Richel A (2021) Biomass-to-hydrogen: a review of main routes production, processes evaluation and techno-economical assessment. Biomass Bioenergy 144:105920. https://doi.org/10.1016/j.biombioe.2020.105920
Lin P-Y, Whang L-M, Wu Y-R, Ren W-J, Hsiao C-J, Li S-L, Chang J-S (2007) Biological hydrogen production of the genus Clostridium: metabolic study and mathematical model simulation. Int J Hydrogen Energy 32(12):1728–1735. https://doi.org/10.1016/j.ijhydene.2006.12.009
Lin C-Y, Chang C-C, Hung C-H (2008) Fermentative hydrogen production from starch using natural mixed cultures. Int J Hydrogen Energy 33:2445–2453. https://doi.org/10.1016/j.ijhydene.2008.02.069
Lipscomb GL, Conway JM, Blumer-Schuette SE, Kelly RM, Adams MWW (2016) A highly thermostable kanamycin resistance marker expands the tool kit for genetic manipulation of Caldicellulosiruptor bescii. Appl Environ Microbiol 82(14):4421–4428. https://doi.org/10.1128/AEM.00570-16
Liu X, Zhu Y, Yang ST (2006) Construction and characterization of ack deleted mutant of Clostridium tyrobutyricum for enhanced butyric acid and hydrogen production. Biotechnol Prog 22(5):1265–1275. https://doi.org/10.1021/bp060082g
Lu Y, Koo J (2019) O(2) sensitivity and H(2) production activity of hydrogenases-a review. Biotechnol Bioeng 116(11):3124–3135. https://doi.org/10.1002/bit.27136
Lucas S, Toffin L, Zivanovic Y, Charlier D, Moussard H, Forterre P, Prieur D, Erauso G (2002) Construction of a shuttle vector for, and spheroplast transformation of, the hyperthermophilic archaeon Pyrococcus abyssi. Appl Environ Microbiol 68(11):5528–5536. https://doi.org/10.1128/AEM.68.11.5528-5536.2002
Manisalidis I, Stavropoulou E, Stavropoulos A, Bezirtzoglou E (2020) Environmental and health impacts of air pollution: a review. Front Public Health 8:14. https://doi.org/10.3389/fpubh.2020.00014
Martínez-Jaramillo JE, Arango-Aramburo S, Giraldo-Ramírez DP (2019) The effects of biofuels on food security: a system dynamics approach for the Colombian case. Sustain Energy Technol Assess 34:97–109. https://doi.org/10.1016/j.seta.2019.05.009
Martins F, Felgueiras C, Smitkova M, Caetano N (2019) Analysis of fossil fuel energy consumption and environmental impacts in European countries. Energies 12(6):964. https://doi.org/10.3390/en12060964
Miller MA, Petrasch J, Randhir K, Rahmatian N, Klausner J (2021) Thermal, mechanical, and hybrid chemical energy storage systems/chapter 5 - chemical energy storage. Acad Press. https://doi.org/10.1016/C2019-0-00430-X
Minnan L, Jinli H, Xiaobin W, Huijuan X, Jinzao C, Chuannan L, Fengzhang Z, Liangshu X (2005) Isolation and characterization of a high H2-producing strain Klebsiella oxytoca HP1 from a hot spring. Res Microbiol 156(1):76–81. https://doi.org/10.1016/j.resmic.2004.08.004
Nanqi R, Wanqian G, Bingfeng L, Guangli C, Jie D (2011) Biological hydrogen production by dark fermentation: challenges and prospects towards scaled-up production. Curr Opin Biotechnol 22(3):365–70. https://doi.org/10.1016/j.copbio.2011.04.022
Niu K, Zhang X, Tan W-S, Zhu M-L (2010) Characteristics of fermentative hydrogen production with Klebsiella pneumoniae ECU-15 isolated from anaerobic sewage sludge. Int J Hydrogen Energy 35(1):71–80. https://doi.org/10.1016/j.ijhydene.2009.10.071
Oh S-E, Zuo Y, Zhang H, Guiltinan MJ, Logan BE, Regan JM (2009) Hydrogen production by Clostridium acetobutylicum ATCC824 and megaplasmid-deficient mutant M5 evaluated usinga large headspace volume technique. Int J Hydrogen Energy 34:9347–9353. https://doi.org/10.1016/j.ijhydene.2009.09.084
Okolie JA, Patra BR, Mukherjee A, Nanda S, Dalai AK, Kozinski JA (2021) Futuristic applications of hydrogen in energy, biorefining, aerospace, pharmaceuticals and metallurgy. Int J Hydrogen Energy 46:8885–8905. https://doi.org/10.1016/j.ijhydene.2021.01.014
Olguin-Maciel E, Singh A, Chable-Villacis R, Tapia-Tussell R, Ruiz HA (2020) Consolidated bioprocessing, an innovative strategy towards sustainability for biofuels production from crop residues: an overview. Agronomy 10:1834. https://doi.org/10.3390/agronomy10111834
Palone O, Gagliardi GG, Mechelli M, Cedola L, Borello D (2023) Techno-economic analysis of sustainable methanol and ammonia production by chemical looping hydrogen generation from waste plastic. Energy Convers Manag 292:117389. https://doi.org/10.1016/j.enconman.2023.117389
Periyasamy S, Isabel B, Kavitha S, Karthik D, Mohamed BA, Gizaw DG, Sivashanmugam P, Aminabhavi TM (2023) Recent advances in consolidated bioprocessing for conversion of lignocellulosic biomass into bioethanol – a review. Chem Eng J 453:139783. https://doi.org/10.1016/j.cej.2022.139783
Schroder C, Selig M, Schonheit P (1994) Glucose fermentation to acetate, CO2 and H 2 in the anaerobic hyperthermophilic eubacterium Thermotoga maritima: involvement of the Embden-Meyerhof pathway. Arch Microbiol 161:460–470. https://doi.org/10.1007/BF00307766
Schut GJ, Adams MW (2009) The iron-hydrogenase of Thermotoga maritima utilizes ferredoxin and NADH synergistically: a new perspective on anaerobic hydrogen production. J Bacteriol 191(13):4451–4457. https://doi.org/10.1128/JB.01582-08
Scott IM, Rubinstein GM, Poole FL 2nd, Lipscomb GL, Schut GJ, Williams-Rhaesa AM, Stevenson DM, Amador-Noguez D, Kelly RM, Adams MWW (2019) The thermophilic biomass-degrading bacterium Caldicellulosiruptor bescii utilizes two enzymes to oxidize glyceraldehyde 3-phosphate during glycolysis. J Biol Chem 294(25):9995–10005. https://doi.org/10.1074/jbc.RA111.007120
Seppälä JJ, Puhakka JA, Yli-Harja O, Karp MT, Santala V (2011) Fermentative hydrogen production by Clostridium butyricum and Escherichia coli in pure and cocultures. Int J Hydrogen Energy 36(17):10701–10708
Servé WM, Kengen AJMS (1994) Growth and energy conservation in batch cultures of Pyrococcus furiosus. FEMS Microbiol Lett 117:305–309. https://doi.org/10.1111/j.1574-6968.1994.tb06784.x
Shahbeik H, Peng W, Panahi HKS, Dehhaghi M, Guillemin GJ, Fallahi A, Amiri H, Rehan M, Raikwar D, Latine H, Pandalone B, Khoshnevisan B, Sonne C, Vaccaro L, Nizami A-S, Gupta VK, Lam SS, Pan J, Luque R, Sels B, Tabatabaei M, Aghbashlo M (2022) Synthesis of liquid biofuels from biomass by hydrothermal gasification: a critical review. Renew Sust Energ Rev 167:112833. https://doi.org/10.1016/j.rser.2022.112833
Singh R, Tevatia R, White D, Demirel Y, Blum P (2019) Comparative kinetic modeling of growth and molecular hydrogen overproduction by engineered strains of Thermotoga maritima. Int J Hydrogen Energy 44:7125–7136. https://doi.org/10.1016/j.ijhydene.2019.01.124
Soboh B, Linder D, Hedderich R (2004) A multisubunit membrane-bound [NiFe] hydrogenase and an NADH-dependent Fe-only hydrogenase in the fermenting bacterium Thermoanaerobacter tengcongensis. Microbiology (reading) 150(Pt 7):2451–2463. https://doi.org/10.1099/mic.0.27159-0
Song Y, Liu M, Xie L, You C, Sun J, Zhang YPJ (2019) A recombinant 12-His tagged Pyrococcus furiosus soluble [NiFe]-hydrogenase I overexpressed in Thermococcus kodakarensis KOD1 facilitates hydrogen-powered in vitro NADH regeneration. Biotechnol J 14(4):e1800301. https://doi.org/10.1002/biot.201800301
Straub CT, Bing RG, Otten JK, Keller LM, Zeldes BM, Adams MWW, Kelly RM (2020) Metabolically engineered Caldicellulosiruptor bescii as a platform for producing acetone and hydrogen from lignocellulose. Biotechnol Bioeng 117(12):3799–3808. https://doi.org/10.1002/bit.27529
Tanisho S (1998) Hydrogen production by facultative anaerobe Enterobacter aerogenes, BioHydrogen. Springer, Boston, MA https://doi.org/10.1007/978-0-585-35132-2_35
Vancanneyt M, De Vos P, Maras M, De Ley J (1990) Ethanol production in batch and continuous culture from some carbohydrates with Clostridium thermosaccharolyticum LMG 656. Syst Appl Microbiol 13:382–387. https://doi.org/10.1016/S0723-2020(11)80237-7
Wagner M, van Wolferen M, Wagner A, Lassak K, Meyer BH, Reimann J, Albers SV (2012) Versatile genetic tool box for the crenarchaeote Sulfolobus acidocaldarius. Front Microbiol 3:214. https://doi.org/10.3389/fmicb.2012.00214
White D (2012) The physiology and biochemistry of prokaryotes. Oxford University Press
Willquist K, Zeidan AA, van Niel EW (2010) Physiological characteristics of the extreme thermophile Caldicellulosiruptor saccharolyticus: an efficient hydrogen cell factory. Microb Cell Fact 9:89. https://doi.org/10.1186/1475-2859-9-89
Yan Liu PY, Song X, Yinbo Qu (2008) Hydrogen production from cellulose by co-culture of Clostridium thermocellum JN4 and Thermoanaerobacterium thermosaccharolyticum GD17. Int J Hydrogen Energy 33(12):2927–2933. https://doi.org/10.1016/j.ijhydene.2008.04.004
Yilmazel YD, Duran M (2021) Biohydrogen production from cattle manure and its mixtures with renewable feedstock by hyperthermophilic Caldicellulosiruptor bescii. J Clea Prod 292:125969. https://doi.org/10.1016/j.jclepro.2021.125969
Yoshida A, Nishimura T, Kawaguchi H, Inui M, Yukawa H (2006) Enhanced hydrogen production from glucose using ldh- and frd-inactivated Escherichia coli strains. Appl Microbiol Biotechnol 73(1):67–72. https://doi.org/10.1007/s00253-006-0456-9
Zafar A, Aftab MN, Asif A, Karadag A, Peng L, Celebioglu HU, Afzal MS, Hamid A, Iqbal I (2021) Efficient biomass saccharification using a novel cellobiohydrolase from Clostridium clariflavum for utilization in biofuel industry. RSC Adv 11(16):9246–9261. https://doi.org/10.1039/d1ra00545f
Zhang K, Zhao W, Rodionov DA, Rubinstein GM, Nguyen DN, Tanwee TNN, Crosby J, Bing RG, Kelly RM, Adams MWW, Zhang Y (2021) Genome-scale metabolic model of Caldicellulosiruptor bescii reveals optimal metabolic engineering strategies for bio-based chemical production. mSystems 6(3):e0135120. https://doi.org/10.1128/mSystems.01351-20
Zheng T, Huang Q, Zhang C, Ni J, She Q, Shen Y (2012) Development of a simvastatin selection marker for a hyperthermophilic acidophile, Sulfolobusislandicus. Appl Environ Microbiol 78(2):568–574. https://doi.org/10.1128/AEM.06095-11
Funding
This study was supported by the Korea Institute of Planning and Evaluation for Technology in Food, Agriculture and Forestry (IPET) and Korea Smart Farm R&D Foundation (KosFarm) through the Smart Farm Innovation Technology Development Program, funded by the Ministry of Agriculture, Food and Rural Affairs (MAFRA), MSIT, and Rural Development Administration (RDA) (No. 421045–03). Also, this research was carried out with the support of the Cooperative Research Program for Agriculture Science and Technology Development (PJ01708203) through the Rural Development Administration and the “Regional Innovation Strategy (RIS)” through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (MOE) (2021RIS-002). This research was also supported by the Chung-Ang University Graduate Research Scholarship (Academic Scholarship for the College of Biotechnology and Natural Resources) in 2023.
Author information
Authors and Affiliations
Contributions
M Cha and JK Kim surveyed the literature and wrote the manuscript. SJ Kim and M Cha designed the structure of the manuscript. WH Lee, H song, TG Lee, and SK Kim reviewed and revised the mini-review. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cha, M., Kim, J.K., Lee, WH. et al. Metabolic engineering of Caldicellulosiruptor bescii for hydrogen production. Appl Microbiol Biotechnol 108, 65 (2024). https://doi.org/10.1007/s00253-023-12974-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00253-023-12974-7