Abstract
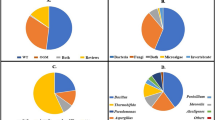
Nicotine-degrading microorganisms (NDMs) are a special microbial group which can use nicotine as the sole carbon and nitrogen source for growth. Since the 1950s, the bioconversion of nicotine by microbes has received increasing attention, and several NDMs have been identified, such as Arthrobacter nicotinovorans, Microsporum gypseum, Pellicularia filamentosa JTS-208, and Pseudomonas sp. 41. In recent years, increasing numbers of NDMs have been isolated and identified from tobacco plantation soil, leaf, and tobacco waste. Meanwhile, the metabolic pathway and degradation mechanism of nicotine have been elucidated in several NDMs, such as A. nicotinovorans, Agrobacterium tumefaciens S33, Aspergillus oryzae, and Pseudomonas putida S16. Moreover, several NDMs have been used in improving the quality of cigarettes, treating tobacco waste, and producing valuable intermediates of nicotine. Here, we summarize the diversity, phylogenetic analysis, and potential applications of NDMs.

Similar content being viewed by others
References
Akcay G, Yurdakoc K (2008) Removal of nicotine and its pharmaceutical derivatives from aqueous solution by raw bentonite and dodecylammoniumbentonite. J Sci Ind Res 67:451–454
Brandsch R (2006) Microbiology and biochemistry of nicotine degradation. Appl Microbiol Biotechnol 69:493–498
Brandsch R, Decker K (1984) Isolation and partial characterization of plasmid DNA from Arthrobacter oxidans. Arch Microbiol 138:15–17
Brandsch R, Faller W, Schneider K (1986) Plasmid pAO1 of Arthrobacter oxidans encodes 6-hydroxy-d-nicotine oxidase: cloning and expression of the gene in Escherichia coli. Mol Gen Genet 202:96–101
Buerge IJ, Kahle M, Buser HR, Müller MD, Poiger T (2008) Nicotine derivatives in wastewater and surface waters: application as chemical markers for domestic wastewater. Environ Sci Technol 42:6354–6360
Chen CM, Li XM, Yang JK, Gong XW, Li X, Zhang KQ (2008) Isolation of nicotine-degrading bacterium Pseudomonas sp. Nic22, and its potential application in tobacco processing. Int Biodeterior Biodegrad 62:226–231
Chen C, Ma GH, Lei LP, Zhou W, Shen XJ, Yang JK (2012) Isolation, identification and characteristics of nicotine-degrading bacterium strain 5-28. Tob Sci Technol 298:74–78
Chiribau CB, Mihasan M, Ganas P, Igloi GL, Artenie V, Brandsch R (2006) Final steps in the catabolism of nicotine. FEBS J 273:1528–1536
Civilini M, Domenis C, Sebastianutto N, de Bertoldi M (1997) Nicotine decontamination of tobacco agro-industrial waste and its degradation by microorganisms. Waste Manag Res 15:349–335
Cobzaru C, Ganas P, Mihasan M, Schleberger P, Brandsch R (2011) Homologous gene clusters of nicotine catabolism, including a new ω-amidase for α-ketoglutaramate, in species of three genera of Gram-positive bacteria. Res Microbiol 162:285–291
de Franco MA, da Silva WL, Bagnara M, Lansarin MA, dos Santos JH (2014) Photocatalytic degradation of nicotine in an aqueous solution using unconventional supported catalysts and commercial ZnO/TiO2 under ultraviolet radiation. Sci Total Environ 494–495:97–103
Decker K, Gries FA, Brühmüller M (1961) Über den Abbau des Nicotins durch Bakterienenzyme: III. Stoffwechselstudien an zellfreien Extrakten. Hoppe Seylers Z Physiol Chem 323:249–263
Eberwein H, Gries FA, Decker K (1961) Über den Abbau des Nicotins durch Bakterienenzyme: II. Isolierung und Charakterisierung eines nicotin-abbauenden Bodenbakteriums. Hoppe Seylers Z Physiol Chem 323:236–248
Ganas P, Sachelaru P, Mihasan M, Igloi GL, Brandsch R (2008) Two closely related pathways of nicotine catabolism in Arthrobacter nicotinovorans and Nocardioides sp. strain JS614. Arch Microbiol 189:511–517
Gong XW, Yang JK, Duan YQ, Dong Y, Zhe W, Wang L, Li QH, Zhang KQ (2009) Isolation and characterization of Rhodococcus sp. Y22 and its potential application to tobacco processing. Res Microbiol 160:200–204
González Alonso S, Valcárcel Y, Montero JC, Catalá M (2012) Nicotine occurrence in bottled mineral water: analysis of 10 brands of water in Spain. Sci Total Environ 416:527–531
Gravely LE, Geiss VL, Gregory CF (1985) Process for reduction of nitrate and nicotine content of tobacco by microbial treatment. US Patent 4557280
Grether-Beck S, Igloi GL, Pust S, Schilz E, Decker K, Brandsch R (1994) Structural analysis and molybdenum-dependent expression of the pAO1-encoded nicotine dehydrogenase genes of Arthrobacter nicotinovorans. Mol Microbiol 13:929–936
Gurusamy R, Natarajan S (2013) Current status on biochemistry and molecular biology of microbial degradation of nicotine. Sci World J 125385
Hochstein LI, Rittenberg CS (1959) The bacterial oxidation of nicotine: I. Nicotine oxidation by cell-free preparations. J Biol Chem 234:151–155
Huang J, Yang J, Duan Y, Gu W, Gong X, Zhe W, Su C, Zhang KQ (2010) Bacterial diversities on unaged and aging flue-cured tobacco leaves estimated by 16S rRNA sequence analysis. Appl Microbiol Biotechnol 88:553–562
Hylin JW (1959) The microbial degradation of nicotine: II. The mode of action of Achromobacter nicotinophagum. Arch Biochem Biophys 83:528–537
Igoli GL, Brandsch R (2003) Sequence of the 165-kilobase catabolic plasmid pAO1 from Arthrobacter nicotinovorans and identification of a pAO1-dependent nicotine uptake system. J Bacteriol 185:1976–1986
Jiang HJ, Ma Y, Qiu GJ, Wu FL, Chen SL (2011) Biodegradation of nicotine by a novel strain Shinella sp. HZN1 isolated from activated sludge. J Environ Sci Health B 46:703–708
Knezevich A, Muzic J, Hatsukami DK, Hecht SS, Stepanov I (2013) Nornicotine nitrosation in saliva and its relation to endogenous synthesis of N’-nitrosonornicotine in humans. Nicotine Tob Res 15:591–595
Lazarevic N, Adnadjevic B, Jovanovic J (2011) Adsorption of nicotine from aqueous solution onto hydrophobic zeolite type USY. Appl Surf Sci 257:8017–8023
Lei LP, Xia ZY, Guo RJ, Wu YP, Cui GM, Liao DZ (2008) Reduction of nicotine in tobacco leaf treated with Arthrobacter spp. Tob Sci Technol 248:56–58
Lei LP, Zhang W, Wei HL, Xia ZY, Liu XZ (2009) Characterization of a novel nicotine-degrading Ensifer sp. strain N7 isolated from tobacco rhizosphere. Ann Microbiol 59:247–252
Li HJ, Li XM, Duan YQ, Zhang KQ, Yang JK (2010) Biotransformation of nicotine by microorganism: the case of Pseudomonas spp. Appl Microbiol Biotechnol 86:11–17
Li HJ, Duan YQ, Ma GH, Lei LP, Zhang KQ, Yang JK (2011) Isolation and characterization of Acinetobacter sp. ND12 capable of degrading nicotine. Afr Microbiol Res 5:1335–1341
Li HL, Xie KB, Huang HY, Wang SN (2014) 6-Hydroxy-3-succinoylpyridine hydroxylase catalyzes a central step of nicotine degradation in Agrobacterium tumefaciens S33. PLoS One 9:e103324
Ma GH, Lei LP, Xia ZY, Gong XW, Zhou W, Yang JK (2012) Diversity and phylogenetic analyses of nicotine degrading bacteria isolated from tobacco plantation soils. Afr Microbiol Res 6:6392–6398
Madigan M, Martinko J ed (2005) Brock biology of microorganisms, 11th edn. Prentice Hall. ISBN 0-13-144329-1
Meher KK, Panchwagh AM, Rangrass S, Gollakota KG (1995) Biomethanation of tobacco waste. Environ Pollut 90:199–202
Meng XJ, Lu LL, Gu GF, Xiao M (2010) A novel pathway for nicotine degradation by Aspergillus oryzae 112822 isolated from tobacco leaves. Res Microbiol 161:626–633
Mihasan M, Brandsch R (2013) pAO1 of Arthrobacter nicotinovorans and the spread of catabolic traits by horizontal gene transfer in Gram-positive soil bacteria. J Mol Evol 77:22–30
Nakano H, Wieser M, Hurh B, Kawai T, Yoshida T, Yamane T, Nagasawa T (1999) Purification, characterization and gene cloning of 6-hydroxynicotinate 3-monooxygenase from Pseudomonas fluorescens TN5. Eur J Biochem 260:120–126
Newton RP, Geiss VL, Jewell JN, Gravely LE (1977) Process for reduction of nicotine content of tobacco by microbial treatment. US Patent 4037609
Novotny TE, Zhao F (1999) Consumption and production waste: another externality of tobacco use. Tob Control 8:75–80
Passananti M, Temussi F, Iesce MR, Previtera L, Mailhot G, Vione D, Brigante M (2014) Photoenhanced transformation of nicotine in aquatic environments: involvement of naturally occurring radical sources. Water Res 55:106–114
Payne RB, May HD, Sowers KR (2011) Enhanced reductive dechlorination of polychlorinated biphenyl impacted sediment by bioaugmentation with a dehalorespiring bacterium. Environ Sci Technol 45:8772–8779
Piotrowska-Cyplik A, Olejnik A, Cyplik P, Dach J, Czarnecki Z (2009) The kinetics of nicotine degradation, enzyme activities and genotoxic potential in the characterization of tobacco waste composting. Bioresour Technol 100:5037–5044
Qiu J, Ma Y, Chen L, Wu L, Wen Y, Liu W (2011) A sirA-like gene, sirA2, is essential for 3-succinoyl-pyridine metabolism in the newly isolated nicotine-degrading Pseudomonas sp. HZN6 strain. Appl Microbiol Biotechnol 92:1023–1032
Raman G, Mohan KN, Manohar V, Sakthivel N (2014) Biodegradation of nicotine by a novel nicotine-degrading bacterium, Pseudomonas plecoglossicida TND35 and its new biotransformation intermediates. Biodegradation 25:95–107
Rathbone DA, Bruce NC (2002) Microbial transformation of alkaloids. Curr Opin Microbiol 5:274–281
Ruan AD, Min H, Peng X, Huang Z (2005) Isolation and characterization of Pseudomonas sp. strain HF-1, capable of degrading nicotine. Res Microbiol 156:700–706
Schenk S, Hoelz A, Krass B, Decker K (1998) Gene structures and properties of enzymes of the plasmid-encoded nicotine catabolism of Arthrobacter nicotinovorans. J Mol Biol 284:1323–1329
Schievelbein H (1982) Nicotine, resorption and fate. Pharmacol Ther 18:233–248
Sguros PL (1955) Microbial transformations of the tobacco alkaloids: I. Cultural and morphological characteristics of a nicotinophile. J Bacteriol 69:28–37
Sindelar RD, Rosasza JP, Barfknecht CF (1979) N-demethylation of nicotine and reduction of nicotine-1′-N-oxide by Microsporum gypseum. Appl Environ Microbiol 38:836–839
Spande TF, Garraffo HM, Edwards MW, Yeh HJC, Pannell L, Daly JW (1992) Epibatidine: a novel (chloropyridyl)-azabicycloheptane with potent analgesic activity from an Ecuadoran poison frog. J Am Chem Soc 114:3475–3478
Su C, Gu W, Zhe W, Zhang KQ, Duan YQ, Yang JK (2011) Diversity and phylogeny of bacteria on Zimbabwe tobacco leaves estimated by 16S rRNA sequence analysis. Appl Microbiol Biotechnol 92:1033–1044
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Tang H, Wang S, Ma L, Meng X, Deng Z, Zhang D, Ma C, Xu P (2008) A novel gene, encoding 6-hydroxy-3-succinoylpyridine hydroxylase, involved in nicotine degradation by Pseudomonas putida strain S16. Appl Environ Microbiol 74:1567–1574
Tang H, Wang L, Meng X, Ma L, Wang S, He X, Wu G, Xu P (2009) Novel nicotine oxidoreductase-encoding gene involved in nicotine degradation by Pseudomonas putida strain S16. Appl Environ Microbiol 75:772–778
Tang HZ, Yao YX, Zhang DK, Meng XZ, Wang LJ, Yu H, Ma LY (2011) A novel NADH-dependent and FAD-containing hydroxylase is crucial for nicotine degradation by Pseudomonas putida. J Biol Chem 286:39179–39187
Tang H, Yao Y, Wang L, Yu H, Ren Y, Wu G, Xu P (2012) Genomic analysis of Pseudomonas putida: genes in a genome island are crucial for nicotine degradation. Sci Rep 2:377
Tang H, Wang L, Wang W, Yu H, Zhang K, Yao Y, Xu P (2013) Systematic unraveling of the unsolved pathway of nicotine degradation in Pseudomonas. PLoS Genet 9:e1003923
Uchida S, Maeda S, Kisaki T (1983) Conversion of nicotine into nornicotine and N-methylmysomine by fungi. Agric Biol Chem 47:1949–1953
Valcárcel Y, González Alonso S, Rodríguez-Gil JL, Gil A, Catalá M (2011) Detection of pharmaceutically active compounds in the rivers and tap water of the Madrid Region (Spain) and potential ecotoxicological risk. Chemosphere 84:1336–1348
Wada E, Yamasaki K (1953) Degradation of nicotine by soil bacteria. Science 117:152–153
Wang SN, Xu P, Tang HZ, Meng J, Liu XL, Huang J, Chen H, Du Y, Blankespoor HD (2004) Biodegradation and detoxification of nicotine in tobacco solid waste by a Pseudomonas sp. Biotechnol Lett 26:1493–1496
Wang SN, Xu P, Tang HZ, Meng J, Liu XL, Ma CQ (2005) ‘Green’ route to 6-hydroxy-3-succinoyl-pyridine from (S)-nicotine of tobacco waste by whole cells of a Pseudomonas sp. Environ Sci Technol 39:6877–6880
Wang SN, Liu Z, Tang HZ, Meng J, Xu P (2007) Characterization of environmentally friendly nicotine degradation by Pseudomonas putida biotype a strain S16. Microbiology 153:1556–1565
Wang M, Yang G, Min H, Lv ZM, Jia X (2009a) Bioaugmentation with the nicotine-degrading bacterium Pseudomonas sp. HF-1 in a sequencing batch reactor treating tobacco wastewater: degradation study and analysis of its mechanisms. Water Res 43:4187–4196
Wang SN, Liu Z, Xu P (2009b) Biodegradation of nicotine by a newly isolated Agrobacterium sp. strain S33. J Appl Microbiol 107:838–847
Wang MZ, Yang GQ, Wang X, Yao YL, Min H, Lu ZM (2011) Nicotine degradation by two novel bacterial isolates of Acinetobacter sp. TW and Sphingomonas sp. TY and their responses in the presence of neonicotinoid insecticides. World J Microbiol Biotechnol 27:1633–1640
Wang MZ, Zheng X, He HZ, Shen DS, Feng HJ (2012a) Ecological roles and release patterns of acylated homoserine lactones in Pseudomonas sp. HF-1 and their implications in bacterial bioaugmentation. Bioresour Technol 125:119–126
Wang SN, Huang HY, Xie KB, Xu P (2012b) Identification of nicotine biotransformation intermediates by Agrobacterium tumefaciens strain S33 suggests a novel nicotine degradation pathway. Appl Microbiol Biotechnol 95:1567–1578
Wang HJ, He HZ, Wang MZ, Wang S, Zhang J, Wei W, Xu HX, Lv ZM, Shen DS (2013a) Bioaugmentation of activated sludge with Acinetobacter sp. TW enhances nicotine degradation in a synthetic tobacco wastewater treatment system. Bioresour Technol 142:445–453
Wang X, Tang L, Yao YL, Wang HX, Min H, Lv ZM (2013b) Bioremediation of the tobacco waste-contaminated soil by Pseudomonas sp. HF-1: nicotine degradation and microbial community analysis. Appl Microbiol Biotechnol 97:6077–6088
Wang L, Tang H, Yu H, Yao Y, Xu P (2014a) An unusual repressor controls the expression of a crucial nicotine-degrading gene cluster in Pseudomonas putida S16. Mol Microbiol 91:1252–1269
Wang MZ, Zheng X, Zhang K, Ding YC, He HZ, Shen DS, Feng HJ (2014b) A new method for rapid construction of a Pseudomonas sp. HF-1 bioaugmented system: accelerating acylated homoserine lactones secretion by pH regulation. Bioresour Technol 169:229–235
Wei X, Deng X, Cai D, Ji Z, Wang C, Yu J, Li J, Chen S (2014) Decreased tobacco-specific nitrosamines by microbial treatment with Bacillus amyloliquefaciens DA9 during the air-curing process of burley tobacco. J Agric Food Chem 62:12701–12706
Xia ZY, Lei LP, Wu YP, Guo RJ (2006) Isolation and identification of degrading nicotine bacteria — Arthrobacter nicotianae strain K9. Chin Tob Sci 2:1–4
Yao YX, Tang HZ, Ren HX, Yu H, Wang LJ, Xu P (2012) Genome sequence of a nicotine-degrading strain of Arthrobacter. J Bacteriol 194:5714–5715
Yu H, Tang H, Wang L, Yao Y, Wu G, Xu P (2011) Complete genome sequence of the nicotine-degrading Pseudomonas putida strain S16. J Bacteriol 193:5541–542
Yu H, Hausinger RP, Tang HZ, Xu P (2014a) Mechanism of the 6-hydroxy-3-succinoyl-pridine 3-monooxygenase flavoprotein from Pseudomonas putida S16. J Biol Chem 289:29158–29170
Yu H, Li Y, Tang H, Xu P (2014b) Genome sequence of a newly isolated nicotine-degrading bacterium, Ochrobactrum sp. SJY1. Genome Announc 2(4)
Yu H, Tang HZ, Xu P (2014c) Green strategy from waste to value-added-chemical production: efficient biosynthesis of 6-hydroxy-3-succinoyl-pyridine by an engineered biocatalyst. Sci Rep 4:5397
Yu H, Tang H, Zhu X, Li Y, Xu P (2015) Molecular mechanism of nicotine degradation by a newly isolated strain Ochrobactrum sp. SJY1. Appl Environ Microbiol 81:272–281
Yuan YJ, Lu ZX, Wu N, Huang LJ, Lü FX, Bie XM (2005) Isolation and preliminary characterization of a novel nicotine-degrading bacterium, Ochrobactrum intermedium DN2. Int Biodeterior Biodegrad 56:45–50
Yuan YJ, Lu ZX, Huang LJ, Li Y, Lu FX, Bie XM, Teng YQ, Lin Q (2007) Biodegradation of nicotine from tobacco waste extract by Ochrobactrum intermedium DN2. J Ind Microbiol Biotechnol 34:567–570
Zhang K, Zheng X, Shen DS, Wang MZ, Feng HJ, He HZ, Wang S, Wang JH (2014) Evidence for existence of quorum sensing in a bioaugmented system by acylated homoserine lactone-dependent quorum quenching. Environ Sci Pollut Res Int. doi:10.1007/s11356-014-3795-6
Zhao L, Zhu CJ, Gao Y, Wang C, Li XZ, Shu M, Shi YP, Zhong WH (2012) Nicotine degradation enhancement by Pseudomonas stutzeri ZCJ during aging process of tobacco leaves. World J Microbiol Biotechnol 28:2077–2086
Zhong WH, Zhu CJ, Shu M, Sun KD, Zhao L, Wang C, Ye ZJ, Chen JM (2010) Degradation of nicotine in tobacco waste extract by newly isolated Pseudomonas sp. ZUTSKD. Bioresour Technol 101:6935–6941
Acknowledgments
We are grateful to Prof. Jianping Xu of the Dept. of Biology, McMaster University for valuable comments and critical discussions. This work was supported by the Science and Technology Program of Chongqing Tobacco Company (approval no. NY20120802070020), the National Natural Science Foundation of China (31060012), and the Program for Excellent Young Talents of Yunnan University (To Jinkui Yang). We also thank the anonymous reviewers for their useful suggestions.
Author information
Authors and Affiliations
Corresponding author
Additional information
Jinkui Yang holds a Ph.D., Yunnan University.
Rights and permissions
About this article
Cite this article
Liu, J., Ma, G., Chen, T. et al. Nicotine-degrading microorganisms and their potential applications. Appl Microbiol Biotechnol 99, 3775–3785 (2015). https://doi.org/10.1007/s00253-015-6525-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-015-6525-1