Abstract
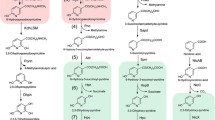
Several bacterial species are adapted to nicotine, the main alkaloid produced by the tobacco plant, as growth substrate. A general outline of nicotine catabolism by these bacteria is presented, followed by an emphasis on new insights based on molecular biology and biochemical work obtained with the catabolic plasmid pAO1 of Arthrobacter nicotinovorans. Its 165-kb sequence revealed the genetic structure of nicotine catabolism and allowed the assignment of new enzyme activities to specific gene products, which extends the known biochemical steps of this pathway. Potential implications of the progress in our understanding of bacterial breakdown of nicotine for biotechnological applications are discussed.


Similar content being viewed by others
References
Andreesen JR, Fetzner S (2002) The molybdenum-containing hydroxylases of nicotinate, isonicotinate, and nicotine. Met Ions Biol Syst 39:405–430
Armstrong DW, Wang X, Ercal N (1998) Enantiomeric composition of nicotine in smokeless tobacco, medicinal products, and commercial reagents. Chiriality 10:587–591
Armstrong DW, Wang X, Lee J-T, Liu Y-S (1999) Enantiomeric composition of nornicotine, anatabine, and anabasine in tobacco. Chiriality 11:82–84
Baitsch D, Sandu C, Brandsch R, Igloi G (2001) Gene cluster on pAO1 of Arthrobacter nicotinovorans involved in degradation of the plant alkaloid nicotine: cloning, purification, and characterization of 2,6-dihydroxypyridine 3-hydroxylase. J Bacteriol 183:5262–5267
Bergeron F, Otto A, Blache P, Day R, Denoroy L, Brandsch R, Bataile D (1998) Molecular cloning and tissue distribution of rat sarcosine dehydrogenase. Eur J Biochem 257:556–561
Bonin I, Martins BM, Purvanov V, Fetzner S, Huber R, Dobbek H (2004) Active site geometry and substrate recognition of the molybdenum hydroxylase quinoline 2-oxidoreductase. Structure 12:1425–1435
Bucherer H (1942) Über den mikrobiellen Abbau von Giftstoffen. I. Mitteilung: Über den mikrobiellen Abbau von Nikotin. Zent Bl Bakteriol 105:166–173
Carl B, Arnold A, Hauer B, Fetzner S (2004) Sequence and transcriptional analysis of a gene cluster of Pseudomonas putida 86 involved in quinoline degradation. Gene 331:177–188
Chiribau C-B, Sandu C, Fraaije M, Schiltz E, Brandsch R (2004) A novel γ-N-methylaminobutyrate demethylating oxidase involved in catabolism of the tobacco alkaloid nicotine by Arthrobacter nicotinovorans pAO1. Eur J Biochem 271:4677–4684
Chiribau C-B, Sandu C, Igloi GL, Brandsch R (2005) Characterization of PmfR, the transcriptional activator of the pAO1-borne purU-mabO-folD operon of Arthrobacter nicotinovorans. J Bacteriol 187:3062–3070
Chlumsky LJ, Zhang L, Jorns MS (1995) Sequence analysis of sarcosine oxidase and nearby genes reveals homologies with key enzymes of folate one-carbon metabolism. J Biol Chem 270:18252–18259
Decker K, Bleeg H (1965) Induction and purification of stereospecific nicotine oxidizing enzymes from Arthrobacter oxidans. Biochem Biophys Acta 105:313–334
Decker K, Eberwein H, Gries FA, Brühmüller M (1961) Über den Abbau des Nicotins durch Bakterienenzyme. IV. l-6-Hydroxy-nicotine als erstes Zwischenprodukt. Biochem Z 334:227–244
Dunn G, Montgomery MG, Mohammed F, Coker A, Cooper JB, Robertson T, Garcia JL, Bugg TD, Wood SP (2005) The structure of the C–C bond hydrolase MhpC provides insights into its catalytic mechanism. J Mol Biol 346:253–265
Eberhardt H-J (1995) The biological degradation of nicotine by nicotinophilic microorganisms. Beitr Tab Forsch Int 16:119–129
Eppink MHM, Schreuder AH, van Berkel WJH (1997) Identification of a novel conserved sequence motif in flavoprotein hydroxylases with a putative dual function in FAD/NAD(P)H binding. Protein Sci 6:2454–2458
Fraaije MW, van Berkel WJH, Benen JAE, Visser J, Mattevi A (1998) A novel oxidoreductase family sharing a conserved FAD-binding domain. Trends Biochem Sci 23:206–207
Freudenberg W, König K, Andreesen J R (1988) Nicotine dehydrogenase from Arthrobacter oxidans: a molybdenum-containing hydroxylase. FEMS Microbiol Lett 52:13–18
Fuhrmann S, Ferner M, Jeffke T, Henne A, Gottschalk G, Meyer O (2003) Complete nucleotide sequence of the circular megaplasmid pHCG3 of Oligotropha carboxidovorans: function in the chemolithoautotrophic utilization of CO, H2 and CO2. Gene 322:67–75
Gauthier JJ, Rittenberg SC (1971) The metabolism of nicotinic acid. I. Purification and properties of 2,5-dihydroxypyridine oxygenase from Pseudomonas putida N-9. J Biol Chem 246:3737–3742
Gherna RL, Richardson SH, Rittenberg SC (1965) The bacterial oxidation of nicotine. VI. The metabolism of 2,6-dihydroxypseudooxynicotine. J Biol Chem 240:3669–3674
Gloger M, Decker K (1969) Zum Mechanismus der Induktion nicotinabbauender Enzyme in Arthrobacter oxydans. Z Naturforsch 24b:1016–1025
Grether-Beck S, Igloi GL, Pust S, Schiltz E, Decker K, Brandsch R (1994) Structural analysis and molybdenum-dependent expression of the pAO1-encoded nicotine dehydrogenase genes of Arthrobacter nicotinovorans. Mol Microbiol 13:929–936
Gries FA, Decker K, Brühmüller M (1961a) Über den Abbau des Nicotins durch Bakterienenzyme. V. Der Abbau des l-6-Hydroxy-nicotins zu [γ-Methylamino-propyl]-[6-hydroxy-pyridyl-(3)]-ketons. Hoppe–Seyler’s Z Physiol Chem 325:229–241
Gries FA, Decker K, Eberwein H, Brühmüller M (1961a) Über den Abbau des Nicotins durch Bakteienenzyme. VI. Die enzymatische Umwandlung des (γ-Methylamino-propyl)-[6-hydroxy-pyridyl-(3)]-ketons. Biochem Z 335:285–302
Hille R (1996) The mononuclear molybdenum enzymes. Chem Rev 96:2757–2816
Hochstein LI, Rittenberg CS (1959) The bacterial oxidation of nicotine. II. The isolation of the first product and its identification as (l)-6-hydroxynicotine. J Biol Chem 234:156–162
Hochstein LI, Rittenberg SC (1960) The bacterial oxidation of nicotine. III. The isolation and identification of 6-hydroxypseudooxynicotine. J Biol Chem 235:795–799
Holmes PE, Rittenberg SC (1972) The bacterial oxidation of nicotine. VII. Partial purification and properties of 2,6-dihydroxypyridine oxidase. J Biol Chem 247:7622–7627
Holmquist M (2000) Alpha/beta-hydrolase fold enzymes: structures, functions and mechanisms. Curr Protein Pept Sci 1:209–235
Hylin JW (1959) The microbial degradation of nicotine. II. The mode of action of Achromobacter nicotinophagum. Arch Biochem Biophys 83:528–537
Igloi GL, Brandsch R (2003) Sequence of the 165-kilobase catabolic plasmid pAO1 from Arthrobacter nicotinovorans and identification of a pAO1-dependent nicotine uptake system. J Bacteriol 185:1976–1989
Kaiser J-P, Feng Y, Bollag J-M (1996) Microbial metabolism of pyridine, quinoline, acridine, and their derivatives under aerobic and anaerobic conditions. Microbiol Rev 60:483–498
Koetter JWA, Schulz G (2005) Crystal structure of 6-hydroxy-d-nicotine oxidase from Arthrobacter nicotinovorans. J Mol Biol 352(2):418–428
Kurihara T, Mihara H, Kato S, Yoshimura T, Esaki N (2003) Assembly of iron–sulfur clusters mediated by cysteine desulfurases, IscS, CsdB and CSD, from Escherichia coli. Biochem Biophys Acta 1647:303–309
Lang H, Polster M, Brandsch R (1991) Dimethylglycine dehydrogenase from rat liver: characterization of a cDNA clone and covalent labeling of the polypeptide with 14C-FAD. Eur J Biochem 198:793–799
Leimkühler S, Klipp W (1999) Role of XDHC in Molybdenum cofactor insertion into xanthine dehydrogenase of Rhodobacter capsulatus. J Bacteriol 181:2745–2751
Malphettes L, Weber CC, El-Baba MD, Schoenmakers RG, Aubel D, Weber W, Fussenegger M (2005) A novel mammalian expression system derived from components coordinating nicotine degradation in Arthrobacter nicotinovorans pAO1. Nucleic Acids Res 33:e107
Menéndez C, Igloi GL, Nick P, Brandsch RJ, Schubach B, Böttcher B, Brandsch RK (1997) Molybdate-uptake genes and molybdopterin-biosynthesis genes on a bacterial plasmid. Eur J Biochem 250:524–531
Meskys R, Harris RJ, Casaite V, Basran J, Scrutton NS (2001) Organization of the genes involved in dimethylglycine and sarcosine degradation in Arthrobacter spp.: implications for glycine betaine catabolism. Eur J Biochem 268:3390–3398
Richardson SH, Rittenberg SC (1961) The bacterial oxidation of nicotine. V. Identification of 2,6-dihydroxypseudooxynicotine as the third oxidation product. J Biol Chem 236:964–967
Sachelaru P, Schiltz E, Igloi GL, Brandsch R (2005) An α/β fold C-C bond hydrolase is involved in a central step of nicotine catabolism by Arthrobacter nicotinovorans. J Bacteriol (in press)
Sandu C, Chiribau C-B, Brandsch R (2003) Characterization of HdnoR, the transcriptional repressor of the 6-hydroxy-d-nicotine oxidase gene of Arthrobacter nicotinovorans pAO1, and ist DNA-binding activity in response to l- and d-nicotine derivatives. J Biol Chem 278:51307–51315
Sandu C, Chiribau C-B, Sachelaru P, Brandsch R (2005) Plasmids for nicotine-dependent and independent gene expression in Arthrobacter nicotinovorans and other Arthrobacter species. Appl Environ Microbiol (in press)
Schenk S, Decker K (1999) Horizontal gene transfer involved in the convergent evolution of the plasmid-encoded enantioselective 6-hydroxynicotine oxidases. J Mol Evol 48:178–186
Schenk S, Hoelz A, Krauß B, Decker K (1998) Gene structure and properties of enzymes of the plasmid-encoded nicotine catabolism of Arthrobacter nicotinovorans. J Mol Biol 284:1322–1339
Schmid A, Dordick JS, Hauer B, Kieners A, Wubbolts M, Witholt B (2001) Industrial biocatalysis today and tomorrow. Nature 409:258–268
Sindelar RD, Rosasza JP, Barfknecht CF (1979) N-demethylation of nicotine and reduction of nicotine-1′-N-oxide by Microsporum gypseum. Appl Environ Microbiol 38:836–839
Uchida S, Maeda S, Kisaki T (1983) Conversion of nicotine into nornicotine and N-methylmyosmine by fungi. Agric Biol Chem 47:1949–1953
Wada E (1957) Microbial degradation of the tobacco alkaloids, and some related compounds. Arch Biochem Biophys 72:145–162
Wada E, Yamasaki K (1953) Mechanism of microbial degradation of nicotine. Science 117:152–153
Zheng L, Cash VL, Flint DH, Dean DR (1998) Assembly of iron–sulfur clusters. Identification of an iscSUA-hscBA-fdx gene cluster from Azotobacter vinelandii. J Biol Chem 273:13264–13272
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Brandsch, R. Microbiology and biochemistry of nicotine degradation. Appl Microbiol Biotechnol 69, 493–498 (2006). https://doi.org/10.1007/s00253-005-0226-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-005-0226-0