Abstract
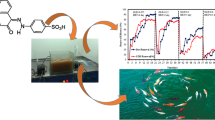
Wastewater effluents from the textile and other dye-stuff industries contain significant amounts of synthetic dyes that require treatment to prevent groundwater contamination. In research aimed at biotechnology for treatment of azo dyes, this study examined 288 strains of azo-dye degrading bacteria to identify efficient strains and determine incubation times required for decolorization. Initial enrichment cultures were carried out using a mixture of four structurally different dyes (Acid Red 88, Reactive Black 5, Direct Red 81, and Disperse Orange 3) as the sole source of C and N to isolate the bacteria from soil, activated sludge, and natural asphalt. Six strains were selected for further study based on their prolific growth and ability to rapidly decolorize the dyes individually or in mixtures. Treatment times required by the most efficient strain, AS96 (Shewanella putrefaciens) were as short as 4 h for complete decolorization of 100 mg l−1 of AR-88 and DR-81 dyes under static conditions, and 6 and 8 h, respectively, for complete decolorization of RB-5 and DO-3. To our knowledge, these bacterial strains are the most efficient azo-dye degrading bacteria that have been described and may have practical application for biological treatment of dye-polluted wastewater streams.





Similar content being viewed by others
References
Alinsafi A, Evenou F, Abdulkarim EM, Pons MN, Zahraa O, Benhammou A, Yaacoubi A, Nejmeddine A (2007) Treatment of textile industry wastewater by supported photocatalysis. Dyes Pigm 74:439–445
Arslan-Alaton I (2007) Degradation of a commercial textile biocide with advanced oxidation processes and ozone. J Env Manage 82:145–154
Bae JS, Freeman HS (2007) Aquatic toxicity evaluation of new direct dyes to the Daphnia magna. Dyes Pigm 73:81–85
Barragan BE, Costa C, Marquez MC (2007) Biodegradation of azo dyes by bacteria inoculated on solid media. Dyes Pigm 75:73–81
Boer CG, Obici L, Souza CG, Peralta RM (2004) Decolourization of synthetic dyes by solid state cultures of Lentinula (Lentinus) edodes producing manganese peroxidase as the main lignolytic enzyme. Bioresour Technol 94:107–112
Chang JS, Lin YC (2001) Decolorization kinetics of recombinant E. coli strain harboring azo dye decolorization determinants for Rhodococcus sp. Biotech Lett 23:631–636
Chen BY (2006) Biologically-feasible screening strategy for optimal decolorization strain to diazo Evercion Red H-E7B. J Chinese Inst Chem Eng 37:117–124
Chung KT, Cerniglia CE (1992) Mutagenicity of azo dyes: structure–activity relationships. Mut Res 277:201–220
Dubrow SF, Boardman GD, Michelsen DL (1996) Chemical pretreatment and aerobic–anaerobic degradation of textile dye wastewater. In: Reife A and Freeman HS (eds) Environmental chemistry of dyes and pigments. Wiley Interscience. pp 75–102
Golab V, Vinder A, Simonic M (2005) Efficiency of the coagulation/flocculation method for the treatment of dye bath effluent. Dyes Pigm 67:93–97
Hao JJ, Song FQ, Huang F, Yang CL, Zhang ZJ, Zheng Y, Tian XJ (2007) Production of laccase by a newly isolated deuteromycete fungus Pestalotiopsis sp. and its decolorization of azo dye. J Industrial Microbiol Biotechnol 34:233–240
Ince H, Tezcanli G (1999) Treatability of textile dye-bath effluents by advanced oxidation: preparation for reuse. Water Sci Technol 40:183–190
Junnarkar N, Murty DS, Bhatt NS, Madamwar D (2006) Decolorization of diazo dye Direct Red 81 by a novel bacterial consortium. World J Microbiol Biotech 22:163–168
Kalme SD, Parshetti GK, Jadhav SU, Govindwar SP (2007) Biodegradation of benzidine based dye Direct Blue-6 by Pseudomonas desmolyticum NCIM 2112. Bioresor Tech 98:1405–1410
Khera MS, Saini HS, Sharma DK, Chadha BS, Chimni SS (2005) Comparative studies on potential of consortium and constituent pure bacterial isolates to decolorize azo dyes. Water Res 39:5135–5141
Kim JS, Crowley DE (2007) Microbial diversity in natural asphalts of the Rancho La Brea tar pits. Appl Env Microbiol 73:4579–4591
Lucas MS, Amaral C, Sampaio A, Peres JA, Dias AA (2006) Biodegradation of the diazo dye Reactive Black 5 by a wild isolate of Candida oleophila. Enzyme Microbiol Tech 39:51–55
O’Neill C, Hawkes FR, Hawkes DL, Lourenco ND, Pinheiro HM, Delee W (1999) Color in textile effluents sources, measurement, discharge consents and simulation: a review. J Chem Technol Biotechnol 74:1009–1018
Ozturk A, Abdullah MI (2006) Toxicological effect of indole and its azo dye derivatives on some microorganisms under aerobic conditions. Sci Total Env 358:137–142
Pandey A, Singh P, Iyengar L (2007) Bacterial decolorization and degradation of azo dyes. Int Biodeter Biodegr 59:73–84
Rajaguru P, Vidya L, Baskarasethupathi B, Kumar PA, Palanivel M, Kalaiselvi K (2002) Genotoxicity evaluation of polluted ground water in human peripheral blood lymphocytes using the comet assay. Mut Res 517:29–37
Saxe JP, Lubenow BL, Chiu PC, Huang CP, Cha DK (2006) Enhanced biodegradation of azo dyes using an integrated elemental iron-activated sludge system: II. Effects of physical–chemical parameters. Water Env Res 78:26–30
Slokar YM, Le Marechal AM (1998) Methods of decoloration of textile wastewater. Dyes Pigm 37:335–356
Sponza DT, Isik M (2002) Decolorization and azo dye degradation by anaerobic/aerobic sequential process. Enzyme Microb Technol. 31:102–110
Stackebrandt E, Goebel BM (1994) A place for DNA-DNA reassociation and 16 S ribosomal-RNA sequence analysis in the present species definition in bacteriology. Int J Syst Bacteriol 44:846–849
Stolz A (2001) Basic and applied aspects in the microbial degradation of azo dyes. Appl Microbiol Biotechnol 56:69–80
Tan NCG, Borger A, Slender P, Svitelskaya AV, Lettinga G, Field JA (2000) Degradation of azo dye Mordant Yellow 10 in a sequential anaerobic and bioaugmented aerobic bioreactor. Water Sci Technol 42:337–344
Umbuzeiro GA, Freeman H, Warren SH, Oliveira DP, Terao Y, Watanabe T, Claxton LD (2005) The contribution of azo dyes to the mutagenic activity of the Cristais River. Chemosphere 60:55–64
Vijaykumar MH, Vaishampayan PA, Shouche YS, Karegoudar TB (2007) Decolourization of naphthalene-containing sulfonated azo dyes by Kerstersia sp strain VKY1. Enzyme Microbial Technol 40:204–211
Wang A, Qu J, Liu H, Ge J (2004) Degradation of azo dye Acid Red 14 in aqueous solution by electrokinetic and electrooxidation process. Chemosphere 55:1189–1196
Xu M, Guo J, Cen Y, Zhong X, Cao W, Sun G (2005) Shewanella decolorationis sp. nov., a dye decolorizing bacterium isolated from activated sludge of a waste-water treatment plant. Int J Syst Evol Microbiol 55:363–368
Zhao X, Hardin IR (2007) HPLC and spectrophotometric analysis of biodegradation of azo dyes by Pleurotus ostreatus. Dyes Pigm 73:322–325
Acknowledgements
Financial support for this study was provided by the Higher Education Commission, Pakistan. We thank Dr. Jason Cantera for assistance in 16S rRNA gene analysis.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Khalid, A., Arshad, M. & Crowley, D.E. Accelerated decolorization of structurally different azo dyes by newly isolated bacterial strains. Appl Microbiol Biotechnol 78, 361–369 (2008). https://doi.org/10.1007/s00253-007-1302-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-007-1302-4