Abstract
Removal of synthetic textile dyes poses a challenge to the textile industry and a threat to the environment’s flora and fauna. These dyes are recalcitrant and not very amenable to physical and chemical techniques of degradation. Hence, several studies on alternative bioremediation methods involving plants, plant roots, single microbes, or a consortium of microbes for the decolorization of dyes have been carried out. In the present study, potent bacteria for dye decolorization were isolated from rhizospheric soil of mangrove plants collected from Kamothe, Navi Mumbai, India. Of the 20 isolates obtained after enrichment, seven isolates were used for further screening of efficient decolorization ability in minimal basal media containing 10% glucose, 2.5% trace metal solution, and 0.1% of Methyl Orange (MO) dye concentration. Physiological parameters to optimize the decolorization of dye at optimum pH, temperature, and incubation time were studied for all the seven isolates. UV–vis and Fourier transform infrared spectroscopy were used to investigate dye decolorization. The seven isolates were characterized morphologically, biochemically, and molecular identification of these bacterial isolates was performed by 16S rRNA sequence analysis. The isolates were identified as Bacillus paramycoides, Pseudomonas taiwanensis, Citrobacter murliniae, Acinetobacter pitti, Exiguobacterium acetylicum, Psychrobacter celer, and Aeromonas taiwanensis. Out of these, Aeromonas taiwanensis has shown exceptional capacity by ~ 100% decolorization of azo dye in minimum time.
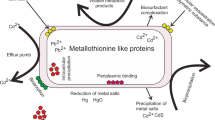
Graphical abstract









Similar content being viewed by others
Data availability
All the data generated during the study has been uploaded on the public platform NCBI under the accession numbers MT256266, MT256267, MT256269, MT256270, MT256271, MT256273, and MT256274.
Abbreviations
- MBM:
-
Minimal basal medium
- CFE:
-
Cell-free extract
- MO:
-
Methyl Orange
- MR:
-
Methyl Red
- CR:
-
Congo Red
References
Abbas SZ, Yee CJ, Hossain K et al (2019) Isolation and characterization of mercury-resistant bacteria from industrial wastewater. Desalin Water Treat 138:128–133. https://doi.org/10.5004/dwt.2019.23279
Akansha K, Chakraborty D, Sachan SG (2019) Decolorization and degradation of methyl orange by Bacillus stratosphericus SCA1007. Biocatal Agric Biotechnol 18:101044. https://doi.org/10.1016/j.bcab.2019.101044
Aruna B, Silviya LR, Kumar ES et al (2015) Decolorization of Acid Blue 25 dye by individual and mixed bacterial consortium isolated from textile effluents. Int J Curr Microbiol App Sci 4:1015–1024
Boontanom P, Chantarasiri A (2020) Short Communication : Diversity of culturable epiphytic bacteria isolated from seagrass ( Halodule uninervis ) in Thailand and their preliminary antibacterial activity. Biodiversitas J Biol Divers 21:2907–2913. https://doi.org/10.13057/biodiv/d210706
Carliell CM, Barclay SJ, Naidoo N, et al (1995) Microbial decolourisation of a reactive azo dye under anaerobic conditions. ISSN 0378–4738=Water SA 21:61–69
Çetin D, Dönmez G (2006) Decolorization of reactive dyes by mixed cultures isolated from textile effluent under anaerobic conditions Demet C. Enzyme Microb Technol 38:926–930. https://doi.org/10.1016/j.enzmictec.2005.08.020
Chantarasiri A (2015) Aquatic Bacillus cereus JD0404 isolated from the muddy sediments of mangrove swamps in Thailand and characterization of its cellulolytic activity. Egypt J Aquat Res 41:257–264. https://doi.org/10.1016/j.ejar.2015.08.003
Chantarasiri A (2020) Klebsiella and Enterobacter isolated from mangrove wetland soils in thailand and their application in biological decolorization of textile reactive dyes. Int J Environ Res Public Health 17:1–21. https://doi.org/10.3390/ijerph17207531
Chantarasiri A (2021) Shewanella baltica strain JD0705 isolated from the mangrove wetland soils in Thailand and characterization of its ligninolytic performance. Biodiversitas 22:354–361. https://doi.org/10.13057/biodiv/d220143
Chantarasiri A, Boontanom P (2017) Decolorization of synthetic dyes by ligninolytic lysinibacillus sphaericus JD1103 isolated from Thai wetland ecosystems. AACL Bioflux 10:814–819
Chaudhary P, Khati P, Chaudhary A et al (2021) Cultivable and metagenomic approach to study the combined impact of nanogypsum and Pseudomonas taiwanensis on maize plant health and its rhizospheric microbiome. PLoS One 16:e0250574. https://doi.org/10.1371/journal.pone.0250574
Chen KC, Wu JY, Liou DJ, Hwang SCJ (2003) Decolorization of the textile dyes by newly isolated bacterial strains. J Biotechnol 101:57–68. https://doi.org/10.1016/S0168-1656(02)00303-6
Chiong T, Lau SY, Lek ZH et al (2016) Enzymatic treatment of methyl orange dye in synthetic wastewater by plant-based peroxidase enzymes. J Environ Chem Eng 4:2500–2509. https://doi.org/10.1016/j.jece.2016.04.030
Daneshvar N, Ayazloo M, Khataee AR, Pourhassan M (2007) Biological decolorization of dye solution containing Malachite Green by microalgae Cosmarium sp. Bioresour Technol. 98:1176–1182. https://doi.org/10.1016/j.biortech.2006.05.025
Fang Z, Li T, Wang Q et al (2011) A bacterial laccase from marine microbial metagenome exhibiting chloride tolerance and dye decolorization ability. Appl Microbiol Biotechnol 89:1103–1110. https://doi.org/10.1007/s00253-010-2934-3
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Int J Org Evol 39:783–791. https://doi.org/10.1111/j.1558-5646.1985.tb00420.x
Ferbiyanto A, Rusmana I, Raffiudin R (2015) Characterization and identification of cellulolytic bacteria from gut of worker macrotermes gilvus. HAYATI J Biosci 22:197–200. https://doi.org/10.1016/j.hjb.2015.07.001
Forgacs E, Cserháti T, Oros G (2004) Removal of synthetic dyes from wastewaters: a review. Environ Int 30:953–971. https://doi.org/10.1016/j.envint.2004.02.001
Fulekar MH, Wadgaonkar SL, Singh A (2013) Decolourization of dye compounds by selected bacterial strains isolated from dyestuff industrial area. Int J Adv Res Technol 2:182–192
Ghodake GS, Kalme SD, Jadhav JP, Govindwar SP (2009) Purification and partial characterization of lignin peroxidase from Acinetobacter calcoaceticus NCIM 2890 and its application in decolorization of textile dyes. Appl Biochem Biotechnol 152:6–14. https://doi.org/10.1007/s12010-008-8258-4
Guo G, Liu C, Hao J et al (2021) Development and characterization of a halo-thermophilic bacterial consortium for decolorization of azo dye. Chemosphere 272:129916. https://doi.org/10.1016/j.chemosphere.2021.129916
Harikumar PS, Joseph L, Dhanya A (2013) Photocatalytic degradation of textile dyes by hydrogel supported titanium dioxide nanoparticles. J Environ Eng Ecol Sci 2:2. https://doi.org/10.7243/2050-1323-2-2
Heer K, Sharma S (2017) Microbial pigments as a natural color: a review. Int J Pharm Sci Res 8:1913–1922
Hidayat A, Tachibana S (2014) Decolorization of azo dyes and mineralization of phenanthrene by Trametes sp. AS03 isolated from Indonesian mangrove forest. Indones J For Res 1:67–75. https://doi.org/10.20886/ijfr.2014.1.1.67-75
Hsueh C, Chen B (2007) Comparative study on reaction selectivity of azo dye decolorization by Pseudomonas luteola. J Hazard Mater 141:842–849. https://doi.org/10.1016/j.jhazmat.2006.07.056
Junnarkar N, Murty DS, Bhatt NS, Madamwar D (2006) Decolorization of diazo dye Direct Red 81 by a novel bacterial consortium. World J Microbiol Biotechnol 22:163–168. https://doi.org/10.1007/s11274-005-9014-3
Kachiprath B, Solomon S, Jayanath G, Philip R (2019) Mangrove microflora as potential source of hydrolytic enzymes for commercial applications. Indian J Geo Mar Sci 48:678–684
Kaewtubtim P, Meeinkuirt W, Seepom S, Pichtel J (2016) Heavy metal phytoremediation potential of plant species in a mangrove ecosystem in Pattani Bay, Thailand. Appl Ecol Environ Res 14:367–382
Kalme SD, Parshetti GK, Jadhav SU, Govindwar SP (2007) Biodegradation of benzidine based dye Direct Blue-6 by Pseudomonas desmolyticum NCIM 2112. Bioresour Technol 98:1405–1410. https://doi.org/10.1016/j.biortech.2006.05.023
Kolekar YM, Pawar SP, Gawai KR et al (2008) Bioresource technology decolorization and degradation of disperse Blue 79 and Acid Orange 10, by Bacillus fusiformis KMK5 isolated from the textile dye contaminated soil. Bioresour Technol 99:8999–9003. https://doi.org/10.1016/j.biortech.2008.04.073
Kumar S, Stecher G, Li M et al (2018) MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Lalnunhlimi S, Veenagayathri K (2016) Decolorization of azo dyes (Direct Blue 151 and Direct Red 31) by moderately alkaliphilic bacterial consortium. Brazilian J Microbiol 47:39–46. https://doi.org/10.1016/j.bjm.2015.11.013
Letunic I, Bork P, Gmbh BS (2021) Interactive Tree Of Life ( iTOL ) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res 49:293–296. https://doi.org/10.1093/nar/gkab301
Meehan C, Banat IM, Mcmullan G et al (2000) Decolorization of Remazol Black-B using a thermotolerant yeast, Kluyveromyces marxianus IMB3. Environ Int 26:75–79. https://doi.org/10.1016/S0160-4120(00)00084-2
Mitsch WJ, Gosselink JG (2015) Wetlands. John Wiley & Sons
Mohan SV, Rao NC, Prasad KK, Karthikeyan J (2002) Treatment of simulated Reactive Yellow 22 ( Azo ) dye effluents using Spirogyra species. Waste Manag 22:575–582. https://doi.org/10.1016/S0956-053X(02)00030-2
Mohandass R, Bhaskar A (2008) Decolorization and biodegradation of Indigo carmine by a textile soil isolate Paenibacillus larva. Biodegradation 19:283–291. https://doi.org/10.1007/s10532-007-9134-6
Morgulis A, Coulouris G, Raytselis Y, et al (2008) Database indexing for production MegaBLAST searches. Bioinformatics 24:1757–1764. https://doi.org/10.1093/bioinformatics/btn322
Ozdemir G, Pazarbasi ÆB, Kocyigit ÆA, Ersoy ÆE (2008) Decolorization of Acid Black 210 by Vibrio harveyi TEMS1 a newly isolated bioluminescent bacterium from Izmir Bay, Turkey. World J Microbiol Biotechnol 24:1375–1381. https://doi.org/10.1007/s11274-007-9619-9
Panswad T, Luangdilok W (2000) Decolorization of reactive dyes with different molecular structures under different environmental conditions. Water Res 34:4177–4184. https://doi.org/10.1016/S0043-1354(00)00200-1
Pearce CI, Lloyd JR, Guthrie JT (2003) The removal of colour from textile wastewater using whole bacterial cells: a review. Dye Pigment 58:179–196. https://doi.org/10.1016/S0143-7208(03)00064-0
Pearce CI, Christie R, Boothman C et al (2006) Reactive azo dye reduction by Shewanella strain J18 143. Biotechnol Bioeng 95:692–773. https://doi.org/10.1002/bit.21021
Pourbabaee AA, Malekzadeh F, Sarbolouki MN, Mohajeri A (2005) Decolorization of Methyl Orange (as a model azo dye) by the newly discovered Bacillus Sp. Mod Multidiscip Appl Microbiol Exploit Microbes Their Interact 494–499. https://doi.org/10.1002/9783527611904.ch87
Preethi S, Pathy R (2020) Exploring prospective forested wetland—Actinomycetes for biodeterioration of genotoxic textile azo dyes. IJEB 58:344–354
Qu W, Liu T, Wang D et al (2018) Metagenomics-based discovery of malachite green-degradation gene families and enzymes from mangrove sediment. Front Microbiol 9:1–11. https://doi.org/10.3389/fmicb.2018.02187
Ramya M, Iyappan SMA, JJS (2010) Biodegradation and decolorization of acid red by Acinetobacter radioresistens. J Bioremediation Biodegrad 01:1–6. https://doi.org/10.4172/2155-6199.1000105
Sahasrabudhe MM, Saratale RG, Saratale GD, Pathade GR (2014) Decolorization and detoxification of sulfonated toxic diazo dye C. I. Direct Red 81 by Enterococcus faecalis YZ 66. J Environ Heal Sci Eng 12:1–13. https://doi.org/10.1186/s40201-014-0151-1
Sandilyan S, Kathiresan K (2014) Decline of mangroves – a threat of heavy metal poisoning in Asia. Ocean Coast Manag 102:161–168. https://doi.org/10.1016/j.ocecoaman.2014.09.025
Saratale RG, Saratale GD, Chang JS, Govindwar SP (2011) Bacterial decolorization and degradation of azo dyes: a review. J Taiwan Inst Chem Eng 42:138–157
Saravanan R, Manoj D, Qin J et al (2018) Mechanothermal synthesis of Ag/TiO2 for photocatalytic methyl orange degradation and hydrogen production. Process Saf Environ Prot 120:339–347. https://doi.org/10.1016/j.psep.2018.09.015
Sarkar S, Banerjee A, Halder U et al (2017) Degradation of synthetic azo dyes of textile industry: a sustainable approach using microbial enzymes. Water Conserv Sci Eng 2:121–131. https://doi.org/10.1007/s41101-017-0031-5
Shah MP, Ka P, Ss N, Am D (2013a) Bioremoval of Azo dye Reactive Red by Bacillus spp . ETL-1982. J Bioremed Biodeg 4:3–7. https://doi.org/10.4172/2155-6199.1000186
Shah MP, Patel KA, Darji AM (2013b) Microbial degradation and decolorization of methyl orange dye by an application of Pseudomonas spp. ETL-1982. Int J Environ Bioremediation Biodegrad 1:26–36
Shaw CB, Carliell CM, Wheatley AD (2002) Anaerobic/aerobic treatment of coloured textile effluents using sequencing batch reactors. Water Res 36:1993–2001. https://doi.org/10.1016/S0043-1354(01)00392-X
Singh K, Arora S (2011) Removal of synthetic textile dyes from wastewaters: a critical review on present treatment technologies. Crit Rev Environ Sci Technol 41:807–878. https://doi.org/10.1080/10643380903218376
Singh M, Singh H, Kumar D et al (2005) Decolorization of various azo dyes by bacterial consortium. Dye Pigment 67:55–61. https://doi.org/10.1016/j.dyepig.2004.10.008
Somasundaram ST (2014) Biodegradation of carcinogenic textile azo dyes using bacterial isolates of mangrove sediment. J Coast Life Med 2:154–162
Sponza DT, Işik M (2002) Decolorization and azo dye degradation by anaerobic / aerobic sequential process. Enzyme Microb Technol 31:102–110. https://doi.org/10.1016/S0141-0229(02)00081-9
Sun C, Zeng X, Lai Q et al (2020) Mangrovibacterium lignilyticum sp. nov., a facultatively anaerobic lignin-degrading bacterium isolated from mangrove sediment. Int J Syst Evol Microbiol 70:4502–4507. https://doi.org/10.1099/ijsem.0.004305
Tamura K, Nei M (1993) Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol Biol Evol 10:512–526. https://doi.org/10.1093/oxfordjournals.molbev.a040023
Tamura K, Stecher G, Kumar S (2021) MEGA11: molecular evolutionary genetics analysis version 11. Mol Biol Evol 38:3022–3027. https://doi.org/10.1093/molbev/msab120
Telke AA, Kalyani DC, Dawkar VV, Govindwar SP (2009) Influence of organic and inorganic compounds on oxidoreductive decolorization of sulfonated azo dye CI Reactive Orange 16. J Hazard Mater 172:298–309. https://doi.org/10.1016/j.jhazmat.2009.07.008
Truu M, Juhanson J, Truu J (2009) Microbial biomass, activity and community composition in constructed wetlands. Sci Total Environ 407:3958–3971. https://doi.org/10.1016/j.scitotenv.2008.11.036
Vijaykumar MH, Vaishampayan PA, Shouche YS, Karegoudar TB (2007) Decolourization of naphthalene-containing sulfonated azo dyes by Kerstersia sp . strain VKY1. Enzyme Microb Technol 40:204–211. https://doi.org/10.1016/j.enzmictec.2006.04.001
Weisburger JH (2002) Comments on the history and importance of aromatic and heterocyclic amines in public health. Mutat Res - Fundam Mol Mech Mutagen 506–507:9–20. https://doi.org/10.1016/S0027-5107(02)00147-1
Yoon S, Ha S, Kwon S, et al (2017) Introducing EzBioCloud : a taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int J Syst Evol Microbiol 1613–1617. https://doi.org/10.1099/ijsem.0.001755
Zollinger H, Morgan TH, The M (2003) Color chemistry: syntheses, properties, and applications of organic dyes and pigments. John Wiley & Sons
Acknowledgements
The author is grateful to the D Y Patil Deemed To Be University’s School of Biotechnology and Bioinformatics in Navi Mumbai, India, for providing research facilities for this study.
Author information
Authors and Affiliations
Contributions
AM: Experiment perform, Compilation of data, Manuscript drafting, Graphical abstract preparation.
SS: Inception of the project, planning and execution of the experiment, Manuscript editing, and reviewing.
NP: Phylogenetic analysis, collection of soil sample.
JP: Inception of the project, planning of the project, collection of soil samples.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Responsible Editor: Diane Purchase
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Highlights
• Collection of soil samples from rhizospheric and non-rhizospheric soil region of the mangroves plant.
• Enrichment of soil sample in media containing azo dye for dye decolorization.
• Screening and isolation of potential dye decolourizers.
• Optimize the condition for dye decolorization.
• FTIR analysis and application-based studies.
• Identification of microbes using 16SrRNA.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Modi, A., Singh, S., Patki, J. et al. Screening and identification of azo dye decolorizers from mangrove rhizospheric soil. Environ Sci Pollut Res 29, 83496–83511 (2022). https://doi.org/10.1007/s11356-022-21610-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-022-21610-2