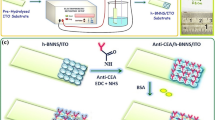
Abstract
To tackle cancer and provide prompt diagnoses and prognoses, the constantly evolving biosensing field is continuously on the lookout for novel markers that can be non-invasively analysed. Extracellular vesicles (EVs) may represent a promising biomarker that also works as a source of biomarkers. The augmented cellular activity of cancerous cells leads to the production of higher numbers of EVs, which can give direct information on the disease due to the presence of general and cancer-specific surface-tethered molecules. Moreover, the intravesicular space is enriched with other molecules that can considerably help in the early detection of neoplasia. Even though EV-targeted research has indubitably received broad attention lately, there still is a wide lack of practical and effective quantitative procedures due to difficulties in pre-analytical and analytical phases. This review aims at providing an exhaustive outline of the recent progress in EV detection using electrochemical and photoelectrochemical biosensors, with a focus on handling approaches and trends in the selection of bioreceptors and molecular targets related to EVs that might guide researchers that are approaching such an unstandardised field.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Liquid biopsy has provided, over the years, pivotal support in the clinical field due to the remarkable advantages that it covers, including the possibility of obtaining cell-specific biomarkers from biological fluids with simple and easily repeatable techniques [1]. Among the variety of molecules that flow in biological fluids, circulating tumour cells (CTCs), microRNAs (miRNAs), long non-coding RNAs (lncRNAs), and vesicles are just a few of the most interesting and commonly studied. Regarding the latter, initially under-appreciated and considered simple debris or ways to dispose of cellular components [2], extracellular vesicles (EVs) are membrane-enclosed cell-secreted nanometric particles now considered biomarkers and, at the same time, sources of biomarkers [3]. Many types of these nanometric particles can be found, but their classification appears not to be unique. One possible distinction is among (i) microvesicles (particles with dimensions around 100–1000 nm, produced due to imbalances in the lipid distribution on plasma membranes), (ii) ectosomes (ubiquitous vesicles generated at the plasma membrane), (iii) shedding vesicles, and (iv) apoptotic bodies (100–5000 nm, secreted by apoptotic cells) (Fig. 1) [4]. Nevertheless, other classifications may be found in the literature, for instance on the basis of their size: (a) large exosomes: 90–120 nm, (b) small exosomes: 60–80 nm, and (c) exomeres (non-membranous EVs): 35 nm [5]. Other sources categorise them by their excretion mechanism; in particular, (I) large EVs or microvesicles are particles that bud from cells’ plasma membrane, whereas (II) small EVs or exosomes are multivesicular endosomes that release intraluminal vesicles upon fusing with the plasma membrane [6]. However, all these categories have evolved over the years and are still confusing.
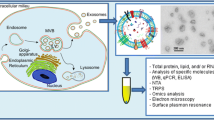
One possible classification of EVs on the basis of their size and biogenesis: apoptotic bodies originate from apoptotic cells, microvesicles bud off the plasma membrane of cells, and exosomes emerge from endosome-multivesicular body (MVB) complexes. Moreover, EVs are characterised by protein markers on their surface, as well as intravesicular nucleic acids, including double-stranded DNA (dsDNA), messenger RNAs (mRNAs), and microRNAs (miRNAs). Reprinted with permission from ref. [9]
Most research papers focus on exosomes. Nonetheless, there are some nomenclature issues regarding these vesicles, and the term “exosome” is often found in the literature to indicate any EV in general [7]. To this end, the MISEV2018 guidelines suggest researchers to use instead the more general term “extracellular vesicles” to avoid misunderstandings in scientific publications or use other parameters or terms to refer to the vesicles analysed, such as descriptors relative to their biochemical composition or their physical characteristics, including size (small, medium, large) or density (low, middle, high) [8]. Hence, to avoid possible confusion and misinterpretations, the generic term “extracellular vesicle” will be used in this manuscript, even when the scientific papers herein reported state otherwise.
In the last decade, multiple techniques have been used to detect and/or characterise EVs, including western blotting, mass spectrometry, flow cytometry, nanoparticles tracking analysis, dynamic light scattering, tunable resistive pulse sensing, transmission electron microscopy, atomic force microscopy, and enzyme-linked immunosorbent assays [10,11,12,13]. However, many of these approaches entail limitations, such as the high cost of the sophisticated instruments required, the incompatibility with in situ analyses, the lack of standardisation in the case of EVs, and the fact that large volumes of samples are often required [14]. An easier and cost-effective solution is lately increasingly being found in the field of biosensors. With their numerous virtues and their versatility toward point-of-care applications [15], many biosensing approaches have been challenged in EV analysis, spanning from (a) optical biosensors, based on colorimetry, fluorescence, chemiluminescence, surface plasmon resonance, surface-enhanced Raman spectroscopy, or photonic structures, to (b) electrochemical and photoelectrochemical biosensors that convert the biorecognition event between a receptor and its ligand into potential-, current-, or impedance-based signals by voltametric/amperometric readouts or electrochemical impedance spectroscopy (EIS). Indeed, (photo)electrochemical approaches are constantly evolving toward portability and miniaturisation [6], and lab-on-a-chip structures and wearable tools, that are compact and portable, might represent the greatest next generation of biosensing devices in the clinical field. Numerous review papers have been published in the last few years covering a summary of the recent strides taken in the field of biosensors using a variety of techniques [6, 16,17,18,19,20,21,22,23,24,25,26,27,28,29]. Herein, instead, the reader will find a focus on electrochemical and photoelectrochemical biosensing strategies, with the aim of giving a guide to the recent procedures found in the literature, as well as precautions and handling of EVs that may be of help to anyone that wants to approach the field, which is indeed constantly evolving and not yet standardised.
Handling extracellular vesicles
Storage conditions
The stability of extracellular vesicles is one of the main hurdles that hinder their analytical application. Moreover, the temperature at which they are stored has important outcomes. However, although new insights on EV preservation are emerging, currently there is a lack of standard criteria for better storage [30]. Storage is pivotal both for the EV-containing matrix, from which EVs are harvested, and for isolated EVs, affecting their stability and the number of vesicles retrievable [31]. Drastic size changes have been observed in EVs subjected to freeze–thaw cycles [32]. Freezing processes can induce the formation and expansion of ice micro- and nano-crystals in the lipid bilayer, thus disrupting the membrane. The value of the zeta potential (ζ) can thus be affected, resulting in its decrease and the formation of multilamellar vesicles and aggregation upon lowering the temperatures [33]. Vesicles stored at a wide range of storage temperatures have been compared, observing alterations in the vesicle’s diameter [17, 33, 34] and major protein and RNA losses at > 0 °C [35]. However, such results do not always arrive at the same conclusions but agree on the fact that, since EVs and their content are relatively unstable, it is recommended to store them below −70 °C. Indeed, more studies need to be conducted in this field, especially regarding short-time effects of sub-zero temperatures [30]. An additional parameter that should be carefully considered is pH. Unfortunately, very little information can be found on this matter in the literature. It has been shown that, upon isolation, a slightly basic pH resulted in a great loss of EV-associated protein expression within 30 min after centrifuging and that low pH values may help reduce such degradation [36]. Nonetheless, more exhaustive information must be acquired for this concern. Another important note is that, upon storage, part of the isolated EVs may adhere to the storage container, resulting in a loss of material [8].
Separation/concentration of EVs
In most literature-reported cases, EVs are usually isolated prior to their utilisation and study. They can be collected from EV-containing matrixes, such as biological fluids, in multiple ways, and many factors can deeply affect the recovery [37]. It is thus important to plan thoroughly this pre-analytical phase, keeping attention to storage and manipulation of the source material. Undoubtedly, a perfect isolation/purification of EVs is unrealistic and other non-vesicular structures may get co-isolated to various degrees [38]. Usually, the term “separation” is preferred to “isolation” and “purification”; other common expressions found in the literature and commercial kits are “enrichment” and “concentration”, which suggest an increase in EV counts relative to the total volume [8]. The decision in regard to what degree one should purify the EVs-containing sample depends on its experimental application. If there is no need to assess the biochemical composition or attribute a biomarker/function to a specific vesicle compared to others or to answer more complex clinical/analytical questions, then less pure EV-preparations can be considered, while focusing on higher recoveries [6, 8].
The methods used to separate EVs rely mainly on their properties, such as density, size, and surface markers. Many researchers use more than one separation/concentration step to obtain the final EV sample. According to the findings of C. Gardiner and colleagues [39], ultracentrifugation (UC) is the most commonly used first-step isolation methodology. Nonetheless, many other techniques can be found in the literature, such as density gradients [40], filtration [41], polymer-based precipitation [42, 43], immunoaffinity-based isolation [44], and size-exclusion chromatography [45], that, at the cost of some output impurities (e.g., co-precipitated proteins), may yield higher EV concentrations [46]. Figure 2 illustrates the various separation techniques often chosen by researchers, with a note on some of the kits and reagents available on the market. Indeed, each different approach presents its advantages and disadvantages. Since a standardisation among them has not yet been provided [47], case-by-case situations should be studied by a researcher that needs to adapt these procedures to the analytical question involved.
Electrochemical detection of extracellular vesicles
When dealing with the detection of EVs, one can be interested in two different information: the total amount of EVs present, useful as an index of the presence of a tumour, or the specific identification of the type of cancer from which EVs originate. Interestingly, the membrane of EVs presents tetraspanins, proteases, transmembrane receptors, and adhesion molecules that may differentiate the vesicles depending on the parent cell. In the case of cancer-originating extracellular vesicles, the antigens present on their surface are considerably enriched, thus representing a great parameter to consider in the (a) detection and (b) identification of EVs. Indeed, one can consider different types of biomarkers that can be broadly categorised into two distinct applications: (a) universal cancer biomarkers, such as proteins abundantly present on the membrane of all EVs (although greatly enriched in cancer-derived EVs), useful when the objective is to quantify the whole set of particles present and detect their upregulation during on-going neoplasia; and (b) cancer-specific antigens, hence molecules that originate from certain tumours and may help identify it or distinguish it from others [48]. Besides membrane-tethered markers, EVs also contain an important cargo that includes a variety of biomolecules, such as proteins and nucleic acids [49]. As will be better discussed in the “Intravesicular biomarkers” section, the analysis of such intra-EV biomarkers is earning extensive attention in biosensing. A number of online-available databases, such as Vesiclepedia, EVpedia, and Exocarta, offer the possibility to check regularly updated information regarding the composition of EVs, thus representing valuable tools for researchers on the lookout for new detectable molecules that may help detect EVs or distinguish among them.
EV detection using universal markers
Since it has been shown that their upregulation occurs in cancer cell lines upon their secretion into the extracellular space, EVs represent a valid diagnostic and prognostic biomarker [50]. The vast majority of research articles present in the literature carry out total EV detection by harnessing the presence of tetraspanins, such as CD9, CD63, CD81, and CD82, although CD81 and CD63 are by far the most exploited. In addition, lipid rafts, including cholesterol, ceramide, and phosphatidylserine, can also be used. Tetraspanins compose a superfamily of proteins characterised by four transmembrane domains. As abovementioned, they usually present widespread tissue distribution, but tissue-specific tetraspanins can also be found [51]. Their wide expression makes them the favourite candidate as the target in bioaffinity assays. Notwithstanding, tetraspanins and other widely expressed surface proteins are indeed often deficient in terms of specificity.
This section will discuss literature-reported examples that tackle total EV detection using ubiquitous markers (e.g., tetraspanins), categorising them by the type of biorecognition elements used for their capture, in label-free or label-based approaches, or even exploiting different capture mechanisms. Table 1 summarises all the crucial aspects behind the realisation of the biosensing platforms discussed in this section. Then, the section that follows will instead focalise on a more targeted recognition exploiting biomarkers that are specifically found in certain types of tumours.
Capture of EVs with antibodies
Although being the simplest and most common biorecognition elements, the scientific community is slowly trying to overcome antibodies due to some of their disadvantages, such as the need for in vivo laborious procedures for their obtainment. Nonetheless, antibodies are still greatly utilised in EV biosensing [52]. Kilic T. and co-workers [53], for instance, proposed a very simple label-free detection scheme involving the biorecognition reaction between an Anti-CD81 antibody and the CD81 present on the lipid membrane of the EVs, followed by EIS measurements. They obtained a LOD of 77 EVs/mL, with a dynamic range between 102 and 109 EVs/mL. Very recently, a similar label-free EIS-based immunosensing approach was studied by Guldin’s group [54].
Magnetic particles (MPs) functionalised with antibodies that recognise tetraspanins are widely available on the market from a variety of sources. Otherwise, researchers can easily functionalise MPs by harnessing a number of interactions (e.g., the streptavidin–biotin interaction, thus immobilising biotinylated antibodies on streptavidin-coated beads). Hence, MPs can be used to achieve a simple and fast pre-concentration of EVs even in complex biological media. Moura and co-workers [55] investigated the electrochemical detection of EVs through a capture based on MPs modified with different Anti-CDX antibodies. In particular, they focused on general tetraspanins (e.g., CD9, CD63, and CD81), but also on tetraspanins more specific in cancer-derived EVs (e.g., CD24, CD44, CD54, CD326, and CD340). The EVs were harvested from three breast cancer cell lines, namely MCF-7, MDA-MB-231, and SKBR3. They noticed that the separation based on Anti-CD81 antibodies was less affected by free receptors potentially present in the sample, hence resulting in an approach that could avoid UC to separate EVs from proteins. Moreover, studying serum samples from patients with breast cancer, they observed better performances when they combined the capture based on a specific cancer receptor (e.g., CD24 and CD340) and the following labelling with a second receptor focused on more general antigens (e.g., CD63). Very recently, the same group [56] proposed for the first time the electrochemical biosensing of EVs by exploiting the intrinsic presence of the enzyme alkaline phosphatase (ALP) on their surface (Fig. 3a). In particular, after capturing the EVs with Anti-CDX antibodies (i.e., CD81, CD63, and CD9), they monitored the activity of ALP by detecting the electroactive product generated through the enzymatic reaction with p-nitrophenyl phosphate (pNPP). Different types of antibodies can also be used in parallel. Revzin’s group [57] managed to simultaneously detect CD63 and CD81 biomarkers by functionalising Au nanoparticles (NPs) with anti-CD63 and anti-CD81 antibodies doped with metal ions (Pb2+ and Cu2+) to generate a redox signal (Fig. 3b). By doing this, they gained a sensitivity expressed with LODs around 105 particles/mL.
a Determination of immunomagnetically separated EVs through a colorimetric and electrochemical assay exploiting the enzymatic activity between the alkaline phosphatase (ALP) intrinsically present on EVs and p-nitrophenyl phosphate (pNPP) to produce electroactive p-nitrophenol (pNP); reprinted with permission from ref. [56]. b Capture of urinary EVs on a Au electrode and multiplexed square-wave voltammetry (SWV) detection by labelling with Au NPs functionalised with anti-CD63 and anti-CD81 antibodies doped with metal ions. Reprinted with permission from ref. [57], Copyright 2021 American Chemical Society. c The aptamer fragment split-a immobilised on a Au electrode recognised the N-glycoprotein on the surface of EVs, whereas a second fragment (split-b) was used for labelling with methylene blue (MB), but such configuration could be destroyed by rinsing the electrode in phosphate buffer saline (PBS) at 37 °C, thus allowing surface regeneration; reprinted with permission from ref. [60]. d EVs were adsorbed on Zr-MOFs-modified paper and then three hairpin probes (HP, HP1, HP2) recognised the CD63 marker on the membrane, triggering an HCR and amplifying the signal generated by using tetramethylbenzidine (TMB); reprinted with permission from ref. [66], Copyright 2021 American Chemical Society
Capture of EVs with nucleic acids
Nucleic acid–based detection uses the functional and structural properties of nucleic acids for the development of interesting approaches in the capture and detection of EVs. Aptamers, for instance, are single-stranded (ss) oligonucleotides whose three-dimensional conformation gives them the ability to bind to specific targets [58]. Indeed, researchers are constantly putting more effort into the development and improvement of aptasensors for the detection of EVs [59, 60] (Fig. 3c). When dealing with nucleic acids, one of the most common forms of signal amplification relies on the hybridisation chain reaction (HCR). It is an isothermal and enzyme-free technique that can be triggered when at two stable DNA monomers (DNA hairpins) in solution gets added a target DNA fragment. Thus, it can be easily initiated by using DNA aptamers when capturing EVs. An et al. [61] captured EVs on a glassy carbon electrode using an aptamer able to recognise CD63 on EVs after incubation at 37 °C for 1 h. To increase the number of immobilised aptamers, the electrode was modified with graphene oxide (GO) and Au NPs. Moreover, the signal was amplified through HCR and the electrochemical method was based on click chemistry, resulting in the ultrasensitive quantification of EVs by monitoring the reduction current of 2,3-diamoniphenazine. The practical applicability of the sensor was successfully tested in real serum samples derived from healthy individuals and breast cancer patients, proving to be resistant to interferences and promising in clinical diagnostics.
The molecular properties of nucleic acids (e.g., self-assembly) can indeed be of use to design biosensing nanostructures. Among these novel nanotechnologies, mainly used for signal amplification, DNA nanotetrahedron-assisted mechanisms have found applications in the field of EVs. Nowadays, DNA walker–based strategies are gaining continuously more momentum in research as a way to amplify the signal. Through strand displacement, nicking endonucleases, or DNAzymes, a DNA walker can be moved along oligonucleotide sequences, producing detectable single-stranded DNA fragments from the destruction of the DNA along its path. Miao P. and Tang Y. [62] managed to immobilise multiple walker strands on a single EV. The captures took place by using Fe3O4@Au magnetic nanoparticles modified with aptamers that recognised CD63 proteins. Such multipedal DNA walker strategy allowed a LOD of 6 particles/µL in buffer solution, using EVs obtained from HeLa cells through conventional centrifugation. HeLa cells were also compared with equal concentrations of MCF-10A cells, which express lower levels of CD63, showing that the system is capable of discriminating between the two.
Capture using both antibodies and nucleic acids
Sandwich-like designs that use both antibodies and aptamers for capture and signal generation can also be found. A sandwich-like structure of these two biorecognition elements was reported in the work of Cao Y. et al. [63]. A first enrichment of the EVs was carried out by using magnetic beads coated with anti-CD63 antibodies at 37 °C for 1 h. Then, the conjugate was recognised by CD63 aptamers in 0.5 h and a cascade toehold-mediated strand displacement reaction (CTSDR) was triggered. Such molecular machine provided a LOD of 1.72 × 104 EVs/mL.
Metal–organic framework–based platforms
Metal–organic frameworks (MOFs) are interesting materials that offer great advantages in analytical chemistry due to the high specific surface area. They are especially useful to augment the number of the platform’s accommodable electroactive molecules, thanks to their high porosity. Zirconium-phosphate (ZIF-67) MOFs are gaining wide attention lately as a coordination chemistry-based interaction with EVs (electrostatic interaction between their phospholipid membrane and Zr4+) in substitution to the classic biological recognition [64]. Sun Y. and colleagues [65] focused on the detection of CD63 proteins on MCF-7 cells-derived EVs. Particularly, they developed a self-calibration sensor based on the assembly of black phosphorus nanosheets and ferrocene (Fc)-doped ZIF-67 MOFs on an indium-tin oxide (ITO) electrode coated with methylene blue–labelled CD63 aptamers. A dual-signal output was thus generated by Fc and methylene blue, allowing a LOD down to 100 EVs/mL. The system was also challenged by analysing clinical plasma samples from breast cancer patients.
Aptamers and antibodies are largely used as capture probes, as already shown. However, some assay schemes could benefit from the absence of capture probes, relying on different forms of immobilisation. For instance, hydrophilic phosphate heads present on the surface of EVs can be cleverly exploited for the direct enrichment of EVs without using other biorecognition elements. It is the case of X. Liu’s work [66], in which the authors used Zr-MOFs-functionalised paper for the coordinate interaction with the hydrophilic phosphate heads onto the surface of EVs (Fig. 3d). After such immobilisation, the signal was generated through the recognition mediated by CD63 aptamers and amplified via HCR, which generated multiple hemin/G-quadruples DNAzymes. Such architecture achieved a LOD of 5 × 103 particles/mL.
EV detection using cancer-specific markers
Surface-embedded biomarkers
Tetraspanins and the other markers mentioned before cannot always give specific information about the cancer involved. However, a wide variety of host cell–dependent proteins are also available on the EVs’ surface and can be of use to gather information on ongoing neoplasia [68]. More biomarkers can be specific for more types of tumours and vice versa [69]. The following section will discuss works that focus on some of the specific cancer markers found on the membrane that encloses the EVs. Table 2 gives a schematic outline of some of the aspects reported in the research papers mentioned, categorising them on the basis of the biomarker and the tumour studied.
Epidermal growth factor receptor (EGFR)
Human EGFR is highly expressed in glioblastoma-derived EVs, thus representing an optimal non-invasive candidate for the early detection of such intracranial tumour. Instead of antibodies and aptamers, anti-EGFR affibodies, namely affinity reagents able to bind to a variety of targets [70], have been successfully used to functionalise microcapillaries and capture EVs harvested from non-small-cell lung cancer (NSCLC) cell lines H1975 in a label-free fashion [71]. Besides avoiding labelling probes and agents, bypassing the need for capture probes is also desirable. As introduced in the previous section, Zr-MOFs show a high affinity to phosphate groups, forming Zr-O-P interactions. Thanks to these bonds, EVs can be captured or recognised without the need for specific probes, but by exploiting the intrinsic phosphate groups present on their surface. Sun Z. et al. [72], for instance, immobilised glioblastoma-derived EVs, obtained by differential centrifugation from U87 cell lines, using a peptide ligand able to selectively bind to the EGFR. Then, the electrochemical signal is generated by the absorption of methylene blue–loaded Zr-MOFs. The researchers obtained a LOD of 7.83 × 103 EVs/µL with this method. Indeed, peptides have shown to be interesting synthetic recognition elements with good binding affinities and low costs [73, 74]. Moreover, their synthesis is easily implemented with unpaired production rates. Their tuneable functional groups can be conveniently used to target multiple biomolecules, hence representing a versatile tool in biosensing. However, to the best of our knowledge, only very few examples of EV detection using peptides can be found in the recent literature.
Epithelial cell adhesion molecule (EpCAM)
EpCAM, sometimes indicated as CD326, is a transmembrane glycoprotein frequently heterogeneously overexpressed in carcinomas, but not in cancers of non-epithelial origin. Chen’s group [75] recently introduced a multidirectional HCR (mHCR) to enhance the detection of EVs from MCF-7 cells using aptamers with an affinity for the EpCAM biomarker for their capture. Notably, to trigger mHCR, they used a cholesterol-modified H shape–like DNA structure able to plunge its cholesterol moiety into the lipid bilayer of EVs, thus getting anchored. The signal, measured via chronoamperometry, was generated by the redox reaction of 3,3′,5,5′-tetramethylbenzidine (TMB) and hydrogen peroxide (H2O2) catalysed by horseradish peroxidase (HRP). The authors reported a LOD of 285 EVs/µL and managed to discriminate between tumorous (MCF-7 cells) and non-tumorous (MCF-10A cells) EVs in human serum. This example highlights the potential of lipid anchors, such as cholesterol, as new biorecognition elements that can be inserted, via hydrophobic interactions, into the lipid bilayer of the EV’s membrane. This is especially interesting since such recognition is not influenced by the number of markers present on the surface of EVs [76].
When dealing with complex biological fluids, the analysis often faces hurdles linked to nonspecific adsorption of unwanted material (e.g., proteins) on the electrode surface, thus limiting the electroactive area available and, therefore, the sensitivity of the assay. This is especially true in the case of non-isolated EVs, which represent a minor subpopulation of the whole set of circulating particles found in such biological media. In this regard, nanoporous structures could be beneficial by limiting the diffusion and adsorption of fouling material [52], as investigated in a recent work that captures prostate cancer EVs from undiluted plasma and urine samples using antibodies (Fig. 4a) [77].
a Signal output and fouling effect between a flat electrode surface and a nanoporous gold (NSG) electrode using tetramethylbenzidine (TMB) as electroactive compound in the analysis of undiluted biological samples; reprinted with permission from ref. [77]. b Dual-aptamer simultaneous recognition of MUC1 (P1) and CD63 (P2) markers on magnetically separated EVs, amplifying the signal through hyperbranched HCR mediated by a cholesterol-modified DNA probe (P3) and DNA tetrahedral nanostructures (DTNH1 and DTNH2); reprinted with permission from ref. [79], Copyright 2021 American Chemical Society. c Sandwich electrochemical immunoassay for the analysis of ovarian cancer cell–derived EVs using CD9 and CA-125 antibodies on paper-based carbon electrodes (PCEs); reprinted with permission from ref. [80]
Mucins
The mucin family is composed of glycosylated macromolecules largely expressed in mammalian epithelial cells [78]. It has been reported that the abnormal expression of Mucin 1 (MUC1), a high molecular weight membrane glycoprotein, can represent a potential cancer marker. Yang L. and colleagues [79] recently proposed a dual-aptamer isolation and recognition of EVs from MCF-7 cell lines exploiting the overexpression of both MUC1 and CD63 markers on their surface (Fig. 5b). After the capture, a cholesterol-modified DNA probe could be anchored onto the EVs and trigger an HCR for signal generation and amplification. Mucin 16, also known as the cancer antigen 125 (CA-125), is the most prominent EV biomarker in the case of ovarian cancer [78]. Very recently, Kasetsirikul S. et al. [80] worked on the development of a paper-based device able to capture and detect the whole bulk of EVs using antibodies that bind to generic markers (e.g., CD9) (Fig. 4c). Then, they used antibodies specific to the CA-125 marker to identify ovarian cancer–positive samples (LOD of 7.1 × 108 EVs/mL).
a Extraction and lysis of EVs to capture intravesicular microRNA-122 using a hairpin DNA (hpDNA) probe immobilised on a Au electrode and triggering HCR using DNA helpers (H1 and H2) to amplify the signal generated by Ru(NH3)6.3+ (RuHex); exonuclease I (Exo I) was used to eliminate false positive signals; reprinted with permission from ref. [91], Copyright 2020 American Chemical Society. b Dual-amplification reactions by target-mediated cyclic strand displacement reaction (TCSDR) and primer exchange DNA amplification reaction (PEDAR) for the detection of miRNA-21 in the presence of primer probes (Pp); template probes (Tp) hybridise with the TCSDR-generated primers and PEDAR continues with the assistance of Bst DNA polymerase; finally, the signal is generated using methylene blue (MB); reprinted with permission from ref. [94]. c Detection of the CD147 biomarker in intact and lysed EVs derived from colorectal cancer (CRC) cell lines with a sandwich of antibodies onto the surface of MPs immobilised on screen-printed carbon electrodes (SPCE), and the subsequent amperometric transduction using hydroquinone (HQ) and H2O2 in the presence of horseradish peroxidase (HRP); reprinted with permission from ref. [97]
Tyrosine kinase–like 7 (PTK7) and prostate-specific membrane antigen (PSMA)
PSMA and PTK7 are two abundantly expressed proteins on the membrane of EVs harvested from LNCaP and HeLa cell lines, respectively. Recently, Yu Y. and co-workers [81] focused on both biomarkers and used them as inputs for an and logic gate assisted by dual-aptamer recognition and cyclic enzymatic signal amplification. With this approach, they managed to improve the specific recognition and lower interferences. In each microliter of sample, they could detect down to 920 EVs obtained from CCRF-CEM cells (human leukemic lymphoblasts) by differential centrifugation.
Intravesicular biomarkers
Indeed, most electrochemical biosensors are used for the detection of surface-embedded protein markers on EVs. However, EVs also inherit important cargo molecules (e.g., intravesicular nucleic acids) that can be of clinical use, keeping in mind that their biochemical composition highly depends on their physiological role, as well as their origin and fate. Many papers that include biorecognition molecules that are used to interrogate EV-derived microRNAs (herein abbreviated as miRNAs or miR) and other nucleic acids have been published. miRNAs from EVs have a role in cancer progression and metastatic events via intercellular communication: such encapsulated RNAs are transferred from cell to cell as a signalling method [84]. High levels of such nucleic acids (miRNA-21 is the most widely studied) may indicate poor survivability rates. Therefore, the analysis of intra-EV miRNAs may represent truly remarkable potential in cancer diagnostics. Interestingly, it is only in 2016 when the first product aimed at isolating and analysing exosomal RNA was released on the market in the USA [85].
The use of detergents and organic extractions are the most common solutions to induce the lysis of the vesicles, hence allowing access to the biomarkers present in the intravesicle space. Many commercial kits are available on the market to meet the need for simple and rapid ways to examine the molecular cargo (nucleic acids and proteins) of pre-enriched EVs. The most widely known is probably the TRIzol™ reagent (Life Technologies), introduced in 1987 by P. Chomczynski and N. Sacchi to isolate RNA from cells and tissues with a single extraction [86]. Similarly, the “Total Exosome RNA & Protein Isolation Kit” by Life Technologies follows three phases based on (i) an organic extraction with acid phenol:chloroform, (ii) a purification step by immobilising RNA on glass-fiber filters, and finally (iii) the elution using a low ionic-strength solution. The whole procedure takes about 30–60 min. Another widely used kit is the “exoRNeasy Midi/Maxi Kit” (Qiagen), which uses a membrane-based affinity binding step to recover the entire spectrum of EVs present in the sample, regardless of particular epitopes. The elution step is, again, based on a phenol and guanidine thiocyanate lysis to collect RNA (and denature RNases), following an organic extraction in chloroform. Table 3 provides a series of information regarding the research articles discussed in this section, including the lysis approach chosen.
Instead of antibodies and aptamers, complementary sequences of DNA are commonly used as bioreceptors in miRNA analysis [87, 88]. Magnetic particles have been applied for years in the field of nucleic acids [89]; thus, it is not unusual to find their implementation in the determination of EV-derived miRNAs [90]. Many approaches are used to enhance the sensitivity of the biosensing scheme, including signal amplification strategies like HCR [91, 92] (Fig. 5a) and molecular tools based on strand displacement reactions [93, 94] (Fig. 5b). Besides DNA sequences, other synthetic receptors are widely used. Peptide nucleic acids (PNA), for instance, are an interesting alternative [95]. The combination of a short electroneutral PNA probe, able to recognise the complementary domain in miRNA-21, with a spherical nucleic acid (SNA) nanoprobe, able to load great quantities of electroactive tags onto the electrode, was reported [96]. Such PNA-miRNA-SNA sandwich allowed the enzyme-free detection of the analyte down to concentrations of 49 aM.
As mentioned in the “Introduction”, the intravesicular cargo comprises multiple macromolecules in addition to nucleic acids. An interesting one, for instance, is the cluster of differentiation 147 (CD147), poorly studied up until now. It plays a crucial role, for instance, in the progression of colorectal cancer. Inspired by its relevance and the lack of articles that explore its detection, the research group of S. Campuzano [97] contributed by designing an electrochemical strategy based on a sandwich of antibodies able to capture the CD147 biomarker in EVs from colorectal cancer cells. In particular, using magnetic beads, they managed to detect such analyte both in intact and in lysed EVs (Fig. 5c), with results in agreement with those obtained by ELISA and Western blot techniques.
Integrated lab-on-a-chip microfluidic platforms
Microfluidics allows the manipulation of small volumes of samples by means of precise fluidic control. The increasing number of integrated analytical platforms that comprise multiple analytical phases, from sample processing to detection, resulted in augmented research around microfluidic approaches to obtain EV separation on a portable scale [100]. After the first successful attempt in 2010 [101], in which it was shown that a microfluidic EV isolation without a previous UC step was possible working in serum, many researchers tried to implement this kind of separation strategy inside their biosensing platforms. Electrochemical sensors are easily integrated with microfluidics, thus obtaining rapid and cost-effective platforms apt for point-of-care testing (POCT) that include all the steps required in EV detection [102].
Among the diverse technologies exploited to isolate EVs, immunoaffinity-based separations can be implemented inside these chips, for instance by modifying the various microchannels with antibodies or inserting MPs functionalised with specific ligands into the flow. Xu H. et al. [103] presented a two-stage microfluidic platform that comprises both an on-chip magnetic isolation and a subsequent electrochemical detection of EVs (Fig. 6a). Interestingly, the capture is accomplished by using MPs modified with Tim4 receptors that bind to the phosphatidylserine expressed on the EVs’ membrane. The various liquids were injected using a programmable syringe pump at constant flow rates. A permanent magnet placed underneath the platform was used to retain the beads in position as the EV-containing solutions flowed. Then, the magnet was moved to the electrode area for the detection (LOD of 4.39 × 103 EVs/mL). The device was able to discriminate between healthy and hepatic carcinoma patients, with a protocol that required 30 µL of clinical blood samples and took about 3.5 h for the response. In many immunoaffinity-based methodologies, the antibodies used to capture EVs are immobilised harnessing the high-affinity binding between biotin and avidin. However, if one needs to release the EVs captured for further downstream analyses, such strong binding is not ideal, since the operations required to break it operate at extreme pH and high temperature [104]. As an alternative to biotin, Nagrath’s group [105] worked with desthiobiotin, an analogue of biotin that presents a lower binding affinity to avidin, making it possible to release the EVs captured onto magnetic beads by EpCAM desthiobiotin-conjugated antibodies inside their microfluidic device.
a Microfluidic platform that integrates magnetic separation of EVs, using Tim4-modified MPs, and electrochemical analysis on an ITO electrode; reprinted with permission from ref. [103], Copyright 2018 American Chemical Society. b Streptavidin-coated MPs modified with cholesterol DNA anchors to capture EVs through hydrophobic interactions, while a sandwich is formed using MUC1 aptamers modified with CuO nanoparticles (NPs) able to generate a change in absorbance and photocurrent response of 10-benzyl-2-amino-acridone (BAA) used as signalling probe; reprinted with permission from ref. [116]. c Au NPs deposited on a MoS2@Ti3C2 nanohybrid–based PEC platform for the analysis of EV-derived miRNA-92a-3p, using 6-mercapto-1-hexanol (MCH) to block nonspecific adsorption sites; reprinted with permission from ref. [120]
Another isolation strategy is accomplished by size-based separations, for instance by adding membranes or nanowires with pore sizes apt for the filtration of particles with desired dimensions. A third option is characterised by the use of external forces between the flow medium and EVs therein contained, thus dynamically concentrating the particles in the light of their size, density, or other properties. Regarding this latter methodology, acoustic waves, electric fields, and centrifugal forces are the most common external inputs to achieve separation [106]. On the other hand, dynamic isolation is also possible relying solely on the hydrodynamic properties generated by the fluidic system. Deterministic lateral displacement (DLD) is indeed the most widespread design among these “passive” dynamic approaches. Briefly, in DLD, different flow laminae separate particles smaller and larger than a defined critical diameter [107]. Indeed, in microfluidic devices, sample mixing is pivotal and can be achieved by playing with the diffusion effect and creating different flows [108]. To enhance flow mixing, increase interactions between analytes and recognition elements, and augment the capture possibility of EVs, herringbone structures are widely used due to the chaotic effect they generate [109]. A geometrically activated surface interaction chip was implemented inside a detachable microfluidic device featuring an asymmetric herringbone structure to enhance the collision between cancerous EVs and biorecognition elements immobilised on a gold electrode [110]. In particular, EpCAM aptamers were used both for capture and labelling, achieving a LOD of 17 EVs/µL, with a dynamic range between 102 and 109 EVs/µL. Moreover, such detachable 3D-printed housing can be separated from the aptasensor, and EVs can be harvested for further downstream analyses. Exploiting a similar herringbone microfluidic mixer, Zhang Y. et al. [109] proposed a chip able to purify pancreatic cancer EVs directly from plasma, focusing on the glypican-1 biomarker. Then, the obtained EVs were also used to identify a miRNA signature in this kind of cancer, finding that miRNA-125b-5p and miRNA-214-3p could be of diagnostic value.
In the last decade, many complete critical reviews discuss thoroughly the application of microfluidics for isolation and analysis of EVs from biological fluids [9, 106, 111, 112]. It is interesting to point out that with such fluidic handling, EVs can maintain their original morphology; this is not always true for UC: the high mechanical force put on the particles tends to aggregate them [113]. Nonetheless, it has to be mentioned that microfluidics has yet to be improved, as for now, it cannot represent a stand-alone approach. As a matter of fact, architectures that run a microfluidic separation of EVs usually also include other techniques for better isolation [6].
Photoelectrochemical detection of extracellular vesicles
With photoelectrochemical (PEC) methods, the advantages listed for electrochemical approaches add up to many others. Indeed, using photoexcitation, PEC schemes couple the merits of electrochemical and optical strategies, resulting in weak background noises, high sensitivities, low costs, and the possibility to be implemented in miniaturised devices with ease [114]. Unfortunately, to the best of our knowledge, very few research articles involving the use of PEC for the detection of EVs are present in the recent literature [115]. In one of the latest published [116], a biotinylated cholesterol DNA anchor was attached to streptavidin-coated MPs to obtain a structure that could act as a capture probe for EVs via hydrophobic interactions between cholesterol and the lipid bilayer of the EVs (Fig. 6b). The modified MPs were thus incubated with EVs derived from MCF-7 and MCF-10A cell lines for 60 min at + 4 °C, continuously shaking gently. Then, an incubation with CuO nanoparticles modified with a thiolated MUC1 aptamer (Cu–S bond) followed, forming a sandwich (MBs-EV-CuO NPs). The subsequent addition of HNO3 caused the hydrolysis of CuO NPs to Cu2+. A 10-benzyl-2-amino-acridone (BAA)-modified ITO electrode was used for photoelectrochemical measurements. Among others, acridone derivatives have photoelectrochemical properties under UV–vis illumination. They found out that Cu2+ inhibited the photocurrent of BAA, producing a signal decrease proportional to the concentration of EVs in the sample (LOD of 1.38 × 103 particles/µL). This phenomenon was also confirmed by UV–vis spectroscopy and a colorimetric approach. Real clinical samples, particularly breast cancer patients’ serum, were also analysed to test the practical capability of the sensor. A similar capture approach based on a cholesterol-labelled DNA probe, but coupled to HCR, was also recently used to design a PEC biosensor able to detect EVs immobilised on magnetic beads modified with CD63 aptamers (LOD of 7.875 × 104 EVs/mL) [117].
On the other hand, PEC analysis of nucleic acids is more robust and common [118]. Thus, compared to the detection of intact EVs, a wider number of research papers that consider the interrogation of intravesicular miRNAs can be found in the literature [119]. With PEC, the key is working with stable photoactive nanomaterials. Indeed, these elements play crucial roles, increasing the sensitivity of the assay and working as photoactive sensitisers. In addition, hybrid structures can be formed to achieve better performances. Very recently, Sun Z. and colleagues [120] fabricated a PEC sensor depositing Au NPs on a MoS2@Ti3C2 nanohybrid (Fig. 6c). By doing this, they immobilised DNA probes to detect miRNA-92a-3p obtained from EVs with the miRNeasy Micro Kit (Qiagen) derived from colorectal cancer patients (LOD of 0.27 fM). Wang Y. et al. [121] focused instead on a heterostructure based on WO3 nanoflakes coated with ZIF-67 MOFs for the PEC detection of miRNA-21 (LOD of 0.5 fM) using the release of hemin to trigger a photocurrent quenching in the presence of the analyte.
Conclusions and future perspectives
The analysis of EVs’ content and surface allows precise conclusions regarding the cells from which they originated, as well as potential pathological conditions. By the examples discussed in the present review, electrochemical biosensors have recently proved their validity toward ultra-low determinations of EVs with the prospect of early-detections in cancer management. Examples of the (photo)electrochemical detection of intravesicular and extravesicular markers have been herein discussed, involving a wide variety of separation procedures, manipulation approaches, signal amplification strategies, and biorecognition elements. Researchers that work in this unstandardised field require constant updates to the recent procedures and their evolutions since EVs are analytes whose utilisation is often not straightforward.
Most scientific papers found in the literature use bioreceptors, e.g., antibodies and aptamers, that bind to the CD63 tetraspanin on EVs’ surface. Regarding aptamers, in particular, it is curious that most researchers use Anti-CD63 sequences and very few Anti-CD81 or Anti-CD9 aptamers, despite the fact that the latter two markers are quite common. When focusing on the molecular cargo of EVs, almost every work found in recent years uses miRNA-21 as model target, probably due to the vast knowledge accumulated over the years on this particular cancer biomarker, even though many other miRNA sequences can be found in EVs. However, the research around EV determination is still in its very first stages. The number of potential clinical biomarkers continuously increases, but their applications in the field are still quite limited. Indeed, many obstacles hamper clinical uses of the wide variety of biomarkers increasingly discovered nowadays, not least the low concentration and stability of these molecules inside biological fluids. More studies in this field will grant better knowledge about EV composition and its variation (e.g., due to the progression of the tumour), thus allowing more precise solutions for isolation and detection. Indeed, one of the main hurdles that has to be addressed in the near future regards the stability of EVs, in order to provide easier and more tailorable analyses. In addition to this, due to the heterogeneity in EV markers, multiplex analyses that focus simultaneously on more EV markers (both outside and inside) are highly desirable to provide accurate profiling of EVs in clinical samples. Furthermore, analytical workflows may be simplified, hence reducing the time required for the overall analysis. It is also crucial to highlight that, nowadays, the conventional methods described in the previous sections are difficult to implement at a clinical level due to their complexity and cost.
In our opinion, next-generation biosensing devices will be based on portable analytical tools and the scientific community is constantly moving toward this objective. After all, the COVID-19 pandemic has shown the true potential of POCT devices in diagnostics. Although not easy, we can envisage more and better-integrated platforms capable of seamlessly performing both separation and detection of EVs in near-patient applications. Magnetic beads could be an interesting tool able to easily perform EV isolation and then be magnetically immobilised onto the electrode for the detection, all in a straightforward workflow.
References
Siravegna G, Marsoni S, Siena S, Bardelli A. Integrating liquid biopsies into the management of cancer. Nat Rev Clin Oncol. 2017;14:531–48. https://doi.org/10.1038/nrclinonc.2017.14.
Johnstone RM. Maturation of reticulocytes: formation of exosomes as a mechanism for shedding membrane proteins. Biochem Cell Biol. 1992;70:179–90. https://doi.org/10.1139/o92-028.
van Niel G, D’Angelo G, Raposo G. Shedding light on the cell biology of extracellular vesicles. Nat Rev Mol Cell Biol. 2018;19:213–28. https://doi.org/10.1038/nrm.2017.125.
Raposo G, Stoorvogel W. Extracellular vesicles: exosomes, microvesicles, and friends. J Cell Biol. 2013;200:373–83. https://doi.org/10.1083/jcb.201211138.
Willms E, Johansson HJ, Mäger I, Lee Y, Blomberg KEM, Sadik M, Alaarg A, Smith CIE, Lehtiö J, EL Andaloussi S, Wood MJA, Vader P (2016) Cells release subpopulations of exosomes with distinct molecular and biological properties. Sci Rep 6:22519. https://doi.org/10.1038/srep22519.
Gaillard M, Thuaire A, Nonglaton G, Agache V, Roupioz Y, Raillon C. Biosensing extracellular vesicles: contribution of biomolecules in affinity-based methods for detection and isolation. Analyst. 2020;145:1997–2013. https://doi.org/10.1039/C9AN01949A.
Gould SJ, Raposo G. As we wait: coping with an imperfect nomenclature for extracellular vesicles. J Extracell Vesicles. 2013;2:20389. https://doi.org/10.3402/jev.v2i0.20389.
Théry C, Witwer KW, Aikawa E, et al. Minimal information for studies of extracellular vesicles 2018 (MISEV2018): a position statement of the International Society for Extracellular Vesicles and update of the MISEV2014 guidelines. J Extracell Vesicles. 7. https://doi.org/10.1080/20013078.2018.1535750.
Contreras-Naranjo JC, Wu HJ, Ugaz VM. Microfluidics for exosome isolation and analysis: enabling liquid biopsy for personalized medicine. Lab Chip. 2017;17:3558–77. https://doi.org/10.1039/c7lc00592j.
van der Vlist EJ, Nolte-’t Hoen ENM, Stoorvogel W, Arkesteijn GJA, Wauben MHM (2012) Fluorescent labeling of nano-sized vesicles released by cells and subsequent quantitative and qualitative analysis by high-resolution flow cytometry. Nat Protoc 7:1311–1326. https://doi.org/10.1038/nprot.2012.065.
Campos-Silva C, Cáceres-Martell Y, Sánchez-Herrero E, Sandúa A, Beneitez-Martínez A, González Á, Provencio M, Romero A, Jara-Acevedo R, Yáñez-Mó M, Valés-Gómez M. A simple immunoassay for extracellular vesicle liquid biopsy in microliters of non-processed plasma. J Nanobiotechnology. 2022;20:72. https://doi.org/10.1186/s12951-022-01256-5.
Dragovic RA, Gardiner C, Brooks AS, Tannetta DS, Ferguson DJP, Hole P, Carr B, Redman CWG, Harris AL, Dobson PJ, Harrison P, Sargent IL. Sizing and phenotyping of cellular vesicles using nanoparticle tracking analysis. Nanomedicine Nanotechnology, Biol Med. 2011;7:780–8. https://doi.org/10.1016/j.nano.2011.04.003.
Imanbekova M, Suarasan S, Lu Y, Jurchuk S, Wachsmann-Hogiu S. Recent advances in optical label-free characterization of extracellular vesicles. Nanophotonics. 2022;11:2827–63. https://doi.org/10.1515/nanoph-2022-0057.
György B, Szabó TG, Pásztói M, Pál Z, Misják P, Aradi B, László V, Pállinger É, Pap E, Kittel Á, Nagy G, Falus A, Buzás EI. Membrane vesicles, current state-of-the-art: emerging role of extracellular vesicles. Cell Mol Life Sci. 2011;68:2667–88. https://doi.org/10.1007/s00018-011-0689-3.
Ligler FS, Gooding JJ. Lighting up biosensors: now and the decade to come. Anal Chem. 2019;91:8732–8. https://doi.org/10.1021/acs.analchem.9b00793.
Liang T-TT, Qin X, Xiang Y, Tang Y, Yang F (2022) Advances in nucleic acids-scaffolded electrical sensing of extracellular vesicle biomarkers. TrAC Trends Anal Chem 148:116532. https://doi.org/10.1016/j.trac.2022.116532.
Lorencova L, Bertok T, Bertokova A, Gajdosova V, Hroncekova S, Vikartovska A, Kasak P, Tkac J. Exosomes as a source of cancer biomarkers: advances in electrochemical biosensing of exosomes. ChemElectroChem. 2020;7:1956–73. https://doi.org/10.1002/celc.202000075.
Bagheri Hashkavayi A, Cha BS, Lee ES, Kim S, Park KS. Advances in exosome analysis methods with an emphasis on electrochemistry. Anal Chem. 2020;92:12733–40. https://doi.org/10.1021/acs.analchem.0c02745.
Vaz R, Serrano VM, Castaño-Guerrero Y, Cardoso AR, Frasco MF, Sales MGF (2022) Breaking the classics: next-generation biosensors for the isolation, profiling and detection of extracellular vesicles. Biosens Bioelectron X 10 https://doi.org/10.1016/j.biosx.2022.100115.
Xie H, Di K, Huang R, Khan A, Xia Y, Xu H, Liu C, Tan T, Tian X, Shen H, He N, Li Z. Extracellular vesicles based electrochemical biosensors for detection of cancer cells: a review. Chinese Chem Lett. 2020;31:1737–45. https://doi.org/10.1016/j.cclet.2020.02.049.
Zhang L, Gu C, Wen J, Liu G, Liu H, Li L. Recent advances in nanomaterial-based biosensors for the detection of exosomes. Anal Bioanal Chem. 2021;413:83–102. https://doi.org/10.1007/s00216-020-03000-0.
Li S, Ma Q (2022) Electrochemical nano-sensing interface for exosomes analysis and cancer diagnosis. Biosens Bioelectron 214:114554. https://doi.org/10.1016/j.bios.2022.114554.
Yu X, Sha L, Dong L, Cao Y, Zhao J (2020) Recent advances in bio-sensing methods for the detection of tumor exosomes. Crit Rev Anal Chem 0:1–19. https://doi.org/10.1080/10408347.2020.1802220.
Malhotra S, Amin ZM, Dobhal G, Cottam S, Nann T, Goreham R V. (2021) Novel devices for isolation and detection of bacterial and mammalian extracellular vesicles. Microchim Acta 188. https://doi.org/10.1007/s00604-021-04790-5.
Bari SMI, Hossain FB, Nestorova GG. Advances in biosensors technology for detection and characterization of extracellular vesicles. Sensors. 2021;21:1–16. https://doi.org/10.3390/s21227645.
Xu L, Shoaie N, Jahanpeyma F, Zhao J, Azimzadeh M, Al−Jamal KT (2020) Optical, electrochemical and electrical (nano)biosensors for detection of exosomes: a comprehensive overview. Biosens Bioelectron 161:112222. https://doi.org/10.1016/j.bios.2020.112222.
Han R, Wang G, Xu Z, Zhang L, Li Q, Han Y, Luo X (2020) Designed antifouling peptides planted in conducting polymers through controlled partial doping for electrochemical detection of biomarkers in human serum. Biosens Bioelectron 164:112317. https://doi.org/10.1016/j.bios.2020.112317.
Zhou S, Yang Y, Wu Y, Liu S (2021) Review: Multiplexed profiling of biomarkers in extracellular vesicles for cancer diagnosis and therapy monitoring. Anal Chim Acta 1175:338633. https://doi.org/10.1016/j.aca.2021.338633.
Zhu X, Chen H, Zhou Y, Wu J, Ramakrishna S, Peng X, Nanda HS, Zhou Y (2021) Recent advances in biosensors for detection of exosomes. Curr Opin Biomed Eng 18:100280. https://doi.org/10.1016/j.cobme.2021.100280.
Cheng Y, Zeng Q, Han Q, Xia W. Effect of pH, temperature and freezing-thawing on quantity changes and cellular uptake of exosomes. Protein Cell. 2019;10:295–9. https://doi.org/10.1007/s13238-018-0529-4.
Erozenci LA, Pham TV, Piersma SR, Dits NFJ, Jenster GW, van Royen ME, Moorselaar RJA, Jimenez CR, Bijnsdorp IV. Simple urine storage protocol for extracellular vesicle proteomics compatible with at-home self-sampling. Sci Rep. 2021;11:20760. https://doi.org/10.1038/s41598-021-00289-4.
Wu Y, Deng W, Klinke DJ II. Exosomes: improved methods to characterize their morphology, RNA content, and surface protein biomarkers. Analyst. 2015;140:6631–42. https://doi.org/10.1039/C5AN00688K.
Maroto R, Zhao Y, Jamaluddin M, Popov VL, Wang H, Kalubowilage M, Zhang Y, Luisi J, Sun H, Culbertson CT, Bossmann SH, Motamedi M, Brasier AR (2017) Effects of storage temperature on airway exosome integrity for diagnostic and functional analyses. J Extracell Vesicles 6. https://doi.org/10.1080/20013078.2017.1359478.
Yu B, Zhang X, Li X. Exosomes derived from mesenchymal stem cells. Int J Mol Sci. 2014;15:4142–57. https://doi.org/10.3390/ijms15034142.
Lee M, Ban J-JJ, Im W, Kim M. Influence of storage condition on exosome recovery. Biotechnol Bioprocess Eng. 2016;21:299–304. https://doi.org/10.1007/s12257-015-0781-x.
Zhao Y, Chen K, Li H, Wu H. Effect of pH on the isolation of urinary exosome. Int Urol Nephrol. 2017;49:165–9. https://doi.org/10.1007/s11255-016-1408-7.
Doyle L, Wang M. Overview of extracellular vesicles, their origin, composition, purpose, and methods for exosome isolation and analysis. Cells. 2019;8:727. https://doi.org/10.3390/cells8070727.
Boriachek K, Islam MN, Möller A, Salomon C, Nguyen N, Hossain MSA, Yamauchi Y, Shiddiky MJA. Biological functions and current advances in isolation and detection strategies for exosome nanovesicles. Small. 2018;14:1702153. https://doi.org/10.1002/smll.201702153.
Gardiner C, Di VD, Sahoo S, Théry C, Witwer KW, Wauben M, Hill AF. Techniques used for the isolation and characterization of extracellular vesicles: results of a worldwide survey. J Extracell Vesicles. 2016;5:32945. https://doi.org/10.3402/jev.v5.32945.
Tauro BJ, Greening DW, Mathias RA, Ji H, Mathivanan S, Scott AM, Simpson RJ. Comparison of ultracentrifugation, density gradient separation, and immunoaffinity capture methods for isolating human colon cancer cell line LIM1863-derived exosomes. Methods. 2012;56:293–304. https://doi.org/10.1016/j.ymeth.2012.01.002.
Kornilov R, Puhka M, Mannerström B, Hiidenmaa H, Peltoniemi H, Siljander P, Seppänen-Kaijansinkko R, Kaur S. Efficient ultrafiltration-based protocol to deplete extracellular vesicles from fetal bovine serum. J Extracell Vesicles. 2018;7:1422674. https://doi.org/10.1080/20013078.2017.1422674.
Coughlan C, Bruce KD, Burgy O, Boyd TD, Michel CR, Garcia‐Perez JE, Adame V, Anton P, Bettcher BM, Chial HJ, Königshoff M, Hsieh EWY, Graner M, Potter H (2020) Exosome isolation by ultracentrifugation and precipitation and techniques for downstream analyses. Curr Protoc Cell Biol 88. https://doi.org/10.1002/cpcb.110.
Paolini L, Zendrini A, Di NG, Busatto S, Lottini E, Radeghieri A, Dossi A, Caneschi A, Ricotta D, Bergese P. Residual matrix from different separation techniques impacts exosome biological activity. Sci Rep. 2016;6:23550. https://doi.org/10.1038/srep23550.
Lane RE, Korbie D, Trau M, Hill MM (2017) Purification protocols for extracellular vesicles. pp 111–130.
Lobb R, Möller A (2017) Size exclusion chromatography: a simple and reliable method for exosome purification. pp 105–110.
Patel GK, Khan MA, Zubair H, Srivastava SK, Khushman M, Singh S, Singh AP. Comparative analysis of exosome isolation methods using culture supernatant for optimum yield, purity and downstream applications. Sci Rep. 2019;9:5335. https://doi.org/10.1038/s41598-019-41800-2.
Kalishwaralal K, Kwon WY, Park KS. Exosomes for non-invasive cancer monitoring. Biotechnol J. 2019;14:1800430. https://doi.org/10.1002/biot.201800430.
Zeng Y, Qiu Y, Jiang W, Shen J, Yao X, He X, Li L, Fu B, Liu X (2022) Biological features of extracellular vesicles and challenges. Front Cell Dev Biol 10. https://doi.org/10.3389/fcell.2022.816698.
Burtenshaw D, Regan B, Owen K, Collins D, McEneaney D, Megson IL, Redmond EM, Cahill PA (2022) Exosomal composition, biogenesis and profiling using point-of-care diagnostics—implications for cardiovascular disease. Front Cell Dev Biol 10. https://doi.org/10.3389/fcell.2022.853451.
Soltész B, Buglyó G, Németh N, Szilágyi M, Pös O, Szemes T, Balogh I, Nagy B. The role of exosomes in cancer progression. Int J Mol Sci. 2021;23:8. https://doi.org/10.3390/ijms23010008.
Jankovičová J, Sečová P, Michalková K, Antalíková J. Tetraspanins, more than markers of extracellular vesicles in reproduction. Int J Mol Sci. 2020;21:7568. https://doi.org/10.3390/ijms21207568.
Sahraei N, Mazloum-Ardakani M, Khoshroo A, Hoseynidokht F, Mohiti J, Moradi A (2022) Electrochemical system designed on a paper platform as a label-free immunosensor for cancer derived exosomes based on a mesoporous carbon foam- ternary nanocomposite. J Electroanal Chem 920:116590. https://doi.org/10.1016/j.jelechem.2022.116590.
Kilic T, Valinhas ATDS, Wall I, Renaud P, Carrara S. Label-free detection of hypoxia-induced extracellular vesicle secretion from MCF-7 cells. Sci Rep. 2018;8:1–9. https://doi.org/10.1038/s41598-018-27203-9.
Suthar J, Prieto-Simon B, Williams GR, Guldin S. Dual-mode and label-free detection of exosomes from plasma using an electrochemical quartz crystal microbalance with dissipation monitoring. Anal Chem. 2022;94:2465–75. https://doi.org/10.1021/acs.analchem.1c04282.
Moura SL, Martín CG, Martí M, Pividori MI (2020) Electrochemical immunosensing of nanovesicles as biomarkers for breast cancer. Biosens Bioelectron 150. https://doi.org/10.1016/j.bios.2019.111882.
Moura SL, Pallarès-Rusiñol A, Sappia L, Martí M, Pividori MI. The activity of alkaline phosphatase in breast cancer exosomes simplifies the biosensing design. Biosens Bioelectron. 2022;198:4–10. https://doi.org/10.1016/j.bios.2021.113826.
Lee S, Crulhas BP, Suvakov S, Verkhoturov SV, Verkhoturov DS, Eller MJ, Malhi H, Garovic VD, Schweikert EA, Stybayeva G, Revzin A. Nanoparticle-enabled multiplexed electrochemical immunoassay for detection of surface proteins on extracellular vesicles. ACS Appl Mater Interfaces. 2021;13:52321–32. https://doi.org/10.1021/acsami.1c14506.
Mascini M, Palchetti I, Tombelli S. Nucleic acid and peptide aptamers: fundamentals and bioanalytical aspects. Angew Chemie Int Ed. 2012;51:1316–32. https://doi.org/10.1002/anie.201006630.
Chang K, Sun P, Dong X, Zhu C, Liu X, Zheng D, Liu C. Aptamers as recognition elements for electrochemical detection of exosomes. Chem Res Chinese Univ. 2022;38:879–85. https://doi.org/10.1007/s40242-022-2088-8.
Liu D, Tang J, Xu H, Yuan K, Aryee AA, Zhang C, Meng H, Qu L, Li Z (2022) Split-aptamer mediated regenerable temperature-sensitive electrochemical biosensor for the detection of tumour exosomes. Anal Chim Acta 1219:340027. https://doi.org/10.1016/j.aca.2022.340027.
An Y, Jin T, Zhu Y, Zhang F, He P (2019) An ultrasensitive electrochemical aptasensor for the determination of tumor exosomes based on click chemistry. Biosens Bioelectron 142. https://doi.org/10.1016/j.bios.2019.111503.
Miao P, Tang Y. A multipedal DNA walker for amplified detection of tumor exosomes. Chem Commun. 2020;56:4982–5. https://doi.org/10.1039/d0cc01817a.
Cao Y, Li L, Han B, Wang Y, Dai Y, Zhao J (2019) A catalytic molecule machine-driven biosensing method for amplified electrochemical detection of exosomes. Biosens Bioelectron 141:111397. https://doi.org/10.1016/j.bios.2019.111397.
Palakollu VN, Chen D, Tang J-N, Wang L, Liu C. Recent advancements in metal-organic frameworks composites based electrochemical (bio)sensors. Microchim Acta. 2022;189:161. https://doi.org/10.1007/s00604-022-05238-0.
Sun Y, Jin H, Jiang X, Gui R. Assembly of black phosphorus nanosheets and MOF to form functional hybrid thin-film for precise protein capture, dual-signal and intrinsic self-calibration sensing of specific cancer-derived exosomes. Anal Chem. 2020;92:2866–75. https://doi.org/10.1021/acs.analchem.9b05583.
Liu X, Gao X, Yang L, Zhao Y, Li F. Metal-organic framework-functionalized paper-based electrochemical biosensor for ultrasensitive exosome assay. Anal Chem. 2021;93:11792–9. https://doi.org/10.1021/acs.analchem.1c02286.
An Y, Li R, Zhang F, He P (2021) A ratiometric electrochemical sensor for the determination of exosomal glycoproteins. Talanta 235:122790. https://doi.org/10.1016/j.talanta.2021.122790.
Wang M, Lin Y, Wu S, Deng Y, Zhang Y, Yang J, Li G (2022) An electrochemical biosensor for PD-L1 positive exosomes based on ultra-thin two-dimensional covalent organic framework nanosheets coupled with CRISPR-Cas12a mediated signal amplification. Sensors Actuators B Chem 362:131813. https://doi.org/10.1016/j.snb.2022.131813.
Hashkavayi AB, Cha BS, Lee ES, Park KS (2022) Dual rolling circle amplification-enabled ultrasensitive multiplex detection of exosome biomarkers using electrochemical aptasensors. Anal Chim Acta 1205:339762. https://doi.org/10.1016/j.aca.2022.339762.
Baydemir G, Bettazzi F, Palchetti I, Voccia D. Strategies for the development of an electrochemical bioassay for TNF-alpha detection by using a non-immunoglobulin bioreceptor. Talanta. 2016;151:141–7. https://doi.org/10.1016/j.talanta.2016.01.021.
Cavallaro S, Horak J, Hååg P, Gupta D, Stiller C, Sahu SS, Görgens A, Gatty HK, Viktorsson K, El Andaloussi S, Lewensohn R, Karlström AE, Linnros J, Dev A. Label-free surface protein profiling of extracellular vesicles by an electrokinetic sensor. ACS Sensors. 2019;4:1399–408. https://doi.org/10.1021/acssensors.9b00418.
Sun Z, Wang L, Wu S, Pan Y, Dong Y, Zhu S, Yang J, Yin Y, Li G. An electrochemical biosensor designed by using Zr-based metal-organic frameworks for the detection of glioblastoma-derived exosomes with practical application. Anal Chem. 2020;92:3819–26. https://doi.org/10.1021/acs.analchem.9b05241.
Sfragano PS, Moro G, Polo F, Palchetti I. The role of peptides in the design of electrochemical biosensors for clinical diagnostics. Biosensors. 2021;11:1–20. https://doi.org/10.3390/bios11080246.
Moro G, Severin Sfragano P, Ghirardo J, Mazzocato Y, Angelini A, Palchetti I, Polo F (2022) Bicyclic peptide-based assay for uPA cancer biomarker. Biosens Bioelectron 213. https://doi.org/10.1016/j.bios.2022.114477.
Wang L, Zeng L, Wang Y, Chen T, Chen W, Chen G, Li C, Chen J (2021) Electrochemical aptasensor based on multidirectional hybridization chain reaction for detection of tumorous exosomes. Sensors Actuators, B Chem 332:129471. https://doi.org/10.1016/j.snb.2021.129471.
Cheng N, Du D, Wang X, Liu D, Xu W, Luo Y, Lin Y. Recent advances in biosensors for detecting cancer-derived exosomes. Trends Biotechnol. 2019;37:1236–54. https://doi.org/10.1016/j.tibtech.2019.04.008.
Sabaté del Río J, Woo HK, Park J, Ha HK, Kim JR, Cho YK (2022) SEEDING to enable sensitive electrochemical detection of biomarkers in undiluted biological samples. Adv Mater 34. https://doi.org/10.1002/adma.202200981.
Lee D-H, Choi S, Park Y, Jin H. Mucin1 and Mucin16: therapeutic targets for cancer therapy. Pharmaceuticals. 2021;14:1053. https://doi.org/10.3390/ph14101053.
Yang L, Yin X, An B, Li F. Precise capture and direct quantification of tumor exosomes via a highly efficient dual-aptamer recognition-assisted ratiometric immobilization-free electrochemical strategy. Anal Chem. 2021;93:1709–16. https://doi.org/10.1021/acs.analchem.0c04308.
Kasetsirikul S, Tran KT, Clack K, Soda N, Shiddiky MJA, Nguyen N (2022) Low-cost electrochemical paper-based device for exosome detection 16–18. https://doi.org/10.1039/d2an00875k.
Yu Y, Guo Q, Jiang W, Zhang H, Cai C. Dual-aptamer-assisted and logic gate for cyclic enzymatic signal amplification electrochemical detection of tumor-derived small extracellular vesicles. Anal Chem. 2021;93:11298–304. https://doi.org/10.1021/acs.analchem.1c02489.
Zhao L, Sun R, He P, Zhang X. Ultrasensitive detection of exosomes by target-triggered three-dimensional DNA walking machine and exonuclease III-assisted electrochemical ratiometric biosensing. Anal Chem. 2019;91:14773–9. https://doi.org/10.1021/acs.analchem.9b04282.
Guo Y, Cao Q, Feng Q. Catalytic hairpin assembly-triggered DNA walker for electrochemical sensing of tumor exosomes sensitized with Ag@C core-shell nanocomposites. Anal Chim Acta. 2020;1135:55–63. https://doi.org/10.1016/j.aca.2020.08.036.
Tkach M, Théry C. Communication by extracellular vesicles: where we are and where we need to go. Cell. 2016;164:1226–32. https://doi.org/10.1016/j.cell.2016.01.043.
Sheridan C. Exosome cancer diagnostic reaches market. Nat Biotechnol. 2016;34:359–60. https://doi.org/10.1038/nbt0416-359.
Chomczynski P, Sacchi N. Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal Biochem. 1987;162:156–9. https://doi.org/10.1016/0003-2697(87)90021-2.
Serrano VM, Silva ISP, Cardoso AR, Sales MGF. Carbon electrodes with gold nanoparticles for the electrochemical detection of miRNA 21–5p. Chemosensors. 2022;10:189. https://doi.org/10.3390/chemosensors10050189.
Saha S, Allelein S, Pandey R, Medina-Perez P, Osman E, Kuhlmeier D, Soleymani L. Two-step competitive hybridization assay: a method for analyzing cancer-related microRNA embedded in extracellular vesicles. Anal Chem. 2021;93:15913–21. https://doi.org/10.1021/acs.analchem.1c03165.
Bettazzi F, Hamid-Asl E, Esposito CL, Quintavalle C, Formisano N, Laschi S, Catuogno S, Iaboni M, Marrazza G, Mascini M, Cerchia L, De Franciscis V, Condorelli G, Palchetti I. Electrochemical detection of miRNA-222 by use of a magnetic bead-based bioassay. Anal Bioanal Chem. 2013;405:1025–34. https://doi.org/10.1007/s00216-012-6476-7.
Tang X, Wang Y, Zhou L, Zhang W, Yang S, Yu L, Zhao S, Chang K, Chen M (2020) Strand displacement-triggered G-quadruplex/rolling circle amplification strategy for the ultra-sensitive electrochemical sensing of exosomal microRNAs. Microchim Acta 187. https://doi.org/10.1007/s00604-020-4143-9.
Guo Q, Yu Y, Zhang H, Cai C, Shen Q. Electrochemical sensing of exosomal microRNA based on hybridization chain reaction signal amplification with reduced false-positive signals. Anal Chem. 2020;92:5302–10. https://doi.org/10.1021/acs.analchem.9b05849.
Zhang J, Hou M, Chen G, Mao H, Chen W, Wang W, Chen J. An electrochemical biosensor based on DNA “nano-bridge” for amplified detection of exosomal microRNAs. Chinese Chem Lett. 2021;32:3474–8. https://doi.org/10.1016/j.cclet.2021.04.056.
Zhang J, Wang LL, Hou MF, Xia YK, He WH, Yan A, Weng YP, Zeng LP, Chen JH. A ratiometric electrochemical biosensor for the exosomal microRNAs detection based on bipedal DNA walkers propelled by locked nucleic acid modified toehold mediate strand displacement reaction. Biosens Bioelectron. 2018;102:33–40. https://doi.org/10.1016/j.bios.2017.10.050.
Wang LL, Chen WQ, Wang YR, Zeng LP, Chen TT, Chen GY, Chen JH (2020) Numerous long single-stranded DNAs produced by dual amplification reactions for electrochemical detection of exosomal microRNAs. Biosens Bioelectron 169:112555. https://doi.org/10.1016/j.bios.2020.112555.
Liu N, Lu H, Liu L, Ni W, Yao Q, Zhang GJ, Yang F. Ultrasensitive exosomal microRNA detection with a supercharged DNA framework nanolabel. Anal Chem. 2021;93:5917–23. https://doi.org/10.1021/acs.analchem.1c00295.
Liu L, Lu H, Shi R, Peng XX, Xiang Q, Wang B, Wan QQ, Sun Y, Yang F, Zhang GJ. Synergy of peptide-nucleic acid and spherical nucleic acid enabled quantitative and specific detection of tumor exosomal microRNA. Anal Chem. 2019;91:13198–205. https://doi.org/10.1021/acs.analchem.9b03622.
Pérez-Ginés V, Torrente-Rodríguez RM, Montero-Calle A, Solís-Fernández G, Atance-Gómez P, Pedrero M, Pingarrón JM, Barderas R, Campuzano S (2022) Tackling CD147 exosome-based cell-cell signaling by electrochemical biosensing for early colorectal cancer detection. Biosens Bioelectron X 11:100192. https://doi.org/10.1016/j.biosx.2022.100192.
Boriachek K, Umer M, Islam MN, Gopalan V, Lam AK, Nguyen NT, Shiddiky MJA. An amplification-free electrochemical detection of exosomal miRNA-21 in serum samples. Analyst. 2018;143:1662–9. https://doi.org/10.1039/c7an01843f.
Luo L, Wang L, Zeng L, Wang Y, Weng Y, Liao Y, Chen T, Xia Y, Zhang J, Chen J (2020) A ratiometric electrochemical DNA biosensor for detection of exosomal MicroRNA. Talanta 207:120298. https://doi.org/10.1016/j.talanta.2019.120298.
Chiriacò M, Bianco M, Nigro A, Primiceri E, Ferrara F, Romano A, Quattrini A, Furlan R, Arima V, Maruccio G. Lab-on-chip for exosomes and microvesicles detection and characterization. Sensors. 2018;18:3175. https://doi.org/10.3390/s18103175.
Chen C, Skog J, Hsu C-H, Lessard RT, Balaj L, Wurdinger T, Carter BS, Breakefield XO, Toner M, Irimia D. Microfluidic isolation and transcriptome analysis of serum microvesicles. Lab Chip. 2010;10:505–11. https://doi.org/10.1039/B916199F.
Wang F, Gui Y, Liu W, Li C, Yang Y. Precise molecular profiling of circulating exosomes using a metal–organic framework-based sensing interface and an enzyme-based electrochemical logic platform. Anal Chem. 2022;94:875–83. https://doi.org/10.1021/acs.analchem.1c03644.
Xu H, Liao C, Zuo P, Liu Z, Ye BC. Magnetic-based microfluidic device for on-chip isolation and detection of tumor-derived exosomes. Anal Chem. 2018;90:13451–8. https://doi.org/10.1021/acs.analchem.8b03272.
Holmberg A, Blomstergren A, Nord O, Lukacs M, Lundeberg J, Uhlén M. The biotin-streptavidin interaction can be reversibly broken using water at elevated temperatures. Electrophoresis. 2005;26:501–10. https://doi.org/10.1002/elps.200410070.
Lo T-W, Zhu Z, Purcell E, Watza D, Wang J, Kang Y-T, Jolly S, Nagrath D, Nagrath S. Microfluidic device for high-throughput affinity-based isolation of extracellular vesicles. Lab Chip. 2020;20:1762–70. https://doi.org/10.1039/C9LC01190K.
Lin B, Lei Y, Wang J, Zhu L, Wu Y, Zhang H, Wu L, Zhang P, Yang C. Microfluidic-based exosome analysis for liquid biopsy. Small Methods. 2021;5:2001131. https://doi.org/10.1002/smtd.202001131.
Hochstetter A, Vernekar R, Austin RH, Becker H, Beech JP, Fedosov DA, Gompper G, Kim S-C, Smith JT, Stolovitzky G, Tegenfeldt JO, Wunsch BH, Zeming KK, Krüger T, Inglis DW. Deterministic lateral displacement: challenges and perspectives. ACS Nano. 2020;14:10784–95. https://doi.org/10.1021/acsnano.0c05186.
Lee C-Y, Chang C-L, Wang Y-N, Fu L-M. Microfluidic Mixing: A Review. Int J Mol Sci. 2011;12:3263–87. https://doi.org/10.3390/ijms12053263.
Zhang Y, Tong X, Yang L, Yin R, Li Y, Zeng D, Wang X, Deng K (2021) A herringbone mixer based microfluidic device HBEXO-chip for purifying tumor-derived exosomes and establishing miRNA signature in pancreatic cancer. Sensors Actuators B Chem 332:129511. https://doi.org/10.1016/j.snb.2021.129511.
Kashefi-Kheyrabadi L, Kim J, Chakravarty S, Park S, Gwak H, Kim S Il, Mohammadniaei M, Lee MH, Hyun KA, Jung H Il (2020) Detachable microfluidic device implemented with electrochemical aptasensor (DeMEA) for sequential analysis of cancerous exosomes. Biosens Bioelectron 169:112622. https://doi.org/10.1016/j.bios.2020.112622.
Yang F, Liao X, Tian Y, Li G. Exosome separation using microfluidic systems: size-based, immunoaffinity-based and dynamic methodologies. Biotechnol J. 2017;12:1600699. https://doi.org/10.1002/biot.201600699.
Abreu CM, Costa-Silva B, Reis RL, Kundu SC, Caballero D. Microfluidic platforms for extracellular vesicle isolation, analysis and therapy in cancer. Lab Chip. 2022;22:1093–125. https://doi.org/10.1039/D2LC00006G.
Cvjetkovic A, Lötvall J, Lässer C. The influence of rotor type and centrifugation time on the yield and purity of extracellular vesicles. J Extracell Vesicles. 2014;3:23111. https://doi.org/10.3402/jev.v3.23111.
Svitkova V, Palchetti I (2020) Functional polymers in photoelectrochemical biosensing. Bioelectrochemistry 136:107590. https://doi.org/10.1016/j.bioelechem.2020.107590.
Pei Y, Ge Y, Zhang X, Li Y. Cathodic photoelectrochemical aptasensor based on NiO/BiOI/Au NP composite sensitized with CdSe for determination of exosomes. Microchim Acta. 2021;188:1–10. https://doi.org/10.1007/s00604-021-04716-1.
Xia Y, Chen T, Chen W, Chen G, Xu L, Zhang L, Zhang X, Sun W, Lan J, Lin X, Chen J (2022) A dual-modal aptasensor based on a multifunctional acridone derivate for exosomes detection. Anal Chim Acta 1191:339279. https://doi.org/10.1016/j.aca.2021.339279.
Qiu Z, Fan D, Xue X, Zhang J, Xu J, Lyu H, Chen Y. Ti 3 C 2 MXene-anchored photoelectrochemical detection of exosomes by in situ fabrication of CdS nanoparticles with enzyme-assisted hybridization chain reaction. RSC Adv. 2022;12:14260–7. https://doi.org/10.1039/d2ra01545e.
Sfragano PS, Pillozzi S, Palchetti I (2021) Electrochemical and PEC platforms for miRNA and other epigenetic markers of cancer diseases: recent updates. Electrochem commun 124:106929. https://doi.org/10.1016/j.elecom.2021.106929.
Pang X, Zhang X, Gao K, Wan S, Cui C, Li L, Si H, Tang B, Tan W. Visible light-driven self-powered device based on a straddling nano-heterojunction and bio-application for the quantitation of exosomal RNA. ACS Nano. 2019;13:1817–27. https://doi.org/10.1021/acsnano.8b07944.
Sun Z, Tong Y, Zhao L, Li J, Gao F, Wang C, Li H, Du L, Jiang Y (2022) MoS2@Ti3C2 nanohybrid-based photoelectrochemical biosensor: a platform for ultrasensitive detection of cancer biomarker exosomal miRNA. Talanta 238:123077. https://doi.org/10.1016/j.talanta.2021.123077.
Wang Y, Yang M, Shi H, Ge S, Wang X, Yu J. Photoelectrochemical detection of exosomal miRNAs by combining target-programmed controllable signal quenching engineering. Anal Chem. 2022;94:3082–90. https://doi.org/10.1021/acs.analchem.1c04086.
Funding
Open access funding provided by Università degli Studi di Firenze within the CRUI-CARE Agreement. This work was supported financially by Fondazione CR Firenze (grant numbers 2020.1662), Regione Toscana Bando Salute 2018 (CUP no. D78D20000870002), and POR Campania ESR 2014–2020 “SATIN” (GC). This work has received funding from the European Union’s Horizon 2020 Research and Innovation Programme under the Marie Sklodowska-Curie projects cONCReTE (872391), PRISAR2 (872860), to GC and to Italian Ministry of Economic Development (MISE) proposal number 529.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Published in the topical collection Electrochemical Biosensors – Driving Personalized Medicine with guest editors Susana Campuzano Ruiz and Maria Jesus Lobo-Castañón.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sfragano, P.S., Pillozzi, S., Condorelli, G. et al. Practical tips and new trends in electrochemical biosensing of cancer-related extracellular vesicles. Anal Bioanal Chem 415, 1087–1106 (2023). https://doi.org/10.1007/s00216-023-04530-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-023-04530-z