Abstract
Key message
Inversions and translocations are the major chromosomal rearrangements involved in Vigna subgenera evolution, being Vigna vexillata the most divergent species. Centromeric repositioning seems to be frequent within the genus.
Abstract
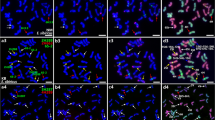
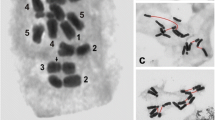
Oligonucleotide-based fluorescence in situ hybridization (Oligo-FISH) provides a powerful chromosome identification system for inferring plant chromosomal evolution. Aiming to understand macrosynteny, chromosomal diversity, and the evolution of bean species from five Vigna subgenera, we constructed cytogenetic maps for eight taxa using oligo-FISH-based chromosome identification. We used oligopainting probes from chromosomes 2 and 3 of Phaseolus vulgaris L. and two barcode probes designed from V. unguiculata (L.) Walp. genome. Additionally, we analyzed genomic blocks among the Ancestral Phaseoleae Karyotype (APK), two V. unguiculata subspecies (V. subg. Vigna), and V. angularis (Willd.) Ohwi & Ohashi (V. subg. Ceratotropis). We observed macrosynteny for chromosomes 2, 3, 4, 6, 7, 8, 9, and 10 in all investigated taxa except for V. vexillata (L.) A. Rich (V. subg. Plectrotropis), in which only chromosomes 4, 7, and 9 were unambiguously identified. Collinearity breaks involved with chromosomes 2 and 3 were revealed. We identified minor differences in the painting pattern among the subgenera, in addition to multiple intra- and interblock inversions and intrachromosomal translocations. Other rearrangements included a pericentric inversion in chromosome 4 (V. subg. Vigna), a reciprocal translocation between chromosomes 1 and 5 (V. subg. Ceratotropis), a potential deletion in chromosome 11 of V. radiata (L.) Wilczek, as well as multiple intrablock inversions and centromere repositioning via genomic blocks. Our study allowed the visualization of karyotypic patterns in each subgenus, revealing important information for understanding intrageneric karyotypic evolution, and suggesting V. vexillata as the most karyotypically divergent species.
Graphical abstract





Similar content being viewed by others
Availability of data and material
All data generated or analyzed during this study are included as supplementary materials.
Code availability
Not applicable.
References
Beliveau BJ, Joyce EF, Apostolopoulos N et al (2012) Versatile design and synthesis platform for visualizing genomes with Oligopaint FISH probes. Proc Natl Acad Sci USA 109:21301–21306. https://doi.org/10.1073/pnas.1213818110
Bi Y, Zhao Q, Yan W et al (2020) Flexible chromosome painting based on multiplex PCR of oligonucleotides and its application for comparative chromosome analyses in Cucumis. Plant J 102:178–186. https://doi.org/10.1111/tpj.14600
Braz GT, He L, Zhao H, Zhang T, Semrau K, Rouillard JM, Torres GA, Jiang J (2018) Comparative oligo-FISH mapping: an efficient and powerful methodology to reveal karyotypic and chromosomal evolution. Genetics 208:513–523. https://doi.org/10.1534/genetics.117.300344
Braz GT, Vale Martins L et al (2020a) A universal chromosome identification system for maize and wild Zea species. Chromosom Res 28:183–194. https://doi.org/10.1007/s10577-020-09630-5
Braz GT, Yu F, Vale Martins L et al (2020b) Fluorescent in situ hybridization using oligonucleotide-based probes. In: Nielsen BS, Jones J (eds) In situ hybridization protocols Methods in molecular biology. Humana, New York, pp 71–83
Braz GT, Yu F, Zhao H, Deng Z, Birchler JA, Jiang J (2021) Preferential meiotic chromosome pairing among homologous chromosomes with cryptic sequence variation in tetraploid maize. New Phytol 229:3294–3302. https://doi.org/10.1111/nph.17098
Carvalho CR, Saraiva LS (1993) An air drying technique for maize chromosomes without enzymatic maceration. Biotech Histochem 68:142–145. https://doi.org/10.3109/10520299309104684
Cheng F, Wu J, Wang X (2014) Genome triplication drove the diversification of Brassica plants. Horticult Res 1:14024. https://doi.org/10.1038/hortres.2014.24
de Oliveira BF et al (2021) Oligo-FISH barcode in beans: a new chromosome identification system. Theor Appl Genet 134:3675–3686. https://doi.org/10.1007/s00122-021-03921-z
Delgado-Salinas A, Thulin M, Pasquet R, Weeden N, Lavin M (2011) Vigna (Leguminosae) sensu lato: the names and identities of the American segregate genera. Am J Bot 98:1694–1715. https://doi.org/10.3732/ajb.1100069
do Vale Martins L et al (2019) Meiotic crossovers characterized by haplotype-specific chromosome painting in maize. Nat Commun 10:1–10. https://doi.org/10.1038/s41467-019-12646-z
do Vale Martins L et al (2021) BAC- and oligo-FISH mapping reveals chromosome evolution among Vigna angularis, V. unguiculata, and Phaseolus vulgaris. Chromosoma 130:133–147. https://doi.org/10.1007/s00412-021-00758-9
Ferguson-Smith MA, Trifonov V (2007) Mammalian karyotype evolution. Nat Rev Genet 8:950–962. https://doi.org/10.1038/nrg2199
Fonsêca A, Ferreira J, dos Santos TRB et al (2010) Cytogenetic map of common bean (Phaseolus vulgaris L.). Chromosom Res 18:487–502. https://doi.org/10.1007/s10577-010-9129-8
Garba M, Pasquet RS (1998) Isozyme diversity in Vigna vexillata (L.) A. Rich (Fabaceae) complex. S Afr J Botan 63(3):163–175. https://doi.org/10.1016/S0254-6299(15)30861-9
Han Y, Zhang Z, Liu C, Liu J, Huang S, Jiang J, Jin W (2009) Centromere repositioning in cucurbit species: implication of the genomic impact from centromere activation and inactivation. Proc Natl Acad Sci USA 106(35):14937–41. https://doi.org/10.1073/pnas.0904833106
Han Y, Zhang T, Thammapichai P, Weng Y, Jiang J (2015) Chromosome-specific painting in Cucumis species using bulked oligonucleotides. Genetics 200:771–779. https://doi.org/10.1534/genetics.115.177642
He L, Braz GT, Torres GA, Jiang J (2018) Chromosome painting in meiosis reveals pairing of specific chromosomes in polyploid Solanum species. Chromosoma 127:505–513
He L, Zhao H, He J et al (2020) Extraordinarily conserved chromosomal synteny of Citrus species revealed by chromosome-specific painting. Plant J 103:2225–2235. https://doi.org/10.1111/tpj.14894
Hedströn I, Thulin M (1986) Pollination by a Hugging Mechanism in Vigna vexillata (Leguminosae-Papilionoideae). Plant Syst Evol 154:274–283
Heslop-Harrison JS, Harrison GE, Leitch IJ (1992) Reprobing of DNA: DNA in situ hybridization preparations. Trends Genet 8:372–373
Hufnagel B, Marques A, Soriano A et al (2020) High-quality genome sequence of white lupin provides insight into soil exploration and seed quality. Nat Commun 11:1–12. https://doi.org/10.1038/s41467-019-14197-9
Iwata-Otsubo A, Lin JY, Gill N, Jackson SA (2016) Highly distinct chromosomal structures in cowpea (Vigna unguiculata), as revealed by molecular cytogenetic analysis. Chromosom Res 24:197–216. https://doi.org/10.1534/g3.115.024984
Jiang J (2019) Fluorescence in situ hybridization in plants: recent developments and future applications. Chromosom Res 27:153–165. https://doi.org/10.1007/s10577-019-09607-z
Kami J, Poncet V, Geffroy V, Gepts P (2006) Development of four phylogenetically-arrayed BAC libraries and sequence of the APA locus in Phaseolus vulgaris. Theor Appl Genet 112:987–998. https://doi.org/10.1007/s00122-005-0201-2
Kang YJ, Kim SK, Kim MY et al (2014) Genome sequence of mungbean and insights into evolution within Vigna species. Nat Commun 5:5443. https://doi.org/10.1038/ncomms6443
Kang YJ, Satyawan D, Shim S et al (2015) Draft genome sequence of adzuki bean, Vigna angularis. Sci Rep 28(5):8069. https://doi.org/10.1038/srep08069
Karuniawan A, Iswandi A, Kale PR, Heinzemann J, Grüneberg WJ (2006) Vigna vexillata (L.) A. Rich. cultivated as a root crop in Bali and Timor. Genet Resour Crop 53:213–217. https://doi.org/10.1007/s10722-005-1654-5
Kirov I, Khrustaleva L, Van LK, Soloviev A, Meeus S, Romanov D, Fesenko I (2017) DRAWID: user-friendly java software for chromosome measurements and idiogram drawing. Comp Cytogenet 11:747–757. https://doi.org/10.3897/compcytogen.v11i4.20830
Kubalová I, Câmara AS, Cápal P et al (2023) Helical coiling of metaphase chromatids. Nucleic Acids Res 51:2641–2654. https://doi.org/10.1093/nar/gkad028
Lewis GP (2005) Tribe Caesalpinieae. In: Lewis G, Schrire B, Mackinder B, Lock M (eds) Legumes of the world. Royal Botanic Gardens, Kew
Li H, Wang W, Lin L, Zhu X, Li J, Zhu X, Chen Z (2013) Diversification of the Phaseoloid legumes: effects of climate change, range expansion and habit shift. Front Plant Sci 4:1–9. https://doi.org/10.3389/fpls.2013.00386
Li G, Zhang T, Yu Z, Wang H, Yang E, Yang Z (2020) An efficient Oligo-FISH painting system for revealing chromosome rearrangements and polyploidization in Triticeae. Plant J 105:978–993. https://doi.org/10.1111/tpj.15081
Liu Y, Wang X, Wei Y, Liu Z, Lu Q, Liu F, Zhang T, Peng R (2020) Chromosome painting based on bulked oligonucleotides in cotton. Front Plant Sci 11:802. https://doi.org/10.3389/fpls.2020.00802
Lonardi S, Muñoz-Amatriaín M, Liang Q et al (2019) The genome of cowpea (Vigna unguiculata [L.] Walp.). Plant J 98:767–782. https://doi.org/10.1111/tpj.14349
LPWG (2017) A new subfamily classification of the Leguminosae based on a taxonomically comprehensive phylogeny – the Legume Phylogeny Working Group (LPWG). Taxon 66:44–77
Lyons E, Pedersen B, Kane J et al (2008) Finding and comparing syntenic regions among Arabidopsis and the outgroups papaya, poplar, and grape: CoGe with rosids. Plant Physiol 148:1772–1781. https://doi.org/10.1104/pp.108.124867
Lysak MA, Fransz PF, Ali HBM, Schubert I (2001) Chromosome painting in Arabidopsis thaliana. Plant J 28(6):689–697. https://doi.org/10.1046/j.1365-313x.2001.01194.x
Lysak MA, Berr A, Pecinka A, Schmidt R, McBreen K, Schubert I (2006) Mechanisms of chromosome number reduction in Arabidopsis thaliana and related Brassicaceae species. Proc Natl Acad Sci 103:5224–5229. https://doi.org/10.1073/pnas.0510791103
Lysak MA, Mandáková T, Schranz ME (2016) Comparative paleogenomics of crucifers: ancestral genomic blocks revisited. Curr Opin Plant Biol 30:108–115. https://doi.org/10.1016/j.pbi.2016.02.001
Mandáková T, Hlousková P, Koch MA, Lysak MA (2020a) Genome evolution in Arabideae was marked by frequent centromere repositioning. Plant Cell 32:650–665. https://doi.org/10.1105/tpc.19.00557
Mandáková T, Hloušková P, Windham MD, Mitchell-Olds T, Ashby K, Price B, Carman J, Lysak MA (2020b) Chromosomal evolution and apomixis in the cruciferous tribe Boechereae. Front Plant Sci 11:514. https://doi.org/10.3389/fpls.2020.00514
Meng Z, Han J, Lin Y et al (2020) Characterization of a Saccharum spontaneum with a basic chromosome number of x = 10 provides new insights on genome evolution in genus Saccharum. Theor Appl Genet 133:187–199. https://doi.org/10.1007/s00122-019-03450-w
Montenegro C, Vale Martins L, Bustamante FO, Brasileiro-Vidal AC, Pedrosa-Harand A (2022) Comparative cytogenomics reveals genome reshuffling and centromere repositioning in the legume tribe Phaseoleae. Chromosome Res. https://doi.org/10.1007/s10577-022-09702-8
Oliveira ARdaS (2020) Breaks of macrosynteny and collinearity among moth bean (Vigna aconitifolia), cowpea (V. unguiculata), and common bean (Phaseolus vulgaris). Chromosom Res 28:293–306. https://doi.org/10.1007/s10577-020-09635-0
Pedrosa A, Sandal N, Stougaard J, Schweizer D, Bachmair A (2002) Chromosomal map of the model legume Lotus japonicus. Genetics 161:1661–1672. https://doi.org/10.1093/genetics/161.4.1661
Pedrosa-Harand A, Kami J, Gepts P, Geffroy V, Schweizer D (2009) Cytogenetic mapping of common bean chromosomes reveals a less compartmentalized small-genome plant species. Chromosom Res 17:405–417. https://doi.org/10.1007/s10577-009-9031-4
Pienaar BJ, Kok PDF (1991) The Vigna vexillata complex (Fabaceae) in southern Africa. S Afr J Bot 57:236–245. https://doi.org/10.1016/S0254-6299(16)30924-3
Piperidis N, D’Hont A (2020) Sugarcane genome architecture decrypted with chromosome-specific oligo probes. Plant J 103:2039–2051. https://doi.org/10.1111/tpj.14881
Rosin LF, Gil J, Drinnenberg IA, Lei EP (2021) Oligopaint DNA FISH reveals telomere-based meiotic pairing dynamics in the silkworm. Bombyx Mori. PLOS Genet 17(7):e1009700. https://doi.org/10.1371/journal.pgen.1009700
Sakai H, Naito K, Ogiso-Tanaka E et al (2015) The power of single molecule real-time sequencing technology in the de novo assembly of a eukaryotic genome. Sci Rep 5:16780. https://doi.org/10.1038/srep16780
Schmutz J, McClean PE, Mamidi S et al (2014) A reference genome for common bean and genome-wide analysis of dual domestications. Nat Genet 46:707–713. https://doi.org/10.1038/ng.3008
Schranz ME, Lysak MA, Mitchell-Olds T (2006) The ABC’s of comparative genomics in the Brassicaceae: building blocks of crucifer genomes. Trends Plant Sci 11:535–542
Schubert I (2018) What is behind “centromere repositioning”? Chromosoma 127:229–234. https://doi.org/10.1038/ng.3008
She CW, Mao Y, Jiang XH, He CP (2020) Comparative molecular cytogenetic characterization of five wild Vigna species (Fabaceae). Comp Cytogenet 14:243–264. https://doi.org/10.3897/CompCytogen.v14i2.51154
Takahashi Y, Somta P, Muto C, Iseki K, Naito K, Pandiyan M, Natesan S, Tomooka N (2016) Novel genetic resources in the genus Vigna unveiled from gene bank accessions. PLoS ONE 11(1):e0147568. https://doi.org/10.1371/journal.pone.0147568
Vasconcelos EV, Fonsêca AFA, Pedrosa-Harand A et al (2015) Intra- and interchromosomal rearrangements between cowpea [Vigna unguiculata (L.) Walp.] and common bean (Phaseolus vulgaris L.) revealed by BAC-FISH. Chromosome Res 23:253–266. https://doi.org/10.1007/s10577-014-9464-2
Wang J, Sun P, Li Y et al (2017) Hierarchically aligning 10 legume genomes establishes a family-level genomics platform. Plant Physiol 174:284–300. https://doi.org/10.1104/pp.16.01981
Wang K, Cheng H, Han J, Esh A, Liu J, Zhang Y, Wang B (2022) A comprehensive molecular cytogenetic analysis of the genome architecture in modern sugarcane cultivars. Chromosom Res 30:29–41. https://doi.org/10.1007/s10577-021-09680-3
Wanzenbock E-M, Schofer C, Schweizer D, Bachmair A (1997) Ribosomal transcription units integrated via T-DNA transformation associate with the nucleolus and do not require upstream repeat sequences for activity in Arabidopsis thaliana. Plant J 11:1007–1016. https://doi.org/10.1046/j.1365-313X.1997.11051007.x
Xia Q, Pan L, Zhang R et al (2019) The genome assembly of asparagus bean. Vigna Unguiculata Ssp Sesquipedalis Sci Data 6:1–10. https://doi.org/10.1038/s41597-019-0130-6
Xin H, Zhang T, Wu Y, Zhang W, Zhang P, Xi M, Jiang J (2020) An extraordinarily stable karyotype of the woody Populus species revealed by chromosome painting. Plant J 101:253–264. https://doi.org/10.1111/tpj.14894
Yang L, Sagar V (2022) Genome evaluation of cucumber in relation to cucurbit family. In: Pandey S, Weng Y, Behera TK, Bo K (eds) The Cucumber genome. Compendium of Plant Genomes. Springer, Cham, pp 105–119
Yang L, Koo D-H, Li D et al (2014) Next-generation sequencing, FISH mapping and synteny-based modeling reveal mechanisms of decreasing dysploidy in Cucumis. Plant J 77:16–30. https://doi.org/10.1111/tpj.12355
Yu F, Zhao XW, Chai J, Ding X, Li XT, Huang YJ, Wang XH, Wu JY, Zhang MQ, Yang QH, Deng ZH, Jiang JM (2022) Chromosome-specific painting unveils chromosomal fusions and distinct allopolyploid species in the Saccharum complex. New Phytol 233:1953–1965. https://doi.org/10.1111/nph.17905
Zhao H, Zeng Z, Koo DH, Gill BS, Birchler JA, Jiang J (2017) Recurrent establishment of de novo centromeres in the pericentromeric region of maize chromosome 3. Chromosome Res 25:299–311. https://doi.org/10.1007/s10577-017-9564-x
Zhao Q, Meng Y, Wang P et al (2021) Reconstruction of ancestral karyotype illuminates chromosome evolution in the genus Cucumis. Plant J 107:1243–1259. https://doi.org/10.1111/tpj.15381
Zhuang W, Chen H, Yang M et al (2019) The genome of cultivated peanut provides insight into legume karyotypes, polyploid evolution and crop domestication. Nat Genet 51:865–876. https://doi.org/10.1038/s41588-019-0402-2
Acknowledgements
We thank Embrapa Meio-Norte, IPK, and NPGS for supplying the seeds. We also thank CAPES (Coordenação de Aperfeiçoamento de Pessoal de Nível Superior, CNPq (Conselho Nacional de Desenvolvimento Científico e Tecnológico), FACEPE (Fundação de Amparo à Ciência e Tecnologia de Pernambuco), and PAPq/UEMG for scholarships and/or financial support.
Funding
This work was supported by CAPES (Coordenação de Aperfeiçoamento de Pessoal de Nível Superior) Grant no. 88881.189152/2018-01; CNPq (Conselho Nacional de Desenvolvimento Científico e Tecnológico) Grant no. 421968/2018-4, 313527/2017-2, 313944/2020-2, 433931/2018-3, and 442019/2019-0, and FACEPE (Fundação de Amparo à Ciência e Tecnologia do Estado de Pernambuco) Grant no. APQ-0390-2.02/19.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by SD, FdOB, LdVM, VAdC, CM, ARdSO, GSdL, GTB, AFdC, and ACB-V. The first draft of the manuscript was written by SD, FdOB, LdVM, and ACBV, and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consented for publication
Not applicable.
Additional information
Communicated by Reyazul Rouf Mir.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Dias, S., de Oliveira Bustamante, F., do Vale Martins, L. et al. Translocations and inversions: major chromosomal rearrangements during Vigna (Leguminosae) evolution. Theor Appl Genet 137, 29 (2024). https://doi.org/10.1007/s00122-024-04546-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00122-024-04546-8