Abstract
Key message
This study reported validation and fine mapping of a Fusarium crown rot resistant locus on chromosome arm 6HL in barley using near isogenic lines, transcriptome sequences, and a large near isogenic line-derived population.
Abstract
Fusarium crown rot (FCR), caused by Fusarium pseudograminearum, is a chronic and serious disease affecting cereal production in semi-arid regions globally. The increasing prevalence of this disease in recent years is attributed to the widespread adoption of minimum tillage and stubble retention practices. In the study reported here, we generated eight pairs of near isogenic lines (NILs) targeting a putative QTL (Qcrs.caf-6H) conferring FCR resistance in barley. Assessing the NILs confirmed the large effect of this locus. Aimed to develop markers that can be reliably used in incorporating this resistant allele into breeding programs and identify candidate genes, transcriptomic analyses were conducted against three of the NIL pairs and a large NIL-derived population consisting of 1085 F7 recombinant inbred lines generated. By analyzing the transcriptomic data and the fine mapping population, Qcrs.caf-6H was delineated into an interval of 0.9 cM covering a physical distance of ~ 547 kb. Six markers co-segregating with this locus were developed. Based on differential gene expression and SNP variations between the two isolines among the three NIL pairs, candidate genes underlying the resistance at this locus were detected. These results would improve the efficiency of incorporating the targeted locus into barley breeding programs and facilitate the cloning of causal gene(s) responsible for the resistance.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Fusarium crown rot (FCR) is a widespread and damaging disease affecting the production of many crop species including wheat, barley, durum, oat, and triticale in semi-arid regions worldwide. The prevalence of FCR has increased in conservation cropping systems in recent years (Paulitz et al. 2002; Smiley et al. 2005; Saremi et al. 2007; Tunali et al. 2008; Chakraborty et al. 2010; Hameed et al. 2012; Li et al. 2016; Xu et al. 2017) likely due to the tight rotations of cereal crops and the retention of stubble (Simpfendorfer et al. 2019). FCR damage in barley has been overlooked due to its less yield loss compared to wheat (Smiley et al. 2005; Daniel and Simpfendorfer 2012). This is likely due to the fact that whiteheads, a commonly used symptom to evaluate FCR severity in wheat, are rarely present in infected barley plants. Despite this, more severe disease symptoms and higher accumulation of Fusarium mycelia were observed in barley when compared to wheat at similar stages of FCR infection (Liu et al. 2012a). Fusarium mycelia can colonize stubble residue and persist for up to three years or more (Burgess 2014), which increases the risk of FCR infection in the following crop seasons. Therefore, developing barley varieties with enhanced FCR resistance would not only reduce grain yield loss in barley itself but might also benefit subsequent crops by reducing the inoculum load in stubbles.
Understanding the genetics of resistant sources is crucial for breeding programs against FCR (Liu and Ogbonnaya 2015). To achieve this, quantitative trait locus (QTL) mapping has been routinely adopted to detect resistance loci. To date, four putative QTL have been detected in barley and their effects were consistently expressed in different genetic backgrounds. They were located on chromosome arms 1HL (Chen et al. 2013b), 3HL (Li et al. 2009), 4HL (Chen et al. 2013a), and 6HL (Gao et al. 2019a; Gao 2020), respectively. However, morphological traits such as plant height and heading date have been found to have a passive contribution to FCR resistance (Liu et al. 2012a, 2012b; Chen et al. 2013b; Bai and Liu 2015). When the effects of these morphological traits were removed by covariance analysis, both LOD values and magnitudes of the QTL were reduced (Chen et al. 2013a; Zheng et al. 2014). As a result, any loci identified from QTL studies must be treated as putative. To accurately assess the effects of a locus and to develop reliable markers targeting an FCR locus, segregation of those undesired characteristics needs to be fixed.
Development of near isogenic lines (NILs) has been considered one of the most effective approaches to achieve fixed genetic backgrounds and has been routinely used to validate FCR resistant loci in wheat and barley by our team (Ma et al. 2012; Habib et al. 2016; Gao et al. 2020; Gao 2020). The difference between the two isolines of a NIL pair is primarily limited in the targeted region, facilitating further investigation on elucidating potential mechanisms via transcriptomic analyses. RNA sequencing (RNA-seq) has become an indispensable tool for transcriptomic profiling (Wang et al. 2009; Stark et al. 2019) and has been employed to understand the mechanisms of various diseases. Compared to traditional studies on genetically distinct genotypes which differ significantly in their genetic backgrounds, analyzing NILs offer huge advantages including minimization of genetic background interference and increased sensitivity and accuracy of transcriptional analyses (Keurentjes et al. 2007). Moreover, populations derived from the NILs primarily segregating at a targeted locus can also convert a quantitative trait into a Mendelian factor facilitating the accurate localization of a locus (Zheng et al. 2015; Jiang et al. 2019; Gao et al. 2020).
In the study reported here, we aimed to validate the effect of the putative QTL (Qcrs.caf-6H) conferring FCR resistance on chromosome arm 6HL in barley (Gao et al. 2019a, b) and explore its underlying mechanism. A few pairs of NILs were developed to minimize genetic background interference and improve the accuracy of the transcriptional analysis. Transcriptomic sequences were obtained from three of these NIL pairs and used to identify single nucleotide polymorphism (SNP) variations and candidate genes. To develop markers tightly linked with this locus, a large NIL-derived population consisting of 1085 F7 lines was then generated and assessed. Taken together, markers co-segregating with this locus were generated and a small number of candidate genes underlying Qcrs.caf-6H were identified in the present study.
Materials and methods
Plant materials
Two segregating populations generated by crossing a resistant donor AWCS799 and two Australian varieties Fleet and Franklin were used in this study (Gao et al. 2019a). AWCS799 was identified from a screening of 1,047 genotypes representing different geographical origins and plant types (Liu et al. 2012b). Combined with a fast generation procedure (Zheng et al. 2013), the method of heterogeneous inbred family (HIF) (Tuinstra et al. 1997) was used to develop NILs in glasshouses at Queensland Bioscience Precinct (QBP), Brisbane. An SSR marker, 6H_497772849 (forward primer GCATTAGTTGTCATAGTAGGTAGCA and reverse primer TTCAAGACCACGACCTTGGG) which was closely linked with Qcrs.caf-6H (Gao et al. 2019a), was used to identify heterozygous plants from each of the two populations. Heterozygous plants identified were self-pollinated, and ten plants derived from each of the heterozygous plants were used for the next round of selection. This process of selecting heterozygous individuals and self-pollination was repeated until F8 generation. Two isolines, one with the resistant allele and the other with the susceptible allele, were then isolated from each of the F8 heterozygous plants and were treated as a pair of putative NIL. Seeds from these putative NIL pairs were then increased at Gatton Research Station for further assessments.
A NIL-derived population consisting of 1085 lines was generated as described above from four different heterozygous F6 plants obtained in generating the 6HL_NIL1 which showed the biggest difference on disease index between R and S isolines. A single-seed-descent approach was used to process this population to F7 generation. Seeds from each of the lines were then increased in glasshouses at QBP.
Assessment of FCR resistance
Previous studies indicated that FCR resistance is unlikely to be species-specific (Van Eeuwijk et al. 1995; Chakraborty et al. 2010; Li et al. 2010b; Ma et al. 2010). Therefore, one highly aggressive isolate of F. pseudograminearum (Fp) CS3096 (Gao 2020) was used to assess FCR resistance in this study. Protocols for inoculum preparation, FCR inoculation, and disease severity measurement were conducted following the methods described by Li et al. (2008). In brief, a piece of infected plant fragment was placed on plates containing 1/2 strength potato dextrose agar (PDA). Inoculated plates were incubated at room temperature for 14 days, and the mycelium was then removed. The plates were further incubated for another 14 days under a combination of cool white and black fluorescent lights with a 12-h photoperiod. Spores were harvested, and the concentration of spore suspension was adjusted to 1 × 106 spores/ml. The spore suspension was stored in a – 20 °C freezer, and Tween 20 (0.1%v/v) was added before use.
Seeds were germinated on two layers of filter paper saturated with water in Petri dishes. The Petri dishes were chilled in a cold room at 4 °C for two days for uniform germination and then, were left at room temperature overnight before transferring into potting mix. The seedlings with similar lengths (~ 0.5 cm) were immersed in the spore suspension for 2 min. Two seedlings were planted into each square punnet (80 cm3) of a 56-well tray (Rite Grow Kwik Pots, Garden City Plastics, Australia) containing sterilized University of California mix C (50% sand and 50% peat v/v). The trays were arranged in a randomized block design and placed in a controlled environment facility (CEF) at QBP. The CEF was set at 25/18 (± 1) °C Day/night temperature, 65/80% (± 5) % day/night relative humidity, and a 14 h photoperiod with 500 µmol m−2 S−1 photon flux density at the level of the plant canopy (Gao 2020). As drought stress would promote FCR development (Liu and Liu 2016), the inoculated seedlings were watered during the assessment only when wilt symptoms were detected.
Four independent trials were conducted to assess FCR resistance against the putative NILs and a subpopulation of 96 NIL-derived lines, while six trials were performed against recombinants identified from the whole NIL-derived population of 1,085 lines. The resistant and susceptible isolines of the NIL_CR6HL_1 were used as positive and negative controls in each of the trials. Each trial contained three replicates and each replicate consisted of 14 seedlings. The flanking markers of the Qcrs.caf-6H locus were determined using the subpopulation and used to identify recombinant lines in the whole NIL-derived population. FCR severity was evaluated four weeks after inoculation using a 0–5 scale, where ‘0’ representing no symptom and ‘5’ standing for whole plant necrotic (Li et al. 2008). Disease indices (DI) were calculated for each line following the formula of DI = (ΣnX /5 N) × 100, where X is the scale value of each plant, n is the number of plants in the category, and N is the total number of plants assessed for each line.
Statistical analysis
Statistical analyses were performed using the SPSS statistics 19.0 for Windows statistical software package (SPSS Inc., Chicago, IL). For each trial, the following mixed-effect model was used: Yij = μ + ri + gj + wij, where: Yij = trait value on the jth genotype in the ith replication; μ = general mean; ri = effect due to ith replication; wij is the error or genotype by replication interaction, where genotype was treated as a fixed effect and that of replicate as random. The Tukey’s HSD test of one-way ANOVA was employed to detect possible differences among the means. The disease ratings from all seedlings for each line of the fine mapping population in each trial were averaged and used to determine whether the line in concern was resistant (< 2.5) or susceptible (> 2.5) to FCR infection (Gao 2020).
RNA extraction and sequencing
Samples for RNA sequencing were collected from three pairs of the NILs (namely 6HL_NIL1, 6HL_NIL2, and 6HL_NIL3). Inoculation was conducted with either the F. pseudograminearum isolate (Fp-inoculation) or distill water (mock) following the protocol described by Gao et al. (2020). Three biological replications were conducted for each of the isolines and each replication contained 14 seedlings. Tissues for RNA extraction were obtained by cutting the shoot bases (2 cm) of 14 seedlings at 4 days post-inoculation (dpi) and snap-frozen in liquid nitrogen and kept at − 80 °C until processed. The time point for sampling was based on previous studies (Habib et al. 2018; Gao et al. 2019b; Gao 2020).
A total of 36 samples were obtained from the six isolines. Samples were crushed into fine powder in a 1.5 μL micro-centrifuge tube and total RNA was extracted using a RNeasy plant mini kit (Qiagen, Hilden, Germany) according to manufacturer’s instructions (including DNase-I digestion). The yield and purity of RNA samples were measured using a Nanodrop-1000 Spectrophotometer. The integrity of all RNA samples was assessed by running the total RNA on 1% agarose gels. RNA sequencing was carried out by the Australian Genome Research Facility Ltd. (Parkville, Victoria, Australia), and 100-bp paired-end reads were produced using the Illumina Hiseq-2000. All 36 RNA-seq libraries were run across four lanes of a HiSeq2000. The RNA sequencing files have been made available at the National Center for Biotechnology Information (NCBI) with the accession number PRJNA922199.
Transcriptomic analysis
Commands used for trimming raw data and analyzing trimmed reads were described by Habib et al. (2018). FastQC (version 0.11.2) was used as a preliminary check for PHRED scores. Raw reads were trimmed using the SolexaQA package (version 2.0.13) with a minimum PHRED quality value of 30 and a minimum final read length of 70 bp. The filtered reads were then mapped to the reference genome of Morex (https://webblast.ipk-gatersleben.de/barley_ibsc/downloads/:150831_barley_pseudomolecules) using BWA (version 0.7.12) (Li and Durbin 2009).
Analysis of differential gene expression
Cufflinks v2.0.2 (Roberts et al. 2011) was used to assemble the mapped reads. Differentially expressed genes (DEGs) were identified with Cuffdiff from the Cufflinks tool package with high-confidence genes annotated from the ‘Morex’ reference. Fragments per kilobase of exon per million mapped reads (FPKM) were applied for each transcript to represent the normalized expression value. The fold change was calculated according to the equation: Fold Change = log2 (FPKMA/FPKMB).
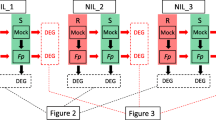
Pairwise comparisons between different treatments for the same genotype and between genotypes under Fp-inoculation and mock-inoculation were conducted (Fig. S1). These are summarized throughout the paper in the following way SM_v_SI, RM_v_RI, SI_v_RI and SM_v_RM, where ‘M’ for ‘Mock’; ‘I’ for Fp-inoculation; ‘S’ for the susceptible isoline; and ‘R’ for the resistant isoline. DEGs were determined with the adjusted p-value threshold of ≤ 0.05 and log2 expression fold change of ≥ 1 or ≤ -1 or ‘inf’ (where the FPKM value in one condition is zero, and the other is not). Venny 2.0 was used for Venn diagram analysis (Oliveros 2007).
SNP calling and marker development
A graphical overview of the experimental design for SNP calling has been provided (Fig. S1). All six sequence files (three biological replicates by two treatments) for each genotype were concatenated after removing low-quality sequences. The files were then aligned to the Morex genome using BWA align. Duplicates were removed by SAMtools rmdup and SNPs were identified using SAMtools mpileup by skipping alignments with mapQ smaller than 20, then SNPs with DP < 4 and SNPs within 3 base pairs of an indel were filtered by SAMtools/BCFtools (version 1.5) (Li 2011). The variant effect of SNPs in candidate genes was evaluated with snpEff (Version 5.1) (Cingolani et al. 2012). Kompetitive allele specific PCR (KASP) markers were then developed based on the SNPs detected. All the primers were designed using the Primer-BLAST (Ye et al. 2012). Linkage analysis was carried out using the computer package JoinMap 4.0 (Van Ooijen 2006).
DNA extraction and genotyping
Leaf tissue from each line of the NIL-derived population was collected and vacuum dried for DNA extraction using the CTAB protocol (Porebski et al. 1997). KASP assay was conducted using 384-well set on the Vii 7 Real-Time PCR system (Applied Biosystems, Foster City, California, USA). One KASP reaction volume consisted of a total volume of 5 μl including 2.0 μl DNA (20 ng/μl), 0.06 μl primer mixture, and 3 μl of KASP Master Mix. On each SNP reaction plate, at least one water sample was included as a no-template control. The PCR profile started with an initial denaturation step at 94 °C for 15 min, continued with 10 touch-down PCR cycles at 94 °C for 20 s, and 60 °C for 1 min with − 0.5 °C/cycle, and then 30 cycles at 94 °C for 20 s, and 55 °C for 1 min. Three replicates for each genotype were performed.
Validation of differentially expressed genes
Expression assessments of five candidate genes in the targeted interval (primer sequences were listed in Table S1) selected from transcriptomic analyses were conducted using quantitative real-time PCR (qRT-PCR) with the actin protein gene as the internal housekeeping reference (forward primer: 5′-GCCGTGCTTTCCCTCTATG-3′; reverse primer 5′-GCTTCTCCTTGATGTCCCTTA-3′). Inoculation, tissue sampling, and RNA extraction were carried out using the aforementioned methods. Three biological replicates, each with two technical replications, were used for each genotype-treatment sample per isoline.
The procedures for synthesizing cDNA and qRT-PCR were described by Gao et al. (2019a, b). The relative fold changes were calculated using the comparative CT method (2−ΔΔCT). Average values of the two technical replications were used to represent the biological replicate for each of the samples.
Results
Near isogenic lines targeting the Qcrs.caf-6H locus
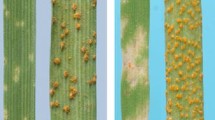
Eleven pairs of putative NILs were initially developed to validate the effect of the 6HL resistant locus based on the marker profiles of 6H_497772849. Among them, eight pairs showed significant differences in disease severity between the two isolines (Fig. 1) with no clear morphologic differences. The resistant isolines performed better on FCR resistance with an average DI value of 34.2 compared with the susceptible isolines with an average DI value of 50.2 (Table 1).
Plants of a pair of the near isogenic lines (NIL_CR6HL_1R and NIL_CR6HL_1S) showing similar morphology of the non-inoculated plants (left) and the difference in resistant to Fusarium pseudograminearum at 28 days post-inoculation (right). R represents lines with the resistant allele, and S represents lines without the resistant allele
DEGs in the Qcrs.caf-6H region
Three of the eight NIL pairs with the largest differences in FCR severity were selected and used to generate RNA-seq. To analyze host response to FCR infection, DEGs were detected between Fp- and mock-inoculated samples of the same isolines. The numbers of up-regulated genes were significantly higher than those of down-regulated ones following Fp-inoculation (Table S2). Of the up-regulated genes, 88 were shared by all the three ‘R’ isolines while 101 were shared by the three ‘S’ isolines (Fig. 2). Of the down-regulated genes, only seven were shared among the three ‘R’ isolines and three among the ‘S’ isolines (Fig. 2).
DEGs for each of the 6HL_NIL pairs following Fp- and mock-inoculation (RM_vs_RI) and SM_vs_SI). Venn diagrams in upper panel show the numbers of up-regulated DEGs in each ‘R’ (left) and ‘S’ (right) isolines. Venn diagrams in lower panel show the numbers of down-regulated DEGs in each ‘R’ (left) and ‘S’ (right) isolines. DEGs were determined with the threshold of FDR ≤ 0.05 and |log2 fold change|≥ 1 or ‘inf’ (one of the comparative objects did not express and the other did)
To assess transcriptomic responses to FCR infection mediated by Qcrs.caf-6H, DEGs between the ‘R’ and ‘S’ isolines were also compared. These comparisons detected a total of 1060 up-regulated genes and 786 down-regulated ones from the Fp-treatment (Table S2). Only five of the up-regulated genes and four of the down-regulated ones were shared by all three NIL pairs, respectively (Fig. 3). Of the DEGs identified from the mock-inoculated samples, there were 1294 up-regulated genes which were more than twice of the down-regulated ones (Table S2). Among them, 36 up- and four down-regulated genes overlapped across all three comparisons (Fig. 3).
DEGs between ‘R’ and ‘S’ isolines under Fp- (RI_vs_SI) or mock-inoculation (RM_vs_SM). Venn diagrams show the numbers of DEGs up-regulated in ‘R’ (left) or ‘S’ (right) isolines under Fp- (up) or mock-inoculation (down). DEGs were determined with the threshold of FDR ≤ 0.05 and |log2 fold change|≥ 1 or ‘inf’ (one of the comparative objects did not express and the other did)
SNPs between the ‘R’ and ‘S’ isolines shared with three 6H_NIL pairs
A total of 37,972 homozygous SNPs were detected between isolines for the three NIL pairs. The number of SNPs detected in the NIL_CR6HL_3 was more than twice compared to those detected from either of the other two NIL pairs (Fig. S2). This number was dramatically reduced to 278 when comparing the R and S isolines among three NIL pairs (Fig. S2) and all of them fell into the distal end of the chromosome arm 6HL obtaining the Qcrs.caf-6H locus spanning a physical interval of ~ 22 Mbp (Fig. S3a). Based on the reference genome of barley cv. Morex, 142 high-confident (HC) genes and 150 low-confident (LC) genes were identified in the common interval across three 6HL NIL pairs. Among these HC genes, 30 contained SNP variants shared by all three NIL pairs and eight were differentially expressed between the isolines for at least one of the NIL pairs (Fig. S3b).
Chromosomal interval containing Qcrs.caf-6H based on the subpopulation
Four polymorphic markers in the targeted interval were developed based on sequence differences between the resistant and susceptible isolines. Together with 6H_497772849, the five markers were used to screen the sub-populations consisting of 96 lines (Fig. 4a). Linkage analysis showed that they spanned a genetic distance of 8.3 cM. Disease ratings of this subpopulation fell into two classes: one rating from 18.4 to 37.2 (resistant) and the other from 54.9 to 78.6 (susceptible). There were no intermediate ratings for any of the lines assessed (Fig. S4). After analyzing the marker profiles and the phenotypic data, Qcrs.caf-6H was mapped into a 3.8 cM interval flanked by CS6HLCR-03 and CS6HLCR-04 (Fig. 4a).
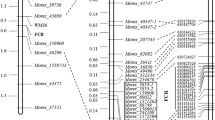
Genetic and physical maps surrounding the Fusarium crown rot resistant locus Qcrs.caf-6H in barley. a The targeted interval based on the assessment of a subpopulation consisting of 96 NIL-derived lines; b The high-density linkage map around the targeted locus based on the analysis of 1,085 NIL-derived lines. Markers co-segregating with the FCR locus were placed in the box; c physical locations of marker on the 6H pseudomolecule and high-confident genes detected in the targeted interval. DEGs detected between ‘R’ and ‘S’ isolines for at least one of the NIL pairs in the region were highlighted in bold
Fine mapping of Qcrs.caf-6H locus using the whole NIL-derived population
A total of 32 recombinants were identified among the whole fine mapping population of 1,085 lines using the two flanking markers. These recombinants were categorized into either resistant or susceptible groups, with a significant difference between the two groups (P < 0.01). Of the 32 recombinants, 15 lines were classified as resistant, while the remaining 17 lines were susceptible (Fig. 5).
Genotypes and phenotypes of the haplotypes identified from the recombinant lines based on markers surrounding the Qcrs.caf-6H locus. The solid regions represent alleles from the resistant parent and the empty ones for the susceptible parent. Phenotypic values (mean ± SE) were calculated based on the disease severity from all trials. The numbers on the chromosome stand for genetic distance (cM)
To further narrow down the candidate region, eleven KASP markers were newly generated based on the SNP calling results and were found to be polymorphic between the two isolines. Genotyping the recombinants revealed that six of the eleven KASP markers co-segregated with the Qcrs.caf-6H locus (Table 2), mapping the locus into a 0.9 cM interval between CS6HLCR-07 and CS6HLCR-13 (Fig. 4b). These markers were found to be highly consistent with their relative physical positions around the targeted region (Fig. 4b, c), with the Qcrs.caf-6H locus falling into a ~ 547 kb interval between 472.58 and 473.13 Mbp on the reference genome of barley cv. Morex (Fig. 4c).
Identification of candidate genes in the targeted region
Ten HC and eight LC genes were identified in the refined interval (Fig. 4c, Table S3). Because the LC genes lacked clear functional annotations, they were not considered in further analysis. Among the HC genes, five of them contain SNP variants between resistant and susceptible isolines shared by all the three NIL pairs (Fig. 4c, Table S4). Expression patterns consistent with the RNA-seq analysis were obtained in the qRT-PCR analysis for each of the five genes assessed (Table S1). Of the five genes, it is of interest to note that only HORVU.MOREX. r2.6HG0503730, coding a U-box domain-containing family protein, harbors three missense variants leading to amino acid changes (Table S4). Moreover, this gene was Fp-induced down-regulated in two of the three analyzed susceptible isolines. These results indicate that the U-box protein coding gene is likely to be a strong candidate underlying Qcrs.caf-6H locus.
Discussion
FCR is a devastating threat to cereal production worldwide, particularly in semi-arid regions. In efforts to address this challenge, breeding varieties with boosted resistance are critical. Working toward this, sources of FCR resistance have been identified and putative QTL conferring resistance was detected. In this study, we aimed to develop markers that can be reliably used to tag the 6HL resistant locus for further incorporating this locus into breeding programs. Resources including eight pairs of NILs, RNA sequences targeting three of the NIL pairs, and a large NIL-derived population were generated and assessed. Using markers developed from the RNA-seq analysis, Qcrs.caf-6H was successfully placed into a 0.9 cM interval covering a physical distance of approximately 547 kb. A candidate gene coding the U-box domain-containing family protein was identified harboring three missense variants between the two isolines, which would be the primary target in further investigations on gene cloning.
In this study, eleven pairs of NILs were initially generated. However, no significant difference in FCR resistance was detected between the two isolines for three of the eleven pairs of putative NILs. Different from the method of using flanking markers for the targeted locus (Pumphrey et al. 2007), we generated NILs using only one linked marker which should reduce the sizes of chromosomal segments differentiating the isolines (Ma et al. 2012; Habib et al. 2016; Gao et al. 2020). However, markers obtained from QTL mapping studies may not be tightly linked with a targeted locus due to the limited resolution (Paterson et al. 1988). It is clear recombination between the linked and its target may occur resulting in false NILs. In the present study, significant variation was also found in the numbers of SNPs and DEGs detected among the three pairs of NILs assessed as different NIL pairs were expected to have different genetic backgrounds which potentially caused such variation and would lead to the difference in FCR development at any given time point. While individual NIL pair contribute large sets of DEGs and SNPs, analysis of multiple NIL pairs could detect conserved SNP differences. This approach, which would not only be effective in narrowing the candidate regions and reducing the number of candidate genes but also uncover genetic markers for fine mapping the targeted loci, has been routinely used for FCR studies in our team (Ma et al. 2014; Habib et al. 2018; Jiang et al. 2019; Gao et al. 2019b, 2020).
Previous studies reported the accuracy of FCR assessment can be affected by several morphologic characteristics including plant height and heading date (Liu et al. 2012a, 2012b; Chen et al. 2013b). Thus, a large NIL-derived population was subsequently developed to minimize the interference from the segregations of these non-targeted characteristics. The individuals in this population mainly differed in the targeted region, allowing precise categorization of FCR severity into either resistant or susceptible. As a result, Qcrs.caf-6H was placed into a refined region containing ten HC genes (Fig. 4). However, none of these genes is a classic NBS-LRR (nucleotide-binding sites and leucine-rich-repeat) resistant gene. Previous studies showed that non-classical NBS-LRR genes confer resistance to Fusarium head blight (FHB), another disease caused by Fusarium pathogens (Moore et al. 2015; Ke et al. 2017; Milne et al. 2019). Similarly, host resistance to FCR and FHB is not pathogen species-specific (Fv et al. 1995; Chakraborty et al. 2010; Li et al. 2010b; Ma et al. 2010). Thus, it is likely that non-classical NBS-LRR genes may also be responsible for FCR resistance.
Of the ten HC genes located within the targeted interval, four were differentially expressed bearing SNP variations between the two isolines among all the three NIL pairs. They were related to either or both biotic and abiotic stresses. Among them, HORVU.MOREX. r2.6HG0503730 coding a U-box domain-containing family protein is the only one containing missense variants, which would form our primary target. The functions of this gene in biotic and abiotic stress are well known (Yan et al. 2003; Zeng et al. 2004) including its role in pathogen recognition and plant immune response to broad-spectrum disease resistance to various pathogens (González-Lamothe et al. 2006; Cheng and Li 2012; Li et al. 2012). Although synonymous variants do not alter the encoded amino acids, they can still alter mRNA structure leading to changes in the rate or efficiency of translation (Hunt et al. 2014) and/or protein production (Komar 2016) and abundance (Maier et al. 2009), which in turn impact fitness (Lebeuf-Taylor et al. 2019; Bailey et al. 2021). Thus, the other three DEGs in the candidate region cannot be ruled out. The second gene HORVU.MOREX. r2.6HG0503700 encodes a basic helix-loop-helix (bHLH) transcription factor which has been functionally characterized in many plant species to contribute to immunity against various biotic and abiotic stresses (Heim et al. 2003; Li et al. 2010a). Previous studies showed that bHLH functions as a transcriptional activator associated with NBS-LRR genes to activate defense responses (Xu et al. 2014). The third gene HORVU.MOREX. r2.6HG0503660 encodes an Alpha/Beta hydrolase fold protein, which has been reported associated with defense against pathogens (Wäspi et al. 1998) and abiotic stress such as salinity and drought (Liu et al. 2014; Akbudak et al. 2019). In addition, it has been reported that the severity of FCR can be strongly affected by drought stress (Liu and Liu 2016). Thus, the fourth gene HORVU.MOREX. r2.6HG0503840 encoding ATPase family AAA domain-containing protein 3 relevant to abiotic stresses (Kim et al. 2021) should be considered as another candidate for FCR resistance at this locus.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author upon reasonable request.
References
Akbudak MA, Filiz E, Uylas S (2019) Identification of O-acetylserine (thiol) lyase (OASTL) genes in sorghum (Sorghum bicolor) and gene expression analysis under cadmium stress. Mol Biol Rep 46:343–354. https://doi.org/10.1007/s11033-018-4477-0
Bai Z, Liu CJ (2015) Histological evidence for different spread of fusarium crown rot in barley genotypes with different heights. J Phytopathol 163:91–97. https://doi.org/10.1111/jph.12283
Bailey SF, Alonso Morales LA, Kassen R (2021) Effects of synonymous mutations beyond codon bias: the evidence for adaptive synonymous substitutions from microbial evolution experiments. Genome Biol Evol 13(9):evab141. https://doi.org/10.1093/gbe/evab141
Blencowe BJ, Ahmad S, Lee LJ (2009) Current-generation high-throughput sequencing: deepening insights into mammalian transcriptomes. Genes Dev 23:1379–1386. https://doi.org/10.1101/gad.1788009
Burgess LW (2014) 2011 McAlpine memorial lecture: a love affair with Fusarium. Australas Plant Pathol 43(4):359–368. https://doi.org/10.1007/s13313-013-0261-8
Chakraborty S, Obanor F, Westecott R, Abeywickrama K (2010) Wheat crown rot pathogens Fusarium graminearum and F. pseudograminearum lack specialization. Phytopathology 100:1057–1065. https://doi.org/10.1094/PHYTO-01-10-0007
Chen G, Liu Y, Ma J, Zheng Z, Wei Y, McIntyre CL, Zheng YL, Liu CJ (2013a) A novel and major quantitative trait locus for Fusarium crown rot resistance in a genotype of wild barley (Hordeum spontaneum L.). PLoS ONE 8:e58040. https://doi.org/10.1371/journal.pone.0058040
Chen G, Liu Y, Wei Y, McIntyre C, Zhou M, Zheng YL, Liu CJ (2013b) Major QTL for Fusarium crown rot resistance in a barley landrace. Theor Appl Genet 126:2511–2520. https://doi.org/10.1007/s00122-013-2151-4
Cheng YT, Li X (2012) Ubiquitination in NB-LRR-mediated immunity. Curr Opin Plant Biol 15:392–399. https://doi.org/10.1016/j.pbi.2012.03.014
Cingolani P, Platts A, le Wang L, Coon M, Nguyen T, Wang L, Land SJ, Lu X, Ruden DM (2012) A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly (austin) 6(2):80–92. https://doi.org/10.4161/fly.19695
Daniel R, Simpfendorfer S (2012) The impact of crown rot on winter cereal yield. In: Working towards reducing crown rot impact on the Australian Grains Industry. Proceedings of the first international crown rot workshop for wheat improvement. Narrabri, Australia, p 20
Fv VE, Mesterhazy A, Kling CI et al (1995) Assessing non-specificity of resistance in wheat to head blight caused by inoculation with European strains of Fusarium culmorum, F. graminearum and F. nivale using a multiplicative model for interaction. Theor Appl Genet 90:221–228. https://doi.org/10.1007/BF00222205
Gao S, Zheng Z, Hu H, Shi H, Ma J, Liu Y, Wei Y, Zheng YL, Zhou M, Liu CJ (2019a) A novel QTL conferring Fusarium crown rot resistance located on chromosome arm 6HL in barley. Front Plant Sci 10:1026. https://doi.org/10.3389/fpls.2019.01206
Gao S, Zheng Z, Powell J et al (2019b) Validation and delineation of a locus conferring Fusarium crown rot resistance on 1HL in barley by analysing transcriptomes from multiple pairs of near isogenic lines. BMC Genom 20:650. https://doi.org/10.1186/s12864-019-6011-8
Gao S, Zheng Z, Hu H et al (2020) Delineating a locus conferring Fusarium crown rot resistance on chromosome arm 1HL in barley by developing and analysing a large population derived from near isogenic lines. Crop J 8(6):1002–1010. https://doi.org/10.1016/j.cj.2020.03.008
Gao S (2020) Genetics of resistance to Fusarium crown rot in barley and pan-genome sequence anchors of this species. Doctoral dissertation, University of Tasmania. https://doi.org/10.25959/100.00034975
González-Lamothe R, Tsitsigiannis DI, Ludwig AA, Panicot M, Shirasu K, Jones JD (2006) The U-box protein CMPG1 is required for efficient activation of defense mechanisms triggered by multiple resistance genes in tobacco and tomato. Plant Cell 18:1067–1083. https://doi.org/10.1105/tpc.106.040998
Habib A, Powell JJ, Stiller J, Liu M, Shabala S, Zhou M, Gardiner DM, Liu CJ (2018) A multiple near isogenic line (multi-NIL) RNA-seq approach to identify candidate genes underpinning QTL. Theor Appl Genet 131:613–624. https://doi.org/10.1007/s00122-017-3023-0
Habib A, Shabala S, Shabala L, Zhou M, Liu C (2016) Near-isogenic lines developed for a major QTL on chromosome arm 4HL conferring Fusarium crown rot resistance in barley. Euphytica 209:555–563. https://doi.org/10.1007/s10681-015-1623-9
Hameed M, Rana R, Ali Z (2012) Identification and characterization of a novel Iraqi isolate of Fusarium pseudograminearum causing crown rot in wheat. Genet Mol Res 11:1341–1348
Heim MA, Jakoby M, Werber M, Martin C, Weisshaar B, Bailey PC (2003) The basic helix–loop–helix transcription factor family in plants: a genome-wide study of protein structure and functional diversity. Mol Biol Evol 20:735–747. https://doi.org/10.1093/molbev/msg088
Hunt RC, Simhadri VL, Iandoli M, Sauna ZE, Kimchi-Sarfaty C (2014) Exposing synonymous mutations. Trends Genet 30:308–321. https://doi.org/10.1016/j.tig.2014.04.006
Jiang Y, Habib A, Zheng Z, Zhou M, Wei Y, Zheng YL, Liu C (2019) Development of tightly linked markers and identification of candidate genes for Fusarium crown rot resistance in barley by exploiting a near-isogenic line-derived population. Theor Appl Genet 132:217–225. https://doi.org/10.1007/s00122-018-3209-0
Ke Y, Deng H, Wang S (2017) Advances in understanding broad-spectrum resistance to pathogens in rice. Plant J 90(4):738–748. https://doi.org/10.1111/tpj.13438
Keurentjes JJ, Bentsink L, Alonso-Blanco C, Hanhart CJ, Blankestijn-De Vries H, Effgen S, Vreugdenhil D, Koornneef M (2007) Development of a near-isogenic line population of Arabidopsis thaliana and comparison of mapping power with a recombinant inbred line population. Genetics 175:891–905. https://doi.org/10.1534/genetics.106.066423
Kim M, Schulz V, Brings L, Schoeller T, Kühn K, Vierling E (2021) mTERF18 and ATAD3 are required for mitochondrial nucleoid structure and their disruption confers heat tolerance in Arabidopsis thaliana. New Phytol 232:2026–2042. https://doi.org/10.1111/nph.17717
Komar AA (2016) The Yin and Yang of codon usage. Hum Mol Genet 25:77–85. https://doi.org/10.1093/hmg/ddw207
Lebeuf-Taylor E, McCloskey N, Bailey SF, Hinz A, Kassen R (2019) The distribution of fitness effects among synonymous mutations in a gene under directional selection. Elife 19(8):e45952. https://doi.org/10.7554/eLife.45952
Li F, Guo S, Zhao Y, Chen D, Chong K, Xu Y (2010a) Overexpression of a homopeptide repeat-containing bHLH protein gene (OrbHLH001) from Dongxiang Wild Rice confers freezing and salt tolerance in transgenic Arabidopsis. Plant Cell Rep 29:977–986. https://doi.org/10.1007/s00299-010-0883-z
Li H (2011) A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27:2987–2993. https://doi.org/10.1093/bioinformatics/btr509
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25:1754–1760. https://doi.org/10.1093/bioinformatics/btp324
Li H, He X, Ding S, Yuan H, Chen L (2016) First report of Fusarium culmorum causing crown rot of wheat in China. Plant Dis 100:2532–2532. https://doi.org/10.1094/PDIS-05-16-0723-PDN
Li HB, Xie GQ, Ma J, Liu GR, Wen SM, Ban T, Chakraborty S, Liu CJ (2010b) Genetic relationships between resistances to Fusarium head blight and crown rot in bread wheat (Triticum aestivum L.). Theor Appl Genet 121:941–950. https://doi.org/10.1007/s00122-010-1363-0
Li HB, Zhou M, Liu CJ (2009) A major QTL conferring crown rot resistance in barley and its association with plant height. Theor Appl Genet 118:903–910. https://doi.org/10.1007/s00122-008-0948-3
Li W, Ahn I-P, Ning Y, Park C-H, Zeng L, Whitehill JG, Lu H, Zhao Q, Ding B, Xie Q (2012) The U-Box/ARM E3 ligase PUB13 regulates cell death, defense, and flowering time in Arabidopsis. Plant Physiol 159:239–250. https://doi.org/10.1104/pp.111.192617
Li X, Liu C, Chakraborty S, Manners JM, Kazan K (2008) A simple method for the assessment of crown rot disease severity in wheat seedlings inoculated with Fusarium pseudograminearum. J Phytopathol 156:751–754. https://doi.org/10.1111/j.1439-0434.2008.01425.x
Liu C, Ogbonnaya FC (2015) Resistance to Fusarium crown rot in wheat and barley: a review. Plant Breed 134:365–372. https://doi.org/10.1111/pbr.12274
Liu D, Wang L, Zhai H, Song X, He S, Liu Q (2014) A novel α/β-hydrolase gene IbMas enhances salt tolerance in transgenic sweetpotato. PLoS ONE 9:e115128. https://doi.org/10.1371/journla.pone.0115128
Liu X, Liu C (2016) Effects of drought-stress on Fusarium crown rot development in Barley. PLoS ONE 11:e0167304. https://doi.org/10.1371/journal.pone.0167304
Liu Y, Ma J, Yan W, Yan G, Zhou M, Wei Y, Zheng Y, Liu C (2012a) Different tolerance in bread wheat, durum wheat and barley to Fusarium crown rot disease caused by Fusarium pseudograminearum. J Phytopathol 160:412–417. https://doi.org/10.1111/j.1439-0434.2012.01920.x
Liu Y, Zheng YL, Wei Y, Zhou M, Liu C (2012b) Genotypic differences to crown rot caused by Fusarium pseudograminearum in barley (Hordeum vulgare L.). Plant Breed 131:728–732. https://doi.org/10.1111/j.1439-0523.2012.02008.x
Ma J, Li HB, Zhang CY, Yang XM, Liu YX, Yan GJ, Liu CJ (2010) Identification and validation of a major QTL conferring crown rot resistance in hexaploid wheat. Theor Appl Genet 120:1119–1128. https://doi.org/10.1007/s00122-009-1239-3
Ma J, Stiller J, Zhao Q, Feng Q, Cavanagh C, Wang P, Gardiner D, Choulet F, Feuillet C, Zheng YL (2014) Transcriptome and allele specificity associated with a 3BL locus for Fusarium crown rot resistance in bread wheat. PLoS ONE 9:e113309. https://doi.org/10.1371/journal.pone.0113309
Ma J, Yan GJ, Liu CJ (2012) Development of near-isogenic lines for a major QTL on 3BL conferring Fusarium crown rot resistance in hexaploid wheat. Euphytica 183:147–152. https://doi.org/10.1007/s10681-011-0414-1
Maier T, Güell M, Serrano L (2009) Correlation of mRNA and protein in complex biological samples. FEBS Lett 583:3966–3973. https://doi.org/10.1016/j.febslet.2009.10.036
Milne RJ, Dibley KE, Schnippenkoetter W, Mascher M, Lui AC, Wang L, Lo C, Ashton AR, Ryan PR, Lagudah ES (2019) The wheat Lr67 gene from the sugar transport protein 13 family confers multipathogen resistance in barley. Plant Physiol 179:1285–1297. https://doi.org/10.1104/pp.18.00945
Moore JW, Herrera-Foessel S, Lan C, Schnippenkoetter W, Ayliffe M, Huerta-Espino J, Lillemo M, Viccars L, Milne R, Periyannan S (2015) A recently evolved hexose transporter variant confers resistance to multiple pathogens in wheat. Nat Genet 47:1494–1498. https://doi.org/10.1038/ng.3439
Paterson AH, Lander ES, Hewitt JD, Peterson S, Lincoln SE, Tanksley SD (1988) Resolution of quantitative traits into Mendelian factors by using a complete linkage map of restriction fragment length polymorphisms. Nature 335(6192):721–726
Paulitz TC, Smiley RW, Cook RJ (2002) Insights into the prevalence and management of soilborne cereal pathogens under direct seeding in the Pacific Northwest, USA. Can J Plant Pathol 24:416–428. https://doi.org/10.1080/07060660209507029
Porebski S, Bailey LG, Baum BR (1997) Modification of a CTAB DNA extraction protocol for plants containing high polysaccharide and polyphenol components. Plant Mol Biol Rep 15:8–15. https://doi.org/10.1007/BF02772108
Pumphrey MO, Bernardo R, Anderson JA (2007) Validating the Fhb1 QTL for Fusarium head blight resistance in near-isogenic wheat lines developed from breeding populations. Crop Sci 47(1):200–206. https://doi.org/10.2135/cropsci2006.03.0206
Roberts A, Pimentel H, Trapnell C, Pachter L (2011) Identification of novel transcripts in annotated genomes using RNA-Seq. Bioinformatics 27:2325–2329. https://doi.org/10.1093/bioinformatics/btr355
Saremi H, Ammarellou A, Jafary H (2007) Incidence of crown rot disease caused by Fusarium pseudograminearum as a new soil born fungal species in North West Iran. Pak J Biol Sci 20:3606–3612. https://doi.org/10.3923/pjbs.2007.3606.3612
Simpfendorfer S, McKay A, Ophel-Keller K (2019) New approaches to crop disease management in conservation agriculture. In: Pratley J and Kirkegaard J (eds) Australian Agriculture in 2020: From Conservation to Automation, 173–188.
Smiley RW, Gourlie JA, Easley SA, Patterson LM, Whittaker RG (2005) Crop damage estimates for crown rot of wheat and barley in the Pacific Northwest. Plant Dis 89:595–604. https://doi.org/10.1094/PD-89-0595
Stark R, Grzelak M, Hadfield J (2019) RNA sequencing: the teenage years. Nat Rev Genet 20(11):631–656. https://doi.org/10.1038/s41576-019-0150-2
Tuinstra M, Ejeta G, Goldsbrough P (1997) Heterogeneous inbred family (HIF) analysis: a method for developing near-isogenic lines that differ at quantitative trait loci. Theor Appl Genet 95:1005–1011. https://doi.org/10.1007/s001220050654
Tunali B, Nicol JM, Hodson D, Uckun Z et al (2008) Root and crown rot fungi associated with spring, facultative, and winter wheat in Turkey. Plant Dis 92:1299–1306. https://doi.org/10.1094/PDIS-92-9-1299
Van Ooijen JW (2006) JoinMap 4, software for the calculation of genetic linkage maps in experimental populations. Kyazma B.V., Wageningen
Wang Z, Gerstein M, Snyder M (2009) RNA-Seq: a revolutionary tool for transcriptomics. Nat Rev Genet 10:57–63. https://doi.org/10.1038/nrg2484
Wäspi U, Misteli B, Hasslacher M, Jandrositz A, Kohlwein SD, Schwab H, Dudler R (1998) The defense-related rice gene Pir7b encodes an α/β hydrolase fold protein exhibiting esterase activity towards naphthol AS-esters. Eur J Biochem 254:32–37. https://doi.org/10.1046/j.1432-1327.1998.2540032.x
Xu F, Kapos P, Cheng YT, Li M, Zhang Y, Li X (2014) NLR-associating transcription factor bHLH84 and its paralogs function redundantly in plant immunity. PLoS Pathog 10:e1004312. https://doi.org/10.1371/journal.ppat.1004312
Xu F, Song Y, Wang J, Liu L, Zhao K (2017) Occurrence of Fusarium crown rot caused by Fusarium pseudograminearum on barley in China. Plant Dis 101:837–837. https://doi.org/10.1094/PDIS-10-16-1436-PDN
Yan J, Wang J, Li Q, Hwang JR, Patterson C, Zhang H (2003) AtCHIP, a U-box-containing E3 ubiquitin ligase, plays a critical role in temperature stress tolerance in Arabidopsis. Plant Physiol 132:861–869. https://doi.org/10.1104/pp.103.020800
Ye J, Coulouris G, Zaretskaya I, Cutcutache I, Rozen S, Madden TL (2012) Primer-BLAST: a tool to design target-specific primers for polymerase chain reaction. BMC Bioinform 13:1–11. https://doi.org/10.1186/1471-2015-13-134
Zeng LR, Qu S, Bordeos A et al (2004) Spotted leaf11, a negative regulator of plant cell death and defense, encodes a U-box/armadillo repeat protein endowed with E3 ubiquitin ligase activity. Plant Cell 16:2795–2808. https://doi.org/10.1105/tpc.104.025171
Zheng Z, Wang HB, Chen GD, Yan GJ, Liu CJ (2013) A procedure allowing up to eight generations of wheat and nine generations of barley per annum. Euphytica 191:311–316. https://doi.org/10.1007/s10681-013-0909-z
Zheng Z, Ma J, Stiller J et al (2015) Fine mapping of a large-effect QTL conferring Fusarium crown rot resistance on the long arm of chromosome 3B in hexaploid wheat. BMC Genom 16:850. https://doi.org/10.1186/s12864-015-2105-0
Acknowledgements
The authors wish to thank Caritta Eliasson for her technical support.
Funding
Open access funding provided by CSIRO Library Services. The study reported here was partially funded by the Grain Research and Development Corporation, Australia (Grant No. CFF00010).
Author information
Authors and Affiliations
Contributions
CL and ZZ conceived the study and designed the experiments. SG, YJ, and ZZ conducted the experiments, collected, and analyzed data. SG, HZ, HL, and ZZ conducted the transcriptomic analysis. SG and ZZ wrote the article with contributions from all the authors. CL, HL, YL, and ZZ supervised and complemented the writing. All authors approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Thomas Miedaner.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gao, S., Jiang, Y., Zhou, H. et al. Fine mapping of a Fusarium crown rot resistant locus on chromosome arm 6HL in barley by exploiting near isogenic lines, transcriptome profiling, and a large near isogenic line-derived population. Theor Appl Genet 136, 137 (2023). https://doi.org/10.1007/s00122-023-04387-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00122-023-04387-x