Abstract
Key message
Two candidate genes (ZmbZIP113 and ZmTSAH1) controlling low-temperature germination ability were identified by QTL-seq and integrative transcriptomic analyses. The functional verification results showed that two candidate genes positively regulated the low-temperature germination ability of IB030.
Abstract
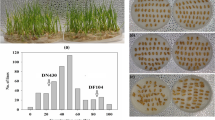
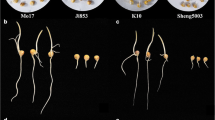
Low-temperature conditions cause slow maize (Zea mays L.) seed metabolism, resulting in slow seedling emergence and irregular seedling emergence, which can cause serious yield loss. Thus, improving a maize cultivar’s low-temperature germination ability (LTGA) is vital for increasing yield production. Wild relatives of maize, such as Z. perennis and Tripsacum dactyloides, are strongly tolerant of cold stress and can thus be used to improve the LTGA of maize. In a previous study, the genetic bridge MTP was constructed (from maize, T. dactyloides, and Z. perennis) and used to obtain a highly LTGA maize introgression line (IB030) by backcross breeding. In this study, IB030 (Strong-LTGA) and Mo17 (Weak-LTGA) were selected as parents to construct an F2 offspring. Additionally, two major QTLs (qCS1-1 and qCS10-1) were mapped. Then, RNA-seq was performed using seeds of IB030 and the recurrent parent B73 treated at 10 °C for 27 days and 25 °C for 7 days, respectively, and two candidate genes (ZmbZIP113 and ZmTSAH1) controlling LTGA were located using QTL-seq and integrative transcriptomic analyses. The functional verification results showed that the two candidate genes positively regulated LTGA of IB030. Notably, homologous cloning showed that the source of variation in both candidate genes was the stable inheritance of introgressed alleles from Z. perennis. This study was thus able to analyze the LTGA mechanism of IB030 and identify resistance genes for genetic improvement in maize, and it proved that using MTP genetic bridge confers desirable traits or phenotypes of Z. perennis and tripsacum essential to maize breeding systems.






Similar content being viewed by others
Data availability
The datasets presented in this study can be found in online repositories and Supplementary Material. The transcriptome data can be found in the National Genomics Data Center (NGDC) database. You can query transcriptome data by visiting the link (https://ngdc.cncb.ac.cn/search/?dbId=gsa&q=CRA008903) (BioProject: PRJCA013187; Accession number: CRA008903).
References
Abe A, Kosugi S, Yoshida K, Natsume S, Takagi H, Kanzaki H, Matsumura H, Yoshida K, Mitsuoka C, Tamiru M, Innan H, Cano L, Kamoun S, Terauchi R (2012) Genome sequencing reveals agronomically important loci in rice using MutMap. Nat Biotechnol 30(2):174–178. https://doi.org/10.1038/nbt.2095
Allam M, Revilla P, Djemel A, Tracy WF, Ordás B (2016) Identification of QTLs involved in cold tolerance in sweet × field corn. Euphytica 208(2):353–365. https://doi.org/10.1007/s10681-015-1609-7
An X, Chen J, Zhang J, Liao Y, Dai L, Wang B, Liu L, Peng D (2015) Transcriptome profiling and identification of transcription factors in ramie (Boehmeria nivea L. Gaud) in response to PEG treatment, using illumina paired-end sequencing technology. Int J Mol Sci 16(2):3493–3511. https://doi.org/10.3390/ijms16023493
Aroca R, Tognoni F, Juan José I, Manuel S-D, Pardossi A (2001) Different root low temperature response of two maize genotypes differing in chilling sensitivity. Plant Physiol Biochem 39(12):1067–1073. https://doi.org/10.1016/S0981-9428(01)01335-3
Båga M, Chodaparambil SV, Limin AE, Pecar M, Fowler DB, Chibbar RN (2007) Identification of quantitative trait loci and associated candidate genes for low-temperature tolerance in cold-hardy winter wheat. Funct Integr Genomics 7(1):53–68. https://doi.org/10.1007/s10142-006-0030-7
Basnet RK, Duwal A, Tiwari DN, Xiao D, Monakhos S, Bucher J, Visser RG, Groot SP, Bonnema G, Maliepaard C (2015) Quantitative trait locus analysis of seed germination and seedling vigor in Brassica rapa reveals QTL hotspots and epistatic interactions. Front Plant Sci 1(6):1032. https://doi.org/10.3389/fpls.2015.01032
Cai W, Yang Y, Wang W, Guo G, Liu W, Bi C (2018) Overexpression of a wheat (Triticum aestivum L.) bZIP transcription factor gene, TabZIP6, decreased the freezing tolerance of transgenic Arabidopsis seedlings by down-regulating the expression of CBFs. Plant Physiol Biochem 124:100–111. https://doi.org/10.1016/j.plaphy.2018.01.008
Cao L, Fu L, Zhang YB, Yan F, Yang ZM (2008) Effects of low temperature on root of maize seedling growth and phosphorus uptake. J Maize Sci 16(4):58–60
Cheng L, Li S, Hussain J, Xu X, Yin J, Zhang Y, Chen X, Li L (2013) Isolation and functional characterization of a salt responsive transcriptional factor, LrbZIP from lotus root (Nelumbo nucifera Gaertn). Mol Biol Rep 40(6):4033–4045. https://doi.org/10.1007/s11033-012-2481-3
Cheng M, Pan Z, Cui K, Zheng J, Luo X, Chen Y, Yang T, Wang H, Li X, Zhou Y, Lei X, Li Y, Zhang R, Iqbal MZ, He R (2022) RNA sequencing and weighted gene co-expression network analysis uncover the hub genes controlling cold tolerance in Helictotrichon virescens seedlings. Front Plant Sci 13:938859. https://doi.org/10.3389/fpls.2022.938859
Cingolani P, Platts A, Wang IE, Coon M, Nguyen T, Wang L, Land SJ, Lu X, Ruden DM (2012) A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of drosophila melanogaster strain w1118; ISO-2; ISO-3. Fly 6(2):80–92. https://doi.org/10.4161/fly.19695
Finkelstein RR, Lynch TJ (2000) The Arabidopsis abscisic acid response gene ABI5 encodes a basic leucine zipper transcription factor. Plant Cell 12(4):599–609. https://doi.org/10.1105/tpc.12.4.599
Fracheboud Y, Ribaut J-M, Vargas M, Messmer R, Stamp P (2022) Identification of quantitative trait loci for cold-tolerance of photosynthesis in maize (Zea mays L.). J Exp Bot 53(376):1967–1977. https://doi.org/10.1093/jxb/erf040
Frascaroli E, Landi P (2017) Registration of maize inbred line Bo23 with high cold tolerance and agronomic performance for early sowing. J Plant Regist 11(2):172–177. https://doi.org/10.3198/jpr2016.05.0029crg
Fujino K, Sekiguchi H, Sato T, Kiuchi H, Nonoue Y, Takeuchi Y, Ando T, Lin SY, Yano M (2004) Mapping of quantitative trait loci controlling low-temperature germinability in rice (Oryza sativa L.). Theor Appl Genet 108(5):794–799. https://doi.org/10.1007/s00122-003-1509-4
Gao SQ, Chen M, Xu Z-S, Zhao C-P, Li L, Xu H-J, Tang Y-M, Zhao X, Ma YZ (2011) The soybean GmbZIP1 transcription factor enhances multiple abiotic stress tolerances in transgenic plants. Plant Mol Biol 75(6):537–553. https://doi.org/10.1007/s11103-011-9738-4
Han Z, Ku L, Zhang Z, Zhang J, Guo SL, Liu H, Zhao R, Ren Z, Zhang L, Su H (2014) QTLs for seed vigor-related traits identified in maize seeds germinated under artificial aging conditions. PLoS ONE 9(3):e92535. https://doi.org/10.1371/journal.pone.0092535
Han Q, Zhu Q, Shen Y, Lee M, Lübberstedt T, Zhao G (2022) QTL mapping low-temperature germination ability in the maize IBM Syn10 DH population. Plants 11(2):214. https://doi.org/10.3390/plants11020214
Hickey LT, Hafeez AN, Robinson H, Jackson SA, Leal-Bertioli SCM, Tester M, Gao C, Godwin ID, Hayes BJ, Wulff BBH (2019) Breeding crops to feed 10 billion. Nat Biotechnol 37(7):744–754. https://doi.org/10.1038/s41587-019-0152-9
Hossain MA, Lee Y, Cho J-L, Ahn C-H, Lee S-K, Jeon J-S, Kang H, Lee C-H, An G, Park PB (2010) The bZIP transcription factor OsABF1 is an ABA responsive element binding factor that enhances abiotic stress signaling in rice. Plant Mol Biol 72(4–5):557–566. https://doi.org/10.1007/s11103-009-9592-9
Hu S, Lübberstedt T, Zhao G, Lee M (2016) QTL mapping of low-temperature germination ability in the maize IBM Syn4 RIL population. PLoS ONE 11(3):e0152795. https://doi.org/10.1371/journal.pone.0152795
Huang J, Zhang J, Li W, Hu W, Duan L, Feng Y, Qiu F, Yue B (2013) Genome-wide association analysis of ten chilling tolerance indices at the germination and seedling stages in maize. J Integr Plant Biol 55(8):735–744. https://doi.org/10.1111/jipb.12051
Hund A, Fracheboud Y, Soldati A, Frascaroli E, Salvi S, Stamp P (2004) QTL controlling root and shoot traits of maize seedlings under cold stress. Theor Appl Genet 109(3):618–629. https://doi.org/10.1007/s00122-004-1665-1
Hund A, Richner W, Soldati A, Fracheboud Y, Stamp P (2007) Root morphology and photosynthetic performance of maize inbred lines at low temperature. Eur J Agron 27(1):52–61. https://doi.org/10.1016/j.eja.2007.01.003
Hund A, Reimer R, Stamp P, Walter A (2012) Can we improve heterosis for root growth of maize by selecting parental inbred lines with different temperature behaviour? Philos Trans R Soc Lond 367(1595):1580–1588. https://doi.org/10.1098/rstb.2011.0242
Hwang I, Manoharan RK, Kang JG, Chung MY, Kim YW, Nou IS (2016) Genome-wide identification and characterization of bZIP transcription factors in Brassica oleracea under cold stress. Biomed Res Int 2016:4376598. https://doi.org/10.1155/2016/4376598
Iqbal MZ, Cheng M, Su Y, Li Y, Jiang W, Li H, Zhao Y, Wen X, Zhang L, Ali A, Rong T, Tang Q (2019) Allopolyploidization facilitates gene flow and speciation among corn Zea Perennis and Tripsacum dactyloides. Planta 249(6):1949–1962. https://doi.org/10.1007/s00425-019-03136-z
Jatimliansky JR, García MD, Molina MC (2004) Response to chilling of Zea mays, Tripsacum dactyloides and their hybrid. Biol Plant 48(4):561–567. https://doi.org/10.1023/B:BIOP.0000047153.23537.26
Jompuk C, Fracheboud Y, Stamp P, Leipner J (2005) Mapping of quantitative trait loci associated with chilling tolerance in maize (Zea mays L.) seedlings grown under field conditions. J Exp Bot 56(414):1153–1163. https://doi.org/10.1093/jxb/eri108
Kim JC, Lee Sh, Cheong YH, Yoo CM, Lee SI, Chun HJ, Yun DJ, Hong JC, Lee SY, Lim CO, Cho MJ (2001) A novel cold-inducible zinc finger protein from soybean, SCOF-1, enhances cold tolerance in transgenic plants. Plant Physiol 25(3):247–259. https://doi.org/10.1046/j.1365-313x.2001.00947.x
Klein H, Xiao Y, Conklin PA, Govindarajulu R, Kelly JA, Scanlon MJ, Whipple CJ, Bartlett M (2018) Bulked-segregant analysis coupled to whole genome sequencing (BSA-Seq) for rapid gene cloning in maize. G3 Genes Genomes Genet 8(11):3583–3592. https://doi.org/10.1534/g3.118.200499
Lakra N, Nutan KK, Das P, Anwar K, Singla-Pareek SL, Pareek A (2015) A nuclear-localized histone-gene binding protein from rice (OsHBP1b) functions in salinity and drought stress tolerance by maintaining chlorophyll content and improving the antioxidant machinery. J Plant Physiol 176:36–46. https://doi.org/10.1016/j.jplph.2014.11.005
Langfelder P, Horvath S (2008) WGCNA: an R package for weighted correlation network analysis. BMC Bioinform 9:559. https://doi.org/10.1186/1471-2105-9-559
Leipner J, Jompuk C, Camp K, Stamp P, Fracheboud Y (2008) QTL studies reveal little relevance of chilling-related seedling traits for yield in maize. Theor Appl Genet 116(4):555–562. https://doi.org/10.1007/s00122-007-0690-2
Li Z, Xu J, Gao Y, Wang C, Guo G, Luo Y, Huang Y, Hu W, Sheteiwy MS, Guan Y, Hu J (2017) The synergistic priming effect of exogenous salicylic acid and H2O2 on chilling tolerance enhancement during maize (Zea mays L.) seed germination. Front Plant Sci 8:1153. https://doi.org/10.3389/fpls.2017.01153
Li X, Wang G, Fu J, Li L, Jia G, Ren L, Lubberstedt T, Wang G, Wang J, Gu R (2018) QTL mapping in three connected populations reveals a st of consensus genomic regions for low temperature germination ability in Zea mays L. Front Plant Sci 9:65. https://doi.org/10.3389/fpls.2018.00065
Li P, Zheng T, Li L, Wang J, Cheng T, Zhang Q (2022a) Genome-wide investigation of the bZIP transcription factor gene family in Prunus mume: classification, evolution, expression profile and low-temperature stress responses. Hortic Plant J 8(2):230–242
Li Z, Fu D, Wang X, Zeng R, Zhang X, Tian J, Zhang S, Yang X, Tian F, Lai J, Shi Y, Yang S (2022b) The transcription factor bZIP68 negatively regulates cold tolerance in maize. Plant Cell 34(8):2833–2851. https://doi.org/10.1093/plcell/koac137
Liu H, Zhang L, Wang J, Li C, Zeng X, Xie S, Zhang Y, Liu S, Hu S, Wang J, Lee M, Lübberstedt T, Zhao G (2017) Quantitative trait locus analysis for deep-sowing germination ability in the maize IBM syn10 DH population. Front Plant Sci 22(8):813. https://doi.org/10.3389/fpls.2017.00813
Liu J, Fernie AR, Yan J (2020) The past, present, and future of maize Improvement: domestication, genomics, and functional genomic routes toward crop enhancement. Plant Commun 1(1):100010. https://doi.org/10.1016/j.xplc.2019.100010
Lu GW, Wu FQ, Wu WX, Wang HJ, Zheng XM, Zhang YH, Chen XL, Zhou KN, Jin MN, Cheng ZJ, Li XY, Jiang L, Wang HY, Wan JM (2014) Rice LTG1 is involved in adaptive growth and fitness under low ambient temperature. Plant Physiol 78(3):468–480. https://doi.org/10.1111/tpj.12487
Lu X, Liu J, Ren W, Yang Q, Chai Z, Chen R, Wang L, Zhao J, Lang Z, Wang H, Fan Y, Zhao J, Zhang C (2018) Gene-indexed mutations in maize. Mol Plant 11(3):496–504. https://doi.org/10.1016/j.molp.2017.11.013
Luang S, Sornaraj P, Bazanova N, Jia W, Eini O, Hussain SS, Kovalchuk N, Agarwal PK, Hrmova M, Lopato S (2018) The wheat TabZIP2 transcription factor is activated by the nutrient starvation-responsive SnRK3/CIPK protein kinase. Plant Mol Biol 96(6):543–561. https://doi.org/10.1007/s11103-018-0713-1
Luo L, Wei D, Li J, Gu WR, He D, Qu TM, Qiao TM, Yang ZF (2014) Effects of low-temperature stress on root system characteristics and electric conductivity of maize seedlings. Chin J Ecol 33(10):2694–2699
Ma H, Liu AC, Li AZ, Ran AQ, Xie AG (2018) ZmbZIP4 contributes to stress resistance in maize by regulating ABA synthesis and root development. Plant Physiol 178(2):753–770. https://doi.org/10.1104/pp.18.00436
Ma L, An R, Jiang L, Zhang C, Li Z, Zou C, Yang C, Pan G, Lübberstedt T, Shen Y (2022a) Effects of ZmHIPP on lead tolerance in maize seedlings: novel ideas for soil bioremediation. J Hazard Mater 430:128457. https://doi.org/10.1016/j.jhazmat.2022.128457
Ma L, Wang C, Hu Y, Dai W, Liang Z, Zou C, Pan G, Lubberstedt T, Shen Y (2022b) GWAS and transcriptome analysis reveal MADS26 involved in seed germination ability in maize. Theor Appl Genet 135(5):1717–1730. https://doi.org/10.1007/s00122-022-04065-4
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, Garimella K, Altshuler D, Gabriel S, Daly M, DePristo MA (2010) The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20(9):1297–1303. https://doi.org/10.1101/gr.107524.110
Najeeb S, Ali J, Mahender A, Pang YL, Zilhas J, Murugaiyan V, Vemireddy LR, Li Z (2020) Identification of main-effect quantitative trait loci (QTLs) for low-temperature stress tolerance germination- and early seedling vigor-related traits in rice (Oryza sativa L.). Mol Breed 40(1):10. https://doi.org/10.1007/s11032-019-1090-4
Ramakrishna C, Singh S, Raghavendrarao S, Padaria JC, Mohanty S, Sharma TR, Solanke AU (2018) The membrane tethered transcription factor EcbZIP17 from finger millet promotes plant growth and enhances tolerance to abiotic stresses. Sci Rep 8(1):2148. https://doi.org/10.1038/s41598-018-19766-4
Revilla P, Malvar RA, Cartea ME, Butrón A, Ordás A (2000) Inheritance of cold tolerance at emergence and during early season growth in maize. Crop Sci 40(6):1579–1585. https://doi.org/10.2135/cropsci2000.4061579x
Revilla P, Rodríguez VM, Ordás A, Rincent R, Charcosset A, Giauffret C, Melchinger AE, Schön CC, Bauer E, Altmann T, Brunel D, Moreno-González J, Campo L, Ouzunova M, Álvarez Á, Ruíz de Galarreta JI, Laborde J, Malvar RA (2016) Association mapping for cold tolerance in two large maize inbred panels. BMC Plant Biol 16(1):127. https://doi.org/10.1186/s12870-016-0816-2
Rodríguez-Zapata F, Barnes AC, Blcher-Juárez KA, Dan JG, Rellán-Lvarez R (2021) Teosinte introgression modulates phosphatidylcholine levels and induces early maize flowering time. Cold Spring Harbor Lab. https://doi.org/10.1101/2021.01.25.426574
Sallam A, Arbaoui M, El-Esawi M, Abshire N, Martsch R (2016) Identification and verification of QTL associated with frost tolerance using linkage mapping and GWAS in winter faba bean. Front Plant Sci 7:1098. https://doi.org/10.3389/fpls.2016.01098
Sasaki T (1979) Studies on breeding for germinability at low temperature of rice varieties adapted to direct sowing cultivation in flooded paddy field in cool region. VII. Effects of breeding conditions on germinability at low temperatures and initial seedling growth in early generations. Bull Hokkaido Prefect 42:11–19
Shi Y, Li G, Tian Z, Wang Z, Wang X, Zhu Y, Chen Y, Guo S, Qi J, Zhang X, Ku L (2016) Genetic dissection of seed vigour traits in maize (Zea mays L.) under low-temperature conditions. J Genet 95(4):1017–1022. https://doi.org/10.1007/s12041-016-0714-2
Takagi H, Abe A, Yoshida K, Kosugi S, Natsume S, Mitsuoka C, Uemura A, Utsushi H, Tamiru M, Takuno S, Innan H, Cano LM, Kamoun S, Terauchi R (2013) QTL-seq: rapid mapping of quantitative trait loci in rice by whole genome resequencing of DNA from two bulked populations. Plant J 74(1):174–183. https://doi.org/10.1111/tpj.12105
Tang QL, He RY, Li XF, Yan X, Li Y, Li Y, Zhao YL, Xie ML (2019) A method for cultivating cold-tolerant maize varieties using maize allopolyploids: CN201710062709.8. University SA, Chengdu
Tian J, Wang C, Xia J, Wu L, Xu G, Wu W, Li D, Qin W, Han X, Chen Q (2019) Teosinte ligule allele narrows plant architecture and enhances high-density maize yields. Science 365(6454):658–664. https://doi.org/10.1126/science.aax5482
Walne CH, Reddy KR (2022) Temperature effects on the shoot and root growth, development, and biomass accumulation of corn (Zea mays L.). Agriculture 12:443
Wang Y, Gao C, Liang Y, Wang C, Yang C, Liu G (2010) A novel bZIP gene from Tamarix hispida mediates physiological responses to salt stress in tobacco plants. J Plant Physiol 167(3):222–230. https://doi.org/10.1016/j.jplph.2009.09.008
Wang B, Zhang Z, Fu Z, Liu Z, Hu Y, Tang J (2016) Comparative QTL analysis of maize seed artificial aging between an immortalized F2 population and its corresponding RILs. Crop J 4(1):30–39. https://doi.org/10.1016/j.cj.2015.07.004
Wang H, Liang Q, Li K, Hu X, Wu Y, Wang H, Liu Z, Huang C (2017a) QTL analysis of ear leaf traits in maize (Zea mays L.)under different planting densities. Crop J 5(5):387–395
Wang L, Cao H, Qian W, Yao L, Hao X, Li N, Yang Y, Wang X (2017b) Identification of a novel bZIP transcription factor in Camellia sinensis as a negative regulator of freezing tolerance in transgenic arabidopsis. Ann Bot 119(7):1195–1209. https://doi.org/10.1093/aob/mcx011
Wang L, Zhu J, Li X, Wang S, Wu J (2018) Salt and drought stress and ABA responses related to bZIP genes from V. radiata and V. angularis. Gene 651:152–160. https://doi.org/10.1016/j.gene.2018.02.005
Wu L, Zhang Z, Zhang H, Wang X-C, Huang R (2008) Transcriptional modulation of ethylene response factor protein JERF3 in the oxidative stress response enhances tolerance of tobacco seedlings to salt, drought, and freezing. Plant Physiol 148(4):1953–1963. https://doi.org/10.1104/pp.108.126813
Yan X, Cheng M, Li Y, Wu Z, Li Y, Li X, He R, Yang C, Zhao Y, Li H, Wen X, Zhang P, Sam E, Rong T, He J, Tang Q (2020) Tripsazea, a novel trihybrid of Zea mays, Tripsacum dactyloides, and Zeaperennis. G3 Genes Genomes Genet 10(2):839–848. https://doi.org/10.1534/g3.119.400942
Ying S, Zhang D-F, Fu J, Shi Y-F, Song Y-C, Wang T-Y, Li Y (2012) Cloning and characterization of a maize bZIP transcription factor, ZmbZIP72, confers drought and salt tolerance in transgenic Arabidopsis. Planta 235(2):253–266. https://doi.org/10.1007/s00425-011-1496-7
Zhang X, Wang L, Meng H, Wen H, Fan Y, Zhao J (2011a) Maize ABP9 enhances tolerance to multiple stresses in transgenic Arabidopsis by modulating ABA signaling and cellular levels of reactive oxygen species. Plant Mol Biol 75(4–5):365–378. https://doi.org/10.1007/s11103-011-9732-x
Zhang XF, Hu B, Jing D (2011b) Effect of exogenous chemical substances pretreatment on maize seed germination under low temperature stress. Heilongjiang Agric Sci 4:69–73
Zhang Z, Xiuling C, Guan X, Liu Y, Chen H, Wang T, Mouekouba LDO, Jingfu L, Wang A (2014) A genome-wide survey of homeodomain-leucine zipper genes and analysis of cold-responsive HD-zip I members’ expression in tomato. Biosci Biotechnol Biochem 78(8):1337–1349. https://doi.org/10.1080/09168451.2014.923292
Zhang L, Zhang L, Xia C, Gao L, Hao C, Zhao G, Jia J, Kong X (2017) A novel wheat C-bZIP gene, TabZIP14-B, participates in salt and freezing tolerance in transgenic plants. Front Plant Sci 8:710. https://doi.org/10.3389/fpls.2017.00710
Zhang H, Zhang J, Xu Q, Wang D, Di H, Huang J, Yang X, Wang Z, Zhang L, Dong L, Wang Z, Zhou Y (2020) Identification of candidate tolerance genes to low-temperature during maize germination by GWAS and RNA-seqapproaches. BMC Plant Biol 20(1):333. https://doi.org/10.1186/s12870-020-02543-9
Zhang Y, Liu P, Wang C, Zhang N, Zhu Y, Zou C, Yuan G, Yang C, Gao S, Pan G, Ma L, Shen Y (2021) Genome-wide association study uncovers new genetic loci and candidate genes underlying seed chilling-germination in maize. Peer J 9:e11707. https://doi.org/10.7717/peerj.11707
Acknowledgements
This research was supported by the National Natural Science Foundation of China (32272035), Sichuan Science and Technology Innovation and Entrepreneurship Seedling Project (key project) (2023JDRC0117), Forage Breeding Projects of Sichuan Province during the 14th Five-Year Plan Period (2021YFYZ0013-3), the Sichuan Corn Innovation Team of National Modern Agricultural Industry Technology System (sccxtd-2020-02), the Science and Technology Project of Sichuan Province (2020YJ0466), and Fundamental Research Funds for the Central Universities (ZYN2022053).
Funding
This study was funded by The National Natural Science Foundation of China (32272035), Sichuan Science and Technology Innovation and Entrepreneurship Seedling Project (key project) (2023JDRC0117), Forage Breeding Projects of Sichuan Province during the 14th Five-Year Plan Period (2021YFYZ0013-3), the Sichuan Corn Innovation Team of National Modern Agricultural Industry Technology System (sccxtd-2020-02), the Science and Technology Project of Sichuan Province (2020YJ0466), and Fundamental Research Funds for the Central Universities (ZYN2022053).
Author information
Authors and Affiliations
Contributions
Q.T, T.R, and R.H conceived the study. J.Z, T.Y, and Z.P supervised the study. R.H, X.L, M.C, Y.Z, Y.C, Y.L, X.N, Y.L, T.Y, and Z.P performed the experiments. R.H and J.Z analyzed the data. R.H, J.Z, Y.L, M.Z.I, and J.H prepared the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no conflicts of interest.
Human and animal rights
This study does not include human or animal subjects.
Additional information
Communicated by Thomas Lubberstedt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
He, RY., Zheng, JJ., Chen, Y. et al. QTL-seq and transcriptomic integrative analyses reveal two positively regulated genes that control the low-temperature germination ability of MTP–maize introgression lines. Theor Appl Genet 136, 116 (2023). https://doi.org/10.1007/s00122-023-04362-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00122-023-04362-6