Abstract
O-GlcNAcylation of proteins is governed by O-GlcNAc transferase (OGT) and O-GlcNAcase (OGA). The homeostasis of O-GlcNAc cycling is regulated during cell cycle progression and is essential for proper cellular division. We previously reported the O-GlcNAcylation of the minichromosome maintenance proteins MCM2, MCM3, MCM6 and MCM7. These proteins belong to the MCM2–7 complex which is crucial for the initiation of DNA replication through its DNA helicase activity. Here we show that the six subunits of MCM2–7 are O-GlcNAcylated and that O-GlcNAcylation of MCM proteins mainly occurs in the chromatin-bound fraction of synchronized human cells. Moreover, we identify stable interaction between OGT and several MCM subunits. We also show that down-regulation of OGT decreases the chromatin binding of MCM2, MCM6 and MCM7 without affecting their steady-state level. Finally, OGT silencing or OGA inhibition destabilizes MCM2/6 and MCM4/7 interactions in the chromatin-enriched fraction. In conclusion, OGT is a new partner of the MCM2–7 complex and O-GlcNAcylation homeostasis might regulate MCM2–7 complex by regulating the chromatin loading of MCM6 and MCM7 and stabilizing MCM/MCM interactions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
O-GlcNAcylation (O-linked β-N-acetylglucosaminylation) of proteins is catalysed by O-GlcNAc transferase (OGT) which uses uridine-diphospho-N-acetylglucosamine (UDP-GlcNAc) to transfer the GlcNAc moiety onto serine or threonine residues of cytosolic, nuclear and mitochondrial proteins [1,2,3,4]. Conversely, O-GlcNAcase (OGA) reverses this abundant post-translational modification by removing the GlcNAc residue [5, 6]. O-GlcNAcylation regulates various cellular processes including transcription, translation, chromatin remodelling and cell cycle progression by modulating protein activity, stability, subcellular localization or protein–protein interaction [4, 7,8,9,10]. Moreover, a crosstalk can occur between O-GlcNAcylation and phosphorylation, either on adjacent sites or at the same sites of the target proteins [7, 9, 11,12,13]. Recently, the motif (pSp/T)P(V/A/T)(gS/gT) has been defined as a very specific and stringent phospho/O-GlcNAc crosstalk motif [14].
In mammalian cells, O-GlcNAcylation levels and the expression of OGT and OGA are highly regulated during the cell cycle, and disruption of O-GlcNAc cycling induces defects in cell cycle progression and mitosis [15,16,17,18,19,20,21,22,23,24]. The loss in O-GlcNAc homeostasis alters the expression of the early-induced transcription factors c-Fos, c-Jun, c-Myc and Sp1 [15] and the expression of cyclin D and p27KIP1 cell-cycle inhibitor [21, 25, 26]. It can also induce abnormal oscillations in the levels of cyclins E, A and B, resulting in aberrant mitotic-specific phosphorylation and defects in cytokinesis [12, 16,17,18, 23, 24]. To identify new targets of OGT during G1/S transition, we previously performed a differential analysis of the O-GlcNAcome of G1- and S phase synchronized human cells and identified the minichromosome maintenance (MCM) proteins MCM2/3/6/7 [22].
MCM2–7 complex is composed of six distinct MCM subunits assembled in a highly defined order: MCM5–MCM3–MCM7–MCM4–MCM6–MCM2. MCM proteins display a well-conserved organization of their functional domains, and possess an AAA+ ATPase activity in their C-terminal domain [27,28,29]. The heterohexameric MCM2–7 complex has a ring-shaped structure to encircle DNA, the MCM2–MCM5 interaction being identified as the gate that promotes the opening of the MCM2–7 complex [29, 30]. Sub-complexes containing MCM2/4/6/7, MCM4/6/7 or MCM3/5 have also been isolated from yeast and mammalian cells [31,32,33,34,35,36]. In vitro, the sub-assembly MCM4/6/7 exhibits the DNA helicase activity [37,38,39,40]. MCM2 which is weakly associated with this sub-complex negatively regulates its helicase activity [36, 37]. The strong interaction between MCM3 and MCM5 serves also negative regulatory function on MCM2–7 ATPase activity [36, 39, 41].
The MCM2–7 complex plays an essential role in the initiation of DNA replication which is a two-step process: licensing and firing. In G1 phase, MCM2–7 takes part in the formation of the pre-replicative complex (pre-RC) which is composed of ORC (origin replication complex, Orc1-6) and the licensing factors Cdc6 and Cdt1. During this licensing step, inactive head-to-head double MCM2–7 hexamers encircle DNA origins. As cells enter the S phase, the firing step requires the recruitment of Cdc45 and GINS to the MCM ring. Within the Cdc45/MCM2–7/GINS (CMG) complex, MCM2–7 exhibits the core replicative helicase activity necessary for parental DNA strands to unwind [29, 42, 43]. Finally, as cells progress in late S phase, MCM2–7 complexes progressively dissociate from chromatin to prevent DNA re-replication [44,45,46,47]. Importantly, most of the origins on which MCM2–7 complexes have been loaded constitute dormant origins that are not used normally in S phase. However, under conditions of replicative stress that induces replication fork stalling, excess MCM2–7 complexes license latent origins as a backup mechanism to prevent under-replication and maintain genome integrity [42, 48,49,50,51].
Phosphorylation of MCM subunits dramatically changes during cell cycle progression to finely regulate the functionality of MCM2–7 complex, from the licensing step to the dissociation of the complex from chromatin in late S phase [44, 52,53,54,55,56,57,58,59,60,61,62]. In this work, we investigate the role of O-GlcNAc post-translational modification on MCM2–7 complex. We show that all the MCM subunits are modified by OGT, mostly in the chromatin-bound fraction. We identify stable interaction between OGT and the MCM3, MCM6 and MCM7 proteins. Moreover, dysregulation of O-GlcNAc cycling by OGT silencing decreases the amount of chromatin-bound MCM2, MCM6 and MCM7 proteins and destabilizes MCM–MCM interactions.
Materials and methods
Antibodies, siRNA and chemicals
Thymidine (T1895), propidium iodide solution (P4864), RNAse A (R4875), complete protease inhibitor cocktail tablets and OGA inhibitor Thiamet G (ThG) were from Sigma-Aldrich (St Quentin Fallavier, France). ThG was used at 1 µM (prepared at 100 mM in DMSO) [63]. The OGT inhibitor acetyl-5S-GlcNAc (5S-G) was kindly provided by Pr. G. W. Hart and used at 50 µM (prepared at 50 mM in DMSO) [64]. GlcNAc was from TCI Chemicals (TCI Europe N.V., Belgium).
The following primary antibodies were used for Western blot: RL2 O-GlcNAc antibody (1:3000, ThermoScientific, Fisher Scientific, France), OGT (DM17 or Ti-14; 1:2000, Sigma-Aldrich), OGA (1:10,000, anti-MGEA5, ab124807, Abcam, Cambridge, UK), GST, MCM2 (PLA0060) and MCM5 (PLA0064) (1:3000, Sigma-Aldrich); MCM3 (N-19), MCM4 (H-300), MCM6 (H-8), MCM7 (141.2), GAPDH (0411) (1:3000, Santa Cruz, Heidelberg, Germany), Cyclin D1 (A-12), Cdt1 (H-300) (1:1000, Santa Cruz), Cdc6 (DCS-180) (1:1000, Merck, Darmstadt, Germany). The same antibodies were used for immunoprecipitation and indirect immunofluorescence or PLA experiments except for immunoprecipitation of MCM7 (D10A11, Cell Signaling Technology, Ozyme, Montigny-le-Bretonneux, France) and detection of MCM3 by PLA (3E1, Abgent, Euromedex, Souffelweyersheim, France). Normal control IgG polyclonal antibodies were used as negative controls for IP experiments (rabbit, mouse, or goat, Santa Cruz). The following secondary antibodies were also used: anti-goat IgG–HRP linked (1:30,000, Santa Cruz), anti-mouse IgG–HRP-linked and anti-rabbit IgG HRP-linked antibodies (1:10,000, GE Healthcare, V.W.R. Fontenay-sous-Bois, France), anti-rabbit IgG Alexa Fluor 488 and anti-mouse IgG Alexa Fluor 568 (1:600, ThermoScientific, Fisher Scientific, France). Control siRNA (siRNA univ. negative control) and siRNA against OGT (GGAGGCUAUUCGAAUCAGU[dT][dT] forward, ACUGAUUCGAAUAGCCUCC[dT][dT] reverse) were purchased from Sigma-Aldrich. siRNA against OGA (siGENOME Human MGEA5 (10724) siRNA SMARTpool) was from Dharmacon (GE Healthcare Europe GmbH, Velizy-Villacoublay, France). The p3XFLAG-OGT-siRNA-resistant vector was generated by directed mutagenesis using Phusion® Hot start (NEB), p3XFLAG-OGT as temple and, 5′-agcagggaaaactgcaggaagctctgatgcattataaagaagcgatcaggatttcccctacctttgctgatgcctactc-3′ and 5′-gagtaggcatcagcaaaggtaggggaaatcctgatcgcttctttataatgcatcagagcttcctgcagttttccctgct-3′ as primers. Prior to transformation in DH5α, template was digested for 2 h at 37 °C by 1 U of DpnI (NEB). Positive clones were screened by sequencing.
Cell culture, transfection and cell cycle synchronization
MCF7 and MDA-MB-231 breast cancer cell lines, and HEK 293T cells were routinely grown at 37 °C in a humidified atmosphere enriched with 5% CO2 in Dulbecco’s modified Eagle’s medium (DMEM) (Lonza, Basel, Switzerland) containing high glucose (4.5 g/L) and glutamine, and supplemented with 10% foetal calf serum (FCS) (Lonza) (complete medium).
For HA-OGT transfection, HEK293T cells were cultured in complete medium (8 × 105 cells/100-mm dish) and when they reached around 60% of confluence, they were transiently transfected with HA-tagged OGT (1.25 µg/100-mm dish) using the Lipofectamine® 2000 (Thermofisher, Fisher Scientific, France), according to the manufacturer’s instructions. HEK293T were harvested 48 h after transfection. For small interfering RNA (siRNA) transfection, MCF7 (1.5 × 106 cells/100-mm dish) and MDA-MB-231 (106 cells/100-mm dish) were reverse-transfected with Lipofectamine® RNAiMAX (Thermofisher, Fisher Scientific, France) according to the manufacturer’s recommendations, using 60 pmol of siControl, siOGA or siOGT. Cells were harvested, respectively, after 60 or 72 h of transfection. To rescue OGT silencing, 1 day later siRNA transfection, MCF7 cells were transfected with the 3XFLAG-OGT-siRNA-resistant plasmid or the 3X-pCMV plasmid as the negative control (250 ng/w), using 2.5 µL Lipofectamine® 2000. For both types of experiments (siRNA ± 3XFLAG-OGT-siRNA), when cell cycle synchronization was needed, cells were serum-starved 24 h after transfection and the synchronization protocol was followed, as mentioned below.
Cell cycle synchronization was performed using starvation followed by serum stimulation [22]. After 24 h in complete medium (DMEM-10% FCS), cell monolayer was rinsed with PBS (Lonza) and placed either 24 h in DMEM-0.5% FCS for MCF7 cells, or 48 h in serum-free medium for MDA-MB-231 cells. Cells were either harvested (time 0 h) or grown in complete medium to release cells in cell cycle. Cells were harvested at different time points, according to cell cycle progression that was systematically monitored by propidium iodide (PI) DNA staining and flow cytometry analysis, as previously described [22]. When mentioned, vehicle (DMSO, 1:1000), acetyl-5S-GlcNAc (5S-G, 50 µM) and ThG (1 µM) were added simultaneously with serum.
Cell lysis and subcellular fractionation
After two washes of cellular layers with ice-cold PBS, whole cellular lysates (WCL) were obtained using RIPA buffer (150 mM NaCl, 50 mM Tris–HCl, pH 7.5, 1% (v/v) Triton X-100, 0.2% (w/v) NaDoc, 0.1% (w/v) SDS, containing 1 mM orthovanadate, 10 mM sodium fluoride and protease inhibitors) and placed on ice for 10 min. The lysate was then clarified by centrifugation at 18,800×g for 15 min and the supernatant was stored at − 20 or − 80 °C before use.
Subcellular fractionation was performed as previously described [55]. Cells were lysed in cytoskeleton extraction buffer (CSK) (100 mM NaCl, 10 mM PIPES, pH 7, 300 mM sucrose, 3 mM MgCl2, 0.1% (v/v) NP-40 with protease inhibitor cocktail and phosphatase inhibitors (10 mM NaF, 1 mM orthovanadate) at 4 °C for 20 min. Lysates were then centrifuged at 300×g for 5 min at 4 °C and supernatants were retrieved. A pellet wash was performed in CSK buffer, and after centrifugation at 300xg for 5 min at 4 °C, supernatants were retrieved. Combined supernatants were then clarified by centrifugation at 18,800×g for 15 min at 4 °C to constitute the soluble fraction (Sol.) containing the nucleocytoplasmic proteins. Finally, pellets were incubated in modified RIPA buffer (350 mM NaCl, 20 mM Tris, pH 7.5, 2.5 mM sodium pyrophosphate, 1% (v/v) Triton X-100) for 20 min on ice, sonicated for 3 min to break DNA, and then centrifuged at 18,800×g for 15 min at 4 °C. The clear supernatant was used as the chromatin-bound protein fraction (Chrom.).
Immunoprecipitation and Western blotting
For co-immunoprecipitation experiments, non-SDS containing lysis and washing buffers were used, and chromatin-bound protein fraction (500 µg) was diluted ½ in NaCl-free modified RIPA-buffer to reduce NaCl concentration to 175 mM. Total, soluble and chromatin-bound protein extracts were precleared with a mix of protein A- and protein G-Sepharose beads (50:50) (GE Healthcare, V.W.R.) in lysis buffer for 2 h at 4 °C (20 µL/mg). After centrifugation (5 min, 5000×g), the supernatant was incubated with primary antibodies for 2 h or overnight at 4 °C (5 µg/mg). Then, protein A-Sepharose (for rabbit IgG) or protein G-Sepharose (for mouse and goat IgG) was added and incubated for an additional 1 h (30 µL/mg). Beads were washed successively three times with RIPA buffer (5 min), once in high-salt containing RIPA buffer (300 mM NaCl), and finally boiled in Laemmli buffer before separation by SDS-PAGE and transfer onto nitrocellulose membranes (Protran supported 0.45 µm NC, GE Healthcare).
For Western blot, membranes were blocked in 5% (w/v) nonfat dry milk in Tris-buffered saline (TBS 10×, Euromedex) with 0.05% (v/v) Tween 20 (TBS-T) and probed with primary antibodies overnight at 4 °C. The membranes were washed three times with TBS-T and incubated with the corresponding HRP-conjugated secondary antibody for 1 h at RT. Membranes were washed three times in TBS-T and immunoblots were developed with enhanced chemiluminescence (ECL prime Reagent, GE Healthcare, Supersignal West Pico Plus or Supersignal West Femto, ThermoScientific, Fisher Scientific, France). Image acquisition was done on a CCD camera (Fusion Solo, Vilber Lourmat, Marne-la-Vallée, France). The membranes were stripped in the antibody stripping buffer (Gene Bio-Application LTD, Euromedex, France) for 15 min at RT, extensively washed in water and TBS-T before reprobing with another antibody.
sWGA lectin chromatography
Soluble and chromatin-bound proteins were enriched for GlcNAc-modified proteins using the GlcNAc-specific lectin succinylated wheat germ agglutinin (sWGA) immobilized on agarose (Vector Laboratories, Clinisciences, Nanterre, France) [65]. sWGA beads were first equilibrated in the twofold diluted RIPA-modified buffer used for chromatin-bound protein extraction (175 mM NaCl, 10 mM Tris, pH 7.5, 1.25 mM sodium pyrophosphate, 0.5% (v/v) Triton X-100). For each fraction (Sol. and Chrom.), the volume corresponding to 1 mg of proteins was adjusted to 500 µL in the initial lysis buffer, and then twofold diluted in PBS to get a final concentration of 1 mg/mL. Each fraction (1 mg) was incubated for 2 h at 4 °C with 50 µL of sWGA-beads. Beads were centrifuged at 1000×g for 3 min, and then washed under vigorous stirring successively, twice with 1 mL ½-diluted RIPA-modified buffer and twice with the same buffer containing 300 mM NaCl. Finally, beads were re-suspended in Laemmli buffer before heating at 95 °C for 7 min and SDS-PAGE. A negative control was performed for each fraction by adding 0.5 M free GlcNAc in the lysate before incubation with sWGA-beads.
GST pull-down assay
Bacterial expression plasmids pGEX-2T for GST and GST–OGT fusion proteins were kindly provided by Drs. D. Leprince and X. Yang, respectively. For GST recombinant protein expression, BL21 DE-3 Escherichia coli were transformed with plasmids and cultured in LB medium containing 50 µg/mL ampicillin. When bacteria reached the exponential growth phase, induction was performed at room temperature with 0.1 mM IPTG for 4 h. Bacteria were centrifuged and pellets were resuspended in PBS containing a cocktail of protease inhibitors (Sigma-Aldrich). Crude lysates were obtained using the high-pressure homogenizer Emulsiflex-C3 (Avestin, Mannheim, Germany) and centrifuged at 10,400×g for 45 min. GST fusion proteins were immobilized on Glutathione Sepharose 4B beads (GE Healthcare) for 2 h at 4 °C under gentle agitation. Beads were successively washed for 5 min by gentle vortex in 20 mM Tris, pH 7.4, with 0.1% (v/v) Triton X-100 (twice) and in the same buffer containing 100 mM NaCl (twice), followed by centrifugation at 500×g for 5 min. For direct elution, beads were equilibrated twice in the elution buffer (50 mM Tris, pH 8, with 0.1% (v/v) Triton X-100) before adding 50 mM reduced glutathione (Sigma-Aldrich) in elution buffer. For GST pull down experiments using human cell lysates, 700 µg of proteins (soluble nucleocytoplasmic and chromatin-bound subcellular fractions) were added in each tube with the beads and incubated overnight at 4 °C with gentle agitation. Beads were successively washed three times in PBS with 0.1% Triton X-100, once in PBS with 0.1% Triton X-100 and 150 mM NaCl, and twice in 50 mM Tris, pH 8, with 0.1% Triton X-100 before elution as described before. Laemmli buffer was added in each eluted fraction, samples were boiled 5 min at 95 °C before SDS-PAGE.
Click chemistry
We used the Click-It O-GlcNAc enzymatic labelling and the Click-It biotin glycoprotein systems (ThermoScientific, Fisher) to enrich O-GlcNAc proteins from cell lysates on streptavidin–agarose beads (Merck), as previously described [66]. Briefly, proteins from the chromatin-bound fraction were first precipitated using chloroform/methanol according to the manufacturer’s protocol, and then solubilized in the presence of 1% (w/v) SDS in 20 mM HEPES (pH 7.9). O-GlcNAcylated proteins were enzymatically labelled with a GalNAz residue using the Y289L galactosyltransferase (GalT) and UDP-GalNAz as the nucleotide sugar. A negative control (time point 18H) in which UDP-GalNAz was omitted, was carried out in parallel and treated exactly in the same conditions than samples. Labelled proteins were then subjected to a click chemistry reaction with a biotin–alkyne probe and enriched on streptavidin–agarose beads. After click-chemistry reaction, 10% of solubilized and Click-It-labelled proteins were removed to attest the presence of MCM proteins in labelled samples (Input Click-It).
We performed O-GlcNAc mass tag labelling to detect and quantify O-GlcNAcylated fraction of MCM proteins using a 4.4 kDa DBCO-PEG mass tag. After using the Click-It O-GlcNAc enzymatic labelling kit following the manufacturer’s instructions, 50 µg of GalNAz-labelled proteins were re-suspended in 1% SDS, 20 mM HEPES (pH 7.9) and incubated for 1 h at room temperature, under a gentle agitation, either with a 4.4 KDa DBCO-PEG at a final concentration of 10 mM (PEG+) or with DMSO (PEG−). Finally, labelled protein samples were precipitated using chloroform/methanol to remove excess of DBCO-PEG and boiled with Laemmli buffer before separation on polyacrylamide gel. For each MCM, electrophoretic Rf values were calculated to estimate the molecular weight of the non-PEGylated and the PEGylated shifted bands and establish how many O-GlcNAc sites are present on MCM proteins. Standard curve was established using the PageRuler Prestained Protein Ladder (10–170 kDa, ThermoScientific, Fisher). In vivo glycosylation stoichiometry was determined by quantifying the relative intensities of each band [67].
Cell cycle and DNA synthesis analysis
Distribution of cells in G0/G1, S and G2/M was routinely determined by DNA staining with PI as previously described [22]. The rate of DNA synthesis was measured using the Click-iT® EdU Flow cytometry assay kit (ThermoScientific, Fisher). After siRNA transfection or treatment with inhibitor and subsequent synchronization, cells were labelled with 10 µM EdU (5-ethynyl-2′-deoxyuridine) for 15 min before harvesting cells. Detection of EdU-positive cells was based on the click reaction (K+) with Alexa Fluor® 647 (AF647) azide fluorescent dye according to the manufacturer’s recommendations. DNA content was further labelled with PI. Several controls were performed to set up the flow cytometry instrument parameters: no labelled cells, PI-labelled cells, Edu-K+-labelled cells and Edu-K−/PI-labelled cells. Cells were then analysed by flow cytometry on a CyAn ADP LX9 instrument using the Summit V4.3.04 software for data acquisition (Beckman Coulter, Life Sciences). AF647 and PI signals were read, respectively, in FL8 channel (laser 635 nm and λem: 665/20 nm) and FL3 channel (laser 488 nm, λem: 613/20 nm). Data were analysed using FlowJo software.
Immunofluorescence and proximity ligation assay
MCF7 cells were grown on glass coverslips for 72 h and washed three times in cold PBS before fixation in 4% paraformaldehyde in PBS at room temperature for 20 min. After three washes in PBS (5 min per wash, at RT), permeabilization of cells was performed either with 0.5% (v/v) Triton X-100 in PBS for 2 min to detect OGT–MCM interaction [68], or 20 min with 0.5% (v/v) Triton X-100 in CSK buffer without NP-40 for the detection of MCM–MCM interactions [47]. This was followed by a quenching with 100 mM glycine (pH 7.4) in PBS for 20 min and three washes in PBS. Coverslips were incubated with blocking buffer (2% (v/v) FCS, 2% bovine serum albumin (w/v), 0.2% (w/v) gelatin in PBS) for 1 h at RT before incubation with the primary antibodies (1:100) diluted in the blocking buffer, overnight at 4 °C. For indirect immunofluorescence, coverslips were washed three times with 0.5% (v/v) Tween 20 in PBS and incubated with Alexa Fluor-conjugated secondary antibodies (1:600 in blocking buffer) for 1 h in the dark, at RT For the Proximity ligation assay (Duolink® in situ kit, Sigma-Aldrich), after incubation with the primary antibodies and two washes in PBS, the coverslips were incubated with PLA PLUS and MINUS probes for mouse and rabbit, respectively, for 1 h, with the ligase for 30 min (ligation step), and with the polymerase for 2 h (amplification step, Duolink in situ detection reagents Green, λex/λem: 495/527 nm) in a humidity chamber at 37 °C. Finally coverslips were washed three times in 0.5% (v/v) Tween 20 in PBS, once in PBS alone, and nuclei were stained with DAPI (50 µg/mL) for 2 min before mounting slides in Mowiol solution (Calbiochem, Merck chemicals, Nottingham, UK). Negative controls were done using only one of the primary antibodies. Immunofluorescence was detected through an inverted Zeiss LSM700 confocal microscope with a 40x oil immersion lens at room temperature and data were collected with the ZEN 2010 software (Zeiss, Oberkochen, Germany). Images from PLA were processed with ImageJ® using a home-made plugin developed by TISBio to detect and quantify the nuclear fluorescent dots in labelled cells. Briefly, for each PLA file (channel 1, DAPI; channel 2, green PLA fluorescence), the nuclei were detected and labelled in channel 1 to define the r.o.i. (regions of interest) that were then applied to channel 2 to measure and quantify the fluorescence inside each labelled nucleus. The mean of fluorescence per cell is the ratio of the integrated density/area measured for each nucleus. Scatter dot plot (median with interquartile range) showing the mean of fluorescence per cell and statistical analysis (one-way ANOVA test, *p < 0.05) were obtained using GraphPad Prism software.
Results
All the MCM subunits are O-GlcNAcylated when they are loaded onto chromatin
We previously reported the O-GlcNAcylation of MCM3, MCM6 and MCM7 in MCF7 human cells after immunoprecipitation of endogenous MCM protein and detection by Western blot of the O-GlcNAc status using the anti-O-GlcNAc RL2 antibody [22]. Using the same approach, we show here that immunoprecipitated MCM2, MCM4 and MCM5 proteins are also O-GlcNAcylated in Thiamet G-treated MCF7 cells (Fig. 1a).
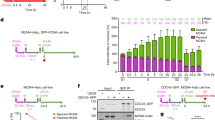
All MCM2–7 subunits are O-GlcNAcylated and mainly found in the chromatin-enriched fraction. a MCF7 cells were treated overnight with 1 µM ThG before lysis and immunoprecipitation of MCM2, MCM4 and MCM5 followed by Western blot analysis. Membranes were first incubated with anti-O-GlcNAc antibody (RL2), stripped and then reprobed with anti-MCM antibodies. b MCF7 cells were synchronized in cell cycle by starvation (time point 0) then released in S phase (15H, 18H). Cell cycle profiles were determined by FACS analysis after DNA staining with PI. Percentage of cells in G0/G1, S and G2/M phases are indicated. c The nucleocytoplasmic soluble fraction (Sol.) and chromatin-bound fraction (Chrom.) were obtained by subcellular fractionation of proteins from synchronized cells. Samples were analysed by Western blot for the indicated proteins (n.s. non-specific band). d O-GlcNAcylated proteins from soluble (Sol.) and chromatin-bound (Chrom.) fractions were enriched on sWGA-agarose beads. Incubation with excess of GlcNAc (0.5 M) was used as negative control (C−). MCM proteins were detected by Western blot before (Inp) and after enrichment on sWGA lectin (sWGA-bound). Equal loading was confirmed by Ponceau staining of the membranes. e O-GlcNAcylated proteins from chromatin-bound proteins of synchronized MCF7 cells were labelled with GalNAz and a biotin–alkyne probe (Input click-it) before enrichment on avidin–agarose beads (avidin bound). Negative controls were done by omitting UDP-GalNAz (C−). Samples were analysed by Western blot for MCM proteins before (Input) and after click-chemistry. (*) these bands correspond to the remnant signal for MCM2, despite membrane stripping. f O-GlcNAc-modified proteins from whole cell extract of MCF7 cells were enzymatically labelled with GalNAz and chemically modified with a 4.4 kDa DBCO-PEG mass tag (PEG+) or incubated with DMSO (PEG−) as negative control. MCM proteins were detected by Western blot and the number of O-GlcNAc sites and O-GlcNAcylation stoichiometry (indicated as the percentage of total MCM protein) were determined as reported in “Materials and methods”
The phosphorylation status of MCM proteins regulates the chromatin loading of MCM2–7 complex [44, 53, 56, 57, 59, 61, 62, 69]. To investigate whether O-GlcNAcylated forms of MCM proteins are differentially distributed in the soluble and the chromatin-bound fractions during cell cycle progression, MCF7 cells were arrested in G0 by serum starvation and released in G1 phase by serum stimulation. Cells were harvested in early (15H) and late (18H) S phase, as measured by FACS (Fig. 1b). Subcellular fractionation from synchronized MCF7 cells was then performed to obtain a nucleocytoplasmic soluble fraction (Sol.) and a chromatin-bound fraction (Chrom.) [55]. Efficiency of fractionation was checked by Western blot using the G1-phase Cyclin D1 and GAPDH which are good markers for the soluble fraction (Fig. 1c). In contrast, as expected, the licensing factors Cdc6 and, to a lesser extent Cdt1, are exclusively detected in the chromatin-bound fraction [70, 71] (Fig. 1c). Interestingly, OGT is more abundant in the chromatin-bound fraction than in the soluble one, while OGA is only detected in the soluble one (Fig. 1c). Each MCM subunit was then detected by Western blot in the two subcellular fractions. Except for MCM2 and MCM7 which are equally detected in both fractions, the four other MCM subunits are preferentially located in the chromatin-bound fraction (Fig. 1d, Input). Then, to assess the O-GlcNAcylation of MCM proteins, we enriched O-GlcNAc proteins on succinylated WGA (sWGA)-agarose beads before Western blotting [65]. Negative controls in the presence of free GlcNAc were performed to confirm the specificity of the binding of GlcNAc-bearing proteins to the lectin (Fig. 1d, lanes C−). Although the six MCM subunits are present in both fractions, sWGA-bound MCM proteins were detected only in the chromatin-bound fraction (Fig. 1d). The same results were obtained with synchronized MDA-MB-231 cells (Suppl. Figure 1a–c). Our results demonstrate that O-GlcNAc-modified MCM proteins are stably loaded onto chromatin.
To ascertain that each chromatin-bound MCM subunit was individually and directly O-GlcNAcylated, we used a click chemistry approach to enrich and detect O-GlcNAcylated proteins from the chromatin fraction of quiescent and S phase-synchronized MCF7 cells. With this approach, SDS (1%) was used in the solubilization buffer, allowing dissociation and denaturation of protein complexes. O-GlcNAcylated proteins were enzymatically labelled with a GalNAz residue before the click chemistry reaction with the biotin–alkyne probe, allowing the enrichment of O-GlcNAcylated proteins on avidin–agarose beads [66]. The presence of MCM proteins in click-labelled samples was confirmed by the signal detected in the input (Input Click-It) (Fig. 1e). Western blotting of the avidin-bound proteins with anti-MCMs antibodies allowed us to confirm that all the six MCM2–7 subunits are strongly O-GlcNAcylated in the chromatin fraction of S phase-synchronized cells in comparison with quiescent cells (18H versus 0) (Fig. 1e).
Then to evaluate the number of O-GlcNAc site and the stoichiometry of O-GlcNAcylated isoforms of MCM proteins [67], we performed the chemoenzymatic labelling of O-GlcNAc proteins with UDP-GalNAz and GalT1 followed by the click reaction to conjugate a 4.4 kDa DBCO-PEG mass tag to the labelled glycoproteins. After SDS-PAGE separation, immunoblotting with MCM antibodies enabled the detection of both the non-glycosylated and glycosylated forms of MCM (indicated with arrows) from the chromatin fraction of asynchronous MCF7 cells (Fig. 1f). Using this approach, we were able to detect two sites on MCM2 and MCM4 (corresponding to a shift of an estimated MW of 10.1 ± 1.5 and 9.3 ± 1.1 kDa, respectively), the glycosylated forms representing less than 3% of both MCMs (Fig. 1f). For MCM6, we observed two distinct glycosylated species with two and three O-GlcNAc sites, representing 10.9 and 2.3% of MCM6 subunit, respectively. For MCM7, only one glycosylated form with three O-GlcNAc sites (ΔMW ≈ 12.3 kDa) was detected, representing nearly 9% of the protein. However, MCM proteins glycosylated at more than two or three sites may also exist in cells. They might be undetectable by immunoblotting due to steric hindrance of PEG molecules which could mask the epitopes and/or are stoichiometrically lower than the limit of detection.
OGT stably interacts with several subunits of the MCM2–7 complex
The progressive and timely regulated assembly of MCM proteins with their partners is very important for the regulation of MCM2–7 stability and chromatin loading [29, 42]. To investigate whether OGT tightly interacts with the MCM2–7 subunits, we first performed a GST pull-down assay, using recombinant GST-tagged OGT immobilized onto Glutathione Sepharose beads as the bait [72]. Soluble and chromatin-bound extracts from asynchronous MCF7 cells were used as the source of prey proteins. The same experiment was conducted with GST as the negative control. After extensive washes and elution of proteins from the matrix with free glutathione, we first checked that GST and GST–OGT were efficiently eluted from Glutathione Sepharose beads using an anti-GST tag antibody, also confirmed by reprobing the membranes with an anti-OGT (Fig. 2a, lower panel). Then MCM proteins were detected by Western blot in the glutathione-eluted fractions; cellular extracts from both fractions were used as positive controls for the detection of endogenous MCM proteins (Fig. 2b, lanes Inp). Except for MCM4 that does not seem to bind to GST–OGT, both soluble and chromatin-bound MCM2/3/5/7 were able to interact with GST–OGT and not GST (Fig. 2b). For MCM6, we could observe a signal in the GST–OGT lane only in the chromatin-bound fraction. However, a faint band was also revealed in the negative control (GST alone), suggesting that MCM6 may also interact weakly with the GST tag in an unspecific manner (Fig. 2b). The same experiment was conducted with MDA-MB-231 cellular extracts in which we could observe the interaction between GST–OGT and MCM3, MCM5 and MCM7 in the two subcellular fractions (Suppl. Figure 1d). When we conducted the same experiment by adding 0.1% SDS in the MCF7 cellular extracts before incubation with beads, no signal was detected for any MCM in the GST–OGT lanes, indicating that SDS may denaturate the recombinant GST–OGT protein (data not shown). As we used here soft experimental conditions for the incubation, washing and elution buffers, we cannot exclude that eluted MCM proteins bind to OGT in an indirect manner due to the presence of MCM2–7 hexameric complexes and MCM proteins sub-complexes [31, 34, 37].
OGT stably interacts with several subunits of MCM2–7 complex. a GST pull-down assay using OGT–GST and GST (negative control) was performed using soluble (Sol.) and chromatin-bound (Chrom.) protein fractions from asynchronous MCF7 cells (Inp). Western blot analysis was performed using anti-GST and anti-OGT antibodies. b Eluted MCM proteins were detected by Western blot after GST pull-down assay. c, d OGT and subunits of MCM2–7 were immunoprecipitated from whole cell extracts of HEK293T cells transiently transfected with HA-OGT. Co-immunoprecipitated proteins were detected by Western blot using the indicated antibodies. e In situ proximity ligation assay was performed in fixed asynchronous MCF7 cells shortly permeabilized with 0.5% T-X100 in PBS to visualize interaction of endogenous OGT with MCM3, MCM4, MCM6 or MCM7. Nuclei were counterstained with DAPI and negative controls were performed by incubating fixed cells with only one of the primary antibodies (MCM-neg and OGT-neg). Quantification of PLA is presented as scatter dot plot; each dot represents the total signal of PLA in the nucleus of a single cell. Bars represents the median with interquartile range for each experience (one-way ANOVA test, *p < 0.05). Scale bar, 30 µM
Then we conducted co-immunoprecipitation experiments to ascertain the interaction of OGT with MCM proteins in living cells. We first tried in MCF7 cells but we could not detect any co-immunoprecipitation of MCM proteins with endogenous OGT, corroborating the low OGT-bound/OGT-unbound MCM ratio that we observed using the GST pull-down approach (Fig. 2b). We used transitory OGT-transfected HEK293T cells and performed immunoprecipitation of either OGT or MCM from whole cell extracts. Western blot analysis showed that MCM3, MCM6 and MCM7 co-immunoprecipitate with OGT (Fig. 2c). Reverse IP allowed us to confirm that OGT interacts with MCM3 and MCM7 and, to a weaker extent, with MCM6 (Fig. 2d).
We next performed Proximity Ligation Assay (PLA) experiments in asynchronous cells, using antibodies against OGT, MCM3, MCM4, MCM6 and MCM7 for which we could validate their use in indirect immunofluorescence (data not shown). The specificity of PLA signal between OGT and MCM was confirmed by doing control experiments with only one of the primary antibodies followed by the incubation with both minus and plus PLA probes (Fig. 2e, lower panel). We observed strong PLA fluorescent signal in nuclei for OGT–MCM3, OGT–MCM6 and OGT–MCM7, in agreement with our GST pull-down and co-IP results (Fig. 2b–d). In contrast, the signal obtained for OGT–MCM4 was not significantly different from the MCM4-negative control (Fig. 2e), indicating that OGT does not stably interact with MCM4, as concluded by our co-IP results (Fig. 2c, d). It is important to note that we had to reduce the time of cell permeabilization to detect OGT–MCM interactions by PLA (2 min in 0.5% Triton X-100 instead of 20 min for the detection of MCM–MCM interactions by PLA, see Fig. 4b). This highlights that OGT is indirectly recruited to the chromatin via stable interaction with DNA-binding factors and chromatin effectors [4, 10, 72], while MCM proteins strongly associate with DNA [28, 29]. Altogether our results indicate that OGT is a new partner of MCM2–7 complex through its direct binding with MCM3, MCM6 and MCM7 subunits.
O-GlcNAcylation does not affect MCM steady-state levels but impacts the loading of MCM2, MCM6 and MCM7 to the chromatin
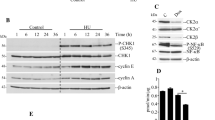
To test whether OGT and O-GlcNAc dynamics could regulate the binding of MCM proteins to chromatin, we induced silencing of either OGT or OGA by small interfering RNA (siRNA). Efficiency of OGT (siOGT) and OGA (siOGA) silencing was determined by Western blot against both enzymes and O-GlcNAcylated proteins (O-GlcNAc), and compared with random silencing (siCtrl) (Fig. 3a, b). As previously reported, OGT silencing induces a strong decrease in OGA protein level. Conversely, OGA silencing induces a moderate decrease in OGT protein level (Fig. 3a, b) [19, 73, 74]. Efficiency of the subcellular fractionation was attested by Western blot with anti-GAPDH antibody as a control for the soluble fraction, and anti-Cdc6 antibody for the chromatin-bound one (Fig. 3b). We did not observe any change in the expression of the six MCM2–7 subunits in the WCL when O-GlcNAcylation levels were disturbed (Fig. 3a–c), indicating that O-GlcNAc homeostasis may not regulate the steady-state level of MCM proteins.
Down-regulation of OGT decreases the chromatin loading of MCM2, MCM6 and MCM7. MCF7 cells were treated with siRNA (Ctrl, OGT, OGA) for 60 h before harvesting. a The whole cellular extracts (WCL) and b the soluble (Sol.) and chromatin-bound (Chrom.) protein fractions were analysed by Western blot using anti-OGT, -OGA and O-GlcNAc proteins antibodies to confirm the efficiency of siRNA transfection. MCM proteins were also detected by Western blot using specific antibodies. Equal loading was confirmed using GAPDH antibodies (WCL) and Ponceau staining of the membranes (Sol./Chrom.). GAPDH and Cdc6 were used as markers for the soluble and chromatin fractions, respectively. c Band intensity was quantitated using Image J; the relative intensity of each MCM protein level in siOGT and siOGA conditions was normalized to that obtained for the siCtrl condition (100%) and depicted as a graph. Statistical analysis was performed by Student’s t test. Values are mean ± SEM of at least 4 independent experiments (**p < 0.05, *p < 0.1)
Western blot analysis of soluble and chromatin-bound fractions showed that chromatin association of MCM3, MCM4, MCM5 was not affected by OGT or OGA silencing (Fig. 3b, c). In contrast, down-regulation of OGT slightly decreased the association of MCM6, MCM7, and to a lesser extent MCM2, with chromatin, while OGA silencing had no significant impact (Fig. 3b, c). For MCM7, this was accompanied by a moderate increase in the soluble fraction, suggesting that OGT may contribute to the loading or the stabilization of MCM7 onto chromatin. This hypothesis is reinforced by the strong interaction between OGT and MCM7 that we showed here for the first time (Fig. 2). We showed here a decrease in the chromatin loading of MCM2 when OGT was silenced, despite that no direct interaction between both proteins could be evidenced (Fig. 2c–e). MCM2 weakly interacts with the other MCM subunits and negatively regulates the helicase activity of the MCM4/6/7 sub-complex in vitro [31, 37, 40, 75]. Our results suggest that OGT might indirectly regulate the MCM2–7 complex via the regulation of MCM2–MCM interactions and the chromatin binding of MCM2.
Perturbation of O-GlcNAc cycling destabilizes MCM2–7 complex
The MCM2–7 helicase complex is a heterohexameric complex but sub-complexes containing MCM2/4/6/7, MCM4/6/7 or MCM3/5 have also been isolated in mammalian cells [31,32,33, 36]. Furthermore, the proper association of MCM subunits is essential for the establishment of active ATPase sites necessary for the helicase activity of the complex [27]. On the other hand, O-GlcNAcylation can alter protein–protein interactions [7]. Then we wondered whether MCM O-GlcNAcylation and/or OGT–MCM interaction could be involved in MCM–MCM interactions. To address this question, O-GlcNAc levels were reduced by OGT silencing, and co-IP and PLA experiments were performed to analyse MCM–MCM interactions in the chromatin-bound protein fraction (Fig. 4). OGT down-regulation induced a slight decrease of co-immunoprecipitation of MCM6 with MCM2, and MCM7 with MCM4 (Fig. 4a). In situ PLA confirmed these results: nuclear PLA signals between MCM2/MCM6 and MCM4/MCM7 were significantly lower in siOGT-transfected cells compared with those in siCtrl-transfected cells (Fig. 4b). In contrast, interaction between MCM4 and MCM6 tended towards a decrease in siOGT-cells, while differences in intensity of both co-IP and PLA fluorescence signals were not statistically significant between siCtrl and siOGT conditions (Fig. 4a, b, middle panels). To confirm that the decrease in MCM/MCM interactions that we observed in siOGT cells was due to the downregulation of OGT, we performed a rescue experiment by transfecting siOGT cells with a plasmid coding for a FLAG-tagged and siRNA-resistant OGT (3X-OGT-RSI). As shown in Fig. 4c, PLA signals obtained for both MCM2/MCM6 and MCM4/MCM7 were significantly restored when OGT was overexpressed in synchronized MCF7 cells, indicating that the presence of OGT may be important to maintain such interactions. To determine whether this could be related to a defect in O-GlcNAcylation of MCM proteins, we next performed immunoprecipitation of these four subunits from the chromatin-bound fractions of synchronized MCF7 cells and revealed their O-GlcNAc status by Western blot. Interestingly, the disturbance in MCM/MCM interactions upon OGT silencing is concomitant with a decrease of the O-GlcNAcylation level of MCM2, MCM4, MCM6 and MCM7 in the chromatin-bound fraction (Fig. 4d).
Silencing of OGT affects MCM2/MCM6 and MCM4/MCM7 interactions. MCF7 cells were transfected with siRNA (siCtrl, siOGT), then synchronized in S phase by stimulation with serum (18H) after serum starvation. a Chromatin-bound protein fraction was used for immunoprecipitation of MCM2 and MCM4 and Western blot analysis of co-immunoprecipitated MCM subunits. Band signal intensity was measured and statistical analysis was performed by Student’s t test. Values are mean ± SEM of three independent experiments (**p < 0.05). b MCM–MCM interactions were detected by in situ PLA and immunofluorescent confocal microscopy after nuclei counterstaining with DAPI. Quantification was performed as in Fig. 2e. Bars represents the median with interquartile range (one-way ANOVA test, *p < 0.05). Scale bar, 15 µM. c pCMV-3X or 3X-OGT-RSI plasmids were transfected in siOGT MCF7 cells for 24 h. Then cells were synchronized in S phase and MCM–MCM interactions were detected by in situ PLA, as described above (t test, ***p < 0.001). Scale bar, 20 µM. d MCM2, MCM4, MCM6 and MCM7 were immunoprecipitated from the chromatin-bound protein fractions and Western blots were revealed using the anti-O-GlcNAc antibody (RL2). Membranes were stripped and reincubated with the corresponding anti-MCM antibodies
Then we perturbed O-GlcNAc cycling in synchronized MCF7 cells using either 5S-G or ThG to inhibit the catalytic activity of OGT or OGA, respectively [63, 64] (Suppl. Figure 2). Surprisingly, we were not able to detect any significant and reproducible changes in MCM2/MCM6 and MCM4/MCM7 interactions in 5S-G-treated cells, as shown by co-IP and PLA experiments (Fig. 5a, b). However, both approaches showed a moderate decrease in MCM2/MCM6 interaction in the chromatin-bound fraction when O-GlcNAc levels were increased through OGA inhibition by ThG treatment (Fig. 5a, b). It is important to note that we could not detect MCM2 when MCM6 was immunoprecipitated (data not shown), probably due to the weak association of MCM2 with MCM6 [40]. Moreover, MCM4/MCM6, MCM4/MCM7, and MCM3/MCM5 interactions did not seem to be sensitive to OGT inhibition (Fig. 5c–e). Co-immunoprecipitation of MCM3 and MCM5 was also not significantly perturbed by ThG (Fig. 5e). However, PLA approach shows that inhibition of OGA induced a moderate decrease in MCM4/MCM7 interactions (Fig. 5c), whereas it tended to increase MCM4/MCM6 co-immunoprecipitation albeit in a non-significant manner (Fig. 5d), that we could confirm by in situ PLA (data not shown).
Hyper-O-GlcNAcylation induced by OGA inhibition alters MCM/MCM interactions. Serum-starved MCF7 cells were released in S phase by serum addition (18H), in presence of DMSO (Ctrl), 5S-G or ThG, before harvesting. a, d MCM proteins were immunoprecipitated from the chromatin-bound fraction and samples were analysed by Western blot using the indicated antibodies. Band signal intensity was measured and statistical analysis was performed by Student’s t test. Values are mean ± SEM of three independent experiments (**p < 0.05). b, c MCM–MCM interactions were detected by in situ PLA and immunofluorescent confocal microscopy. Nuclei were counterstained with DAPI. Quantification was performed as in Fig. 2e. Bars represent the median with interquartile range (one-way ANOVA test, *p < 0.05). Scale bar, 10 µM
Effect of perturbation of O-GlcNAc cycling on S phase progression and DNA replication
We next determined whether dysregulation in O-GlcNAc homeostasis would affect DNA replication and S phase progression. We used synchronized MCF7 cells that were either transfected by siRNA or treated with potent inhibitors of OGT and OGA. After serum stimulation, cells were harvested at different time points and cell cycle progression was analysed by flow cytometry after staining with PI. DNA replication rate was also evaluated by labelling nascent DNA within the last 15 min of serum stimulation with EdU. After click chemistry reaction and PI staining, the percentage of EdU-positive cells was determined by flow cytometry. In parallel, cells were seeded and treated with either siRNA or inhibitors, and counted every day during 5 days.
For each experiment, the efficiency of treatments was systematically confirmed by Western blotting (Suppl. Figure 3). It is noteworthy that the decrease in O-GlcNAcylation levels induced by 5S-G was accompanied by a significant decrease in OGA expression, whereas elevation of O-GlcNAcylation levels induced by ThG treatment was accompanied by a decrease in OGT expression and an increase of OGA protein (Suppl. Figure 3a, see also Suppl. Figure 2). Similarly, as we observed above (Fig. 3), silencing of OGT strongly decreased OGA protein level. Conversely silencing of OGA decreased the levels of OGT, albeit to a lesser extent (Suppl. Figure 3b).
As shown in Fig. 6a, inhibition of OGT by 5S-G induced a moderate decrease in the percentage of MCF7 cells in S phase compared with control condition (35.6 versus 43% at 15H; 32 versus 52% at 24H). However, this was not correlated with changes on the rate of EdU incorporation (Fig. 6b), although the cellular growth of 5S-G-treated cells was significantly reduced by 30% after 5 days of treatment (Fig. 6c). Likewise, inhibition of OGA did not modify the DNA replication rate but tended towards a slowdown in S phase progression; this was accompanied with a decrease of 25% of proliferation after 5 days of treatment (Fig. 6a–c). Surprisingly, S phase progression of siOGT- and siOGA-transfected MCF7 cells was not significantly affected compared with that of control siRNA-transfected cells, nor was the rate of DNA replication (Fig. 6d, e). In agreement with these results, no difference in cellular growth was observed between siCtrl, siOGT and siOGA conditions in MCF7 cells (Fig. 6f).
Perturbation of O-GlcNAc cycling does not affect DNA replication rate in MCF7 cells. Serum-starved MCF7 cells were released into the cell cycle by serum addition for the indicated times in the presence of OGT and OGA inhibitors (5S-G, ThG) or DMSO (Ctrl). a Cell cycle distribution was determined by flow cytometry after DNA staining with PI. Percentage of cells in S phase is indicated for each time point. Results are representative of three independent experiments. b EdU was added during the last 15 min of serum stimulation to label nascent DNA using the Click-It EdU flow cytometry assay, and DNA content was stained with PI before flow cytometry analysis. Percentage of EdU-positive cells is indicated for each time point. Results are representative of three independent experiments. c One day after seeding, MCF7 cells were treated with DMSO (Ctrl), 5S-G or ThG. Cells were counted every day for 5 days and medium was replaced every 2 days (with inhibitors). Values are mean ± SEM of three independent experiments. d MCF7 cells were transfected with siRNA (siCtrl, siOGT, siOGA) and then synchronized as in a. Results are representative of three independent experiments. e MCF7 cells were treated as above. Percentage of EdU-positive cells was obtained as in b. Results are representative of three independent experiments. f MCF7 cells were transfected with siRNA (siCtrl, siOGT, siOGA). Cells were counted every day for 5 days and medium was replaced every 2 days (with siRNA). Values are mean ± S.E.M. of three independent experiments
Discussion
The heterohexameric MCM2–7 helicase complex is crucial for the initiation of DNA replication and is finely regulated by post-translational modifications, including phosphorylation. We previously showed that MCM3, MCM6 and MCM7 were O-GlcNAcylated by OGT [22]. Here we demonstrate that MCM2, MCM4 and MCM5 are also O-GlcNAcylated in human cells (Fig. 1a, b). This result is strengthened by two recent works which have identified MCM2, MCM3, MCM4 and MCM5 as O-GlcNAc-modified proteins using metabolic incorporation of chemical GlcNAc analogue probes combined with click chemistry labelling [76, 77]. Using a mass-tagging strategy, we show that the O-GlcNAcylated MCM proteins are of low stoichiometry (from less than 3% for MCM2 and MCM4, to 13% for MCM6) (Fig. 1f), as it has been evidenced for many O-GlcNAc-modified proteins [78, 79]. We could detect 2 O-GlcNAc sites on MCM2 and MCM4, 2 and 3 O-GlcNAc sites on MCM6, and 3 O-GlcNAc sites on MCM7, although we cannot exclude that some of the O-GlcNAc sites may not have been detectable using this approach. Many phosphorylation sites have been characterized on MCM proteins, especially within the N-terminal extension of MCM2 and MCM4; some of them are known to play crucial roles in regulating MCM2–7 loading and helicase activity [53,54,55,56,57,58, 60,61,62]. It is tempting to speculate that O-GlcNAc modification may cooperate or compete with phosphorylation to regulate MCM2–7 complex. Therefore, we performed the in silico analysis of the potential O-GlcNAc/phosphorylation crosstalk according to the recently published stringent motif (S/T)-P-(V/A/T)-(gT/gS) [14]. We also used the YinOYang1.2 server to predict the O-GlcNAc sites on MCM proteins, by taking into account only the high-scoring potential O-GlcNAc sites (http://www.cbs.dtu.dk/services/YinOYang). Such prediction tool has to be taken with caution and experimental data are required to ascertain their localization. Using the human primary sequences of MCM2 to MCM7 proteins of the UniProtKB database, we found the specific crosstalk motif [611SPVT614] within the sensor 2 subdomain of MCM3, which is adjacent to the potential predicted O-GlcNAc site at Thr610. It is noteworthy that phosphorylation of Ser611 of MCM3 has been identified by mass spectrometry in human leukaemia cells, but to date, no functional role has been assigned to this residue [80]. For MCM4, we found the motif [3SPAS6] in the N-terminal extension of which is known to be phosphorylated at multiple sites (http://www.PhosphoSitePlus) [53, 56]. Interestingly, three highly potential O-GlcNAc sites are located within the same N-terminal region of MCM4, at position Ser2, Ser3 and Thr7, suggesting that the N-terminal tail of MCM4 may be targeted by both kinases and OGT to regulate MCM2–7 helicase complex. On the other hand, for MCM5 subunit, we found the degenerated crosstalk motif [133SPSS136], which is located at the hinge of two structural subdomains of the N-terminal domain (NTD) of MCM5, NTD-A (A subdomain of NTD, amino acids 32–129) and oligonucleotide/oligosaccharide-binding (OB)-fold (amino acids 136–172) (http://www.ebi.ac.uk/interpro/InterPro) [28]. It is of interest that the two predicted O-GlcNAc sites of MCM5, Ser135 and Ser136, are adjacent to Ser133 which is a phosphorylation site with unknown function [81]. Although no such O-GlcNAc-phospho crosstalk motif has been found for MCM2, the potential O-GlcNAc site Thr25 is close to Ser27 which is phosphorylated by Cdc7/Dbf4 and takes part in the initiation of DNA replication [54, 59]. This in silico analysis strongly suggests that a crosstalk may occur between O-GlcNAcylation and phosphorylation on MCM proteins, adding a layer of complexity to finely regulate MCM2–7 complex. Further mass spectrometry analysis is now required to unambiguously identify the glycosylation sites of human MCM proteins and experimentally test this hypothesis.
Our results show that OGT glycosylates the six subunits of MCM2–7 complex when quiescent cells progress normally in S phase, and that O-GlcNAcylated MCM proteins are present nearly exclusively in the chromatin fraction (Fig. 1d, e). This indicates that O-GlcNAcylation of MCM2–7 complex may occur mostly when MCM proteins are loaded onto chromatin. Another possible explanation to the lack of O-GlcNAc-enriched MCM proteins in the soluble fraction is that the turnover of MCM protein O-GlcNAcylation may be higher in the nucleoplasmic soluble fraction in which both OGT and OGA were present, than in the chromatin-bound fraction in which we could not detect OGA (Figs. 1c, 3b, Suppl. Figure 2). O-GlcNAcylation of MCM proteins might help in the recruitment of MCM2–7 complex onto the chromatin, as described for the phosphorylation of MCM2 and MCM3 [54, 60]. In contrast, for MCM4, O-GlcNAc modification could act in an opposite manner to phosphorylation, since highly phosphorylated MCM4 is less tightly bound to chromatin than underphosphorylated form of MCM4 [31].
On the other hand, we demonstrate that OGT strongly interacts with distinct MCM2–7 subunits in human cells. Indeed, biochemical and in situ approaches show that OGT is a new partner of MCM2–7 complex through its direct binding with MCM3, MCM6 and MCM7 subunits (Fig. 2). Our findings are in agreement with previous studies that demonstrate that the recruitment of MCM2–7 partners occurs via their specific interaction with one or two MCM subunits, as it has been reported for the interactions of the licensing factor Cdt1 with MCM6 [82] and Cdc45 with MCM2 and MCM5 [43]. It has also been shown for the DDK subunits: whereas Dbf4 strongly binds to MCM2, Cdc7 interacts with both MCM4 and MCM5 subunits [83]. Our data suggest that OGT might regulate the chromatin loading of MCM6 and MCM7 through this strong interaction. Indeed, down-regulation of OGT protein level by siRNA decreases the chromatin-bound level of MCM6 and MCM7 (Fig. 3b). This hypothesis is reinforced by the destabilization of MCM2/MCM6 and MCM4/MCM7 interactions when OGT is silenced in synchronized MCF7 cells (Fig. 4a–c). Given the fact that we did not observe significant changes in these interactions when OGT is inhibited by 5S-G (Fig. 5a–c) whereas OGT rescue experiment in siOGT-transfected cells could restore them (Fig. 4c), we hypothesize that in normal culture conditions, OGT protein might be more important than its catalytic activity and act as a scaffold protein to regulate such interactions and recruit MCM2–7 complexes to the chromatin through its direct binding to MCM6 and MCM7 subunits. Moreover, our data show that the destabilization of the interactions between MCM2/MCM6 and MCM4/MCM7 are concomitant with a decrease in the O-GlcNAcylation levels of these MCM subunits (Fig. 4d). This suggests that O-GlcNAc modification of MCM proteins may be also involved in the stabilization of MCM2–7 complex onto chromatin. Since this glycosylation is known to stabilize protein–protein interactions [4, 6], we would have expected an elevation of MCM/MCM interactions upon OGA inhibition. But it did not, since the increase of the overall O-GlcNAcylation levels in ThG-treated cells slightly downregulated the binding of MCM2/MCM6 and MCM4/MCM7 in S phase-synchronized MCF7 cells (Fig. 5b, c). Altogether, our data suggest that O-GlcNAc homeostasis might contribute to stabilize MCM–MCM interactions. Further investigations are needed to decipher the molecular mechanisms underlying the regulation of the chromatin loading and stabilization of MCM2–7 complex through the recruitment of OGT and O-GlcNAcylation of the MCM2–7 complex subunits.
Here we perturbed O-GlcNAc cycling using either siRNA or selective and potent inhibitors of the O-GlcNAc enzymes, OGT and OGA. As previously reported, in case of disruption to O-GlcNAc homeostasis, cells adjust OGT and OGA expression to compensate for the changes in O-GlcNAc levels, with OGA protein expression being more sensitive to O-GlcNAc homeostasis than OGT. Reduction in O-GlcNAc levels induces a decrease in OGA protein level and reciprocally, elevation of O-GlcNAcylation induces a decrease in the expression of OGT, (Fig. 3, Suppl. Figures 2, 3) [16, 19, 22, 64, 73, 74, 84,85,86]. Similar results were obtained not only in MDA-MB-231 breast cancer cells, but also in the colorectal cell lines HCT116, HT29 and CCD841CoN (unpublished data). This mutual regulation of OGT and OGA to compensate the loss of O-GlcNAc homeostasis is far to be fully understood. To date, no mechanism has been established that can explain how OGA protein is as much downregulated when OGT is inhibited or silenced. However, Zhang and collaborators have shown that Thiamet G increases the transcription of OGA mRNA [84]. Moreover, it has been recently demonstrated that OGT mRNA levels are controlled by an intron splicing silencer (ISS) that induces the nuclear degradation of the mRNA under high O-GlcNAcylation conditions, thus allowing a decrease in OGT protein levels [85]. This may explain the lower levels of OGT that we and others have observed upon inhibition or silencing of OGA. In contrast, an overall decrease of O-GlcNAcylation upon OGT inhibition induces an efficient splicing of OGT mRNA to produce a cytoplasmic mRNA that will be further translated [85].
Neither silencing nor inhibition of the O-GlcNAc-regulating enzymes perturbs significantly the rate of DNA synthesis in synchronized human MCF7 cells (Fig. 6b, e). Nevertheless, the S phase progression and growth rate slow down when OGT is inhibited and to a lesser extent when OGA is inhibited (Fig. 6a, c). Thus, the delay in S phase entry induced by OGT or OGA inhibition may not be related to a defect in DNA synthesis in MCF7 cells. It could be due to an abnormal activity or expression of cell cycle-related proteins that are known to be directly or indirectly regulated by OGT, like transcription factors, Cyclin/CDK, or cell cycle inhibitors [15, 21, 22, 87]. Collectively, our results indicate that the effects of OGT down-regulation on MCM proteins that we report here for the first time are not sufficient to disrupt S phase progression and DNA synthesis in human MCF7 cells. Although we assessed the DNA helicase activity in an indirect manner through the measurement of Edu incorporation, our results are consistent with previous work demonstrating that an acute down-regulation of each one of the MCM2–7 subunits by silencing approach does not slow down the replication fork speed during DNA elongation in human cells [50]. Therefore, we believe that when cells normally progress in S phase, perturbation of O-GlcNAc cycling may destabilize MCM/MCM interactions but without interfering with MCM2–7 helicase activity and DNA replication. The possible explanation is that a large excess of MCM2–7 complexes is loaded to chromatin during G1 phase to license dormant replication origins that are not used during normal DNA replication but are required under conditions of replicative stress to maintain genome integrity [48,49,50]. This hypothesis is reinforced by recent studies highlighting that OGT relocates to the sites of DNA damage and targets key signalling proteins and DNA polymerase η in response to DNA damage [13, 88, 89]. In conclusion, our work demonstrates that OGT is a new partner of MCM2–7 complex and regulates MCM–MCM interactions. Although further investigations are needed to investigate the molecular mechanisms in detail, it opens up new prospects for the role of OGT and O-GlcNAc post-translational modification in the regulation of DNA replication under conditions of unperturbed replication as well as replicative stress.
References
Kreppel LK, Blomberg MA, Hart GW (1997) Dynamic glycosylation of nuclear and cytosolic proteins. Cloning and characterization of a unique O-GlcNAc transferase with multiple tetratricopeptide repeats. J Biol Chem 272:9308–9315
Love DC, Kochan J, Cathey RL et al (2003) Mitochondrial and nucleocytoplasmic targeting of O-linked GlcNAc transferase. J Cell Sci 116:647–654
Hanover JA, Yu S, Lubas WB et al (2003) Mitochondrial and nucleocytoplasmic isoforms of O-linked GlcNAc transferase encoded by a single mammalian gene. Arch Biochem Biophys 409:287–297
Yang X, Qian K (2017) Protein O-GlcNAcylation: emerging mechanisms and functions. Nat Rev Mol Cell Biol 18:452–465. https://doi.org/10.1038/nrm.2017.22
Gao Y, Wells L, Comer FI et al (2001) Dynamic O-glycosylation of nuclear and cytosolic proteins: cloning and characterization of a neutral, cytosolic beta-N-acetylglucosaminidase from human brain. J Biol Chem 276:9838–9845. https://doi.org/10.1074/jbc.M010420200
Nagel AK, Ball LE (2015) Intracellular protein O-GlcNAc modification integrates nutrient status with transcriptional and metabolic regulation. Adv Cancer Res 126:137–166. https://doi.org/10.1016/bs.acr.2014.12.003
Hart GW, Slawson C, Ramirez-Correa G, Lagerlof O (2011) Cross talk between O-GlcNAcylation and phosphorylation: roles in signaling, transcription, and chronic disease. Annu Rev Biochem 80:825–858. https://doi.org/10.1146/annurev-biochem-060608-102511
Bond MR, Hanover JA (2013) O-GlcNAc cycling: a link between metabolism and chronic disease. Annu Rev Nutr 33:205–229. https://doi.org/10.1146/annurev-nutr-071812-161240
Hardivillé S, Hart GW (2014) Nutrient regulation of signaling, transcription, and cell physiology by O-GlcNAcylation. Cell Metab 20:208–213. https://doi.org/10.1016/j.cmet.2014.07.014
Leturcq M, Lefebvre T, Vercoutter-Edouart A-S (2017) O-GlcNAcylation and chromatin remodeling in mammals: an up-to-date overview. Biochem Soc Trans 45:323–338. https://doi.org/10.1042/BST20160388
Wang Z, Pandey A, Hart GW (2007) Dynamic interplay between O-linked N-acetylglucosaminylation and glycogen synthase kinase-3-dependent phosphorylation. Mol Cell Proteom 6:1365–1379. https://doi.org/10.1074/mcp.M600453-MCP200
Wang Z, Udeshi ND, Slawson C et al (2010) Extensive crosstalk between O-GlcNAcylation and phosphorylation regulates cytokinesis. Sci Signal 3:ra2. https://doi.org/10.1126/scisignal.2000526
Zhong J, Martinez M, Sengupta S et al (2015) Quantitative phosphoproteomics reveals crosstalk between phosphorylation and O-GlcNAc in the DNA damage response pathway. Proteomics 15:591–607. https://doi.org/10.1002/pmic.201400339
Leney AC, El Atmioui D, Wu W et al (2017) Elucidating crosstalk mechanisms between phosphorylation and O-GlcNAcylation. Proc Natl Acad Sci USA. https://doi.org/10.1073/pnas.1620529114
O’Donnell N, Zachara NE, Hart GW, Marth JD (2004) Ogt-dependent X-chromosome-linked protein glycosylation is a requisite modification in somatic cell function and embryo viability. Mol Cell Biol 24:1680–1690
Slawson C, Zachara NE, Vosseller K et al (2005) Perturbations in O-linked beta-N-acetylglucosamine protein modification cause severe defects in mitotic progression and cytokinesis. J Biol Chem 280:32944–32956. https://doi.org/10.1074/jbc.M503396200
Slawson C, Lakshmanan T, Knapp S, Hart GW (2008) A mitotic GlcNAcylation/phosphorylation signaling complex alters the posttranslational state of the cytoskeletal protein vimentin. Mol Biol Cell 19:4130–4140. https://doi.org/10.1091/mbc.E07-11-1146
Sakabe K, Hart GW (2010) O-GlcNAc transferase regulates mitotic chromatin dynamics. J Biol Chem 285:34460–34468. https://doi.org/10.1074/jbc.M110.158170
Kazemi Z, Chang H, Haserodt S et al (2010) O-linked beta-N-acetylglucosamine (O-GlcNAc) regulates stress-induced heat shock protein expression in a GSK-3beta-dependent manner. J Biol Chem 285:39096–39107. https://doi.org/10.1074/jbc.M110.131102
Yang YR, Song M, Lee H et al (2012) O-GlcNAcase is essential for embryonic development and maintenance of genomic stability. Aging Cell 11:439–448. https://doi.org/10.1111/j.1474-9726.2012.00801.x
Olivier-Van Stichelen S, Drougat L, Dehennaut V et al (2012) Serum-stimulated cell cycle entry promotes ncOGT synthesis required for cyclin D expression. Oncogenesis 1:e36. https://doi.org/10.1038/oncsis.2012.36
Drougat L, Olivier-Van Stichelen S, Mortuaire M et al (2012) Characterization of O-GlcNAc cycling and proteomic identification of differentially O-GlcNAcylated proteins during G1/S transition. Biochim Biophys Acta 1820:1839–1848. https://doi.org/10.1016/j.bbagen.2012.08.024
Lanza C, Tan EP, Zhang Z et al (2016) Reduced O-GlcNAcase expression promotes mitotic errors and spindle defects. Cell Cycle 15:1363–1375. https://doi.org/10.1080/15384101.2016.1167297
Tan EP, Duncan FE, Slawson C (2017) The sweet side of the cell cycle. Biochem Soc Trans 45:313–322. https://doi.org/10.1042/BST20160145
Caldwell SA, Jackson SR, Shahriari KS et al (2010) Nutrient sensor O-GlcNAc transferase regulates breast cancer tumorigenesis through targeting of the oncogenic transcription factor FoxM1. Oncogene 29:2831–2842. https://doi.org/10.1038/onc.2010.41
Ding F, Yu L, Wang M et al (2013) O-GlcNAcylation involvement in high glucose-induced cardiac hypertrophy via ERK1/2 and cyclin D2. Amino Acids 45:339–349. https://doi.org/10.1007/s00726-013-1504-2
Bochman ML, Schwacha A (2009) The Mcm complex: unwinding the mechanism of a replicative helicase. Microbiol Mol Biol Rev 73:652–683. https://doi.org/10.1128/MMBR.00019-09
Li N, Zhai Y, Zhang Y et al (2015) Structure of the eukaryotic MCM complex at 3.8 Å. Nature 524:186–191. https://doi.org/10.1038/nature14685
Riera A, Barbon M, Noguchi Y et al (2017) From structure to mechanism-understanding initiation of DNA replication. Genes Dev 31:1073–1088. https://doi.org/10.1101/gad.298232.117
Samel SA, Fernández-Cid A, Sun J et al (2014) A unique DNA entry gate serves for regulated loading of the eukaryotic replicative helicase MCM2–7 onto DNA. Genes Dev 28:1653–1666. https://doi.org/10.1101/gad.242404.114
Musahl C, Schulte D, Burkhart R, Knippers R (1995) A human homologue of the yeast replication protein Cdc21. Interactions with other Mcm proteins. Eur J Biochem 230:1096–1101
Ishimi Y, Ichinose S, Omori A et al (1996) Binding of human minichromosome maintenance proteins with histone H3. J Biol Chem 271:24115–24122
Ishimi Y (1997) A DNA helicase activity is associated with an MCM4, -6, and -7 protein complex. J Biol Chem 272:24508–24513
Ritzi M, Baack M, Musahl C et al (1998) Human minichromosome maintenance proteins and human origin recognition complex 2 protein on chromatin. J Biol Chem 273:24543–24549
Sherman DA, Forsburg SL (1998) Schizosaccharomyces pombe Mcm3p, an essential nuclear protein, associates tightly with Nda4p (Mcm5p). Nucleic Acids Res 26:3955–3960
Lee JK, Hurwitz J (2000) Isolation and characterization of various complexes of the minichromosome maintenance proteins of Schizosaccharomyces pombe. J Biol Chem 275:18871–18878. https://doi.org/10.1074/jbc.M001118200
Ishimi Y, Komamura Y, You Z, Kimura H (1998) Biochemical function of mouse minichromosome maintenance 2 protein. J Biol Chem 273:8369–8375
You Z, Ishimi Y, Masai H, Hanaoka F (2002) Roles of Mcm7 and Mcm4 subunits in the DNA helicase activity of the mouse Mcm4/6/7 complex. J Biol Chem 277:42471–42479. https://doi.org/10.1074/jbc.M205769200
Schwacha A, Bell SP (2001) Interactions between two catalytically distinct MCM subgroups are essential for coordinated ATP hydrolysis and DNA replication. Mol Cell 8:1093–1104
Sakwe AM, Nguyen T, Athanasopoulos V et al (2007) Identification and characterization of a novel component of the human minichromosome maintenance complex. Mol Cell Biol 27:3044–3055. https://doi.org/10.1128/MCB.02384-06
Chuang C-H, Yang D, Bai G et al (2012) Post-transcriptional homeostasis and regulation of MCM2–7 in mammalian cells. Nucleic Acids Res 40:4914–4924. https://doi.org/10.1093/nar/gks176
Fragkos M, Ganier O, Coulombe P, Méchali M (2015) DNA replication origin activation in space and time. Nat Rev Mol Cell Biol 16:360–374. https://doi.org/10.1038/nrm4002
Simon AC, Sannino V, Costanzo V, Pellegrini L (2016) Structure of human Cdc45 and implications for CMG helicase function. Nat Commun 7:11638. https://doi.org/10.1038/ncomms11638
Todorov IT, Attaran A, Kearsey SE (1995) BM28, a human member of the MCM2–3–5 family, is displaced from chromatin during DNA replication. J Cell Biol 129:1433–1445
Krude T, Musahl C, Laskey RA, Knippers R (1996) Human replication proteins hCdc21, hCdc46 and P1Mcm3 bind chromatin uniformly before S-phase and are displaced locally during DNA replication. J Cell Sci 109(Pt 2):309–318
Prasanth SG, Prasanth KV, Siddiqui K et al (2004) Human Orc2 localizes to centrosomes, centromeres and heterochromatin during chromosome inheritance. EMBO J 23:2651–2663. https://doi.org/10.1038/sj.emboj.7600255
Kuipers MA, Stasevich TJ, Sasaki T et al (2011) Highly stable loading of Mcm proteins onto chromatin in living cells requires replication to unload. J Cell Biol 192:29–41. https://doi.org/10.1083/jcb.201007111
Woodward AM, Göhler T, Luciani MG et al (2006) Excess Mcm2–7 license dormant origins of replication that can be used under conditions of replicative stress. J Cell Biol 173:673–683. https://doi.org/10.1083/jcb.200602108
Ge XQ, Jackson DA, Blow JJ (2007) Dormant origins licensed by excess Mcm2–7 are required for human cells to survive replicative stress. Genes Dev 21:3331–3341. https://doi.org/10.1101/gad.457807
Ibarra A, Schwob E, Méndez J (2008) Excess MCM proteins protect human cells from replicative stress by licensing backup origins of replication. Proc Natl Acad Sci USA 105:8956–8961. https://doi.org/10.1073/pnas.0803978105
Courbet S, Gay S, Arnoult N et al (2008) Replication fork movement sets chromatin loop size and origin choice in mammalian cells. Nature 455:557–560. https://doi.org/10.1038/nature07233
Masai H, Matsui E, You Z et al (2000) Human Cdc7-related kinase complex. In vitro phosphorylation of MCM by concerted actions of Cdks and Cdc7 and that of a critical threonine residue of Cdc7 bY Cdks. J Biol Chem 275:29042–29052. https://doi.org/10.1074/jbc.M002713200
Ishimi Y, Komamura-Kohno Y (2001) Phosphorylation of Mcm4 at specific sites by cyclin-dependent kinase leads to loss of Mcm 4,6,7 helicase activity. J Biol Chem 276:34428–34433. https://doi.org/10.1074/jbc.M104480200
Tsuji T, Ficarro SB, Jiang W (2006) Essential role of phosphorylation of MCM2 by Cdc7/Dbf4 in the initiation of DNA replication in mammalian cells. Mol Biol Cell 17:4459–4472. https://doi.org/10.1091/mbc.E06-03-0241
Zhu Y, Ishimi Y, Tanudji M, Lees E (2005) Human CDK2 inhibition modifies the dynamics of chromatin-bound minichromosome maintenance complex and replication protein A. Cell Cycle 4:1254–1263. https://doi.org/10.4161/cc.4.9.1975
Komamura-Kohno Y, Karasawa-Shimizu K, Saitoh T et al (2006) Site-specific phosphorylation of MCM4 during the cell cycle in mammalian cells. FEBS J 273:1224–1239. https://doi.org/10.1111/j.1742-4658.2006.05146
Montagnoli A, Valsasina B, Brotherton D et al (2006) Identification of Mcm2 phosphorylation sites by S-phase-regulating kinases. J Biol Chem 281:10281–10290. https://doi.org/10.1074/jbc.M512921200
Cho W-H, Lee Y-J, Kong S-I et al (2006) CDC7 kinase phosphorylates serine residues adjacent to acidic amino acids in the minichromosome maintenance 2 protein. Proc Natl Acad Sci USA 103:11521–11526. https://doi.org/10.1073/pnas.0604990103
Chuang L-C, Teixeira LK, Wohlschlegel JA et al (2009) Phosphorylation of Mcm2 by Cdc7 promotes pre-replication complex assembly during cell-cycle re-entry. Mol Cell 35:206–216. https://doi.org/10.1016/j.molcel.2009.06.014
Lin DI, Aggarwal P, Diehl JA (2008) Phosphorylation of MCM3 on Ser-112 regulates its incorporation into the MCM2–7 complex. Proc Natl Acad Sci USA 105:8079–8084. https://doi.org/10.1073/pnas.0800077105
Francis LI, Randell JCW, Takara TJ et al (2009) Incorporation into the prereplicative complex activates the Mcm2–7 helicase for Cdc7–Dbf4 phosphorylation. Genes Dev 23:643–654. https://doi.org/10.1101/gad.1759609
Wei Q, Li J, Liu T et al (2013) Phosphorylation of minichromosome maintenance protein 7 (MCM7) by cyclin/cyclin-dependent kinase affects its function in cell cycle regulation. J Biol Chem 288:19715–19725. https://doi.org/10.1074/jbc.M112.449652
Yuzwa SA, Macauley MS, Heinonen JE et al (2008) A potent mechanism-inspired O-GlcNAcase inhibitor that blocks phosphorylation of tau in vivo. Nat Chem Biol 4:483–490. https://doi.org/10.1038/nchembio.96
Gloster TM, Zandberg WF, Heinonen JE et al (2011) Hijacking a biosynthetic pathway yields a glycosyltransferase inhibitor within cells. Nat Chem Biol 7:174–181. https://doi.org/10.1038/nchembio.520
Zachara NE (2009) Detecting the “O-GlcNAc-ome”; detection, purification, and analysis of O-GlcNAc modified proteins. Methods Mol Biol 534:251–279. https://doi.org/10.1007/978-1-59745-022-5_19
Dehennaut V, Slomianny M-C, Page A et al (2008) Identification of structural and functional O-linked N-acetylglucosamine-bearing proteins in Xenopus laevis oocyte. Mol Cell Proteom 7:2229–2245. https://doi.org/10.1074/mcp.M700494-MCP200
Clark PM, Rexach JE, Hsieh-Wilson LC (2013) Visualization of O-GlcNAc glycosylation stoichiometry and dynamics using resolvable poly(ethylene glycol) mass tags. Curr Protoc Chem Biol 5:281–302. https://doi.org/10.1002/9780470559277.ch130153
Scovassi AI, Prosperi E (2006) Analysis of proliferating cell nuclear antigen (PCNA) associated with DNA. Methods Mol Biol 314:457–475
Li J, Deng M, Wei Q et al (2011) Phosphorylation of MCM3 protein by cyclin E/cyclin-dependent kinase 2 (Cdk2) regulates its function in cell cycle. J Biol Chem 286:39776–39785. https://doi.org/10.1074/jbc.M111.226464
Coverley D, Pelizon C, Trewick S, Laskey RA (2000) Chromatin-bound Cdc6 persists in S and G2 phases in human cells, while soluble Cdc6 is destroyed in a cyclin A-cdk2 dependent process. J Cell Sci 113(Pt 11):1929–1938
Ballabeni A, Zamponi R, Caprara G et al (2009) Human CDT1 associates with CDC7 and recruits CDC45 to chromatin during S phase. J Biol Chem 284:3028–3036. https://doi.org/10.1074/jbc.M803609200
Yang X, Zhang F, Kudlow JE (2002) Recruitment of O-GlcNAc transferase to promoters by corepressor mSin3A: coupling protein O-GlcNAcylation to transcriptional repression. Cell 110:69–80
Yehezkel G, Cohen L, Kliger A et al (2012) O-linked β-N-acetylglucosaminylation (O-GlcNAcylation) in primary and metastatic colorectal cancer clones and effect of N-acetyl-β-d-glucosaminidase silencing on cell phenotype and transcriptome. J Biol Chem 287:28755–28769. https://doi.org/10.1074/jbc.M112.345546
Yang YR, Jang H-J, Yoon S et al (2014) OGA heterozygosity suppresses intestinal tumorigenesis in Apc(min/+) mice. Oncogenesis 3:e109. https://doi.org/10.1038/oncsis.2014.24
You Z, Komamura Y, Ishimi Y (1999) Biochemical analysis of the intrinsic Mcm4–Mcm6–mcm7 DNA helicase activity. Mol Cell Biol 19:8003–8015
Li J, Wang J, Wen L et al (2016) An OGA-resistant probe allow specific visualization and accurate identification of O-GlcNAc-modified proteins in cells. ACS Chem Biol. https://doi.org/10.1021/acschembio.6b00678
Chuh KN, Batt AR, Zaro BW et al (2017) The new chemical reporter 6-alkynyl-6-deoxy-GlcNAc reveals O-GlcNAc modification of the apoptotic caspases that can block the cleavage/activation of caspase-8. J Am Chem Soc 139:7872–7885. https://doi.org/10.1021/jacs.7b02213
Rexach JE, Rogers CJ, Yu S-H et al (2010) Quantification of O-glycosylation stoichiometry and dynamics using resolvable mass tags. Nat Chem Biol 6:645–651. https://doi.org/10.1038/nchembio.412
Trinidad JC, Barkan DT, Gulledge BF et al (2012) Global identification and characterization of both O-GlcNAcylation and phosphorylation at the murine synapse. Mol Cell Proteom 11:215–229. https://doi.org/10.1074/mcp.O112.018366
Zhou H, Di Palma S, Preisinger C et al (2013) Toward a comprehensive characterization of a human cancer cell phosphoproteome. J Proteome Res 12:260–271. https://doi.org/10.1021/pr300630k
Mertins P, Yang F, Liu T et al (2014) Ischemia in tumors induces early and sustained phosphorylation changes in stress kinase pathways but does not affect global protein levels. Mol Cell Proteom 13:1690–1704. https://doi.org/10.1074/mcp.M113.036392
Zhang J, Yu L, Wu X et al (2010) The interacting domains of hCdt1 and hMcm6 involved in the chromatin loading of the MCM complex in human cells. Cell Cycle 9:4848–4857. https://doi.org/10.4161/cc.9.24.14136
Ramer MD, Suman ES, Richter H et al (2013) Dbf4 and Cdc7 proteins promote DNA replication through interactions with distinct Mcm2–7 protein subunits. J Biol Chem 288:14926–14935. https://doi.org/10.1074/jbc.M112.392910
Zhang Z, Tan EP, VandenHull NJ et al (2014) O-GlcNAcase expression is sensitive to changes in O-GlcNAc homeostasis. Front Endocrinol 5:206. https://doi.org/10.3389/fendo.2014.00206
Park S-K, Zhou X, Pendleton KE et al (2017) A conserved splicing silencer dynamically regulates O-GlcNAc transferase intron retention and O-GlcNAc homeostasis. Cell Rep 20:1088–1099. https://doi.org/10.1016/j.celrep.2017.07.017
Burén S, Gomes AL, Teijeiro A et al (2016) Regulation of OGT by URI in response to glucose confers c-MYC-dependent survival mechanisms. Cancer Cell 30:290–307. https://doi.org/10.1016/j.ccell.2016.06.023
Wells L, Slawson C, Hart GW (2011) The E2F-1 associated retinoblastoma-susceptibility gene product is modified by O-GlcNAc. Amino Acids 40:877–883. https://doi.org/10.1007/s00726-010-0709-
Chen Q, Yu X (2016) OGT restrains the expansion of DNA damage signaling. Nucleic Acids Res 44:9266–9278. https://doi.org/10.1093/nar/gkw663
Ma X, Liu H, Li J et al (2017) Polη O-GlcNAcylation governs genome integrity during translesion DNA synthesis. Nat Commun 8:1941. https://doi.org/10.1038/s41467-017-02164-1
Acknowledgements
We are grateful to Drs. Xiaoyong Yang (Yale University School of Medicine, New Haven, USA) and Dominique Leprince (IBL, Lille, France) for providing, respectively, the GST–OGT construct and the GST plasmid, Prof. Gerald W. Hart (Johns Hopkins University School of Medicine, Baltimore, USA) for providing acetyl-5S-GlcNAc, and Dr. T. Issad (INSERM U1016, CNRS UMR8104, Institut Cochin, Paris, France) for the HA-tagged OGT plasmid. We also thank Dr. Olivier Vidal (UMR CNRS 8576) for advices in GST–OGT production, Dr. Corentin Spriet (TISBio core facility, FRABIO FR 3688 CNRS, Lille) for helping in PLA analysis with ImageJ, Nicolas Vossart and Lucie Hurtevent for their technical assistance, and James Biwi (UMR 8576, UGSF) for critical reading of this manuscript. ML was a recipient of a fellowship from the Ministère de l’Enseignement Supérieur et de la Recherche. This work was supported by la Ligue Nationale Contre le Cancer (Comités du Nord et de l’Aisne) and the Centre National de la Recherche Scientifique (CNRS).
Author information
Authors and Affiliations
Contributions
ASVE conceived the project. ML, TL and ASVE designed experiments. ML, MM, CS, SH and ASVE performed experiments. ML, SH, ASVE analysed data. ML and ASVE wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Figure Captions: Suppl. Figure
1 O-GlcNAcylated MCM2-7 subunits are mainly found in the chromatin-enriched fraction in MDA-MB-231 cells. Serum-starved MDA-MB-231 cells were released into the cell cycle by serum addition for the indicated times. a Cell cycle distribution was determined by flow cytometry after DNA staining with PI. Percentage of cells in G0/G1, S and G2/M phases are indicated. b The nucleocytoplasmic soluble (Sol.) and chromatin-bound fractions (Chrom.) were analysed by Western blot for the indicated proteins. c O-GlcNAcylated proteins from both fractions were enriched on sWGA-agarose beads. Incubation with excess of GlcNAc (0.5 M) was used as negative control (C−). MCM proteins were detected by Western blot before (Input) and after enrichment on sWGA lectin (sWGA-bound). d Recombinant GST-tagged OGT (OGT–GST) and GST alone (GST) were immobilized onto Glutathione Sepharose beads. GST pull-down assay was performed using soluble (Sol.) and chromatin-bound (Chrom.) protein fractions from asynchronous MDA-MB-231 cells (Inp). Eluted MCM proteins were detected by Western blot and anti-GST antibodies were used to confirm the presence of the recombinant proteins after elution. Suppl. Figure 2 Efficiency of inhibition of OGT and OGA in MCF7 cells. Serum-starved cells were released in S phase by serum addition for 18 h in presence of DMSO (Ctrl), 5S-G (50 µM) or ThG (1 µM), before harvesting. Proteins from subcellular fractions (Sol. and Chrom.) were separated by SDS-PAGE and analysed by Western blot for the indicated proteins. Equal loading was confirmed by Ponceau staining of the nitrocellulose membrane. Suppl. Figure 3 Serum-starved MCF7 were released in S phase by serum addition for the indicated times. a DMSO (1/1000, Ctrl), 5S-G (50 µM) or ThG (1 µM) was added at the same time as serum. b cells were transfected with siRNA 24 h before serum starvation. Whole cell lysates were resolved by SDS-PAGE and analysed by Western blot for the indicated proteins. (PPTX 1094 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Leturcq, M., Mortuaire, M., Hardivillé, S. et al. O-GlcNAc transferase associates with the MCM2–7 complex and its silencing destabilizes MCM–MCM interactions. Cell. Mol. Life Sci. 75, 4321–4339 (2018). https://doi.org/10.1007/s00018-018-2874-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-018-2874-0