Abstract
To gain new insights into the complex evolutionary history of Brassica rapa, we investigated the genetic diversity of wild and domesticated populations from an unexplored area in its center of origin (Mediterranean basin). These populations were collected in Algeria, which presents a wide ecogeographic range and an exceptional bioclimatic gradient. We wanted to answer the following questions: (1) Were the local landraces domesticated from the local wild forms? (2) Do these populations offer a new diversity never described? (3) How is this diversity related to B. napus? Morphological traits and SSR markers were analysed to explore the genetic diversity among 18 Algerian B. rapa accessions that were compared to those previously analysed in this species and to B. napus varieties. Among worldwide B. rapa diversity, the wild and cultivated Algerian groups showed the highest allelic richness, suggesting that breeding has not significantly eroded genetic diversity in Algerian local landraces. Wild and cultivated B. rapa accessions from Algeria formed two clusters regardless of their local geographic origin and were distinct from all the B. rapa groups already described. Surprisingly, B. napus accessions clustered either with wild Algerian group or with the group containing mainly Asian accessions. This new diversity will be of high interest for B. rapa and B. napus breeding.

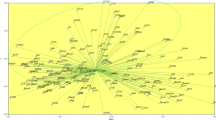
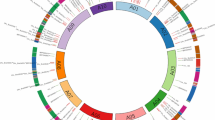
The bioclimatic zones are those established by Stewart (1974). (Color figure online)





Similar content being viewed by others
References
Aïssiou F, Chèvre AM, Abrous O, Hadj-Arab H (2018) Variation in floral organs in wild populations of Brassica rapa L. Acta Hortic 1202. In: ISHS 2018 proceedings VII international symposium on Brassicas, pp 69–74. https://doi.org/10.17660/ActaHortic.2018.1202.10
Allender CJ, King GJ (2010) Origins of the amphiploid species Brassica napus L. investigated by chloroplast and nuclear molecular markers. BMC Plant Biol 10:54. https://doi.org/10.1186/1471-2229-10-54
Andersen NS, Poulsen G, Andersen BA, Kiær LP, D’Hertefeldt T, Wilkinson MJ, Jørgensen RB (2009) Processes affecting genetic structure and conservation: a case study of wild and cultivated Brassica rapa. Genet Resour Crop Evol 56:189–200. https://doi.org/10.1007/s10722-008-9354-6
Annisa, Chen S, Cowling WA (2013) Global genetic diversity in oilseed Brassica rapa. Crop Pasture Sci 64:993–1007. https://doi.org/10.1071/CP13206
Bird KA, An H, Gazave E, Gore MA, Pires JC, Robertson LD, Labate JA (2017) Population structure and phylogenetic relationships in a diverse panel of Brassica rapa L. Front Plant Sci 8:321. https://doi.org/10.3389/fpls.2017.00321
Celucia SU, De la Peña RC, Villa NO (2009) Genetic characterization of Brassica rapa chinensis L., B. rapa parachinensis (L. H. Bailey) Hanelt, and B. oleracea alboglabra (L. H. Bailey) Hanelt using simple sequence repeat markers. Philipp J Sci 138:141–152
Chatterji S, Pachter L (2006) Reference based annotation with GeneMapper. Genome Biol 7:R29. https://doi.org/10.1186/gb-2006-7-4-r29
Chen ZJ, Pikaard CS (1997) Epigenetic silencing of RNA polymerase I transcription: a role for DNA methylation and histone modification in nucleolar dominance. Genes Dev 11:2124–2136. https://doi.org/10.1101/gad.11.16.2124
Chen S, Zou J, Cowling WA, Meng J (2010) Allelic diversity in a novel genepool of canola-quality Brassica napus enriched with alleles from B. rapa and B. carinata. Crop Pasture Sci 61:483–492. https://doi.org/10.1071/CP09327
Chen S, Wan Z, Nelson MN, Chauhan JS, Redden R, Burton WA, Lin P, Salisbury PA, Fu T, Cowling WA (2013) Evidence from genome-wide simple sequence repeat markers for a polyphyletic origin and secondary centers of genetic diversity of Brassica juncea in China and India. J Hered 104:416–427. https://doi.org/10.1093/jhered/est015
Cheng F, Sun R, Hou X, Zheng H, Zhang F, Zhang Y, Liu B, Liang J, Zhuang M, Liu Y, Liu D, Wang X, Li P, Liu Y, Lin K, Bucher J, Zhang N, Wang Y, Wang H, Deng J, Liao Y, Wei K, Zhang X, Fu L, Hu Y, Liu J, Cai C, Zhang S, Zhang S, Li F, Zhang H, Zhang J, Guo N, Liu Z, Liu J, Sun C, Ma Y, Zhang H, Cui Y, Freeling MR, Borm T, Bonnema G, Wu J, Wang X (2016) Subgenome parallel selection is associated with morphotype diversification and convergent crop domestication in Brassica rapa and Brassica oleracea. Nat Genet 48:1218–1224. https://doi.org/10.1038/ng.3634
Chevalier A (1932) Les productions végétales du Sahara et de ses confins Nord et Sud. Revue de Botanique appliquée et d’agriculture coloniale. 133–134:669–924. https://doi.org/10.3406/jatba.1932.5282
Crouch JH, Lewis BG, Lydiate DJ, Mithen R (1995) Genetic diversity of wild, weedy and cultivated forms of Brassica rapa. Heredity 74:491–496. https://doi.org/10.1038/hdy.1995.69
De Candolle A (1886) Origin of cultivated plants. (Hafner, New York, 1967). English translation of the second edition, pp 316–321. https://doi.org/10.1017/CBO9781139107365
Del Carpio DP, Basnet RK, De Vos RC, Maliepaard C, Visser R, Bonnema G (2011) The patterns of population differentiation in a Brassica rapa core collection. Theor Appl Genet 122:1105–1118. https://doi.org/10.1007/s00122-010-1516-1
Dice LR (1945) Measures of the amount of ecologic association between species. Ecology 26:297–302. https://doi.org/10.2307/1932409
Doyle J, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Gerlach WL, Bedrook JR (1979) Cloning and characterization of ribosomal RNA genes wheat from and barley. Nucleic Acids Res 7:1869–1885. https://doi.org/10.1093/nar/7.7.1869
Gomez-Campo C, Prakash S (1999) Origin and domestication. In: Gomez-Campo C (ed) Biology of Brassica Coenospecies. Elsevier, Hoboken, pp 33–58. https://doi.org/10.1016/S0168-7972(99)80003-6
Guo Y, Chen S, Li Z, Cowling WA (2014) Center of origin and centers of diversity in an ancient crop, Brassica rapa (Turnip rape). J Hered 105:555–565. https://doi.org/10.1093/jhered/esu021
Hasterok R, Wolny E, Hosiawa M, Kowalczyk M, Kulak-Ksiazczyk S, Ksiazczyk T, Heneen WK, Maluszynska J (2006) Comparative analysis of rDNA distribution in chromosomes of various species of Brassicaceae. Ann Bot 97:205–216. https://doi.org/10.1093/aob/mcj031
Husson F, Josse J, Le S, Mazet J (2008) Factor analysis and data mining with R. R package version 1.10
IPGRI (1990) Descriptors for Brassica and Raphanus. International Plant Genetic Resources Institute, Rome
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, Thierer T, Ashton B, Meintjes P, Drummond A (2012) Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649. https://doi.org/10.1093/bioinformatics/bts199
Ksiazczyk T, Kovarik A, Eber F, Huteau V, Khaitova L, Tesarikova Z, Coriton O, Chèvre AM (2011) Immediate unidirectional epigenetic reprogramming of NORs occurs independently of rDNA rearrangements in synthetic and natural forms of a polyploid species Brassica napus. Chromosoma 120:557–571. https://doi.org/10.1007/s00412-011-0331-z (epub 2011 Jul 23)
Kumar R, Nawroth PP, Tyedmer J (2016) Prion aggregates are recruited to the insoluble protein deposit (IPOD) via myosin 2-based vesicular transport. PLoS Genet 12:e1006324. https://doi.org/10.1371/journal.pgen.1006324
Li CW (1982) The origin, evolution, taxonomy and hybridization of Chinese cabbage. In: Talekar NS, Griggs TD (eds) Chinese cabbage. In: Proceedings of the 1st international AVRDC symposium, Taiwan, pp 1–10
Li P, Zhang S, Li F, Zhang S, Zhang H, Wang X, Sun R, Bonnema G, Borm TJA (2017) A phylogenetic analysis of chloroplast genomes elucidates the relationships of the six economically important Brassica species comprising the triangle of U. Front Plant Sci 8:e111. https://doi.org/10.3389/fpls.2017.00111
Mei J, Fu Y, Qian L, Xu X, Li J, Qian W (2011) Effectively widening the gene pool of oilseed rape (Brassica napus L.) by using Chinese B. rapa in a ‘virtual allopolyploid’ approach. Plant Breed 130:333–337. https://doi.org/10.1111/j.1439-0523.2011.01850
Ofori A, Becker HC, Kopisch-Obuch FJ (2008) Effect of crop improvement on genetic diversity in oilseed Brassica rapa (turnip-rape) cultivars, detected by SSR markers. J Appl Genet 49:207–212. https://doi.org/10.1007/BF03195615
Pang W, Li X, Choi SR, Dhandapani V, Im S, Park MY, Jang CS, Yang MS, Ham IK, Lee EM, Kim W, Lee SS, Bonnema G, Park S, Piao Z, Lim YP (2015) Development of a leafy Brassica rapa fixed line collection for genetic diversity and population structure analysis. Mol Breed New Strateg Plant Improv 35:1–15. https://doi.org/10.1007/s11032-015-0221-9
Perrier X, Jacquemoud-Collet JP (2006) DARwin software. http://darwin.cirad.fr/darwin.cirad.fr/darwin, 5th edn. Cirad, Montpellier. Accessed June 2016
Pinnegar A, Astley D, Dehmer K, Graichen ML, Wilner E, Rosa E, Aires A, Paula R, Amando OP, Cartea González ME (2004) Brassica collections for broadening agricultural use: rESGEN CT99 109/112 Brassica rapa subgroup. Crucif Newsl 25:113–114
Piquemal J, Cinquin E, Couton F, Rondeau C, Seignoret E, Doucet I, Perret D, Villeger MJ, Vincourt P, Blanchard P (2005) Construction of an oilseed rape (Brassica napus L.) genetic map with SSR markers. Theor Appl Genet 111(8):1514–1523
Posada D (2008) jModelTest: phylogenetic model averaging. Mol Biol Evol 25:1253–1256. https://doi.org/10.1093/molbev/msn083
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Qi X, An H, Ragsdale AP, Hall TE, Gutenkunst RN, Pires JC, Barker MS (2017) Genomic inferences of domestication events are corroborated by written records in Brassica rapa. Mol Ecol 100:1–16. https://doi.org/10.1111/mec.14131
Qian W, Meng J, Li M, Frauen M, Sass O, Noack J, Jung C (2006) Introgression of genomic components from Chinese Brassica rapa contributes to widening the genetic diversity in rapeseed (B. napus L.), with emphasis on the evolution of Chinese rapeseed. Theor Appl Genet 113:49–54. https://doi.org/10.1007/s00122-006-0269-3
Redden R, Vardy M, Edwards D, Raman H, Batley J (2009) Genetic and morphological diversity in the Brassicas and wild relatives. In: Proceedings of the 16th Australian research assembly on Brassicas
Rouxel T, Kollmann A, Boulidard L, Mithen R (1991) Abiotic elicitation of indole phytoalexins and resistance to Leptosphaeria maculans within Brassiceae. Planta 184:271–278. https://doi.org/10.1007/BF00197957
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Schuelke M (2000) An economic method for the fluorescent labeling of PCR fragments. Nat Biotechnol 18(2):223–234. https://doi.org/10.1038/72708
Soengas P, Cartea ME, Francisco M, Lema M, Velasco P (2011) Genetic structure and diversity of a collection of Brassica rapa subsp. rapa L. revealed by simple sequence repeat markers. J Agric Sci 149:617–624. https://doi.org/10.1017/S002185961100013X
Song KM, Osborn TC, Williams PH (1988a) Brassica taxonomy based on nuclear restriction fragment length polymorphism (RFLPs). 2. Preliminary analysis of sub-species within B. rapa (syn. campestris) and B. oleracea. Theor Appl Genet 76:593–600. https://doi.org/10.1007/BF00260914
Song KM, Osborn TC, Williams PH (1988b) Brassica taxonomy based on nuclear restriction fragment length polymorphism (RFLPs). 1. Genome evolution of diploid and amphidiploid species. Theor Appl Genet 75:784–794. https://doi.org/10.1007/BF00265606
Song KM, Osborn TC, Williams PH (1990) Brassica taxonomy based on nuclear restriction fragment length polymorphisms (RFLPs). 3. Genome relationships in Brassica and related genera and the origin of B. oleracea and B. rapa (syn. campestris). Theor Appl Genet 79:497–506. https://doi.org/10.1007/BF00226159
Stewart P (1974) Un nouveau climagramme pour l’Algérie et son application au barrage vert. Bull Soc Hist Nat Afrique du Nord 65:239–248
Suwabe K, Iketani H, Nunome T, Kage T, Hirai M (2002) Isolation and characterization of microsatellites in Brassica rapa L. Theor Appl Genet 104:1092–1098. https://doi.org/10.1007/s00122-002-0875-7
Takahashi Y, Yokoi S, Takahata Y (2016) Genetic divergence of turnip (Brassica rapa L. em. Metzg. subsp. rapa) inferred from simple sequence repeats in chloroplast and nuclear genomes and morphology. Genet Resour Crop Evol 63:869–879. https://doi.org/10.1007/s10722-015-0290-y
Takuno S, Kawahara T, Ohnishi O (2007) Phylogenetic relationships among cultivated types of Brassica rapa L. em. Metzg. As revealed by AFLP analysis. Genet Resour Crop Evol 54:279–285. https://doi.org/10.1007/s10722-005-4260-7
Tanhuanpää P, Erkkilä M, Tenhola-Roininen T, Tanskanen J, Manninen O (2016) SNP diversity within and among Brassica rapa accessions reveals no geographic differentiation. Genome 59:11–21. https://doi.org/10.1139/gen-2015-0118
U N (1935) Genome analysis in Brassica with special reference to the experimental formation of B. napus and peculiar mode of fertilization. Jpn J Bot 7:389–452
Vellvé R (1993) The decline of diversity in European agriculture. Ecologist 23:64–69
Zhang N, Zhao J, Lens F, De Visser J, Menamo T, Fang W, Xiao D, Bucher J, Basnet RK, Lin K, Cheng F, Wang X, Bonnema G (2014) Morphology, carbohydrate composition and vernalization response in a genetically diverse collection of Asian and European turnips (Brassica rapa subsp. rapa). PLoS ONE 9:e114241. https://doi.org/10.1371/journal.pone.0114241
Zhao J, Wang X, Deng B, Lou P, Wu J, Sun R, Xu Z, Vromans J, Koornneef M, Bonnema G (2005) Genetic relationships within Brassica rapa as inferred from AFLP fingerprints. Theor Appl Genet 110:1301–1314. https://doi.org/10.1007/s00122-005-1967-y
Zhao J, Paulo MJ, Jamar D, Lou P, Van Eeuwijk F, Bonnema G, Vreugdenhil D, Koornneef M (2007) Association mapping of leaf traits, flowering time, and phytate content in Brassica rapa. Genome 50:963–973. https://doi.org/10.1139/G07-078
Zhao YG, Ofori A, Lu CM (2009) Genetic diversity of European and Chinese oilseed Brassica rapa cultivars from different breeding periods. Agric Sci China 8:931–938. https://doi.org/10.1016/S1671-2927(08)60297-7
Zhao J, Artemyeva A, Del Carpio DP, Basnet RK, Zhang N, Gao J, Li F, Bucher J, Wang X, Visser RG, Bonnema G (2010) Design of a Brassica rapa core collection for association mapping studies. Genome 53:884–898. https://doi.org/10.1139/G10-082
Acknowledgements
The authors thank the University of Science and Technology Houari Boumediene (USTHB), Faculty of Biological Sciences, the team Biosystematics, Genetics and Evolution, Algiers and the French National Research Institute for Agronomy (INRA). The present work was financially supported by Algerian Ministry of Higher Education and Scientific Research in the framework of the Accord-Programme of the Algerian-French Convention CMEP-TASSILI (16MDU952). The work described in this manuscript was performed partly within the framework of the project COREBRAS funded by Promosol. We acknowledge the Genetic Resource Center (BrACySol, UMR IGEPP, Ploudaniel, France) for providing seeds and the staff for their technical assistance in greenhouses and in the lab (especially L. Charlon, P. Rolland, J.P. Constantin, J.M. Lucas and F. Letertre, S. Guichard). We thank O. Kherabi for assistance in collecting plants and seeds from Algerian localities. We also thank the UMR INRA 1095 ‘GENTYANE platform’ (Clermont-Ferrand, France, http://gentyane.clermont.inra.fr/) for the generation of SSR genotyping data.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
10681_2018_2318_MOESM1_ESM.docx
Table S1 B. rapa accessions corresponding to the worldwide set, defined as set 2 in the “Materials and methods” part. These accessions correspond to the core collection defined by Zhao et al. (2005) in order to represent the Brassica rapa genetic diversity. The geographic origin (America, Europe, West Asia, North Africa, Central Asia, East Asia, South East Asia) as well as the country of origin of each accession are presented. (DOCX 41 kb)
10681_2018_2318_MOESM4_ESM.pptx
Figure S1 FISH analyses of somatic metaphase chromosomes revealing 10 or 8 35S loci from cultivated accessions (BRY (A), BRL (B) and wild populations (BRC (C), BRE (D)), respectively. FISH was carried out using 35S rDNA (red signals and orange stars). Chromosomes were counterstained with DAPI (white). Scale bar 5 µm. Abbreviations of the accessions are given in Table 1. (PPTX 17052 kb)
10681_2018_2318_MOESM5_ESM.pptx
Figure S2 Molecular phylogeny of Brassiceae based on the chloroplast ndhC-trnV intergenic spacer (data matrix: 866 bp) using the maximum Likelihood method (Tamura 3 parameter) with MEGA 7.0 (Kumar et al. 2016). The chloroplast data sequences obtained in this study for sets 1, 3 and 4 are indicated in bold and are underlined. The other sequences were retrieved from the chloroplast genome sequences of the accessions described in Li et al. (2017). The bootstrap percentages (1000 replicates) are shown above the branches. The tree is rooted using B. carinata and B. nigra that belongs to the nigra clade. (PPTX 83 kb)
10681_2018_2318_MOESM6_ESM.pptx
Figure S3 Plot of first and third principal components showing relationships between all B. rapa Algerian accessions tested on the basis of the morphological traits. (A) Phenotypic variation of the wild and cultivated accessions. (B) Contribution of the different traits to the phenotypic variation. Algerian B. rapa wild (BRA, BRC, BRD, BRE, BRF, BRJ, BRH, BRO, BRZ) and cultivated (BRI, BRL, BRN, BRM, BRR, BRT, BRY) accessions are represented in green and blue, respectively. (PPTX 87 kb)
Rights and permissions
About this article
Cite this article
Aissiou, F., Laperche, A., Falentin, C. et al. A novel Brassica rapa L. genetic diversity found in Algeria. Euphytica 214, 241 (2018). https://doi.org/10.1007/s10681-018-2318-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-018-2318-9