Abstract
Cultivation of spinach requires high amounts of nitrogen (N), which puts a strain on the environment. A sustainable solution to this problem is to breed for crops with higher N use efficiency (NUE). The aim of this study was to provide tools for molecular breeding and to elucidate the genetic variation of factors contributing to NUE in spinach. A cross was made between two F1 hybrid cultivars contrasting in NUE. Several F1 progeny were self-pollinated and based on evaluation of the F2 generation, a mapping F2 population (335 individuals) of a single F1 was selected. SNP markers for the genetic map were discovered by RNA sequencing of the two parent cultivars, and 283 SNP markers were used to produce a genetic map comprising of six linkage groups (P01–P06), ranging in size from 46 to 116 cM. NUE related traits were determined for a set of F2:3 families grown under low and high N conditions in a hydroponics system under an Ingestad N-addition model. Interval mapping analysis detected 39 trait-specific QTLs, with several QTLs accumulating on P01 and P02 of the linkage map. The QTLs and in particular the P01 and P02 regions provide potential targets for the improvement of NUE in spinach.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nitrogen is the nutrient that most frequently limits plant growth (Fageria and Baligar 2005). In most current crop production systems plants rely on mineral fertilizers to meet their N demand. The high levels of N fertilization are often associated with environmental problems such as eutrophication of soil and surface water and emission of greenhouse gases (Lawlor et al. 2001). The environmental pollution together with human health issues has resulted in strict regulation of nitrogen fertilization in Europe (European Commission 2010). This legislation has a large impact on the cultivation of vegetable crops, in particular of spinach. Mitigation of the negative effects of lowering N input on productivity of spinach therefore is a major challenge. This holds true in particular for production systems that require low external input, such as organic agriculture, in which N inputs would preferably be reduced from 150 kg/ha to approx. 100 kg N/ha or less (Fageria and Baligar 2005). A long term sustainable strategy contributing to cultivating spinach with less nitrogen is the genetic improvement of its nitrogen use efficiency (NUE), resulting in cultivars that can realize an economically acceptable yield even under low input conditions.
Nitrogen efficiency is a complex trait that is the resultant of two component traits: nitrogen uptake efficiency and nitrogen utilisation efficiency (Hirel et al. 2007). Insight in the physiological processes governing NUE under low N conditions and the genetic basis for NUE is essential for efficient breeding for this complex trait. A number of studies were conducted on N use of spinach, but these were mainly focused on maximizing spinach yield to get close to the yield potential of the cultivars under study (e.g. Smolders and Merckx 1992; Biemond 1995; Biemond et al. 1996), and on NO3 − accumulation in relation to consumption quality (Breimer 1982; Steingröver 1986). These studies revealed that sufficient N must be available at the start of growth to realize optimal growth in spinach (Biemond 1995) and that spinach can acquire nutrients better with improved root systems (Smolders and Merckx 1992, Hirel et al. 2007). Biemond et al. (1996) found that with increased N availability the total green leaf area increased through a higher leaf expansion rate, emphasizing the strong dependence of spinach growth and yield on N availability.
To assess the potential of breeding cultivars with improved NUE it is necessary to (i) determine the genetic variation present in spinach germplasm for traits that govern NUE and (ii) develop a methodological setup that enables accurate quantification of and selection for the physiological and growth parameters that contribute to improved NUE (Baligar et al. 2001). Chan-Navarrete et al. (2014) evaluated traits related to growth and photosynthesis and their correlation with NUE in a hydroponics system. They reported that leaf area as well as specific leaf area (SLA) were strong determinants of variation for NUE in spinach.
Insight in the genetics of a complex trait such as NUE can be gained by Quantitative Trait Locus (QTL) analysis of a dedicated segregating population. Spinach is a dioecious species with separate male and female plants and occasional monoecious plants (Correll et al. 2011). Spinach is diploid with 2n = 12 chromosomes and has a genome size of 989 Mb (Arumuganathan and Earle 1991). The genome sequence is not available yet, but the spinach mitochondrial and chloroplast genomes were sequenced (327 and 150 kb, respectively) (Stern and Palmer 1986; Schmitz-Linneweber et al. 2001). QTL analysis of a segregating population requires a genetic linkage map with sufficient molecular markers distributed over the genome. For spinach only one genetic linkage map with a limited number of simple sequence repeat (SSR) and amplified fragment length polymorphism (AFLPs) markers was published until now (Khattak et al. 2006). It was used to analyse genetic variation related to sex expression. Onodera et al. (2011) used the same molecular marker data to map genes for dioecism and monoecism in spinach.
The aim of the current study is to provide tools for molecular breeding for NUE in spinach, and elucidate the genetic factors determining the variation in NUE and traits contributing to NUE. NUE is a quantitative plant trait with a polygenic inheritance, which is influenced by N availability (Hirel et al. 2001). The latter is difficult to control under field conditions (Fageria and Baligar 2005) and this will complicate genetic analysis of NUE variation. Hence, a hydroponics system providing controlled evaluation conditions thus reducing the environmental effects as described by Chan-Navarrete et al. (2014) was used in this study.
A dedicated F2:3 mapping population derived from a cross between heterozygous parents was evaluated on a hydroponics system. The parents were shown to differ highly for NUE under low N conditions (Chan-Navarrete et al. 2014). A genetic linkage map was constructed using a selected set of gene-based Single Nucleotide Polymorphisms (SNP) markers to enable a QTL analysis of the NUE evaluation data of the F2:3 lines. The analysis revealed several regions in the spinach genome with clustered QTLs that contribute to improve NUE, providing potential targets for molecular breeding for NUE in spinach.
Materials and methods
Plant material and mapping population
The mapping population consisted of a random set of F2:3 lines derived from a single monoecious F1-plant that resulted from a cross between the hybrid spinach cultivars Ranchero (Enza Zaden) and Marabu (Rijk Zwaan). The parents were selected because they differed strongly in NUE (Chan-Navarrete et al. 2014). Ranchero is a cultivar with an upright growth habit and many dark oval-shaped leaves, suitable for spring, autumn and winter cultivation. Marabu is a cultivar with thick, dark green leaves suitable for cultivation in spring and early autumn. Ranchero had high NUE while NUE of Marabu was relatively low under low N conditions (Chan-Navarrete et al. 2014).
With a few exceptions that were strictly male the F2 plants were monoecious, and 320 of these F2 plants were grown and self-pollinated to generate F2:3 lines. In parallel leaf material of each plant was sampled for DNA analysis and molecular genotyping with Single Nucleotide Polymorphisms (SNPs). The F2:3 lines were grown on hydroponics for phenotypic evaluation of NUE.
Discovery of gene-based markers
Leaf material of 10 young plants of Ranchero and Marabu, grown under optimal N conditions, was bulked. From both bulks, total RNA was extracted using the RNeasy Plant Mini Kit (QIAGEN). The RNA was quantified using Qubit (Invitrogen) and checked on a 1 % agarose gel. After library preparation, the samples were sequenced on a HiSeq 2000 sequencer (Illumina, Varshney et al. 2009), obtaining 2 × 100 bp paired-end reads. For Marabu, the overlapping ends of the paired-end reads were first merged with FLASH (http://ccb.jhu.edu/software/FLASH/, Magoč and Salzberg 2011), resulting in 72 % merged read-pairs, with an average merged length of 140 bases. After merging, the merged and non-merged paired-end reads were quality-trimmed using PRINSEQ (http://prinseq.sourceforge.net/index.html, Schmieder and Edwards 2011). Bases with a PHRED Q-value lower than Q20 were trimmed, poly-A trailing bases longer than 20 bp were removed, and remaining sequences shorter than 50 bp were discarded. Low complexity regions were filtered with the ‘dust’ option, and duplicate reads were removed. To remove chloroplast reads, the sequences were mapped against the chloroplast genome of spinach (AJ400848) using Bowtie2 (http://bowtie-bio.sourceforge.net/bowtie2/index.shtml, (Langmead and Salzberg 2012), and mapped reads were excluded from further analysis. The remaining reads were extracted using SAMtools (http://samtools.sourceforge.net/, Li et al. 2009) and used for a de novo transcriptome assembly using Trinity (http://trinityrnaseq.sourceforge.net/, Grabherr et al. 2011). The Trinity assembly was performed with minimal k-mer coverage of 2, generating 80,483 transcripts from 45,900 components, with an N50 of 2004 bp. A component could be related to a ‘unigene’ and a component can have multiple transcript types (isoforms). To avoid redundancy, only one transcript of each component was used for marker development. The abundance of each isoform was determined with RSEM (RNA-Seq by Expectation–Maximization, included in the trinity distribution), and for each component, the most abundant transcript was kept in the final transcript fasta sequence file. Finally, a set of 45,900 transcripts with an N50 of 1491 was used as transcriptome reference sequence. The RNAseq reads from both genotypes were mapped to the reference transcriptome sequence using Bowtie2. On the resulting SAM file, SNPs were called using QualitySNPng (http://www.bioinformatics.nl/QualitySNPng/, Nijveen et al. 2013).
The screening resulted in discovery of 12,477 candidate SNPs with a non-polymorphic flanking region of 50 bp on both sides of the SNP. SNPs polymorphic between the parents, but homozygous in Marabu or Ranchero were selected. This reduced the number of SNPs to 704, present in 506 unique transcripts. Of these SNPs, 419 were used to genotype the F2 plants.
Genotyping
The sequence information of 419 SNPs was used to develop KASPar assays for genotyping 335 F2 plants of the mapping population. The Competitive Allele Specific PCR (KASPar) platform (http://www.kbioscience.co.uk) is a PCR-based novel homogeneous fluorescent SNP genotyping system. The array included seven technical DNA replicates of each of the parents. The DNA was extracted with a modified C-TAB (Steward and Via 1993).
Construction of linkage map
High quality and informative molecular markers (no segregation distortion and less than 15 % missing values) were used to generate a linkage map using software package JoinMap 4.1 (van Ooijen 2006) with the maximum-likelihood option for ordering markers within linkage groups. The Haldane mapping function was used to calculate the final maps (Haldane and Waddington 1931). Possible segregation distortion was determined by testing the actual against the expected segregation ratio of 1:2:1, using the χ2 test of goodness of fit with two degrees of freedom.
Experimental setup
A randomly selected set of 94 F2:3 lines was evaluated for NUE using a hydroponics system in a temperature-controlled sunlit greenhouse compartment at UNIFARM (Wageningen, The Netherlands) as described in detail by Chan-Navarrete et al. (2014). The system consisted of six units, each having 16 containers with 24 plant positions. N application could be separately controlled for each of the six units. The daily application of N aimed at a stable relative plant growth rate (RGR) of either 0.10 or 0.18 g·g−1·day−1 based on the Ingestad (1982) model to acquire a steady-state N-nutrition level. We further refer to the 0.10 and 0.18 g·g−1·day−1 N-rate as low N and high N, respectively.
Evaluation of F2:3 lines
The hydroponics experiment to evaluate the F2:3 mapping population for NUE and related traits was executed over a period of 35 days. The measurements were done at the end of the trial, except for non-destructive chlorophyll content measurements. Every measurement was done on individual plants except for NUE measurements, which were based on bulked samples of 4 plants (in total 3 samples per line for both N treatments).
The plants were patted dry at harvest with industrial paper tissue and divided in a root and shoot fraction. The shoot fraction was weighed immediately to determine shoot fresh weight (SFW). The shoot and root fraction of each plant were dried for two days at 70 °C to get measures for plant shoot dry weight (SDW) and root dry weight (RDW). The plant root-to-shoot ratio (R:S) was determined as R:S = RDW·SDW−1. Other traits evaluated at harvest time were dry matter percentage (DM %) (g·g−1·100), which is the percentage of the dry mass from the fresh weight of the shoot; leaf area (LA) (cm2) determined with a Licor Leaf Area Scanner (LI-3100C) directly after harvest; specific leaf area (SLA) (cm2·g−1) calculated as SLA = LA/SDW; Leaf number (LN) scored the day before harvest time; Stem length (SL) (cm) measured with a metric ruler; Flowering (Fl), scored as presence (1) or absence (0) of flower structures at harvest time. Chlorophyll content (CC) (SPAD units) was measured with a SPAD 502 meter (Konica Minolta, Osaka, Japan) 21 (CCi) and 28 (CCh) days after transplanting of seedlings on hydroponics. SPAD values were collected on the first appearing pair of leaves of each plant, one measurement per plant. Nitrogen Use Efficiency (NUE) (g SDW g−1 N) was calculated as the SDW divided by the N content in SDW (determined with a Kjeldahl analysis). The latter measurements were performed pooled samples of 4 plants per line-treatment combination, resulting in 3 pooled samples per combination.
Statistical analysis
Descriptive statistics were applied to estimate the overall effects of the two N treatments and to quantify the variation present in the mapping population. The relationship between traits was studied with a correlation analysis. Analyses of variance (ANOVAs) were carried out for each N treatment separately using GenStat 16th; each having a randomized block design. Each block consisted of four adjacent containers within one of the units of the hydroponics system. In all, the experiment contained 24 blocks, i.e. 6 (units) × 4 (blocks/unit). To each of the 96 plant positions available within a block a single plant was randomly assigned from each of the 94 F2:3 lines to be evaluated, and the 2 parental hybrids. Broad sense heritabilities (h 2m ) were calculated for means over replicates of F2:3 lines (n = 12), the genotypic variance (σ 2g ) and the experimental variance (σ 2e ) with the following formula:
For the analysis between N treatments the experimental design was evaluated as a split-plot design. The six hydroponics units were grouped in three subsets of two adjacent units. The two N-treatment levels were than assigned to one of the two units within a subset.
QTL analysis
QTL analysis of the phenotypic data was performed with the mapping software available in GenStat 16th Edition (VSN International 2013). Separate single trait association analyses (also known as linkage disequilibrium mapping) were performed separately for low and high N using the F2:3 line means for each phenotypic trait, the F2 linkage map and the marker data of the F2 progenitors of the F2:3 lines. A genome-wide permutation test according to Li and Ji (2005) was performed to calculate the threshold for QTLs. Only QTLs with significance level >95 % were taken into consideration. The linkage maps with QTLs were displayed with MapChart (Voorrips 2002).
Results
Molecular marker discovery
SNP discovery was done by RNA sequencing of spinach cultivars Marabu and Ranchero. The Marabu sequence was used as a reference to detect SNPs. The total number of Marabu reads was 23,755,238, which after processing (see Materials and methods) resulted in 45,872 unique transcripts. The Ranchero sequence reads were mapped against this final transcript set and with the use of QualitySNPng (Nijveen et al. 2013) 27,499 SNPs were identified. Of these, only 1351 SNPs were polymorphic between but homozygous within Ranchero and Marabu, while 11,781 SNPs were heterozygous in one parent only and 14,317 SNPs were heterozygous in both parents.
The SNPs that were homozygous in both parents but polymorphic between parents were preferred for genetic map construction, as these were all expected to segregate in the F2 population. After filtering for insufficient flanking sequence information for assay development, a set of 704 SNPs in 506 unique transcripts was available for genotyping.
Genotyping
Genotyping of the mapping population was performed with KASPar assays using a selection of 419 SNPs from the set of 506 candidate SNP markers. Of these, 136 markers were not informative because of a lack of signal (59), a high percentage of missing values (19), distorted segregation (17) or simply lack of segregation (41). The 283 informative markers showed a 1:2:1 segregation ratio and were used to generate a genetic linkage map. To this end the marker data of 320 out of 335 genotyped F2 plants were used; data of 15 genotypes were omitted because they had a high number of markers with missing values.
Linkage map
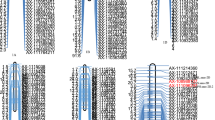
The genetic linkage map was constructed using JoinMap 4.1 (van Ooijen 2006) with 283 informative markers and 320 genotypes. All the markers were incorporated in a map with a total size of 433.6 cM, divided over six linkage groups (Fig. 1). This is in accordance with the basic number of spinach chromosomes. The linkage groups were coded P01–P06 (P for putative) as no chromosomal anchors were available to link the linkage groups to chromosomes. The linkage groups varied in size from 46.7 to 116.2 cM. Some linkage groups contained relatively large gaps (20 cM (46.51–66.08 cM) in P02 and 35 cM (24.17–59.19 cM) in P05). Substantial clustering of markers was observed as well.
Phenotypic evaluation of the mapping population
A random set of 94 F2:3 lines from the mapping population was evaluated for NUE-related traits on hydroponics at low N and high N. The results are summarized in Table 1 and visualized for selected traits in Fig. 2. The N treatment affected plant growth considerably. In general, shoot fresh weight (SFW), shoot dry weight (SDW), Leaf Area (LA) and NUE were reduced at low N compared to high N conditions (Table 1), and dry matter percentage (DM %) and root to shoot ratio (R:S) were increased. N levels had no effect on root dry weight (RDW), and Flowering (Fl) was increased at low N conditions. Chlorophyll Content (CC) was slightly but significantly negatively affected. Plants grown at high N conditions were bigger than plants grown at low N and had larger leaves with a relatively dark green colour.
The F2:3 line means for all traits determined at low and high N showed a continuous and in most cases normal distribution, as exemplified in Fig. 2. Analyses of variance of the traits at low as well as at high N demonstrated highly significant F2:3 line-to-line variation (p < 0.001). There were large differences between the most extreme lines for all traits (Table 1). The heritability estimates (h 2m ) for all traits were intermediate to high, but varied considerably from trait to trait (Table 1). The heritability estimates of each trait at low and high N were mostly similar. The heritability estimates for R:S, SLA and DM % were relatively low as these traits represent ratios of measured traits.
Relationship between NUE-related traits
To get insight in the relationships between NUE and NUE-related traits two separate correlation analyses were carried out: one with line means from low N and one with line means from high N. The results are summarized in Fig. 3. The values on the diagonal represent the trait-specific coefficients of correlation between trait values of plants grown at low and high N. The correlation between N treatments for NUE (r = 0.42) was moderate, reflecting the significant line × N interaction for this trait (Table 1). At both N treatments NUE was highly correlated with SDW and not surprisingly also with SFW, LA and RDW (0.65–0.74 for low N and 0.78–0.89 for high N). The most interesting traits with a moderate negative correlation with NUE were the physiological traits SLA and R:S. Fl and SL are highly correlated but no obvious relation with any of the other traits including NUE was found. Remarkably, chlorophyll content at 28 days (CCh) showed no significant correlation to NUE at low N while at high N it was significantly correlated with NUE as well as with SFW, SDW, RDW and LA.
Correlations between the phenotypic line means for twelve NUE-related traits determined at Low and High N. The coefficients above the diagonal refer to traits determined at low N and the ones below the diagonal to the corresponding traits from the high N treatment. The values on the diagonal (black cells) represent coefficients of correlation between line means determined at low and high N. The colour scale (depicted below the correlation table) indicates the correlation strength. Correlation coefficients with |r| > 0.20 were significant for a p < 0.001
QTL mapping for NUE-related traits
QTL analysis was performed to discover chromosomal regions that contribute to the variation observed within the mapping population grown under high N and low N conditions. Separate QTL analyses were performed with GenStat 16th (VSN International 2013) using the F2:3 line mean values from the phenotypic evaluation at low and high N. The genome-wide LOD score threshold was calculated to be 2.97. The QTLs for the NUE-related traits are summarized in Table 2 and are graphically represented in Fig. 4. Nineteen and 20 QTLs for high and low N, respectively, were detected. The alleles from the male parent Marabu had a favourable effect for a relatively large number of QTLs. The QTLs for which Ranchero contributed the favourable allele were HN_R:S_2, HN_SLA_1, LN_NUE_1, LN_R:S_1, LN_SFW_2 and all the QTLs related to CC (Table 2). The Marubu allele of the QTL for LA at P01 (3.8 cM) showed a differential response to N level with a relatively strong favourable effect at high N (9.11) and unfavourable effect at low N (3.79).
A number of QTLs co-localized at the top of P01 and at the bottom of P02. The QTLs related to Fl co-localized with multiple other QTLs at P01 and P02. Flowering is a developmental process that impacts N metabolism and therefore may be affecting several other traits. To investigate to what extent the QTLs were independent of flowering, Fl was used as a covariate in QTL mapping using the software MapQTL 6.0 (van Ooijen and Kyazma 2009). The QTLs were confirmed, indicating that these QTLs are not only dependent of Fl.
Biomass QTLs
Two QTLs were determined for SDW, one at low N and one at high N. Both of them were detected in the multiple-QTL region on P01, with a peak at 3.8 cM and 15.1 % explained variation for low N and 13.9 % for high N. Under N limiting conditions, two QTLs were found, at P01 and P05 that explained 29.8 % of the variation. For LN_SFW_1 the favourable allele is donated by Marabu, but for LN_SFW_2 by Ranchero. At high N, a single QTL was identified for this trait, explaining 18.7 % of the variation. For high N conditions no QTL was found for DM %, but for low N conditions a QTL was found on P05 that explained 14.3 % of the phenotypic variation. Two QTLs were detected for R:S for high N and one QTL for low N with an explained variation of 33.0 and 14.0 %, respectively. Three QTLs were determined for LN at both low N and high N. At high N all three QTLs were found in P01, but at low N there were two QTLs at P01 and one at P02.
Leaf QTLs
For LA at high N conditions, the two QTLs explained 16.7 and 17.2 % of the phenotypic variation and in low N conditions a single QTL that explained 14.3 % of the variation was detected. A QTL for SLA was found only at high N conditions with a LOD-value of 5.56. The explained variation is 22.9 % and high values were driven by the Ranchero allele. The chlorophyll content measurements were performed at an intermediate time point (CCi) and at harvest (CCh). Multiple QTLs were observed for CCi that explained most of the phenotypic variation. For CCh, two QTLs were identified with in total 59.8 % phenotypic variation for high N explained, and a unique QTL for low N was found that explained 31.2 % of the variation. For all the QTLs found for CC, the dominant alleles were coming from Ranchero. Three QTLs were detected for LN at high N of which HN_LN_3 (in P02) explained 24.6 % of the phenotypic variation. The QTLs found at low N co-localized with the ones detected at high N. LN_LN_1 on P01 explained under this condition most of the phenotypic variation (30.2 %).
Flowering
Three QTLs were found at low N (two at P01 and one at P02) for Fl and also at high N, and these together explained most of the variation between lines. HN_Fl_3 and LN_Fl_1 co-localized with strong QTLs for SL at high and low N, respectively, in line with the high correlation between these traits.
NUE
A QTL controlling NUE was found at P01 at low N conditions. This QTL explained 17.0 % of the phenotypic variation with a LOD value of 3.82. This QTL co-localized with multiple QTLs for SDW, LA, SFW, CCh, CCi, Fl, LN and SL (Fig. 4). At high N conditions no QTL was detected.
Discussion
Cultivation of spinach, like many other leafy vegetables, requires high amounts of nitrogen. This puts a strain on the environment, consumes large amounts of fossil energy for the synthesis of synthetic N, and limits the opportunities for spinach as an organically produced crop. This study provides a first step towards a sustainable solution to this problem by providing molecular tools for breeders through identification of genetic factors governing NUE. Improved NUE is beneficial under high N conditions as it helps limiting the amount of nitrogen that leaches into the environment, but is at least equally important at low N availability as it enables the plant to produce more biomass and yield when N availability is limiting. We created a genetic linkage map and several QTL regions were identified that can be breeding targets for improvement of NUE in spinach.
QTL analysis of complex traits (such as NUE) is often hampered by a low heritability of the stress-related traits detected under stress conditions (Lafitte et al. 2004). It is likely that often at least in part this is due to the lack of control over the stress conditions. Our results showed substantial variation under low N for most of the traits, and the heritability was high. This may be partly due to the experimental setup, which involved a hydroponics system and N addition rates according to the Ingestad model. The plants were thus exposed to a highly controlled steady state N stress in a controlled root environment, reducing the environmental variation and improving detection of heritable selection traits under low N availability. This set-up is particularly useful for a genetic study aiming at the traits contributing to N utilisation efficiency.
The genetic map presented in this study was based on SNP markers identified in actively transcribed genes. The map, composed of close to 283 markers in expressed genes divided over 6 linkage groups (matching the basic chromosome number in spinach), presents a significant improvement over the currently available published map by Khattak et al. (2006) with 110 markers and 7 linkage groups. Moreover, the Khattak map contained a considerable number of AFLP markers, which are not easily usable as allelic bridges between different molecular maps. The available sequence information of the SNP markers of the here reported map will enable integration with other maps and physical mapping using the spinach genome sequence, which is expected to become available soon. The current linkage map still contains several gaps and clusters of markers. This may be related to the fact that the markers originate from coding regions of the genome, which are mostly present in the euchromatin. Marker clustering within linkage maps is not restricted to gene-based markers, and not uncommon for random DNA-based markers as well (Qi et al. 1998; Haanstra et al. 1999; Vuylsteke et al. 1999; Young et al. 1999; Jeuken et al. 2001). Brugmans et al. (2002) suggested that the ideal distribution of markers to find functional or causal genes for a trait of interest is not necessarily regular spacing across the whole genome, but rather a concentration of markers in the coding regions of the genome, suggesting that the map presented here is well-suited for QTL analysis.
Gaps and clusters may also result from a lack of homology between the genomes of the parents, reducing recombination frequency. The pedigree of the female parent of the mapping population map includes germplasm from the wild species S. turkestanika (van Diemen, personal communication 2013). However, this was also the case for the parents of the Khattak mapping population, but the markers on this map are not clustered as much as the markers on our map. Another explanation for the clustering and presence of gaps could be linked to the size and structure of the spinach chromosomes. A cytogenetic study of Ramanna (1976) showed that four of the six spinach chromosomes had a low short-to-long arm ratio, which is normally associated with a high number of rod bivalents at the late prophase I or metaphase I of meiosis (for a review, see Jones 1987). Rod bivalents are due to the absence of chiasmata between the chromatids of the short arms of two homologous chromosomes and are therefore indicative for a low degree of recombination (Sanchez-Moran et al. 2002). Since spinach chromosomes are quite small, the overall number of chiasmata that occur within single chromosome pairs may be low, which may at least partly account for the clustering and gaps in our map. It is important to note that this may severely complicate breeding for traits for which the genes are located on the short arms of the spinach chromosomes.
Several phenotypic traits that were measured in this study may be determined by the same biological processes or even a single gene, which is exemplified by the accumulation of QTLs at two locations, i.e. on linkage groups P01 and P02 (Fig. 4). The QTL effects may be strongly influenced by differences in flowering time of the plants; the response to abiotic stress is influenced by maturity of the plants (Vargas et al. 2006, Reynolds and Tuberosa 2008). Flowering influences N use efficiency because the fruits and flowers are strong N sinks (Schieving et al. 1992). Genotypic differences in flowering time may therefore directly affect traits related to plant growth and N use efficiency. Flowering differences were evaluated by measuring presence/absence of flowers (Fl), leaf number (LN) and indirectly, stem length (SL). The six QTLs for Fl co-localized with QTLs for biomass, suggesting that the genetic variation for biomass may be partly determined by flowering traits. However, separate QTL analyses with flowering as a covariate resulted in the same QTLs for biomass traits with similar LOD scores, indicating that the biomass trait QTLs on linkage groups P01 and P02 were to a larger extent determined by other factors than flowering.
The detected flowering QTLs together explained a large part of the total variation both under high and low N conditions, indicating that the majority of the genomic regions controlling flowering in this population were identified. Two other traits directly associated with flowering (stem length and leaf number) showed QTL co-localization with the flowering QTLs, emphasizing the significance of these QTL regions. Together with studies of Khattak et al. (2006), Onodera et al. (2011) and Yamamoto et al. (2014) related to sex expression, these results present a basis for understanding the genetic control of flower development in spinach and similar species.
Nitrogen nutrition plays a crucial role in determining plant photosynthetic capacity in both natural and agricultural environments (Abrol et al. 1999). Because the photosynthetic apparatus utilizes a large part of the available N in the plant, N availability is a key external factor for photosynthetic capacity and plant growth. Photosynthetic capacity is dependent on leaf area and on chlorophyll content of the leaf. A number of studies have demonstrated the correlation of chlorophyll content with N availability. In maize, Crafts-Brandner and Poneleit (1992) found a correlation between Rubisco content, chlorophyll content and photosynthetic activity, and Hageman and Lambert (1988) linked photosynthetic activity to leaf N concentration. SPAD readings were suggested to give good estimations of NO3 − N concentrations in spinach, assisting in the evaluation of N availability in fields and assessment of optimal harvest time (Liu et al. 2006). In cereals, leaf chlorophyll content was decreased under low N conditions (Muchow and Davis 1988; Sinclair and Vadez 2002), and in spinach under suboptimal N conditions a considerable reduction of chlorophyll content (CC) was observed (Evans and Terashima 1987). In most cases, the decrease in CC under low N availability is directly related to stress-induced senescence. In our study CC was not affected by N level, seemingly contradicting results from others. However, our experiments used the N addition model of Ingestad (1982), and N is provided to the plants in an exponential and progressive manner according to a specific relative growth rate (0.10 and 0.18). At the low N addition rate, the plants are able to adapt to the low N availability (Gutshick 1999). The plants do grow slower, but these are likely able to balance photosynthetic capacity and growth to the available N, and the leaves hardly senesce (Chan-Navarrete et al. 2014). The CC measurements at harvest therefore may be indicative for physiological aging of the measured leaves, which was affected by low N availability. This is in agreement with the co-localization of CC QTLs at high and low N. The HN_CCh_1 and LN_CCh_1 QTLs localized in the QTL hotspot at P01, linking CC to biomass production both under low and high N conditions. The Ranchero allele linked to delayed senescence (higher chlorophyll content at end harvest) located on P01 may be an interesting quality target for spinach breeding.
Nitrogen use efficiency was highly correlated with SFW, SDW, RDW and LA under both low and high N conditions. Under low N conditions, leaf area is typically limited, balancing it with the limited N availability (Evans and Terashima 1987, Evans 1989). The genotypes that can retain a relatively high LA while balancing nitrogen and carbon generally would be expected to have higher NUE, which is exemplified by the positive correlation between LA and NUE (Fig. 3) and the colocalization of QTLs for LA, NUE and biomass traits under high N conditions on P01 and P02, and under low N conditions at P1.
R:S was negatively correlated with NUE under high N conditions and DM % did not show any correlation. The lower (negative) correlation between R:S and NUE at low N was likely caused by prioritizing an investment in roots over shoots (the NUE measurements are based on leaf N content). The R:S QTLs HN_RS_2 and LN_RS_1 co-located on P06 at 29.6 cM, indicating that this QTL at last partly controls partitioning of assimilates independent of N-availability. Under low N conditions, DM % increased and a low correlation with NUE was detected, as was also observed by Elia et al. (1998). In addition, we identified QTLs for SLA and LA at P02 under high N conditions, but these QTLs were not found under low N conditions. This points to genotypic differences in how assimilate usage is adapted at low N, and may reflect variation in changing from producing N-rich proteins and maximizing leaf surface area under high N to production and accumulation of carbohydrates (Blaby et al. 2013), production of low-N containing structural components, or production of osmolytes to adapt osmotic balance of the cells to cope with a reduction of the water content. The here observed reduction in LA and prioritization of investment in roots at low N conditions may reflect a response aimed at maintaining the balance between carbon and nitrogen. This is in line with the fact that plants possess an intricate regulatory mechanism that coordinates N metabolism with C metabolism (Nunes-Nesi et al. 2010). Nitrogen availability is directly linked to photosynthetic capacity because the proteins of the Calvin cycle and thylakoids represent the majority of leaf nitrogen (Evans 1989). Limited N availability increases the C/N ratio, with less N available for photosynthetic proteins, and the resulting accumulation of carbohydrates feeds back negatively on photosynthesis (Noguchi and Terashima 2006). The C/N balance can be further restored by a reduction of LA, decreasing photosynthetic capacity, and an increase in the thickness of the leaf, represented by a reduction of SLA (Kant et al. 2011). Indeed we find that SLA is inversely correlated with NUE (−0.40 and −0.31 under low and high N conditions, respectively), indicating that in the plant material evaluated, the restoration of the C/N balance as described above positively affects NUE. We also found that leaf area was highly positively correlated with NUE (0.69 at low N and 0.85 at high N). Moreover, under high N conditions, SLA and RDW are correlated (0.52), while under low N conditions, this correlation disappears. This may imply that with ample N available, investment in roots is accompanied by large and thin leaves, matching increased N uptake capacity with higher photosynthetic surface. Under low N, SLA and RDW are no longer correlated: this may imply that plants grown under low N conditions tend to prioritize roots over shoots (R:S is increased), but this investment in roots is under these conditions not associated with higher SLA. The investment in roots does not compensate for the lower levels of N in the root environment, N-uptake is decreased, and therefore C/N needs to be adapted as well. The strong positive correlation of LA with NUE under low N indicates that there is substantial variation for selecting NUE related traits in the spinach population under study to optimize NUE under N-limiting conditions.
Concluding remarks
The results presented in this study provide a first step towards molecular breeding for complex traits in spinach, in particular for nitrogen use efficiency and adaptation to growth under low input conditions. The identified QTLs may be targets for breeding programs aimed at improving NUE, bith under high N and N limiting condition, thus providing tools to increase yields under low N input conditions, and decrease environmental strain at high N input. However, it should be taken into account that the approach chosen in this study, i.e. the use of a hydroponics system and N addition according to the Ingestad model, focused on QTL detection for traits related mostly to N utilization under highly controlled N stress conditions. The relevance of the detected QTLs under field conditions still needs to be proven.
References
Abrol YP, Chatterjee SR, Kumar PA, Jain V (1999) Improvement in nitrogen use efficiency: physiological and molecular approaches. Curr Sci 76:1357–1364
Arumuganathan K, Earle ED (1991) Nuclear DNA content of some important plant species. Plant Mol Biol Report 9:208–218
Baligar VC, Fageria NK, He ZL (2001) Nutrient use efficiency in plants. In: J.A. Delgado (ed.). Special issue communication soil science plant analysis 32:921–950
Biemond H (1995) Effects of nitrogen on development and growth of the leaves of vegetables. 3. Appearance and expansion growth of leaves of spinach. Neth J Agric Sci 43:247–260
Biemond H, Vos J, Struik PC (1996) Effects of nitrogen on accumulation and partitioning of dry matter and nitrogen of vegetables. 3. Spinach. Neth J Agric Sci 44:227–239
Blaby IK, Glaesener AG, Mettler T, Fitz-Gibbon ST, Gallaher SD, Liu B, Merchant SS (2013) Systems-level analysis of nitrogen starvation-induced modifications of carbon metabolism in a chlamydomonas reinhardtii starchless mutant. Plant Cell Online 25:4305–4323
Breimer T (1982) Environmental factors and cultural measures affecting the nitrate content in spinach. Fertil Res 3:191–292
Brugmans B, Carmen A, Bachem C, Os H, Eck H, Visser R (2002) A novel method for construction of genome-wide transcriptome maps. Plant J 31:211–222
Chan-Navarrete R, Kawai A, Dolstra O, Lammerts van Bueren ET, van der Linden CG (2014) Genetic diversity for nitrogen use efficiency in spinach (Spinacia oleracea L.) cultivars using the ingestad model on hydroponics. Euphytica 199:155–166
Correll JC, Bluhm BH, Feng C, Lamour K, du Toit LJ, Koike ST (2011) Spinach: better management of downy mildew and white rust through genomics, European. J Plant Pathol 129:193–205
Crafts-Brandner SJ, Poneleit CG (1992) Selection for seed growth characteristics: effect on leaf senescence in maize. Crop Sci 32(1):127–131
Elia A, Santamaria P, Serio F (1998) Nitrogen nutrition, yield and quality of spinach. J Sci Food Agric 76(3):341–346
European Commission (2010). European Commission: Report from the commission to the Council and the European Parliament on implementation of the Council Directive 91/676/EEC concerning the protection of water against pollution caused by s from agricultural sources for the period 2004–2007 SEC(2010)118, COM(2007)47 final/2, Brussels, 2011
Evans JR (1989) Photosynthesis and nitrogen relationships in leaves of C3 plants. Oecologia 78:9–19
Evans JR, Terashima I (1987) Effects of nitrogen nutrition on electron transport components and photosynthesis in spinach. Funct Plant Biol 14:59–68
Fageria NK, Baligar VC (2005) Enhancing nitrogen use efficiency in crop plants. Adv Agron 88:97–185
Grabherr MG, Haas BJ, Yassour M, Levin JZ, Thompson DA, Amit I, Regev A et al (2011) Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotech 29:644–652
Gutshick VP (1999) Biotic and abiotic consequences of differences in leaf structure. New Phytol 143:3–18
Haanstra J, Wye C, Verbakel H, Meijer-Dekens F, van den Berg P, Odinot P, van Heusden AW, Tanksley S, Lindhout P, Pelemen J (1999) An integrated high-density RFLP-AFLP map of tomato based on two Lycopersicum esculentum × L. pennellii F2 populations. Theor Appl Genet 99:254–271
Hageman RH, Lambert RJ (1988) The use of physiological traits for corn improvement. Corn Corn Improv 3:431–461
Haldane BS, Waddington CH (1931) Inbreeding and linkage. Genetics 16:357–374
Hirel B, Bertin P, Quilléré I, Bourdoncle W, Attagnant C, Dellay C, Gouy A, Cadiou S, Retailliau C, Falque M, Gallais A (2001) Towards a better understanding of the genetic and physiological basis for nitrogen use efficiency in maize. Plant Physiol 125:1258–1270
Hirel B, Le Gouis J, Ney B, Gallais A (2007) The challenge of improving nitrogen use efficiency in crop plants: towards a more central role for genetic variability and quantitative genetics within integrated approaches. J Exp Bot 58:2369–2387
Ingestad T (1982) Relative addition rate and external concentration; driving variables used in plant nutrition research. Plant Cell Environ 5:443–453
Jeuken M, Van Wijk R, Peleman J, Lindhout P (2001) An integrated interspecific AFLP map of lettuce (Lactuca) based on two L. sativa × L. saligna F2 populations. Theor Appl Genet 103:638–847
Jones GH (1987) Chiasmata. In: Moens P (ed) Meiosis. Academic Press Inc, New York, pp 213–238
Kant S, Bi YM, Rothstein SJ (2011) Understanding plant response to nitrogen limitation for the improvement of crop nitrogen use efficiency. J Exp Bot 62:1499–1509
Khattak JZK, Torp AM, Andersen SB (2006) A genetic linkage map of Spinacia oleracea and localization of a sex determination locus. Euphytica 148:311–318
Lafitte HR, Price AH, Courtois B (2004) Yield response to water deficit in an upland rice mapping population: associations among traits and genetic markers. Theor and Appl Genet 109(6):1237–1246
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9(4):357–359
Lawlor DW, Lemaire G, Gastal F (2001) Nitrogen, plant growth and crop yield. In: Lea PJ, Morot-Gaudry J-F (eds) Plant nitrogen. Springer, Berlin, pp 343–367
Li J, Ji L (2005) Adjusting multiple testing in multilocus analyses using the eigenvalues of a correlation matrix. Heredity 95(3):221–227
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Durbin R (2009) The sequence alignment/map format and SAM tools. Bioinformatics 25(16):2078–2079
Liu Y, Tong Y, Zhu H, Ding E, Smith A (2006) Leaf chlorophyll readings as an indicator for spinach yield and nutritional quality with different nitrogen fertilizer applications. J Plant Nutr 29:1207–1217
Magoč T, Salzberg SL (2011) FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27(21):2957–2963
Muchow RC, Davis R (1988) Effect of nitrogen supply on the comparative productivity of maize and sorghum in a semi-arid tropical environment II. Radiation interception and biomass accumulation. Field Crops Res 18:17–30
Nijveen H, van Kaauwen M, Esselink DG, Hoegen B, Vosman B (2013) QualitySNPng: a user-friendly SNP detection and visualization tool. Nucleic Acids Res. doi:10.1093/nar/gkt333
Noguchi K, Terashima I (2006) Responses of spinach leaf mitochondria to low N availability. Plant Cell Environ. 29:710–719
Nunes-Nesi A, Fernie AR, Stitt M (2010) Metabolic and signaling aspects underpinning the regulation of plant carbon nitrogen interactions. Mol Plant 3(6):973–996
Onodera Y, Itaru Y, Hiroki M, Tanaka A, Niikura S, Yamazaki S, Mikami T (2011) Mapping of the genes for dioecism and monoecism in Spinacia oleracea L.: evidence that both genes are closely linked. Plant Cell Reports 30:965–971
Qi X, Stam P, Lindhout P (1998) Use of locus-specific AFLP markers to construct a high-density molecular map in barley. Theor App Genet 96:376–384
Ramanna MS (1976) Are there heteromorphic sex chromosomes in spinach (Spinacia oleracea L.)? Euphytica 25:277–284
Reynolds M, Tuberosa R (2008) Translational research impacting on crop productivity in drought-prone environments. Curr Opin Plant Biol 11:171–179
Sanchez-Moran E, Armstrong SJ, Santos JL, Franklin FCH, Jones GH (2002) Variation in chiasma frequency among eight accessions of Arabidopsis thaliana. Genetics 162:1415–1422
Schieving F, Werger MJA, Hirose T (1992) Canopy structure, nitrogen distribution and whole canopy photosynthetic carbon gain in growing and flowering stands of tall herbs. Vegetatio 102:173–181
Schmieder R, Edwards R (2011) Quality control and preprocessing of metagenomic datasets. Bioinformatics 27(6):863–864
Schmitz-Linneweber C, Maier RM, Alcaraz JP, Cottet A, Herrmann RG, Mache R (2001) The plastid chromosome of spinach (Spinacia oleracea): complete nucleotide sequence and gene organization. Plant Mol Biol 45:307–315
Sinclair TR, Vadez V (2002) Physiological traits for crop yield improvement in low N and P environments. Plant Soil 245:1–15
Smolders E, Merckx R (1992) Growth and shoot:root partitioning of spinach plants as affected by nitrogen supply. Plant Cell Environ 15:795–807
Steingrover EG (1986) Nitrate accumulation in spinach. Ph.D. thesis, RU Groningen
Stern DB, Palmer JD (1986) Tripartite mitochondrial genome of spinach: physical structure, mitochondrial gene mapping, and locations of transposed chloroplast DNA sequences. Nucleic Acids Res 14:5651–5666
Steward CN, Via LE (1993) A rapid CTAB DNA isolation technique useful for RAPD fingerprinting and other PCR applications. Biotechniques 14:748–750
van Ooijen JW (2006) JoinMap 4, software for the calculation of genetic linkage maps in experimental populations. Kyazma B.V., Wageningen
van Ooijen JW, Kyazma BV (2009) MapQTL 6. Software for the mapping of quantitative trait loci in experimental populations of diploid species. Kyazma B.V., Wageningen
Vargas M, Van Eeuwijk FA, Crossa J, Ribaut JM (2006) Mapping QTLs and QTL environment interaction for CIMMYT maize drought stress program using factorial regression and partial least squares methods. Theor Appl Genet 112:1009–1023
Varshney RK, Nayak SN, May GD, Jackson SA (2009) Next-generation sequencing technologies and their implications for crop genetics and breeding. Trends Biotech 27:522–530
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Heredity 93:77–78
VSN International (2013) Genstat for Windows, 16th edn. VSN International, Hemel Hempstead
Vuylsteke M, Mank R, Antonise R, Bastiaans E, Senior ML, Stuber CW, Melchinger AE, Luebberstedt T, Xia XC, Stam P (1999) Two high-density AFLP® linkage maps of Zea mays L.: analysis of distribution of AFLP markers. Theor Appl Genet 99:921–935
Yamamoto K, Oda Y, Haseda A, Fujito S, Mikami T, Onodera Y (2014) Molecular evidence that the genes for dioecism and monoecism in Spinacia oleracea L. are located at different loci in a chromosomal region. Heredity 112:317–324
Young WP, Schupp JM, Keim P (1999) DNA methylation and AFLP marker distribution in the soybean genome. Theor Appl Genet 99:785–790
Acknowledgments
The authors would like to acknowledge the technical assistance of Degife Zebire, Hanneke van der Schoot, and Elly Janssen. Marcela Viquez-Zamora is acknowledged for her help with the linkage map construction. We thank Maarten Peters and Geurt Versteeg for technical assistance with the hydroponics system. This work was performed in collaboration with Enza Zaden, Vitalis Organic Seeds, Bayer CropScience Vegetable Seeds, Rijk Zwaan, and Pop Vriend Seeds. The research was funded by the Ministry of Economic Affairs of the Netherlands under the Groene Veredeling (Green Breeding) Program.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Chan-Navarrete, R., Dolstra, O., van Kaauwen, M. et al. Genetic map construction and QTL analysis of nitrogen use efficiency in spinach (Spinacia oleracea L.). Euphytica 208, 621–636 (2016). https://doi.org/10.1007/s10681-015-1618-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-015-1618-6