Abstract
Deoxynivalenol (DON) is the major mycotoxin produced by Fusarium fungi in grains. Food and feed contaminated with DON pose a health risk to humans and livestock. The risk can be reduced by enzymatic detoxification. Complete mineralization of DON by microbial cultures has rarely been observed and the activities turned out to be unstable. The detoxification of DON by reactions targeting its epoxide group or hydroxyl on carbon 3 is more feasible. Microbial strains that de-epoxidize DON under anaerobic conditions have been isolated from animal digestive system. Feed additives claimed to de-epoxidize trichothecenes enzymatically are on the market but their efficacy has been disputed. A new detoxification pathway leading to 3-oxo-DON and 3-epi-DON was discovered in taxonomically unrelated soil bacteria from three continents; the enzymes involved remain to be identified. Arabidopsis, tobacco, wheat, barley, and rice were engineered to acetylate DON on carbon 3. In wheat expressing DON acetylation activity, the increase in resistance against Fusarium head blight was only moderate. The Tri101 gene from Fusarium sporotrichioides was used; Fusarium graminearum enzyme which possesses higher activity towards DON would presumably be a better choice. Glycosylation of trichothecenes occurs in plants, contributing to the resistance of wheat to F. graminearum infection. Marker-assisted selection based on the trichothecene-3-O-glucosyltransferase gene can be used in breeding for resistance. Fungal acetyltransferases and plant glucosyltransferases targeting carbon 3 of trichothecenes remain promising candidates for engineering resistance against Fusarium head blight. Bacterial enzymes catalyzing oxidation, epimerization, and less likely de-epoxidation of DON may extend this list in future.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mycotoxins are fungal metabolites toxic to animals. When mycotoxin-producing fungi infect crop plants or colonize stored agricultural commodities, contamination with mycotoxins creates a health risk for humans and livestock. Trichothecenes, which are often found in grains, are sesquiterpenoid mycotoxins with a 12,13-epoxy-trichothec-9-ene skeleton produced fungal genera Fusarium, Myrothecium, Stachybotrys, Trichoderma, and Trichothecium. From an economic perspective, the most important trichothecene producers are those Fusarium species that cause Fusarium Head Blight (FHB) in small-grain cereals and Gibberella Ear Rot (GER) in maize (Bottalico and Perrone 2002). Trichothecenes dominant in grains are deoxynivalenol (DON), nivalenol, and their acetylated derivatives (Foroud and Eudes 2009).
Trichothecenes exert multiple toxic effects on eukaryotic cells, including inhibition of protein synthesis, cytotoxicity, and apoptosis (Rocha et al. 2005; Rotter et al. 1996). Comparison of the effect of trichothecenes on different cell lines confirmed that DON possessed a relatively low toxicity; however, the hazard posed on human health may be underestimated in common tests based on lymphoid cells (Nielsen et al. 2009). The ingestion of DON with contaminated feed and food leads to reduced immunity, anemia, headache, nausea, and abdominal pain in humans and to food refusal, vomiting, growth retardation, and reproductive disorders in animals (Rocha et al. 2005; Sobrova et al. 2010; Pestka 2010). Human and animal exposure to trichothecenes in years with weather conditions conducive to FHB and GER may exceed tolerable limits.
The permissible quantity of DON in harvested grain, food products, and feedstuff is limited by regulations in most countries. Fungicides targeting Fusarium spp. in cereal plants have been available since the 1990s and breeding has gradually improved the resistance of commercial varieties. In spite of these efforts, contamination of grain with DON continues to pose a threat to grain production. The chemical detoxification of DON in solution and on/in contaminated grains by ozone (Young et al. 2006), ammonia, chlorine, hydrogen peroxide, sodium bisulfite (Young et al. 1986), sodium carbonate (Abramson et al. 2005), and chlorine dioxide (Wilson et al. 2005) has been demonstrated, but its application on a large scale has been hampered by costs, safety concerns, and the negative impact on grain quality. The most promising technique for chemical detoxification of DON is reaction with sodium metabisulfite (Young et al. 1987). The process possesses potential for large-scale on-farm decontamination (Dänicke et al. 2010), but is prohibited for food-grade grains in Europe.
Enzymatic detoxification of mycotoxins in situ has been discussed since the 1970s as a strategy for the elimination of mycotoxins from grain, feed, and food. The research was also fuelled by military interest in trichothecenes allegedly used in chemical warfare (Wannemacher and Wiener 1997; Tucker 2001). Although the toxicity of trichothecenes is relatively low, they can be easily manufactured by the fermentation of cheap substrates. Culture supernatants amended with a detergent and dimethyl sulfoxide for improved adherence and skin penetration are chemical weapons ready for delivery. Extensive research on the biological decontamination of trichothecenes was carried out in military facilities and by extramural military-funded research in the 1980s and 1990s (Meister 1987). Military-funded research has not yielded useful agents for biological detoxification (e.g., Beeton and Bull 1989), but it contributed to our understanding of detoxification processes (e.g., Visconti and Mirocha 1985; Chatterjee et al. 1986; Swanson et al. 1987a, b).
Several reviews on the biological detoxification of mycotoxins have been published (Karlovsky 1999; Bata and Lasztity 1999; Styriak and Conkova 2002; Jouany 2007; Halász et al. 2009; Awad et al. 2010), but because of their wide scope, the space devoted to trichothecenes was small. Two reviews focusing on trichothecenes are available (Zhou et al. 2008; He et al. 2010), but they provide a limited coverage of epimerization, de-epoxidation, and glycosylation, as well as of the application of acetylation in plant biotechnology.
Scope of the review
The targets for the detoxification of DON are shown in Fig. 1. Major recent breakthroughs were:
-
(a)
Isolation of pure bacterial cultures that de-epoxidize DON
-
(b)
Genetic dissection of the glycosylation of DON in plants and elucidation of its role in the resistance of wheat to FHB
-
(c)
Engineering of plants for the acetylation of DON
These developments will be covered in depth. The recent discovery of the epimerization of DON on C3 will be described. The reports of complete mineralization of DON will be summarized, and the records of transformation processes by unknown mechanisms will be listed. The use of enzymatic detoxification of DON in plant biotechnology including intellectual property issues will be covered. The controversy about the activity of commercially available feed additives claimed to enzymatically detoxify DON will be discussed.
Except for acetylation of the hydroxyl group on C3, acylation and deacylation will not be covered. Glucuronidation of DON in mammals and observations of a gradual loss of DON in plants in the field, in stored grains, and in the silage will not be treated. The review will also not cover strategies different from enzymatic modification of the toxin molecule such as modification of the binding site on the ribosome and active export.
Destruction of the epoxide group of DON
Function and stability of the epoxide group
A characteristic feature of trichothecenes is an epoxide on C12/C13 (Fig. 1). Since the 1970s, researchers have understood that the epoxide is the primary toxicity determinant of DON (Sato and Ueno, 1977; Eriksen et al. 2004). That the epoxide group is necessary for the inhibition of protein biosynthesis, which is the major mode of action of trichothecenes, was reported in a seminal work by Yoshio Ueno (Ueno et al. 1973) and elucidated in a series of unsurpassed biochemical studies in the 1970s (Cundliffe et al. 1974; Wei et al. 1974; Cundliffe and Davies 1977). Structural data concerning the complex of DON with the ribosome are not available, but the structural diversity of toxic trichothecenes indicates that the toxin binds to the ribosome regardless of the variations in trichothecene structure as long as the skeleton and the epoxide group are preserved. Differences in the ability of natural trichothecenes to cross phospholipid membranes, differences in glucuronidation, as well as different efficiencies of active extrusion of these toxins from the cytoplasm by membrane transporters are likely to contribute to the differences in the toxicity of natural trichothecenes. The modes of action other than inhibition of protein synthesis, such as disruption of cellular membranes by trichothecenes with hydrophobic substituents (Kim et al. 1991), are believed to play only a secondary role in trichothecene toxicity.
The tetrahydropyran and cyclopentane rings of trichothecenes share carbon atoms 2, 12, and 5 (Fig. 1), which force the conformation of the molecule into a rigid fold. Although the spiro attachment of the epoxide group in the oxabicyclo[3.2.1]octane system appears to facilitate nucleophilic attacks leading to hydrolysis, the epoxide group of trichothecenes is stable under neutral and slightly acidic conditions and survives boiling, baking, and even steaming at 135 °C (Pronyk et al. 2006) and extrusion at 150 °C (Bullerman and Bianchini 2007). Trichothecenes carrying intact epoxide group can therefore be found in bread and pastries, pasta, beer, breakfast cereals, and other products.
Two mechanisms leading to destruction of the epoxide group of trichothecenes have been reported: reductive de-epoxidation leading to olefin and hydrolytic de-epoxidation generating two vicinal hydroxyls. The possibility of nucleophilic attack of the epoxide group by thiols in plants was suggested (Subramanian 2002) but not supported by data.
Reductive de-epoxidation
De-epoxidation by mixed cultures
Bacteria in the digestive system of animals are able to reduce the epoxide group of trichothecenes, generating 9,12-diene derivatives. The structure of the product of the de-epoxidation of DON was elucidated nearly three decades ago (Yoshizawa et al. 1983; King et al. 1984). Since then, the de-epoxidation of trichothecenes by mixed populations of ruminal and intestinal bacteria has been extensively documented (Yoshizawa et al. 1985, Swanson et al. 1987a; Lake et al. 1987; Worrell et al. 1989, He et al. 1993; Kollarczik et al. 1994). Negative results reported by some authors (He et al. 1992; Swanson et al. 1987a, b; Munger et al. 1987) may be accounted for by the intestinal or ruminal microbes having not been previously exposed to trichothecenes and therefore having lacked the necessary adaptation. In support of this explanation, Hedman and Pettersson (1997) reported that neither DON nor nivalenol was detoxified in pig feces unless the pigs were fed with a diet containing trichothecenes. The experiments confirming this observation for chickens are described in a recent patent application by Zhou et al. (2010).
The isolation of pure cultures that can detoxify DON from ruminal and intestinal content has proven difficult because these bacteria are strictly anaerobic and have fastidious nutritional demands. Fourteen years after the discovery of trichothecene de-epoxidation, anaerobic bacteria, able to hydrolyze ester bonds of trichothecenes but incapable of de-epoxidation, were isolated from the digestive system of animals (e.g., Visconti and Mirocha 1985; Westlake et al. 1987; Matsushima et al. 1996).
De-epoxidation by pure cultures
The first pure culture that could de-epoxidize DON and other trichothecenes was isolated by Binder et al. (1997). The culture originated from the rumen of a cow. The strain, which was designated Eubacterium sp. BBSH 797, de-epoxidized both type A and type B trichothecenes in vitro under anaerobic conditions (Fuchs et al. 2002a, b). In 1998, the researchers who isolated the strain applied for patent protection, which was awarded (e.g., Binder and Binder 2004). De-epoxidation activity of Eubacterium sp. BBSH 797 was also demonstrated in dissected pig intestine (Schatzmayr et al. 2006).
It took 10 years since the discovery of the strain BBSH 797 to isolate other bacteria that could de-epoxidize trichothecenes. The two strains of unknown taxonomic affiliation, isolated from chicken intestines and designated LS100 and S33, were able to de-epoxidize DON anaerobically (Young et al. 2007). In a follow-up publication, ten bacterial strains belonging to four taxonomically distinct groups that de-epoxidized DON were described, all of them purified from the same source (Yu et al. 2010). Two of the strains (LS-100 and SS-3) appear to be identical to previously described strains (Young et al. 2007). The isolation of pure cultures with de-epoxidation activity from the intestinal content was facilitated by a treatment with antibiotic cocktails that reduced the diversity of the microbial population. Bacteria that survived the treatment were serially diluted and their activity toward DON was determined. The diversity of bacterial populations during the selection was monitored by denaturing gradient gel electrophoresis. The strategy proved successful, yielding pure strains of DON-de-epoxidizing bacteria belonging to Clostridiales, Anaerofilum sp., Collinsella sp., and Bacillus spp. (Yu et al. 2010). In their application for a patent (Zhou et al. 2010), the authors documented that the Bacillus isolate was able to detoxify DON in moldy maize used to feed pigs and that the treatment significantly improved the feed intake, weight gain, and feed efficiency of the animals.
Specificity and biological function of de-epoxidation
Ivie (1976) described de-epoxidation activity of ruminal microflora toward insect juvenile hormone mimics and the insecticide dieldrin before the de-epoxidation of trichothecenes was discovered. He concluded that the activity was nonspecific and speculated that it may serve a nutritional function in the reduction of epoxides in plant fatty acids and cutin. If Ivie was correct, de-epoxidation activities should be widespread among herbivores. Most herbivores in farm fields are insects. The selection of de-epoxidation activities towards fatty acid epoxides of plant origin is therefore likely to take place in herbivorous insects. Furthermore, strong selection pressure for the development of protection mechanisms against trichothecenes is acting on invertebrates that feed on fungi (Rohlfs and Churchill 2011). Insects and other invertebrates are therefore promising candidates for trichothecene detoxification. Transformation activities of insects and insect symbionts toward trichothecenes have been studied (Dowd 1992; Dowd and van Middlesworth 1989), but de-epoxidation has not been observed, indicating that insects may use other mechanisms to protect themselves against trichothecenes.
While enzymatic hydrolysis of the epoxide group of trichothecenes is rare or impossible (see below), reductive de-epoxidation brought about by ruminal and intestinal microflora is common. Because the de-epoxidation enzymes possess little substrate specificity (Ivie 1976) and because their activity is expressed by distantly related bacterial genera (Yu et al. 2010; Fuchs et al. 2002a), it is conceivable that enzymatically catalyzed reductive de-epoxication takes place in anaerobic environments other than the digestive system of mammals. Such environments my serve as a source of new strains suitable for the de-epoxidation of DON.
Can de-epoxidation occur under aerobic conditions?
Most life on Earth depends on oxygen. The de-epoxidation activities toward trichothecenes described so far occur under strictly anaerobic conditions, limiting their practical application. Until recently, aerobic de-epoxidation of trichothecenes had not been observed. Chatterjee et al. (1986) reported the production of de-epoxidized diacetoxyscirpenol by aerobic Fusarium culture, which was (probably incorrectly) characterized as Fusarium graminearum. We hypothesize that the de-epoxy-trichothecene resulted from a defect in biosynthesis, preventing the epoxidation of trichothecene intermediate trichodiene, rather than from de-epoxidation.
A recent finding indicates that some of the anaerobic bacterial strains de-epoxifying DON may be facultative aerobes. The partial 16S rRNA sequence of one of the bacterial isolates from chicken digesta that detoxify DON under anaerobic conditions shared 99% similarity to the 16S sequence of Bacillus arbutinivorans (Yu et al. 2010). Anaerobic Bacillus species are common opportunistic pathogens of mucosa but most species related to B. arbutinivorans are aerobic, plant growth-promoting soil microbes. It is tempting to speculate that the strain discovered by Yu et al. (2010) may turn out to be a facultative anaerobe, which would likely improve its survival in feed additives.
The possibility of de-epoxidation of DON under aerobic conditions may also be inferred from the study reported by Guan et al. (2009), who investigated the transformation of DON by fish digesta. The availability of oxygen was not specified in the publication, but the description of media, culture conditions, and screening for DON transformation contains no indication that the incubations were performed under anaerobic conditions. From one of 62 fish investigated, the authors isolated a mixed microbial culture, designated C133, that de-epoxidized DON over a surprisingly wide range of pH (from 4.5 to 10.4). If the incubation was performed under aerobic conditions and if a pure active culture can be isolated from this source, the aerobic de-epoxidation of trichothecenes would be a fundamental breakthrough for the development of biotechnological applications. Another advantage of the culture C133 relative to previously described cultures is that it has de-epoxidation activity at low temperatures (as low as 4 °C).
Hydrolytic de-epoxidation
Epoxides are susceptible to hydrolysis and other transformations initiated by nucleophilic attack on one of their carbon atoms. The epoxide group of DON, however, is resistant to hydrolysis. An Austrian patent (Erber 1996) claimed that a strain of the yeast Saccharomyces telluris was able to hydrolyze the epoxide group of trichothecenes, but to our knowledge no data supporting the claim are available. We incubated a number of strains of Arxiozyma telluris (syn. of S. telluris) with DON but detected no transformation (unpublished results). Theisen and Berger (2005) tested 30 strains of bacteria, yeast, and fungi known to produce epoxide hydrolases but could not detect any de-epoxidation of DON. Nakamura et al. (1977) found that trichothecenes were resistant to epoxide hydrolase from rat liver. We also incubated a series of commercially available epoxide hydrolases with DON under conditions recommended by the manufacturers but did not detect any transformation activity (unpublished results).
He et al. (2010) remarked that epoxides can be transformed enzymatically by the epoxide hydrolase and that the biochemical details of the pathways leading from trichothecenes to their de-epoxide forms are uncertain. The negative results of the screening projects attempted so far indicate that such a pathway may not exist in nature.
Conjugation of epoxide group to glutathione
Foster et al. (1975) pointed out that the thiol group of glutathione may act as a nucleophile, attacking the epoxide group of trichothecenes. The reaction is an important detoxification mechanism for the epoxides of aflatoxins (Hayes et al. 1991). The first attempt to detoxify trichothecenes by conjugation with glutathione, without enzymatic catalysis as well as with rat liver glutathione transferase, was performed in the laboratory of Yoshio Ueno (Nakamura et al. 1977). The attempt failed. Optimization of the activity of plant glutathione-S-transferases towards trichothecenes by DNA shuffling was envisioned in a patent awarded to Maxygen (Subramanian 2002). Analysis of barley genes induced by DON treatment and observation that cystein increased the resistance of yeast to DON indicate that glutathione-S-transferase might be involved in DON detoxification in plants (Gardiner et al. 2010). The authors demonstrated the formation of at least three conjugates of DON with glutathione in vitro. The structures of the conjugates have not been established conclusively but the conjugation did not appear to involve the epoxide group (Gardiner et al. 2010).
Modification of the hydroxyl on C3
Role of the hydroxyl on C3 in toxicity
In addition to the epoxide group of trichothecenes, the hydroxyl on carbon 3 plays a major role in toxicity. Temporary acetylation of this hydroxyl during biosynthesis of trichothecenes protects the fungus from toxic effects of the intermediates (Kimura et al. 1998a, b). The use of the enzyme responsible for the acetylation of DON in genetically engineered crops (reviewed by Alexander 2008, see the “Industrial applications of the biological detoxification of deoxynivalenol” or the “Genetically modified crops” section below) can therefore be regarded as an example of pathogen-derived resistance, which has been mainly applied to resistance to plant viruses (Johnson and Sanford 1998).
Acetylation
Gene Tri101 encoding trichothecene-3-O-acetyltransferase from F. graminearum was characterized in the RIKEN laboratory in Japan (Kimura et al. 1998a, b). At the same time, a homologous gene from Fusarium sporotrichioides was cloned at the USDA ARS in Peoria (USA) (McCormick et al. 1999).
The three-dimensional structures and kinetic properties of trichothecene-3-O-acetyltransferases from F. graminearum and F. sporotrichioides have been studied in detail (Garvey et al. 2008). Khatibi et al. (2011) expressed, purified, and compared the properties of trichothecene-3-O-acetyltransferases from seven Fusarium species with the goal of identifying an optimal source of the enzyme for biotechnological applications.
The esterification of C3-OH of DON decreased the toxicity in rabbit reticulocytes by two orders of magnitude (Kimura et al. 1998a) and in rabbit kidney cells by a factor of 20 (Rotter et al. 1993). Acetylation of C3-OH is therefore regarded as detoxification, though the acetyl group may be liberated by hydrolysis in the digestive system of animals. The effect of acetylation on DON on phytotoxicity is less pronounced. Wang and Miller (1988) reported that DON and 3-acetyl-DON had similar effects on wheat tissue. The growth of wheat coleoptiles was also inhibited by both compounds to a similar extent (Bruins et al. 1993). In Arabidopsis thaliana, esterification of C3-OH diminished the toxicity in some cases but increased or left the toxicity unchanged in other cases (Desjardins et al. 2007). Considering these findings, an expectation that the resistance to FHB in cereal crops expressing trichothecene-3-O-acetylase will increase dramatically might be overly optimistic (see the “Genetically modified crops” section below).
Glycosylation
The conjugation of DON with glucose was suggested in 1986 for a wheat suspension culture (Miller and Arnison 1986) and later proven for a maize suspension culture (Sewald et al. 1992). The latter authors determined the structure of the conjugation product to be 3-β-d-glucopyranosyl-4-deoxynivalenol. It took a decade to demonstrate the production of the conjugate in intact plants (A. thaliana; Poppenberger et al. 2003); the authors applied for patent protection (Glossl et al. 2005; Poppenberger et al. 2006). Co-localization of the ability to convert DON to DON-3-glucoside and resistance to FHB in wheat proved that the glycosylation of DON is a major resistance factor against FHB in wheat (Lemmens et al. 2005). Although DON-3-glucoside represents up to 46% of the total amount of DON in infected wheat and maize varieties (Berthiller et al. 2009), cloning the responsible glucosyltransferase proved difficult because of the high number of UDP-glycosyltransferase genes in the genomes of cereal plants. The wheat gene TaUGT3 was suggested as a candidate (Lulin et al. 2010) but it did not protect yeast from DON, while the HvUGT13248 gene from barley did (Schweiger et al. 2010). Furthermore, DON was converted to DON-3-glucoside by yeast cultures expressing HvUGT13248.
Overexpression of a suitable UDP-glycosyltransferase in transgenic wheat or maize appears to be a promising strategy for increasing the resistance against FHB and GER. Alternatively, markers coupled to wheat genes encoding suitable UDP-glucosyltransferases genes may be used for marker-assisted selection in breeding for resistance.
Oxidation and epimerization
Active cultures from three continents
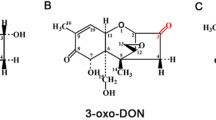
The oxidation of DON to 3-keto-DON was reported for three bacterial cultures: a Gram-negative bacterium isolated in Japan and belonging to the Agrobacterium-Rhizobium group (Shima et al. 1997); a mixed culture D107 isolated in Germany (Volkl et al. 2004); and a bacterial strain isolated in Canada of unknown taxonomic affiliation and dubbed Barpee (unpublished results by He and Zhou, quoted in Zhou et al. 2008). The epimerization of C3-OH of DON was also reported for three cultures: a Gram-positive bacterium Nocardioides sp. WSN05-2 isolated in Japan (Ikunaga et al. 2011); an α-Proteobacterium isolated from a mixed culture D107, related to Devosia riboflavina, and designated HOH107 (Volkl and Karlovsky, unpublished); and the bacterium Barpee from Canada (see above). The time course of the concentrations of DON, 3-oxo-DON, and 3-epi-DON during the transformation of DON by strain HOH107 revealed the sequence of conversions shown in Fig. 2. The same sequence may account for the activity of all of these cultures, provided the ratio of the rates of oxidation and reduction is high for the bacterium Nocardioides sp. WSN05-2 so that the intermediate does not accumulate. A summary of the published data concerning these transformations and assuming that 3-oxo-DON is always an intermediate is shown in Table 1.
Why does epimerization of DON appear to be irreversible?
The oxidation of alcohols and the reduction of ketones are reversible reactions. An intriguing question arises: why do concentrations of DON and 3-epi-DON not reach equilibrium in cultures that can oxidize DON to 3-oxo-DON as well reduce 3-oxo-DON to 3-epi-DON? The difference between the free energy of the two compounds due to steric constraints and possibly a hydrogen bond formed between the hydroxyl of 3-epi-DON and the epoxide group (Nagy et al. 2005) is an unlikely explanation of why essentially all DON is converted to 3-epi-DON. We speculate that the completeness of the conversion results from the coupling of the transformation to another pathway that drives it away from equilibrium. For example, DON may be oxidized to 3-oxo-DON in a reversible reaction catalyzed by an alcohol dehydrogenase that is specific for the R configuration of the hydroxyl on C3. The product may serve as electron acceptor for a membrane-bound component of the respiration chain that reduces the ketone to hydroxyl in the S configuration. Coupling the reduction of 3-oxo-DON with electron transmission along the respiration chain renders the process irreversible, driving the conversion of DON to 3-epi-DON away from equilibrium.
How common is DON epimerization in nature?
Another intriguing question is whether epimerization of DON is common or rare in nature. Conversion of DON to 3-epi-DON requires two enzymatic activities of partially overlapping substrate specificity, occurring at the same time and space (Fig. 2). This may be a general reason why isomerizations are rarely observed in screening for biological degradation. Regarding trichothecenes, the only isomerization reported apart from the conversion of DON to 3-epi-DON is the epimerization of a secondary alcohol group on the macrocyclic ring of type D trichothecenes in Baccharis sp. (Jarvis et al. 1981). On the other hand, bacterial cultures that can epimerize DON have been found on three continents (Table 1). The chromatographic properties of DON and 3-epi-DON are similar so that the epimerization may have been overlooked in screening programs relying on high-throughput, low-resolution thin-layer chromatography. Elucidation of the reaction mechanisms and identification of the genes encoding enzymes involved in the process will help answer the question asked in the subheading by enabling researchers to search for homologous genes in the genomes of soil bacteria as well as in the growing number of available soil metagenomes.
Complete mineralization of DON
Mineralization by mixed cultures
The large amounts of DON are produced in plant debris remaining on the soil surface after harvest yet no accumulation of DON in soil or surface water has been observed, indicating that the trichothecene is being biodegraded. Screening of bacterial populations from soil and surface water for DON mineralization, however, regularly failed to detect biodegradation. Negative results of such projects were only published in cases when successful detoxification of another mycotoxin was achieved (Beeton and Bull 1989; unpublished results of He and Zhou cited in Zhou et al. 2008). Personal communications with colleagues in academia and industry indicate that the number of screening projects to identify microbial cultures detoxifying DON is much higher than is known from literature; most of these attempts failed and were not published.
The simplest explanation of why it has been difficult to demonstrate mineralization of DON by soil microflora is the inability of the active organisms to grow in laboratory media. Even if the organisms are culturable, the media used for screening may be unsuitable. Guan et al. (2009) reported that medium FM, which was previously used in the successful screening of soil microbes for DON transformation (Volkl et al. 2004), was the most suitable among seven media tested for DON transformation by culture C133 from fish digesta. No transformation was observed in minimal medium and in the common bacterial medium LB. Only low conversion was achieved in media with a single nitrogen source such as tryptone, peptone, yeast extract, (NH4)2HPO4, or NH4NO3. Therefore, the screening for DON mineralization should use different media in parallel.
Mineralization of DON by pure cultures
Several bacterial strains capable of complete removal of DON from the medium by mineralization have been described; unfortunately, most of these activities turned out to be unstable. The first pure culture reported to mineralize trichothecenes was Curtobacterium sp. strain 114–2, which was isolated from soil in the laboratory of Yoshio Ueno from soil (Ueno et al. 1983). The complete mineralization of trichothecenes was not rigorously proven but the growth of the strain with nivalenol as a sole carbon source and the fact that the epoxide group was lost after transformation strongly indicated that the trichothecene skeleton was destroyed. The strain lost its activity (Y. Ueno, personal communication).
The symbiotic yeast Pseudotaphrina kochii, which was isolated from an insect, was reported to utilize DON as the only source of carbon (Dowd and Shen 1991; Shen and Dowd 1991), but we were unable to detect any activity of the strain toward DON (Volkl and Karlovsky, unpublished results).
The soil bacterium E3-39, which was assigned to the Agrobacterium-Rhizobium group, completely removed DON from the medium after converting it to the intermediate 3-oxo-deoxynivalenol (Shima et al. 1997). Surprisingly, the DON-oxidizing activity was present in culture supernatant but not in sonicated cells, indicating that an extracellular enzyme was responsible. No follow-up publication or patent on the strain appeared since 14 years; we therefore assume that the activity has been lost.
Strain WSN05-2 (Ikunaga et al. 2011) completely removed DON from the medium, producing 3-epi-DON as an intermediate (see part of the “Oxidation and epimerization” section). This strain is currently the only available organism that is able to completely catabolize DON (Table 2).
Some reports of mineralization can likely be accounted for by adsorption
The utilization of DON as a carbon source by Rhizopus oryzae and Aspergillus oryzae was suggested in a recent report (Garda-Buffon and Badiale-Furlong 2010), but the data presented in the publication may be explained by adsorption of DON onto the mycelium of the fungus. Similarly, the reduction of DON concentration by up to 37% after incubation with yeast cultures (Styriak et al. 2001) and the loss of DON in fodder inoculated with several yeast genera reported by Paskevicius et al. (2006) may have been caused by adsorption. In our screening for DON transformation, we often observed loss of DON from the culture medium, but subsequent treatment of pelleted cells with proteinase K, detergents, and chaotropic agents led to the release of DON, i.e., DON was not degraded but was adsorbed or taken up by the cells (Volkl and Karlovsky, unpublished).
DON transformation by unknown mechanisms
Several published reports described microbial transformations of DON without revealing chemical structures of the products. Although it is not known whether these processes lead to detoxification, the reports identify interesting candidates for further studies.
Filamentous fungi have often been reported to transform DON and other trichothecenes into products with different chromatographic behaviors and/or different toxicities. The conversion of DON by Aspergillus tubingensis was reported by He et al. (2009). The molecular weight of the product was 18.1 kDa larger than that of DON, and the product was fluorescent and eluted from a reverse-phase column earlier than DON. Alternaria alternata transformed DON into a single more polar product that retained the epoxide group (Theisen and Berger 2005). Jesenska and Sajbidorova (1991) described transformations of T-2 toxin by several filamentous fungi leading to products with reduced toxicity.
Roots of sweet potato transformed DON into unknown products (Fujita and Yoshizawa 1989 and Fujita and Yoshizawa 1990). According to what we learned from Arabidopsis and wheat (see the “Modification of the hydroxyl on C3” section, subtitle “Glycosylation”), these products were likely glucoconjugates. The same mechanism likely explains the detoxification of DON by FHB-resistant wheat described by Yao et al. (1996).
Cheng et al. (2010) described removal of DON as determined by ELISA after incubation with Bacillus licheniformis and Bacillus subtilis strains under anaerobic conditions. Interestingly, DON transformation activity was associated with culture supernatants rather than with pelleted cells and was thermolabile, indicating that the loss of DON cannot be accounted for by adsorption. The character of the transformation remains unknown. To demonstrate the suitability of these strains for practical applications, the authors incubated ground wheat and maize grains with the strains and determined that B. subtilis removed 80% of DON.
Flesch and Vogt-Scheuermann (1994) reported that incubation of trichothecin with Saccharomyces cerevisiae lead to the destruction of its epoxide group and rearrangement of the skeleton. Trichothecin is a type B trichothecene similar to DON and is produced by Trichothecium roseum and, according to a single report, by a particular strain of F. graminearum (Combrinck et al. 1988). Because several laboratories studied the effect of trichothecenes on S. cerevisiae without observing any transformation of the toxin, and because no follow-up report from the Flesch/Vogt-Scheuermann laboratory appeared, S. cerevisiae does not seem to be a promising detoxification agent for trichothecenes.
Industrial applications of the biological detoxification of deoxynivalenol
Microorganisms that can detoxify DON can be used as feed additives, and the genes encoding detoxification activities can be used in the genetic engineering of crops. Both strategies are being pursued by industry.
Feed additives
A proof of concept for the decontamination of grain containing trichothecenes was provided long before pure cultures of bacteria that de-epoxidize trichothecenes became available. He et al. (1993) reported that more than 50% of the DON in moldy maize was degraded when the maize was incubated with chicken intestine contents under anaerobic conditions. The ratio of chicken digesta to maize grain was large (1:1), but the result motivated further research.
A feed additive containing microorganisms that is claimed to detoxify DON by enzymatic de-epoxidation is on the market since two decades but its efficacy has often been questioned. Results of animal trials both supporting and disputing the claims of the manufacturer have been published. While a team associated with the company producing the additive reported positive effects (Plank et al. 2009), another group found that the additive decreased livestock performance in a manner independent of mycotoxin concentration in the feeds (Dänicke et al. 2003). As pointed out by Döll and Dänicke (2004), most published studies investigating feed additives that are claimed to biologically detoxify mycotoxins suffer from inappropriate experimental design and from the failure to monitor relevant parameters. So far, most in vivo studies that used an appropriate experimental design and that monitored relevant specific parameters did not find measurable detoxifying effects of commercial feed additives (Dänicke et al. 2004). To determine whether the ability of the product to biologically degrade trichothecenes can be proven in vitro, Döll et al. (2004) incubated the additive with DON in a model system simulating the conditions of the porcine gastrointestinal tract. They did not detect any transformation of DON.
Since 1998, a new DON-detoxifying strain Eubacterium BBSH 797 has been incorporated into the product in some countries (Biomin 2011; see the “Reductive de-epoxidation” or the “De-epoxidation by pure cultures” section). The strain is able to de-epoxidize DON in pure culture (Fuchs et al. 2002a, b) as well as in dissected pig intestine under anaerobic conditions (Schatzmayr et al. 2006). Ingested with feed, the strain should unfold its activity in the digestive system of animals. Because it is a strict anaerobe, formulation securing the survival of the strain during storage and transportation of the product is a prerequisite for the success of the strategy.
New trichothecene-detoxifying strains (Young et al. 2007, Guan et al. 2009, Yu et al. 2010) and particularly strains active under aerobic conditions (if they ever become available) may improve the efficacy of feed additives intended to protect livestock against trichothecenes by enzymatic de-epoxidation. Such products should be evaluated by independent institutions according to a complete two by two factorial design as recommended by Döll and Dänicke (2004).
Genetically modified crops
Equipping crops with DON detoxification activities may reduce the mycotoxin concentration in grain and also increase plant resistance against infection. The idea stems from the fact that trichothecenes act as virulence factors, facilitating colonization of some plant hosts by certain trichothecene-producing Fusarium species. The role of DON in infection was proven when host plants were experimentally inoculated with fungal strains in which trichothecene synthesis was inactivated failed to infect (Proctor et al. 1995; Maier et al. 2006). The role of DON in infection was also supported by research demonstrating that a major QTL responsible for the resistance of wheat to FHB co-segregated with the ability to detoxify DON by glycosylation (Lemmens et al. 2005).
With the availability of genes encoding suitable enzymes, it has become possible to commercialize transgenic crops detoxifying trichothecenes. The RIKEN group who isolated the first Tri101 gene encoding trichothecene-3-acetylase applied for a patent only in Japan (Yamaguchi et al. 2000), which allowed Syngenta to pursue worldwide patent protection (Hohn et al. 2000). In the USA, Syngenta readily obtained a patent with 390 claims covering the use of any trichothecene-3-O-acetylase with a homology to Tri101 protein (Hohn et al. 2002). In Europe, Syngenta limited the claims to Tri101 from F. sporotrichioides. The arguments between the company and the European Patent Office (EPO) lasted for 6 years. In 2007, the EPO was ready to grant a patent but Syngenta abandoned the application. Maxygen Inc. received legal protection for the improvement of the activity of acetyl-O-transferase for trichothecenes by DNA shuffling (Subramanian 2002).
Before the patent battle in Europe was finished, Syngenta carried out field trials with wheat expressing Tri101 in Canada, USA, Argentina, and three European countries. Because of a pending patent application, the company did not reveal the nature of the genetic modification to the public. However, technical details regarding plasmid constructs used for wheat transformation in the published version of the application for approval of the release of transgenic wheat in Germany indicate that the wheat expressed the trichothecene-3-O-acetylase gene from F. sporotrichioides under the control of maize ubiquitin promoter and nopaline synthase terminator. Unfortunately, the field trials were vandalized, preventing the evaluation of the effect of the modification on the tolerance of plants to infection with trichothecene-producing Fusarium species.
Independently from Syngenta, scientists at the USDA/ARS (USA) constructed transgenic wheat using the same Tri101 gene from F. sporotrichioides but a different selection marker (Okubara et al. 2002; Alexander 2008). They reported only a moderate increase in resistance to F. graminearum infection. Tri101 was also expressed in Arabidopsis, rice and barley (Table 3). In Arabidopsis and rice, only the effect of pure toxin has been tested; infection with trichothecene-producing Fusarium species has not been investigated. In barley, no effect of DON-detoxifying activity on resistance to fungal infection in field trials was observed (Manoharan et al. 2006). The outcome was in line with the data of Jansen et al. (2005) and Maier et al. (2006) who reported that DON is not a virulence factor in barley.
A likely reason for the limited success of the wheat trial described in the previous paragraph was that the gene from F. sporotrichioides was not well suited to the purpose. Enzymatic characterization of purified trichothecene-3-O-acetylases from F. graminearum and F. sporotrichioides revealed that the former enzyme is more promising. Affinity for DON is 130 times greater with the F. graminearum enzyme than with the F. sporotrichioides enzyme, and catalytic activity is 70 times greater with the F. graminearum enzyme than with the F. sporotrichioides enzyme (Garvey et al. 2008). Enzymes from other Fusarium species (Khatibi et al. 2011) or enzymes optimized by DNA shuffling (Subramanian 2002) are other alternatives to the F. sporotrichioides enzyme. Testing Tri101 from F. graminearum in wheat is still worth trying but Syngenta no longer seems to be pursuing this strategy; the company abandoned its European application for a pertinent patent.
Outlook
Mechanisms involved in the anaerobic de-epoxidation of DON and in the oxidation and epimerization of the hydroxyl on C3 are being studied in several laboratories. Technical barriers, however, may prevent the use of these activities in transgenic plants. De-epoxidation is likely to require strictly anaerobic conditions, and oxidation/epimerization may be catalyzed by membrane-bound systems that are difficult to express and assemble in plants. The use of intact microorganisms in feed additives appears more feasible. Proofs of concept for de-epoxidation as well as epimerization of DON in grain by bacterial cultures have been provided. The exploitation of these microbes in feed additives appears to be limited only by the development of suitable formulations.
Acetylation and glycosylation of C3-OH, which are well understood and achievable in transgenic plants, are promising activities for plant biotechnology. Acetyltransferase from F. sporotrichioides, which was probably selected in the past because of intellectual property issues rather than for scientific reasons, is likely to be replaced by more active enzymes. Molecular markers associated with genes encoding plant glucosyltransferases that detoxify DON will help breeders develop new wheat varieties resistant against FHB. In the future, the efficiency of detoxification may be improved by overexpressing the enzyme in transgenic plants, ideally under the control of a promoter induced by infection. Optimization of DON acetylases and glycosylases by DNA shuffling or by structure-aided design may improve their kinetic properties for the next generation of transgenic crops.
Mineralization is the most enigmatic of the detoxification processes studied so far. Although the catabolism of DON is likely to be widespread in arable soils, the isolation of active pure cultures has proven to be extremely difficult. When such attempts have succeeded, the cultures have tended to lose their activity in the laboratory. In spite of these difficulties, the high potential of DON catabolism for biotechnology is likely to encourage laboratories to conduct new screening and enrichment programs for DON degradation in the future.
References
Abramson D, House JD, Nyachoti CM (2005) Reduction of deoxynivalenol in barley by treatment with aqueous sodium carbonate and heat. Mycopathologia 160:297–301
Alexander N (2008) The TRI101 story: engineering wheat and barley to resist Fusarium head blight. World Mycotoxin J 1:31–37
Anonymous (2004) Fungal resistant wheat Germany 2004. Report of EC Joint Research Centre. http://gmoinfo.jrc.ec.europa.eu/finalreports/B-DE-03-151-Final-Report.pdf. Accessed 19 March 2011
Awad WA, Ghareeb K, Bohm J, Zentek J (2010) Decontamination and detoxification strategies for the Fusarium mycotoxin deoxynivalenol in animal feed and the effectiveness of microbial biodegradation. Food Addit Contam A 27:510–520
Bata A, Lasztity R (1999) Detoxification of mycotoxin-contaminated food and feed by microorganisms. Trend Food Sci Technol 10:223–228
Beeton S, Bull AT (1989) Biotransformation and detoxification of T-2 oxin by soil and freshwater bacteria. Appl Environ Microbiol 55:190–197
Berthiller F, Dall’asta C, Corradini R, Marchelli R, Sulyok M, Krska R, Adam G, Schuhmacher R (2009) Occurrence of deoxynivalenol and its 3-beta-D-glucoside in wheat and maize. Food Addit Contam A 26:507–511
Binder EM, Binder J (2004) Strain of Eubacterium that detoxifies trichothecenes. US Patent 6:794–175
Binder J, Horvath EM, Schatzmayr G, Ellend N, Danner H, Krska R, Braun R (1997) Screening for deoxynivalenol-detoxifying anaerobic rumen microorganisms. Cereal Res Commun 25:343–346
Biomin (2011) Biomin writes history. http://www.biomin.net/en/company/milestones. Accessed 16 March 2011
Bottalico A, Perrone G (2002) Toxigenic Fusarium species and mycotoxins associated with head blight in small-grain cereals in Europe. Eur J Plant Path 108:611–624
Bruins MBM, Karsaï I, Schepers J, Snijders CHA (1993) Phytotoxicity of deoxynivalenol to wheat tissue with regard to in vitro selection for Fusarium head blight resistance. Plant Sci 94:195–206
Bullerman LB, Bianchini A (2007) Stability of mycotoxins during food processing. Int J Food Microbiol 119:140–146
Chatterjee K, Pawlosky RJ, Mirocha CJ, Zhu TX (1986) Production of deepoxy-diacetoxyscirpenol in a culture of Fusarium graminearum. Appl Environ Microbiol 52:311–313
Cheng BC, Wan CX, Yang SL, Yu HY, Wei H, Liu JS, Tian WH, Zeng M (2010) Detoxification of deoxynivalenol by Bacillus strains. J Food Saf 30:599–614
Combrinck S, Gelderblom WC, Spies HS, Burger BV, Thiel PG, Marasas WF (1988) Isolation and characterization of trichothecin from corn cultures of Fusarium graminearum MRC 1125. Appl Environ Microbiol 54:1700–1702
Cundliffe E, Davies JE (1977) Inhibition of initiation, elongation and termination of eukaryotic protein synthesis by tricothecene fungal toxins. Antimicrob Agent Chemother 11:491–499
Cundliffe E, Cannon M, Davies JE (1974) Mechanism of inhibition of eukaryotic protein synthesis by trichothecene fungal toxins. Proc Natl Acad Sci USA 71:30–34
Dänicke S, Matthes S, Halle I, Ueberschär KH, Döll S, Valenta H (2003) Effects of graded levels of Fusarium toxin-contaminated wheat and of a detoxifying agent in broiler diets on performance, nutrient digestibility and blood chemical parameters. Br Poult Sci 44:113–326
Dänicke S, Valenta H, Döll S, Ganter M, Flachowsky G (2004) On the effectiveness of a detoxifying agent in preventing fusario-toxicosis in fattening pigs. Anim Feed Sci Technol 114:141–157
Dänicke S, Pahlow G, Beyer M, Goyarts T, Breves G, Valenta H, Humpf HU (2010) Investigations on the kinetics of the concentration of deoxynivalenol (DON) and on spoilage by moulds and yeasts of wheat grain preserved with sodium metabisulfite (Na2S2O5, SBS) and propionic acid at various moisture contents. Arch Anim Nutr 64:190–203
Desjardins AE, McCormick SP, Appell M (2007) Structure–activity relationships of trichothecene toxins in an Arabidopsis thaliana leaf assay. J Agric Food Chem 55:6487–6492
Döll S, Dänicke S (2004) In vivo detoxification of Fusarium toxins. Arch Anim Nutr 58:419–441
Döll S, Dänicke S, Valenta H, Flachowsky G (2004) In vitro studies on the evaluation of mycotoxin detoxifying agents for their efficacy on deoxynivalenol and zearalenone. Arch Anim Nutr 58:311–324
Dowd PF (1992) Insect fungal symbionts: a promising source of detoxifying enzymes. J Indust Microbiol 9:149–161
Dowd PF, Shen SK (1991) Biological pure culture of yeast strain used for the microbial detoxification of xenobiotics. US Patent 4:968–620
Dowd PF, van Middlesworth FL (1989) In vitro metabolism of the trichothecene 4-monoacetoxyscirpenol by fungus- and non-fungus-feeding insects. Experientia 45:393–395
Erber E (1996) Patent application WO96/12414
Eriksen SG, Pettersson H, Lundh T (2004) Comparative cytotoxicity of deoxynivalenol, nivalenol, their acetylated derivatives and de-epoxy metabolites. Food Chem Toxicol 42:619–624
Flesch P, Vogt-Scheuermann I (1994) About the decomposition of trichothecin and iso-trichothecin during the alcoholic fermentation of grape juice. Vitic Enol Sci 49:180–184
Foroud NA, Eudes F (2009) Trichothecenes in cereal grains. Int J Mol Sci 10:147–173
Foster PMD, Slater TF, Patterson DSP (1975) A possible enzymic assay for trichothecene mycotoxins in animal feedstuffs. Biochem Soc Trans 3:875–878
Fuchs E, Binder EM, Heidler D, Krska R (2002a) Characterization of metabolites after the microbial degradation of A- and B-trichothecenes by the bacterial strain BBSH 797. Mycotoxin Research 16(S1):66–69
Fuchs E, Binder EM, Heidler D, Krska R (2002b) Structural characterization of metabolites of the microbial degradation of type A trichothecenes by the bacterial strain BBSH797. Food Addit Contam 19:379–386
Fujita M, Yoshizawa T (1989) Induction of phytoalexins by various mycotoxins and metabolism of mycotoxins in sweet potato root tissues. J Food Hyg Soc Jpn 30:501–505
Fujita M, Yoshizawa T (1990) Matabolism of deoxynivalenol, a trichothecene mycotoxin, in sweet potato root tissues. J Food Hyg Soc Jpn 31:474–478
Garda-Buffon J, Badiale-Furlong E (2010) Kinetics of deoxynivalenol degradation by Aspergillus oryzae and Rhizopus oryzae in submerged fermentation. J Braz Chem Soc 21:710–714
Gardiner SA, Boddu J, Berthiller F, Hametner C, Stupar RM, Adam G, Muehlbauer GJ (2010) Transcriptome analysis of the barley-deoxynivalenol interaction: evidence for a role of glutathione in deoxynivalenol detoxification. Mol Plant Microbe Interact 23:962–976
Garvey GS, McCormick SP, Rayment I (2008) Structural and functional characterization of the TRI101 trichothecene 3-O-acetyltransferase from Fusarium sporotrichioides and Fusarium graminearum: kinetic insights to combating Fusarium head blight. J Biol Chem 283:1660–1669
Glossl J, Ruckenbauer P, Schuhmacher R, Luschnig C, Kuchler K, Adam G, Poppenberger B, Sieberer T, Lucyshyn D, Berthiller F (2005) Method for detoxification of mycotoxins. Eur Pat Appl EP 1:510–573
Guan S, He JW, Young JC, Zhu HH, Li XZ, Ji C, Zhou T (2009) Transformation of trichothecene mycotoxins by microorganisms from fish digesta. Aquaculture 290:290–295
Halász A, Lásztity R, Abonyi T, Bata A (2009) Decontamination of mycotoxin-containing food and feed by biodegradation. Food Rev Int 25:284–298
Hayes JD, Judah DJ, McLellan LI, Neal GE (1991) Contribution of the glutathione S-transferases to the mechanisms of resistance to aflatoxin B1. Pharmacol Ther 50:443–472
He P, Young LG, Forsberg C (1992) Microbial transformation of deoxynivalenol (vomitoxin). Appl Environ Microbiol 58:3857–3863
He P, Young LG, Forsberg C (1993) Microbially detoxified vomitoxin-contaminated corn for young pigs. J Anim Sci 71:963–967
He C, Fan Y, Liu G, Zhang H (2009) Isolation and identification of a strain of Aspergillus tubingensis with deoxynivalenol biotransformation capability. Int J Mol Sci 9:2366–2375
He J, Zhou T, Young JC, Boland GJ, Scott PM (2010) Chemical and biological transformations for detoxification of trichothecene mycotoxins in human and animal food chains: a review. Trend Food Sci Technol 21:67–76
Hedman R, Pettersson H (1997) Transformation of nivalenol by gastrointestinal microbes. Arch Tierernahr 50:321–329
Higa A, Kimura M, Mimori K, Ochiai-Fukuda T, Tokai T, Takahashi-Ando N, Nishiuchi T, Igawa T, Fujimura M, Hamamoto H, Usami R, Yamaguchi I (2003) Expression in cereal plants of genes that inactivate Fusarium mycotoxins. Biosci Biotechnol Biochem 67:914–918
Hohn TM, Peters C, Salmeron JM, Reed JN, Dawson JL (2000) Transgenic plant and methods. International patent application WO 00/60061
Hohn TM, Peters C, Salmeron JM (2002) Trichothecene-resistant transgenic plants. US Patent 6:346–655
Ikunaga Y, Sato I, Grond S, Numaziri N, Yoshida S, Yamaya H, Hiradate S, Hasegawa M, Toshima H, Koitabashi M, Ito M, Karlovsky P, Tsushima S (2011) Nocardioides sp. strain WSN05-2, isolated from a wheat field, degrades deoxynivalenol, producing the novel intermediate 3-epi-deoxynivalenol. Appl Microbiol Biotechnol 89:419–427
Ivie GW (1976) Epoxide to olefin: a novel biotransformation in the rumen. Science 4230:959–961
Jansen C, von Wettstein D, Schäfer W, Kogel KH, Felk A, Maier FJ (2005) Infection patterns in barley and wheat spikes inoculated with wild-type and trichodiene synthase gene disrupted Fusarium graminearum. Proc Natl Acad Sci USA 102:16892–16897
Jarvis BB, Midiwo JO, Tuthill D, Bean GA (1981) Interaction between the antibiotic trichothecenes and the higher plant Baccharis megapotamica. Science 214:460–462
Jesenska Z, Sajbidorova I (1991) T-2 toxin degradation by micromycetes. J Hyg Epidemiol Immunol 35:41–49
Johnson SA, Sanford SA (1998) Parasite-derived resistance. U S Patent 5:840–481
Jouany JP (2007) Methods for preventing, decontaminating and minimizing the toxicity of mycotoxins in feeds. Anim Feed Sci Technol 137:342–362
Karlovsky P (1999) Biological detoxification of fungal toxins and its use in plant breeding, feed and food production. Nat Toxins 7:1–23
Khatibi PA, Newmister SA, Rayment I, McCormick SP, Alexander NJ, Schmale DG (2011) Bioprospecting for trichothecene 3-O-acetyltransferases in the fungal genus Fusarium yields functional enzymes with different abilities to modify the mycotoxin deoxynivalenol. Appl Environ Microbiol 77:1162–1170
Kim YW, Sharma RP, Eisner Y (1991) Effects of T-2 toxin and its congeners on membrane functions of cultured human fibroblasts. Mycotoxin Res 7:19–28
Kimura M, Kaneko I, Komiyama M, Takatsuki A, Koshino H, Yoneyama K, Yamaguchi I (1998a) Trichothecene 3-O-acetyltransferase protects both the producing organism and transformed yeast from related mycotoxins. Cloning and characterization of Tri101. J Biol Chem 273:1654–1661
Kimura M, Shingu Y, Yoneyama K, Yamaguchi I (1998b) Features of Tri101, the trichothecene 3-O-acetyltransferase gene, related to the self-defense mechanism in Fusarium graminearum. Biosci Biotechnol Biochem 62:1033–1036
King RR, McQueen RD, Levesque D, Greenhalgh R (1984) Transformation of deoxynivalenol (vomitoxin) by rumen microorganisms. J Agric Food Chem 32:1181–1183
Kollarczik B, Gareis M, Hanelt M (1994) In vitro transformation of the Fusarium mycotoxins deoxynivalenol and zearalenone by the normal gut microflora of pigs. Nat Toxins 2:105–110
Lake BG, Phillips JC, Walters DG, Bayley DL, Cook MW, ThomasLV GJ, Startin JR, Baldwin NCP, Bycroft BW, Dewick PM (1987) Studies on the metabolism of deoxynivalenol in the rat. Food Chem Toxicol 25:589–592
Lemmens M, Scholz U, Berthiller F, Dall’Asta C, Koutnik A, Schuhmacher R, Adam G, Buerstmayr H, Mesterházy A, Krska R, Ruckenbauer P (2005) The ability to detoxify the mycotoxin deoxynivalenol colocalizes with a major quantitative trait locus for Fusarium head blight resistance in wheat. Mol Plant Microbe Interact 18:1318–1324
Lulin M, Yi S, Aizhong C, Zengjun Q, Liping X, Peidu C, Dajun L, Xiu-E W (2010) Molecular cloning and characterization of an up-regulated UDP-glucosyltransferase gene induced by DON from Triticum aestivum L. cv. Wangshuibai. Mol Biol Rep 37:785–795
Maier FJ, Miedaner T, Hadeler B, Felk A, Salomon S, Lemmens M, Kassner H, Schäfer W (2006) Involvement of trichothecenes in fusarioses of wheat, barley and maize evaluated by gene disruption of the trichodiene synthase (Tri5) gene in three field isolates of different chemotype and virulence. Mol Plant Pathol 7:449–461
Manoharan M, Dahleen LS, Hohn TM, Neate SM, Yu XH, Alexander NJ, McCormick SP, Bregitzer P, Schwarz PB, Horsley RD (2006) Expression of 3-OH trichothecene acetyltransferase in barley (Hordeum vulgare L.) and effects on deoxynivalenol. Plant Sci 171:699–706
Matsushima T, Okamoto E, Miyagawa E, Matsui Y, Shimizu H, Asano K (1996) Deacetylation of diacetoxyscirpenol to 15-acetoxyscirpenol by rumen bacteria. J Gen Appl Microbiol 42:225–234
McCormick SP, Alexander NJ, Trapp SE, Hohn TM (1999) Disruption of TRI101, the gene encoding trichothecene 3-O-acetyltransferase, from Fusarium sporotrichioides. Appl Environ Microbiol 65:5252–5256
Meister A (1987) Detoxification of mycotoxins and other compounds of military interest. U.S. Army Medical Research and Development Command Annual Reports, Fort Detrick
Miller JD, Arnison PG (1986) Degradation of deoxynivalenol by suspension cultures of the Fusarium head blight resistant wheat cultivar Frontana. Can J Plant Pathol 8:147–150
Muhitch MJ, McCormick SP, Alexander NJ, Hohn TM (2000) Transgenic expression of the TRI101 or PDR5 gene increases resistance of tobacco to the phytotoxic effects of the trichothecene 4,15-diacetoxyscirpenol. Plant Sci 157:201–207
Munger CE, Ivie GW, Christopher RJ, Hammock BD, Phillips TD (1987) Acetylation deacetylation reactions of T-2, acetyl T-2, HT-2, and acetyl HT-2 toxins in bovine rumen fluid in vitro. J Agric Food Chem 35:354–358
Nagy CM, Fejer SN, Berek L, Molnar J, Viskolcz B (2005) Hydrogen bondings in deoxynivalenol conformations—a density functional study. J Mol Struct THEOCHEM 726:55–59
Nakamura Y, Ohta M, Ueno Y (1977) Reactivity of 12, 13-epoxytrichothecenes with epoxide hydrolase, glutathione-S-transferase and glutathione. Chem Pharm Bull 25:3410–3414
Nielsen C, Casteel M, Didier A, Dietrich R, Märtlbauer E (2009) Trichothecene-induced cytotoxicity on human cell lines. Mycotoxin Res 25:77–84
Ohsato S, Ochiai-Fukuda T, Nishiuchi T, Takahashi-Ando N, Koizumi S, Hamamoto H, Kudo T, Yamaguchi I, Kimura M (2007) Transgenic rice plants expressing trichothecene 3-O-acetyltransferase show resistance to the Fusarium phytotoxin deoxynivalenol. Plant Cell Rep 26:531–538
Okubara PA, Blechl AE, McCormick SP, Alexander NJ, Dill-Macky R, Hohn TM (2002) Engineering deoxynivalenol metabolism in wheat through the expression of a fungal trichothecene acetyltransferase gene. Theor Appl Genet 106:74–83
Paskevicius A, Bakutis B, Baliukoniene V, Sakalyte J (2006) The search for ecologically safe means of mycotoxin detoxification in fodder. Ekologija 3:128–131
Pestka JJ (2010) Deoxynivalenol: mechanisms of action, human exposure, and toxicological relevance. Arch Toxicol 84:663–679
Plank B, Schuh M, Binder EM (2009) Investigations on the effect of two feed additives, Biomin BBSH 797 and Mycofix Plus (R) 3.E, as detoxificants of DON contaminated feed of piglets. Wien Tierarztl Monatssch 96:55–71
Poppenberger B, Berthiller F, Lucyshyn D, Sieberer T, Schuhmacher R, Krska R, Kuchler K, Glossl J, Luschnig C, Adam G (2003) Detoxification of the Fusarium mycotoxin deoxynivalenol by a UDP-glucosyltransferase from Arabidopsis thaliana. J Biol Chem 278:47905–47914
Poppenberger B, Adam G, Berthiller F, Krska R, Kuchler K, Luschnig C, Glossl J, Lucyshyn D, Schuhmacher R, Sieberer T (2006) Method for detoxification of mycotoxins. US Patent Application 0183202
Proctor RH, Hohn TM, McCormick SP (1995) Reduced virulence of Gibberella zeae caused by disruption of a trichothecene toxin biosynthetic gene. Mol Plant Microbe Interact 8:593–601
Pronyk C, Cenkowski S, Abramson D (2006) Superheated steam reduction of deoxynivalenol in naturally contaminated wheat kernels. Food Control 17:789–796
Rocha O, Ansari K, Doohan FM (2005) Effects of trichothecene mycotoxins on eukaryotic cells: a review. Food Addit Contam Part A 22:369–378
Rohlfs M, Churchill AC (2011) Fungal secondary metabolites as modulators of interactions with insects and other arthropods. Fungal Genet Biol 48:23–34
Rotter B, Thompson BK, Clarkin S, Owen TC (1993) Rapid colorimetric bioassay for screening of Fusarium mycotoxins. Nat Toxins 1:303–307
Rotter B, Prelusky DB, Pestka JJ (1996) Toxicology of deoxynivalenol (vomitoxin). J Toxicol Env Health 48:1–34
Sato N, Ueno A (1977) Comparative toxicities of trichothecenes. In: Rodricks JV, Hesseltine CW, Mehlman MA (eds) Mycotoxins in human and animal health. Pathotox Publishers, Park Forest South, IL, pp 295–307
Schatzmayr G, Zehner F, Täubel M, Schatzmayr D, Klimitsch A, Loibner AP, Binder EM (2006) Microbiologicals for deactivating mycotoxins. Mol Nutr Food Res 50:543–551
Schweiger W, Boddu J, Shin S, Poppenberger B, Berthiller F, Lemmens M, Muehlbauer GJ, Adam G (2010) Validation of a candidate deoxynivalenol-inactivating UDP-glucosyltransferase from barley by heterologous expression in yeast. Mol Plant Microbe Interact 23:977–986
Sewald N, JvG L, Schuster M, Müller G, Aplin RT (1992) Structure elucidation of a plant metabolite of 4-desoxynivalenol. Tetrahedron-Asymmetry 3:953–960
Shen SK, Dowd PF (1991) Detoxification spectrum of the cigarette beetle symbiont Symbiotaphrina kochii in culture. Entomol Exp Appl 60:51–59
Shima J, Takase S, Takahashi Y, Iwai Y, Fujimoto H, Yamazaki M, Ochi K (1997) Novel detoxification of the trichothecene mycotoxin deoxynivalenol by a soil bacterium isolated by enrichment culture. Appl Environ Microbiol 63:3825–3830
Sobrova P, Adam V, Vasatkova A, Beklova M, Zeman L, Kizek R (2010) Deoxynivalenol and its toxicity. Interdiscip Toxicol 3:94–99
Styriak I, Conkova E (2002) Microbial binding and biodegradation of mycotoxins. Vet Hum Toxicol 44:358–361
Styriak I, Conkova E, Kmec V, Böhm J, Razzazi E (2001) The use of yeast for microbial degradation of some selected mycotoxins. Mycotoxin Res 17:24–27
Subramanian V (2002) DNA shuffling to produce nucleic acids for mycotoxin detoxification. US Patent 6:500–639
Swanson SP, Nicoletti J, Rood HD Jr, Buck WB, Cote LM, Yoshizawa T (1987a) Metabolism of three trichothecene mycotoxins, T-2 toxin, diacetoxyscirpenol and deoxynivalenol, by bovine rumen microorganisms. J Chromatogr 414:335–342
Swanson SP, Rood HD Jr, Behrens JC, Sanders PE (1987b) Preparation and characterization of the deepoxy trichothecenes: deepoxy HT-2, deepoxy T-2 triol, deepoxy T-2 tetraol, deepoxy 15-monoacetoxyscirpenol, and deepoxy scirpentriol. Appl Environ Microbiol 53:2821–2826
Theisen S, Berger S (2005) Screening of epoxide hydrolase producing microorganisms for biotransformation of deoxynivalenol. Mycotoxin Res 21:71–73
Tucker (2001) The “Yellow Rain” controversy: lessons from the arms control compliance. The Nonproliferation Rev Spring 2001:25–42
Ueno Y, Nakajima M, Sakai K, Ishii K, Sato N (1973) Comparative toxicology of trichothec mycotoxins: inhibition of protein synthesis in animal cells. J Biochem 74:285–296
Ueno Y, Nakayama K, Ishii K, Tashiro F, Minoda Y, Omori T, Komagata K (1983) Metabolism of T-2 toxin in Curtobacterium sp. strain 114–2. Appl Env Microbiol 46:120–127
Visconti A, Mirocha C (1985) Identification of various T-2 toxin metabolites in chicken excreta and tissues. Appl Environ Microbiol 49:1246
Volkl A, Vogler B, Schollenberger M, Karlovsky P (2004) Microbial detoxification of mycotoxin deoxynivalenol. J Basic Microbiol 44:147–156
Wang YZ, Miller JD (1988) Effects of Fusarium graminearum metabolites on wheat tissue in relation to Fusarium head blight resistance. J Phytopath 122:118–125
Wannemacher RW, Wiener SL (1997) Potential military chemical/biological agents and compounds. In: Army Field Manual No 3–9. U.S. Government Printing Office, Washington, pp. 655–676
Wei CM, Hansen BS, Vaughan MH Jr, McLaughlin CS (1974) Mechanism of action of the mycotoxin trichodermin, a 12,13-epoxytrichothecene. Proc Natl Acad Sci USA 71:713–717
Westlake K, Mackie RI, Dutton MF (1987) Effects of several mycotoxins on specific growth rate of Butyrivibrio fibrisolvens and toxin degradation in vitro. Appl Environ Microbiol 53:613–614
Wilson SC, Brasel TL, Martin JM, Wu C, Andriychuk L, Douglas DR, Cobos L, Straus DC (2005) Efficacy of chlorine dioxide as a gas and in solution in the inactivation of two trichothecene mycotoxins. Int J Toxicol 24:181–186
Worrell NR, Mallett AK, Cook WM, Baldwin NC, Shepherd MJ (1989) The role of gut microorganisms in the metabolism of deoxynivalenol administered to rats. Xenobiotica 19:25–32
Yamaguchi I, Kimura M, Takatsuki A, Hiroyuki E, Katsumi Y (2000) Trichothecene 3-O-acetyltransferase gene. Japanese Patent No. 200032985
Yao Q, Liu Z, Zeng Y (1996) Detoxification of deoxynivalenol by scab resistant wheat and the bioactivities of the product. Acta Mycol Sin 15:59–64
Yoshizawa T, Takeda H, Ohi T (1983) Structure of a novel metabolite from deoxynivalenol, a trichothecene mycotoxin, in animals. Agric Biol Chem 47:2133–2135
Yoshizawa T, Sakamoto T, Kuwamura K (1985) Structures of deepoxytrichothecene metabolites from 3′-hydroxy HT-2 toxin and T-2 tetraol in rats. Appl Environ Microbiol 50:676–679
Young JC, Subryan LM, Potts D, McLaren ME, Gobran FH (1986) Reduction in levels of deoxynivalenol in contaminated wheat by chemical and physical treatment. J Agric Food Chem 34:461–465
Young JC, Trenholm HL, Friend DW, Prelusky DB (1987) Detoxification of deoxynivalenol with sodium bisulfite and evaluation of the effects when pure mycotoxin or contaminated corn was treated and given to pigs. J Agric Food Chem 35:259–261
Young JC, Zhu H, Zhou T (2006) Degradation of trichothecene mycotoxins by aqueous ozone. Food Chem Toxicol 44:417–424
Young JC, Zhou T, Yu H, Zhu H, Gong J (2007) Degradation of trichothecene mycotoxins by chicken intestinal microbes. Food Chem Toxicol 45:136–143
Yu H, Zhou T, Gong J, Young C, Su X, Li XZ, Zhu H, Tsao R, Yang R (2010) Isolation of deoxynivalenol-transforming bacteria from the chicken intestines using the approach of PCR-DGGE guided microbial selection. BMC Microbiol 10:182
Zhou T, He J, Gong J (2008) Microbial transformation of trichothecene mycotoxins. World Mycotox J 1:23–30
Zhou T, Gong J, Yu H, Li XZ (2010) Bacterial isolate and methods for detoxification of trichothecene mycotoxins. US Patent application no. 12/671,514
Acknowledgment
The author is indebted to numerous colleagues for stimulating discussions and sharing unpublished results, and to the reviewers for the valuable comments.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Karlovsky, P. Biological detoxification of the mycotoxin deoxynivalenol and its use in genetically engineered crops and feed additives. Appl Microbiol Biotechnol 91, 491–504 (2011). https://doi.org/10.1007/s00253-011-3401-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-011-3401-5